Abstract
Off-target evaluation is essential in preclinical safety assessments of novel biotherapeutics, supporting lead molecule selection, endpoint selection in toxicology studies, and regulatory requirements for first-in-human trials. Off-target interaction of a therapeutic antibody and antibody derivatives has been historically assessed via the Tissue Cross-Reactivity (TCR) study, in which the candidate molecule is used as a reagent in immunohistochemistry (IHC) to assess binding of the candidate molecule to a panel of human tissue sections. The TCR approach is limited by the performance of the therapeutic as an IHC reagent, which is often suboptimal to outright infeasible. Furthermore, binding of the therapeutic in IHC conditions typically has poor in vitro to in vivo translation and lacks qualitative data of the identity of putative off-targets limiting the decisional value of the data. More recently, cell-based protein arrays (CBPA) that allow for screening against a large portion of the human membrane proteome and secretome have emerged as a complement, and likely a higher value alternative, to IHC-based off-target assessment. These arrays identify specific protein interactions and may be useful for testing nontraditional antibody-based therapeutic formats that are unsuitable for TCR studies. This article presents an overview of CBPA technologies in the context of TCR and off-target assessment studies. Selected case examples and strategic considerations covering a range of different modalities are presented.
Keywords
Introduction
The assessment of antibodies and antibody derivatives specificity is recommended by regulatory agencies prior to human clinical trials. Tissue cross-reactivity (TCR) studies are typically required, with the exception for serious and life-threatening oncology indications, for which alternative evaluations should be considered. While biologics are typically considered to be highly specific to their target(s), evidence has demonstrated a need to better characterize the potential for off-target effects as part of a comprehensive safety assessment. Structural components of biologics that may contribute to off-target interactions include antibody fragment antigen binding (Fab)-, single-chain fragment variable (scFv)- and fragment crystallizable (Fc)-mediated interactions.1,27
While there is no prescribed technique required for off-target assessment, the most common approach is the TCR study. TCR studies use the candidate therapeutic molecule as a reagent in an IHC assay to identify potential target organs based on the binding of the therapeutic agent to specific cell or tissue components.18,25 For many years, the value of TCR studies in predicting toxicity to humans that would not otherwise have been detected by in vivo pharmacology and toxicology animal studies has been a subject of debate. Industry surveys and publications have highlighted the challenges associated with TCR as well as limited to no value in supporting a candidate molecule safety assessment.5,25,31 TCR methods depend on the ability of the candidate molecule to perform as a suitable IHC reagent, which can be technically challenging, time-consuming, or unfeasible to develop, especially for novel or unconventional formats. The availability and suitability of robust positive controls is also essential. Furthermore, TCR results do not provide information on the specific proteins being bound, an essential detail for proper hazard and risk assessment of the off-target profile of a therapeutic. Given the limitations of TCR assay development and interpretation, CBPA technologies have emerged as a complement and, in some cases, an alternative to TCR studies. 31 Increasingly, CBPA studies are being used to inform lead selection, improve identification of molecule specificity, and increase the efficiency of identifying off-target binding. In this article, we will provide an overview of CBPA applications, including case examples, and present considerations for derisking off-target interactions.
Regulatory Guidance
The pioneering guidance mandating TCR studies (“Points to Consider in the Manufacture and Testing of Monoclonal Antibody Products For Human Use”) (PTC) was issued by the U.S. Food and Drug Administration (FDA) in 1983 and was last updated in 1997. 45 Further guidance from the International Council on Harmonization (ICH) were issued, focusing on biopharmaceuticals in “ICH S6(R1)” (1997, updated 2011) and anticancer pharmaceuticals in “ICH S9” (2010) and “ICH S9 Q&A” (2018).19-21
Both the PTC and ICH S6(R1) guidelines acknowledge that sponsors may employ alternative methods for assessing the specificity of monoclonal antibodies or related therapeutics. The ICH S6(R1) document mentions “Other technologies can be employed in place of IHC techniques to demonstrate target/binding site distribution.” Similarly, the PTC document advises, “Appropriate newer technologies should be employed as they become available and validated.”
In addition, the ICH S9 Q&A document clarified that, for complementary-determining region (CDR) containing anticancer biopharmaceuticals with preclinical pharmacologic activity, “tissue cross reactivity studies have little utility and are not needed with the initial first-in-human study or later in development, unless there is a specific cause for concern. In cases where there are no pharmacologically relevant species, human tissue cross reactivity or alternative methods should be considered for the first-in-human study”.
Recently, the FDA provided nonbinding recommendations for chimeric antigen receptor (CAR)-T cell therapies, including considerations to evaluate their specificity using methods like TCRs and protein arrays. 44 Contributors’ experience with regulatory interactions and submissions will be discussed below in the case summaries and discussion.
