Abstract
Existing methods for voxelwise transient dopamine (DA) release detection rely on explicit kinetic modeling of the [11C]raclopride PET time activity curve, which at the voxel level is typically confounded by noise, leading to poor performance for detection of low-amplitude DA release-induced signals. Here we present a novel data-driven, task-informed method—referred to as Residual Space Detection (RSD)—that transforms PET time activity curves to a residual space where DA release-induced perturbations can be isolated and processed. Using simulations, we demonstrate that this method significantly increases detection performance compared to existing kinetic model-based methods for low-magnitude DA release (simulated +100% peak increase in basal DA concentration). In addition, results from nine healthy controls injected with a single bolus of [11C]raclopride performing a finger tapping motor task are shown as proof-of-concept. The ability to detect relatively low magnitudes of dopamine release in the human brain using a single bolus injection, while achieving higher statistical power than previous methods, may additionally enable more complex analyses of neurotransmitter systems. Moreover, RSD is readily generalizable to multiple tasks performed during a single PET scan, further extending the capabilities of task-based single-bolus protocols.
Introduction
Dopamine release in PET
Neurotransmitter release can be detected in vivo using dynamic (4 D) positron emission tomography (PET) imaging. To date, the most common mechanism for detection relies on competition for binding between endogenous neurotransmitter and a PET ligand; 1 during release, a localized time-varying increase in endogenous neurotransmitter concentration results in a decrease in ligand binding in that region, ultimately causing a decrease in the PET time activity curve (TAC).
The majority of research thus far has focused on the neurotransmitter dopamine (DA), due to the important role DA plays in healthy brain functioning and to the favorable binding properties of the relatively commonly available PET ligand [11C]raclopride (RAC), a D2/D3 receptor antagonist. RAC DA-detection frameworks mostly confine their search to the D2-rich regions of the striatum where the signal-to-noise ratio (SNR) is adequate for reliable detection. DA is involved in motor function, executive function, and reward,2 –4 making it a useful research target for neurodegenerative diseases, neuropsychiatric disorders, and neuropsychological tasks in the healthy and diseased brain.
In its earliest form, this competition mechanism was exploited in a double scan protocol (double RAC), where one scan occurred with the subject in the baseline condition and a second scan occurred with a sustained stimulus evoking DA release.
5
The change in binding potential estimate from the baseline to stimulus scan is a proxy for the evaluation of the magnitude of DA release. However, such a protocol has several disadvantages: scan-to-scan variability in the binding potential confounds stimulus-evoked changes; increased commitment and radiation dose is required for study participants; and the release magnitude is assumed to be constant over time for analysis purposes, which may not be true. More advanced double RAC analysis methods leveraged the residual behavior between the baseline and activated scans by use of singular value decomposition (SVD) to accurately determine DA release at varied task timings,
6
however this method is practically limited since
Transient release modeling
More recently, several competition models have been suggested to quantify the transient kinetics of neurotransmitter release within a single scan: the subject is scanned in baseline condition during a first part of the scan that includes tracer uptake, then the stimulus is presented later in the scan to observe and quantify any transient responses.8 –10 Such frameworks use compartmental models that include time-varying kinetic parameters to model the dynamic competition between the tracer and endogenous neurotransmitter.
Stimulus-evoked endogenous DA changes often occur over second or sub-second intervals. 11 However, the clearance rate of RAC is on the order of minutes, meaning high frequency endogenous DA changes are mapped to low frequency RAC changes. This is similar—conceptually—to the hemodynamic response in functional magnetic resonance imaging (fMRI): neuronal activation occurs on the order of milliseconds, but the measured hemodynamic response via the Blood Oxygen Level Dependent (BOLD) signal evolves on the order of seconds.12,13
Transient release detection
An initial voxelwise model comparison framework was formalized and optimized to detect and localize transient DA release.14,15 The framework employed a voxelwise F-test between the multilinear reference tissue method (MRTM)
16
—incapable of modeling competition—and a method that models the competition, linear parametric neurotransmitter PET (lp-ntPET),
10
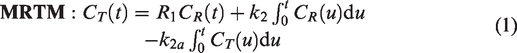
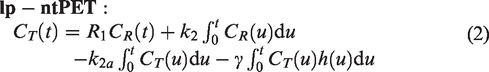
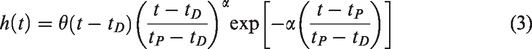
where
More recently several additional approaches have been developed in an effort to increase the accuracy and reliability of detection: direct reconstruction with lp-ntPET, 20 a machine learning denoising approach using a feed-forward perceptron neural network, 21 a machine learning detection method that trains personalized neural networks on simulated lp-ntPET data, 22 a Monte Carlo extension to the F-test approach, 23 and a Bayesian framework for lp-ntPET parameter estimation. 24 At the core, though, all of these methods rely on the accuracy of a kinetic model—namely lp-ntPET—and a reliable fit of this model to the data.
Disadvantages of kinetic model-based detection
The lp-ntPET model incorporates a family of basis functions that describe physiologically-plausible release timecourses. Thus, the number of degrees of freedom used to parametrize the F-distribution used for significance testing needs to be modified based on the number of basis functions chosen; failing to do so will alter the expected false positive rate (FPR). 25 Additionally, the F-test itself assumes that the data are normally distributed, but image noise in PET is best described by a gamma distribution. 26 Furthermore, detection of release is greatly improved by significant post-processing, 15 which presumably further violates the normality assumption of the F-test.
While the model comparison approach has produced high detection sensitivity in simulations of relatively high levels of DA release expected in pharmacological challenges, such as cigarette smoking,14,15 model comparison in simulations of lower DA release levels—which typically are present in motor, cognitive, or neuropsychological tasks—11,27,28 proved less successful.
23
The major goals of this work are thus three-fold:
Develop a DA release detection method that does not rely on direct kinetic modeling. Improve detection sensitivity for lower levels of DA release. Demonstrate feasibility of the method for a single bolus injection of RAC.
Kinetic models like lp-ntPET were initially developed to accurately quantify release and not specifically to be embedded within a detection framework. 10 Thus, we hypothesize that decoupling kinetic modeling from detection will increase the reliability and localization of DA release, after which lp-ntPET or a similar model may be used to accurately quantify the release profiles at these locations. If our hypothesis is verified—especially for lower levels of DA release—the ability of PET to detect transient fluctuations in DA levels in response to motor, cognitive, or neuropsychological tasks will enhance studies of the dopaminergic system.
Isolating DA-induced perturbations via residuals analysis
As an initial effort to decouple detection from kinetic modeling, we propose to shift detection to a residual space where DA-induced perturbations to the TAC are isolated. The percentage difference between an observed voxel TAC and a data-driven hypothetical baseline TAC define the voxelwise residuals. Any change to the observed TAC during the task will then appear as structured deviations in the residual space. This mapping to the residual space has conceptual similarities to previous work, 29 and acts to remove the confound of baseline pharmacokinetics and provide a constant signal during baseline. A model comparison via F-test is thus no longer required and simpler detection methods—like those used in fMRI—may be applied. We hypothesize that the isolation of DA-induced perturbations will increase detection sensitivity compared to the model comparison approach by overcoming some of its shortcomings. In principle, this further enables the detection of multiple-task, condition-specific DA release using residual analysis.
Due to typically high levels of noise in voxel-level TACs it is necessary to apply denoising to the data before applying subsequent analyses on voxel residuals. To best characterize the voxel residuals we rely on a combination of novel denoising methods, namely HYPR4D kernelized reconstruction,30,31 along with denoising using IHYPR4D, 32 both of which extend the HYPR framework to the temporal domain (Figure 1). HYPR4D kernelized reconstruction has been shown to outperform the standard and current state-of-the-art clinical reconstructions in terms of 4 D noise reduction while preserving spatiotemporal patterns within data. IHYPR4D has been shown to provide improvements over HYPR3D in joint noise and bias reduction, and inherently bypasses limitations imposed by HYPR3D; for specific details see literature30 –32.
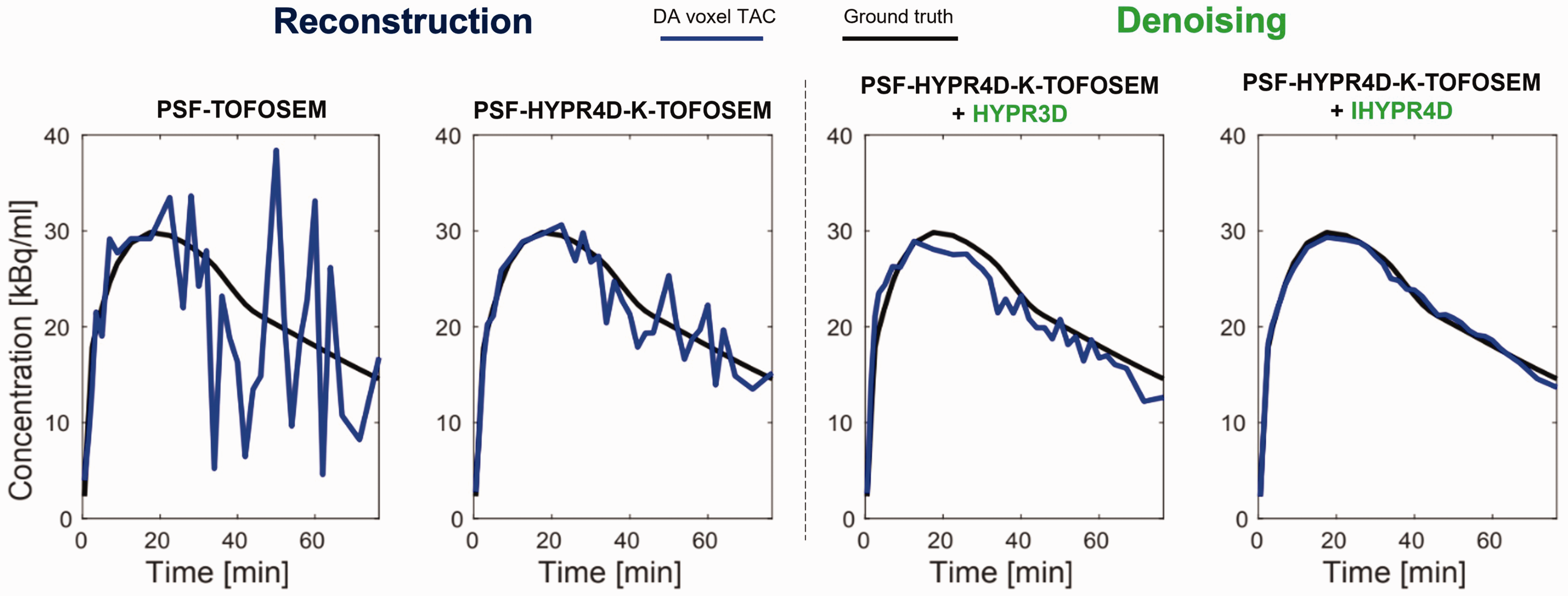
Simulated noisy voxel TACs with different reconstruction and denoising approaches. Gaussian noise is added to simulated single-task release ground truth voxel TACs, with noise levels matching real noise measured on the GE SIGNA PET/MR for each reconstruction protocol. Results in this paper use PSF-HYPR4D-K-TOFOSEM+IHYPR4D (right panel).
This paper proceeds as follows. We first outline how to calculate the residuals from acquired data in the context of a DA release experiment. We then propose a detection method in residual space inspired by fMRI 33 : using a simpler General Linear Model (GLM) rather than kinetic modeling in image space. Collectively, we refer to this process as Residual Space Detection (RSD). Next, we test and optimize RSD on simulated single-task DA release experiments under the conditions of homogeneous and heterogeneous RAC binding; previous simulation studies assume homogenous RAC baseline binding across the striatum, which may not be valid in diseased populations that are of interest for DA studies, such as Parkinson’s disease (see Supplemental Fig. 2). Finally, the optimized RSD method is applied to single-task human scans as a proof-of-concept. Simulation and human results from RSD are compared with results from the traditional lp-ntPET model comparison method.
Theory
Transforming time activity data to a residual space
The first step of this work is to define a hypothetical baseline TAC for each voxel that may be observed in the absence of DA release. DA release in response to non-pharmacological interventions is likely to be localized, 34 and of relatively low amplitude, meaning that a significant fraction of striatal voxel TACs are unlikely to contain DA-induced perturbations. Therefore, on first approximation it is reasonable to assume that the mean striatal TAC appears mostly devoid of DA-induced perturbations and can approximate baseline conditions.
Under the assumption that the mean striatal TAC and measured voxel TACs behave very similarly prior to a DA release-inducing task, a first attempt to define the voxel-level baseline TAC is to use the mean striatal TAC and fit it to a given voxel TAC for timepoints prior to task initiation. To attain a full scan-length voxel-level baseline TAC, the scaling factor from this pre-task fit can be applied to the entire mean striatal TAC, hence extrapolating baseline behavior to later time points. Comparing this extrapolated fit with the measured voxel TAC yields structured residuals in voxels where DA release is present (Figure 3).
Two immediate issues arise when attempting this protocol: (i) TACs of voxels near the boundaries of the striatum have poor fits to the mean striatum TAC due to partial volume effects, and (ii) in the presence of heterogeneous tracer binding within the striatum, defining a single mean striatal TAC as a regressor will result in biased fits and extrapolations for voxels that have higher or lower non-displaceable binding potentials (
The first issue is remedied by extracting the mean TAC from a reference region devoid of specific binding, in the case of RAC using the cerebellum, and adding this as a regressor to the pre-task fit. The second issue may not arise in general as the density of D2 receptors in the healthy striatum is relatively uniform
35
; this is however not always true, particularly for conditions affecting the striatum such as Parkinson's disease (PD). To resolve this, we propose applying k-means clustering to pre-task voxel TACs normalized by their peak value, with
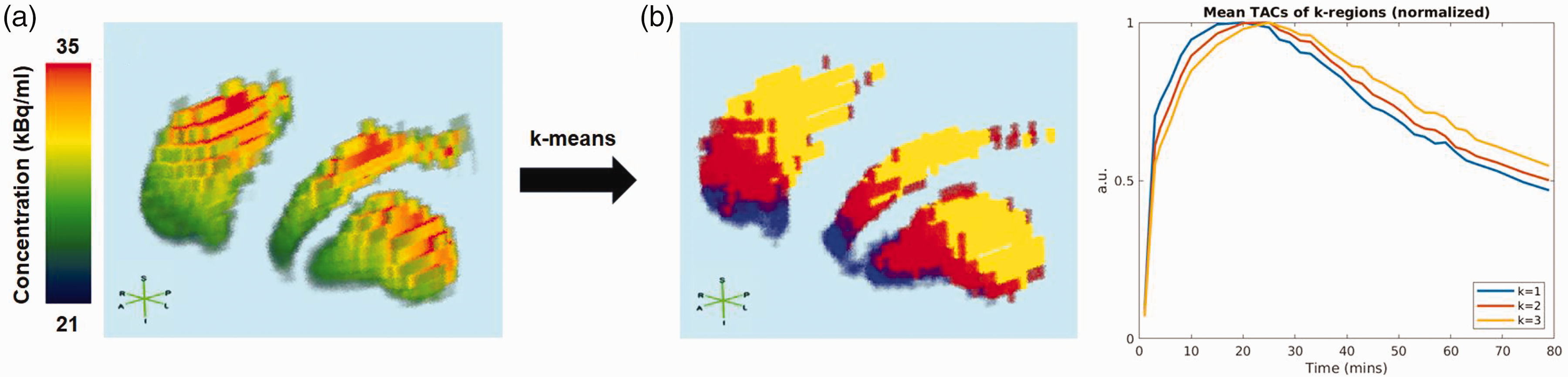
K-means results for noisy simulated data with an anterior-posterior/superior-inferior Bmax gradient. (a) Concentration of a simulated noise-free PET frame (frame 10, t = 25 min) displaying the effect of a simulated Bmax gradient in the striatum. (b) Spatial k-regions found by running k-means on pre-task normalized TACs from a noisy simulation. Normalized mean TACs of each k-region are displayed on the plot, demonstrating spatially-varied binding characteristics. These TACs, along with the mean cerebellum TAC, are used to define the voxelwise set of baseline TACs. Voxel and TAC colors are consistent.
Before the commencement of a DA release-inducing task, a striatal voxel TAC, 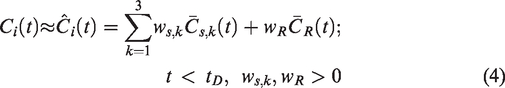
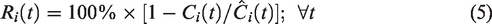
Note, by this construction, a negative DA-induced perturbation in the voxel TAC post-intervention,
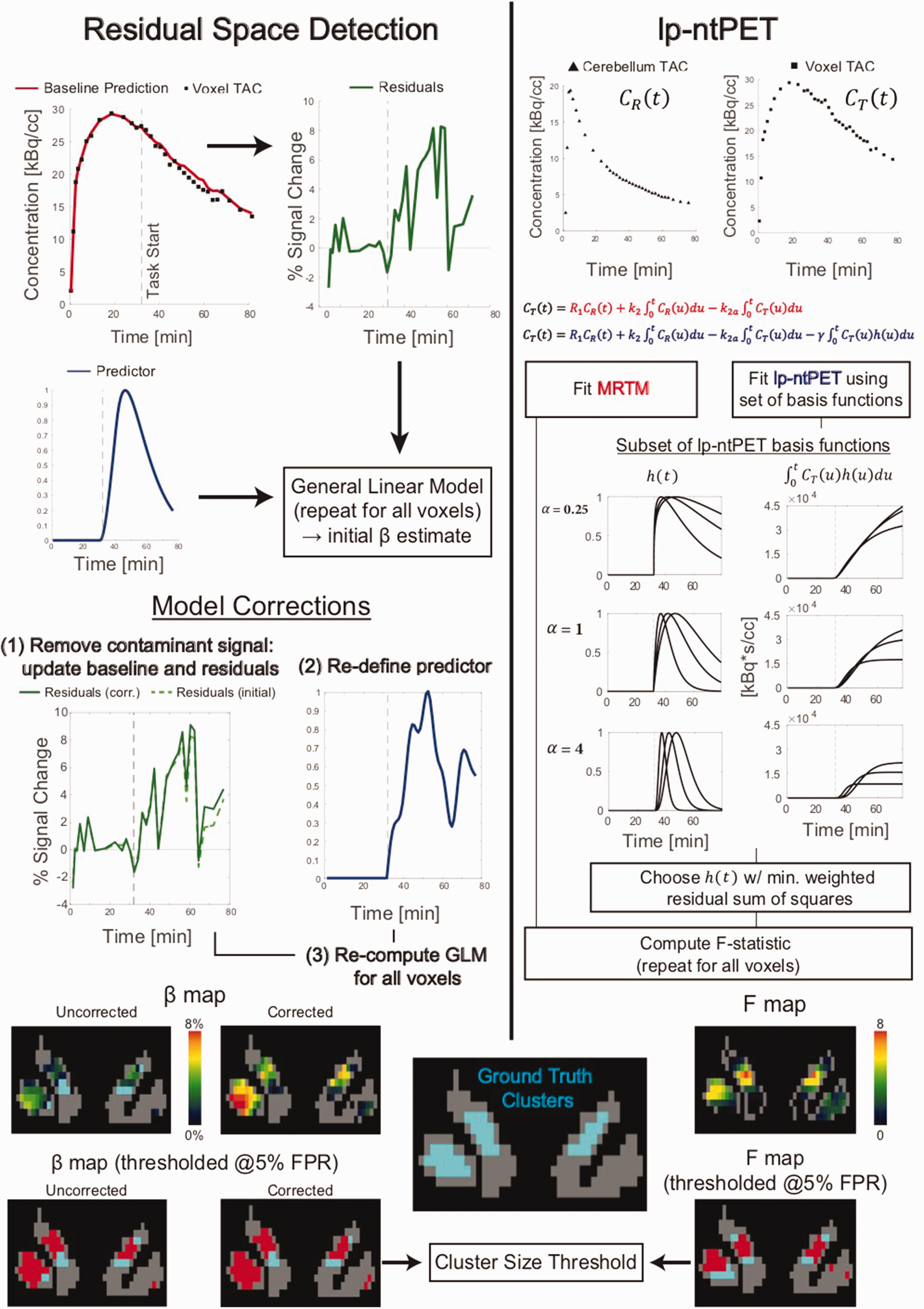
Summary of the Residual Space Detection (RSD) and lp-ntPET methods.
Residual space detection (RSD)
Transforming the PET TACs to residual space allows for analysis and subsequent detection of voxel TACs that deviate from the extrapolated mean baseline behavior as a consequence of task-evoked response. The basis of RSD is to compare residuals to a predictor,
Inspired by fMRI detection approaches—where task events/blocks are convolved with a hemodynamic response function to yield predictors—we first define the predictor as the convolution between an event vector, 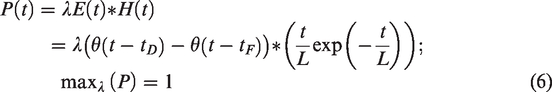
A simple means of comparing residuals to predictors is to employ GLM.
36
GLM in the case of RSD is written as

Finally, to minimize bias and increase accuracy in our results, we shift the method to being more data-driven by iteratively (i) redefining baseline TACs to remove possible contamination from DA release-related signals, and (ii) redefining predictors to reduce possible bias introduced by an incorrectly assumed response function (Figure 3). Baseline TACs, and thus residuals, are iteratively redefined following an initial determination of 
Data-driven predictors are then defined using
From simulations it was observed that only one iterative update of this protocol was required to achieve predictions of baseline voxel
Methods and materials
Simulation design
To test our method in comparison to the lp-ntPET method, we simulated 79-minute single-task RAC scans on the GE SIGNA PET/MR system (GE Healthcare, Chicago, IL, USA). A T1 MRI from a healthy human subject scanned on the SIGNA was segmented using FreeSurfer 37 to generate an anatomical reference for simulating regions with pre-determined RAC kinetics; the segmented image was then resampled to the PET voxel dimensions (1.39 × 1.39 × 2.78 mm) using nearest-neighbor interpolation. The striatum was defined using bilateral ROIs from the putamen, caudate, and nucleus accumbens.
Noiseless TACs were simulated following the ntPET model, 9 using the same approach as Wang et al. 15 Briefly, the arterial input function was modeled as in literature 38 to match concentration levels observed in 15 mCi RAC injections on the GE SIGNA PET/MR scanner at our centre. Kinetic parameters and modeling from literature 39 were used alongside pre-specified DA concentration increases and timings to obtain simulated TACs.
DA release clusters were placed in the left caudate (cluster size: 118 voxels), in the right putamen (160 voxels), and in the right caudate (104 voxels) to simulate a possible response to a single 10-minute functional task block starting at 32 minutes, involving motor and cognitive processing. The peak simulated DA release magnitude was set to 100% increase from basal DA concentration (for full details see Supplemental Material and Supplemental Figure 1); this is relatively low compared to previous studies which simulated peak DA increases of 200% or more relative to baseline.14,15
To ensure our method applies more generally, we then performed identical simulations in the presence of spatially heterogeneous RAC binding to assess the performance of the k-means methodology. Motivated by observed binding heterogeneities in subjects with PD (Supplemental Figure 2) we simulated a gradient in the binding potential (BP), accomplished by modulating 
The TACs were framed 4 × 1, 3 × 2, 4 × 5 minutes for estimation of the baseline portion of the scan, then 20 × 2 minutes for higher temporal sampling during DA release, and finally 1 × 4, and 2 × 5 minutes at the end of the scan for better counting statistics. The dynamic images were then spatially filtered with a 1.8 × 1.8 × 3.6 mm FWHM Gaussian to simulate the SIGNA point spread function (PSF). 40 We generated 100 noisy realizations (NRs) using scaled Gaussian noise, 14 before post-processing with IHYPR4D (7.2 mm, two frames spatiotemporal kernel, one iteration) 30 ; the scaling coefficient was set so that the noise level after applying IHYPR4D matched human RAC scan noise levels reconstructed with PSF-HYPR4D-K-TOFOSEM (5.6 mm, ten frames spatiotemporal kernel, 10 iterations) 31 and post-processed with IHYPR4D.
Implementation of RSD
Cluster size thresholding
To control for the multiple comparisons problem, clusters of predicted DA release voxels in the binary maps were subject to a cluster size threshold (CST), where clusters smaller than the CST were rejected as DA release clusters. 15 In our simulations the left putamen was kept devoid of release, therefore we used this region to define the CST. The CST was set for each simulated dataset and methodology independently and defined so that 90% of noisy realizations resulted in the left putamen being completely free of false positives.
Comparison to model-based lp-ntPET method
We compare our proposed RSD method to the existing model-based lp-ntPET method as described in literature.
15
Both MRTM and lp-ntPET (equations (1) and (2) were fit to each voxel TAC in the striatum. Parameters for lp-ntPET basis functions,
Weighted residual sum of squares (WRSS) was calculated for lp-ntPET and MRTM fits, which permitted the calculation of the F-statistic. Voxels with an F-statistic greater than a critical value were accepted as release voxels. The critical value was varied similarly to the thresholding performed in RSD to generate ROC curves for each simulated release cluster, and eventually chosen to have a matching FPR to that of RSD. Note that this is in contrast to previous studies with lp-ntPET that used a fixed threshold for the F-statistic,14,15 corresponding to a theoretical p-value < 0.05 as defined by the degrees of freedom in the models. The purpose of variable thresholding employed in this work is to enable a comparison of detection sensitivity between the two methods at matched FPRs. F-statistic thresholding defined binary maps for each noisy realization, which then went through the same in-plane edge voxel removal and CST processing as RSD.
Performance metrics
To determine the performance of our method in comparison with the lp-ntPET method we used two sensitivity measures as defined in literature.
15
(1)
Human scans
Scan protocol
As proof-of-concept, we present results from a simple finger tapping motor task study. Nine healthy controls (mean age = 64.8, SD = 7.6, range = 47–73) were scanned on the GE SIGNA PET/MR scanner. All participants were right-handed. The UBC Research Ethics Board (following ethical standards of the Helsinki Declaration of 1975 and as revised in 1983) approved this study (certificate number: H19-03166), and written informed consent was obtained for each subject prior to all study procedures.
At 36 minutes post bolus injection of RAC (550 MBq; 14.86 mCi), subjects tapped fingers of their dominant hand for 10 minutes, in blocks of 2 minutes with 15 seconds of rest between blocks. The task involved serial tapping of the index finger, middle finger, ring finger, and little finger, then reversed (‘1-2-3-4, 4-3-2-1’…), at the subject’s own pace.
Image processing
Raw PET data were reconstructed using PSF-HYPR4D-K-TOFOSEM, 31 with attenuation, scatter, randoms, and normalization corrections, after which PET images underwent frame realignment (i.e. inter-frame motion correction). Dynamic PET images were denoised using IHYPR4D, with the same reconstruction and post-processing parameters as previously mentioned. Masks of the striatum and cerebellum were extracted from subject-specific FreeSurfer segmentations of structural T1 images for use in RSD and lp-ntPET. To create group-level images, subject T1 images were normalized to the Neurodevelopmental MRI Database (age 65-69) atlas with a 2 mm isotropic voxel size, 41 using ANTs normalization software. 42
Image analysis
RSD and lp-ntPET were separately performed on subject-space dynamic PET images. RSD
Results
Simulations
Release in the presence of homogeneous binding
Figure 4 displays voxelwise prediction maps calculated from the set of binary classification maps (thresholded at 5% FPR) across 100 NRs for the lp-ntPET + F-test method vs. the proposed RSD method, in addition to ROC curves for the cognitive clusters (bilateral caudate) and motor cluster (right putamen). CST was determined for each method from the set of binary maps and was subsequently applied to each individual binary map; the resulting voxelwise sensitivity values after CST application are given in Table 1.
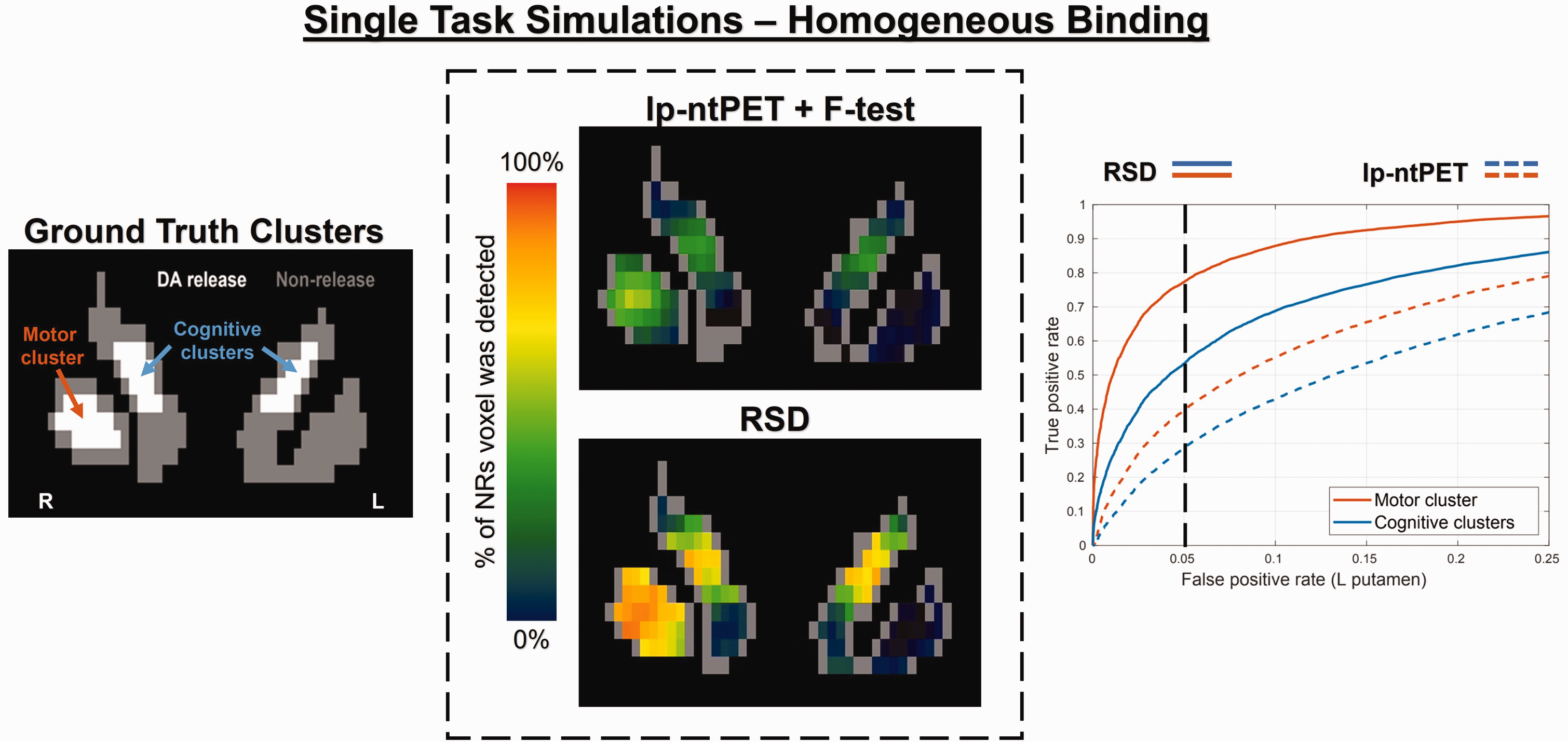
Simulated single-task release – homogeneous RAC binding.
Comparison between RSD and lp-ntPET, with sensitivities and CST for both simulation sets.
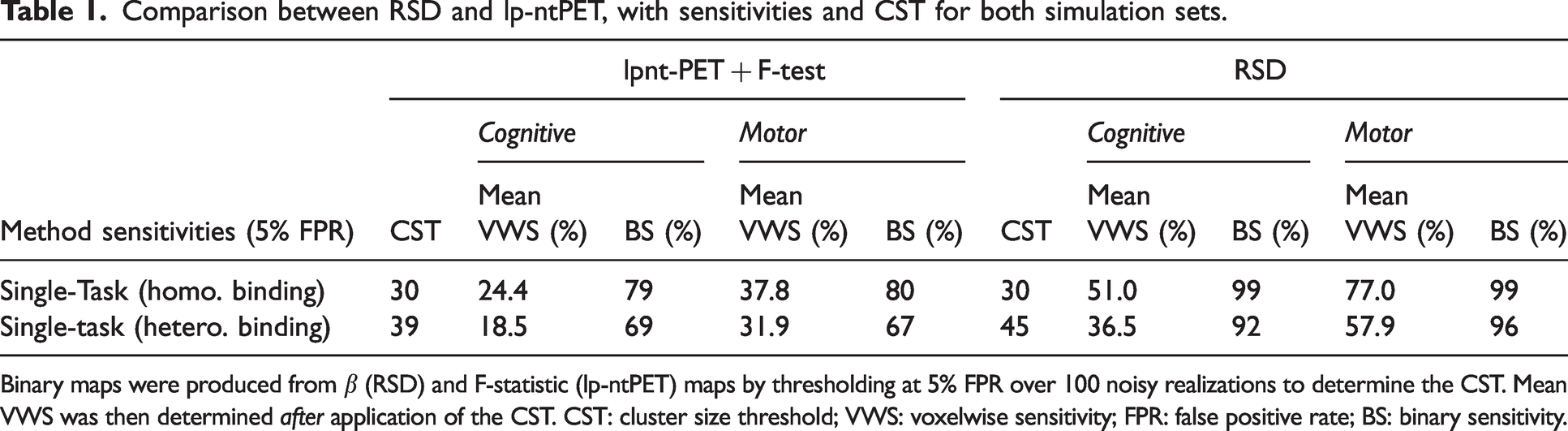
Binary maps were produced from
As seen in the ROC curves, RSD drastically outperforms lp-ntPET across all FPRs for this low-magnitude single-task release (
Release in the presence of heterogeneous binding
Results for the heterogeneous binding simulations prior to application of CST are displayed in Figure 5, while Table 1 summarizes the heterogeneous binding simulation results after application of the CST. Voxelwise sensitivity is reduced in both methods and clusters when compared with the results for homogeneous binding, with the RSD method still considerably outperforming lp-ntPET. Binary sensitivity of RSD remains quite high in these clusters, resulting from relatively high peak
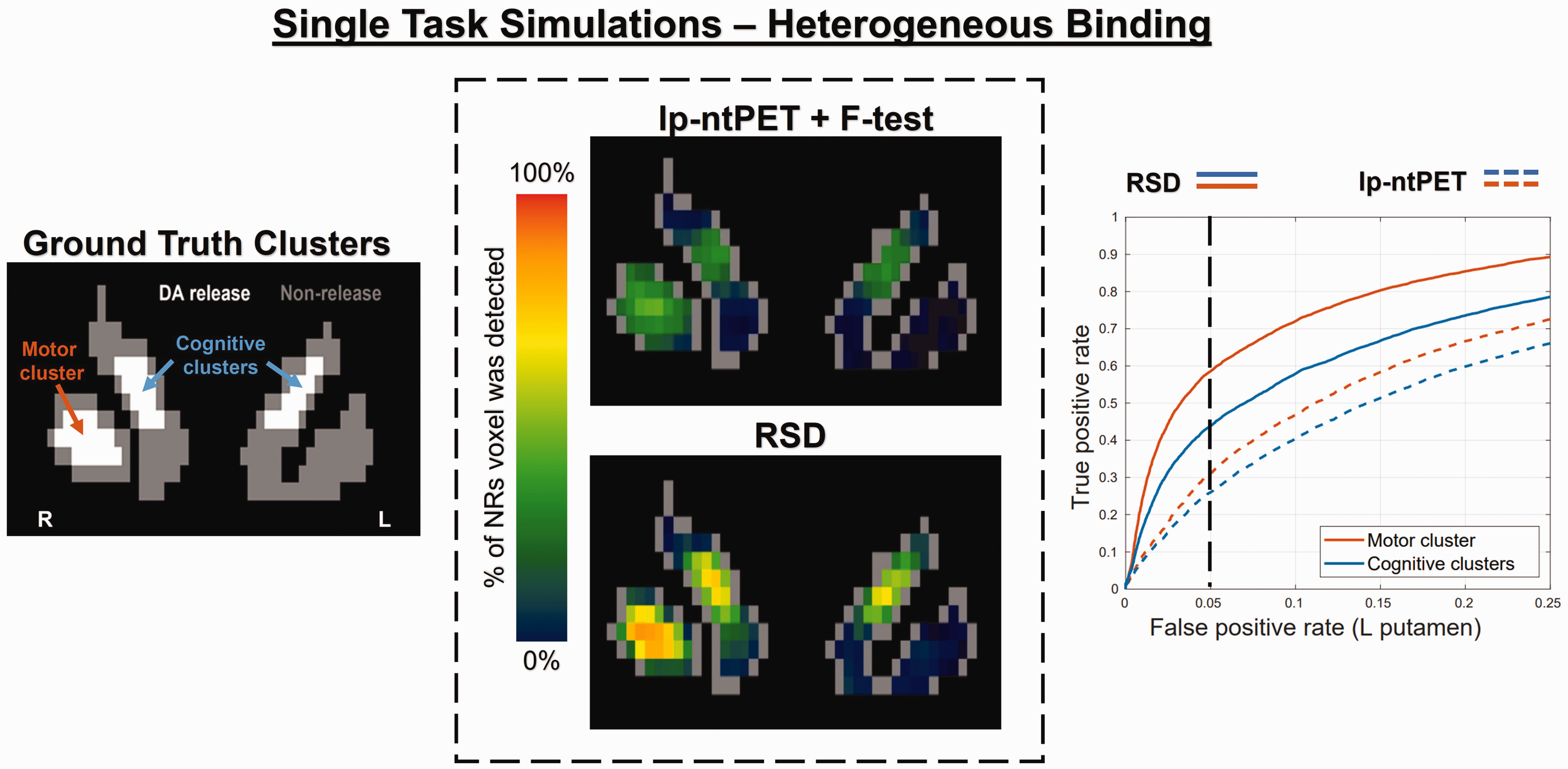
Simulated single-task release – heterogeneous RAC binding.
Human scans
Results from the proof-of-concept study with 9 healthy control subjects are shown in Figure 6, displaying statistical parametric maps from RSD and lp-ntPET. Due to the intrinsically high variability of lp-ntPET
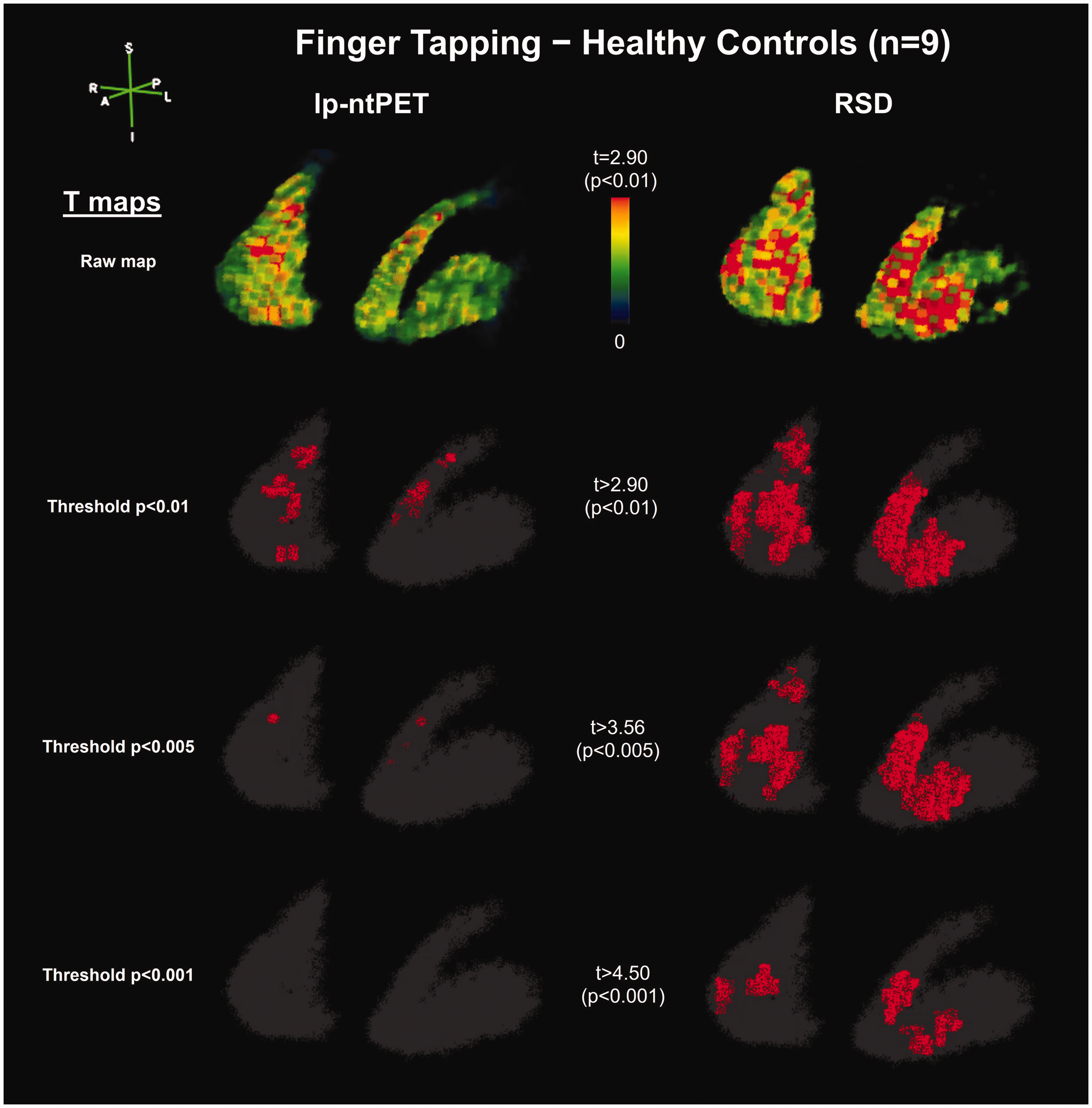
Motor task human results – statistical parametric maps. Subject-level
Discussion
This work presents a novel detection framework for PET-based DA release detection, most advantageous for spatially localized and low-magnitude responses which are inherently difficult to detect. Such release may be expected in motor, cognitive, and/or neuropsychological reward-based tasks rather than pharmacological interventions,11,27,28 where the spatial extent and magnitude of release is much greater. 43 Given the increasing incidence of addiction and neurodegenerative diseases, which alter dopaminergic function, the ability to detect dopamine release in response to non-pharmacological stimuli is becoming increasingly more important.
Our detection framework has two major differences compared to existing methods which primarily incorporate significance testing between models to detect release:
We have demonstrated that the RSD method improves detection sensitivity of single-task protocols in the presence of both homogeneous and heterogeneous RAC binding. While we did not test these methods on non-denoised data, we can confidently claim that the denoising algorithms employed significantly aided DA release detection, as has been shown in previous work with HYPR-based denoising applied to voxelwise DA release detection. 15
Simulations
In this work we simulated two separate datasets: single-task release with homogeneous RAC binding and with heterogeneous RAC binding. In both simulations, RSD outperformed existing model-based methods employing MRTM/lp-ntPET fitting alongside an F-test. RSD provides higher binary and voxelwise sensitivities for clusters of low-amplitude DA release when compared to lp-ntPET at equivalent FPR, exhibiting improved robustness to noise and greater ability to distinguish release from non-release.
In the presence of positive noise spikes prior to task initiation and/or negative noise spikes during the task, lp-ntPET is especially prone to predicting release when it does not exist. In these scenarios lp-ntPET overfits to the noise-altered TAC by tweaking any of its four free parameters (
Heterogeneous binding
When observing scans of patients with PD, it was clear that the binding homogeneity assumption was not always satisfied (see Supplemental Material), and thus using a single mean striatal TAC as regressor in RSD was insufficient; when the mean striatal TAC (mid-BP) is fit pre-task to a relatively low-BP voxel TAC, the resulting baseline prediction will consistently overshoot the measured voxel TAC, resulting in large positive residuals post-task that are consistently predicted as DA release. To remedy this, we introduced a k-means step that supplies further baseline regressors in a data-driven manner, which partially accounted for this issue by removing these structured biases.
Results from RSD for single-task simulations with heterogeneous binding provided moderate to high detection sensitivity as shown in Figure 5 and Table 1 at 5% FPR, albeit with diminished performance relative to the homogeneous binding simulations. This was to be expected to some degree; as implemented, RSD made use of
Human scans
To test the performance of RSD relative to lp-ntPET, we performed scanning and analysis on a cohort of nine healthy controls. RSD found significantly more consistent results across subjects as demonstrated in Figure 6, while lp-ntPET showed highly variable output, with several subjects’
Statistical parametric maps were computed by means of one-sample t-test (Figure 6), using the
Limitations
Limitations of RSD include the possible presence of TAC shapes that cannot be described accurately by the data-derived baseline TAC regressors. This may occur in the case of extreme binding heterogeneities; though if such heterogeneities are observed, the k-means protocol may be adapted to attempt to account for its presence. Another possibility is that the localization assumption is violated for physiological tasks, and the majority of the striatum is activated by DA release such that the baseline regressors are unavoidably contaminated. In this scenario RSD may fail to detect the full extent of release and may be limited to detecting the regions of highest TAC deflection.
It has been demonstrated that [11C]raclopride TACs may also be sensitive to task-induced blood flow changes, though this perturbation has opposite polarity and a different temporal shape to that from task-induced DA release,
39
which would result in negative
Additionally, the predictors presented in this work are by construct only sensitive to task-induced DA release close to task initiation. This may be a limitation if there is a reason to suspect significant DA release before task initiation, such as potentially present in a task anticipatory stage. We simulated cases where the predictor timing is partially offset from the true release time (up to ±5 minutes) and found that RSD sensitivity diminishes, but still outperforms lp-ntPET + F-test (shown in Supplemental Figure 8). RSD somewhat accounts for this possibility in its iteratively updated predictor by incorporating residuals up to 5 minutes prior to task initiation, though careful evaluation of the TAC behavior and parameter choice needs to be performed to fully describe such behavior.
From our simulations, the accuracy of the shape of the predictor is less important for detection than accurate quantification; RSD assigns
Conclusion
In the presence of low-magnitude DA release in response to a task-based single-bolus PET protocol we have shown that our proposed method, Residual Space Detection (RSD), is capable of achieving high detection sensitivity at low FPR, outperforming the traditional lp-ntPET + F-test method for localized, low-amplitude release. As shown using simulations and proof-of-concept human scanning, RSD achieves superior performance as a result of its robustness to noise and data-driven methodology. Future work will explore the extension of RSD to two-task release in attempts of differentiating function-specific (e.g. cognitive, reward) release from confounding motor release in cohorts of healthy and diseased individuals.
Supplemental Material
sj-pdf-1-jcb-10.1177_0271678X231214823 - Supplemental material for Novel voxelwise residual analysis of [11C]raclopride PET data improves detection of low-amplitude dopamine release
Supplemental material, sj-pdf-1-jcb-10.1177_0271678X231214823 for Novel voxelwise residual analysis of [11C]raclopride PET data improves detection of low-amplitude dopamine release by Connor WJ Bevington, Jordan U Hanania, Giovanni Ferraresso, Ju-Chieh (Kevin) Cheng, Alexandra Pavel, Dongning Su, A Jon Stoessl and Vesna Sossi in Journal of Cerebral Blood Flow & Metabolism
Footnotes
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Natural Sciences and Engineering Research Council grant (240670-13) and by the Pacific Parkinson’s Research Institute. TRIUMF is supported by the National Research Council Canada.
Acknowledgements
The authors thank TRIUMF staff for preparation of the radiotracers and the UBC PET/MRI Imaging Centre nurses and technologists for their assistance during the human scans.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Authors’ contributions
Supplementary material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