General Selectivity Assessment of Novel Biologic Therapeutics
Biologic therapeutics should ideally bind specifically to the intended target in the “diseased” tissue. It is therefore important to assess whether the candidate cross-reacts with other proteins (off-target/polyspecificity) but also whether it recognizes its target protein(s) in a normal, “healthy” tissue. A lack of selectivity can be associated with safety and potential pharmacokinetic consequences.12,27,29,31,48 Finally, biologics can exhibit unwanted low-affinity binding to extracellular matrix and/or membrane. This chemical “stickiness,” also called “polyreactivity” is related to some physicochemical properties of antibodies such as excessive positive charge or hydrophobicity.2,9 It can impact both the pharmacokinetic profile and bioavailability. Various approaches to detect and avoid polyreactivity at an early stage of the development process have been reviewed elsewhere and will not be discussed further. 9
Understanding therapeutic target expression is essential for putative liability identification that can result from on-target interactions, due to target epitope expression and localization in both normal and disease states. This knowledge is typically derived from target tissue expression profiling (TEP) studies. TEP studies use various methods to characterize spatial and cellular target distribution in human and preclinical toxicology species tissues. In situ hybridization (ISH; to localize genes or RNA) and immunohistochemistry (IHC; to localize protein) are the prototypic assays in a standard TEP data package, as they provide spatially resolved data with microanatomical context. Public databases exist (eg, GTEx and Human Protein Atlas) and can be complemented with Sponsor-directed ad hoc assays such as flow cytometry, western blot, and polymerase chain reactions (PCR). These public and proprietary data are integrated with literature, target biology, in silico, and data science deliverables. Cumulatively, this data forms the basis for target risk assessment, toxicology species selection, study design, and biomarker selection in preclinical and clinical studies. TEP assessments are detailed further in the literature. 36 Current strategies to evaluate off-target interactions include TCR studies, and protein and CBPA studies. Affinity approaches, followed via mass spectrometry, can also help characterize off-target identification for biologics. However, this requires knowledge of tissue/cells where unexpected binding is observed and therefore is not used as a screening tool.
TCR Studies
The original method to determine specificity of a therapeutic monoclonal antibody was guided by the 1983-1997 “Points to consider” documents from FDA on the conduct of TCR studies. These tissue-based assays involve the ex vivo IHC staining of a large panel of frozen normal tissues from humans and, as warranted, relevant toxicology species. TCR studies are filed with IND/CTA (investigational new drug application/clinical trial application) to support first-in-human (FIH) clinical trials. Regulatory agencies have provided guidance on the number of donors/panels and the tissue list to be investigated as well as some quality aspects (e.g. tissue integrity markers). 45 Additional considerations on the design and technical aspects are covered in the literature. 25 While TCR has been a mainstay for evaluation of on- and off-target binding potential, several publications have noted the challenges of this technology, notably IHC method development and limited ability to detect some secreted or circulating proteins lost during processing. If necessary, the need for additional molecular and biochemical efforts may be required to identify the off-target protein(s) binding.5,25,31
Cell-based Protein Array Studies
Two types of in vitro arrays exist for the evaluation of protein-protein interactions: peptide/protein arrays and cellular protein arrays.13,30,32,33,35,38,41 Peptide and protein arrays use matrix-associated antigens. Despite incremental improvements to minimize nonspecific binding and to enhance protein quality and conformation, proteins are expressed outside of their cellular context and proper physiological epitope presentation cannot be guaranteed, notably for proteins with multiple transmembrane domains. Peptide/protein assays will not be discussed further in this article, beyond case 2.
Cellular-based protein presentation assays are a more recent development. These arrays use cell lines transfected with a library of vectors encoding for proteins which are expressed in a more physiologically relevant state. The libraries are enriched for plasma membrane proteins and, more recently, for secreted proteins. Two commercially available platforms, the Retrogenix Human Cell Microarray (Charles River Labs) and the Membrane Proteome Array (MPA; Integral Molecular), will be discussed in more detail below.
Considerations for Cell-based Protein Arrays: Advantages and Limitations
CBPA offer several benefits such as higher-throughput capacity and reduced amount of required test article compared with TCR. Consequently, these assays can be deployed earlier in the drug development process. For instance, some sponsors are using those assays during lead selection to deprioritize nonspecific candidates, thus contributing to the reduction in animal testing, including nonhuman primates. These assays also allow for the identification of specific protein-protein interactions which can be essential for off-target hazard and risk assessment. In contrast to TCR, cellular arrays are also more likely to work with various biologic modalities. The CBPA platforms have been used to determine the binding interactions of a diverse range of test articles (TAs) including human and nonhuman antibodies, antibody-drug conjugates (ADC) and antibody fragments such as scFvs, Fabs, and VHH (variable domain of heavy-chain antibody) nanobodies.3,8,10,13,15,16,18,26,28,34,39,40,47 Peptides, proteins, and DARPins (designed ankyrin repeat proteins) have also been profiled as well as viruses or whole cells such as CAR-T cells.3,13,16,18,43 This was made possible by adaptation of the screening methodologies. For example, CAR-T cells can be labeled with a fluorescent dye prior to incubation on array slides allowing cell-to-protein interactions to be detected directly. However, individual authors found that profiling CAR-T cells in the array screens was more challenging, whereas profiling CAR-T targeting moieties (e.g., scFvs, VHHs) proved to be a more feasible and reproducible approach that has been accepted by regulators. Often, the specificity of CAR-Ts can be supplemented with additional functional assays to assess CAR-T cell activation or cytolytic activity in co-culture with target cells that lack the intended antigen or that express closely related proteins.
There are several limitations of CBPAs to consider. Data generated lack spatially relevant target information in the context of normal tissue architecture, cellular diversity, and extracellular matrix organization. The expression levels are typically several fold greater than typical endogenous protein level expression. Therefore, the biological relevance of the confirmed interactions must be considered in the context of endogenous protein expression levels on primary cells and distribution across a potentially diverse set of cell types. Another aspect to consider is the quality of the libraries. While both commercial cellular arrays claim to cover ≥90% of membrane proteins, this figure does not account for all existing protein isoforms. Although libraries often include multiple isoforms of a protein, they may not encompass all isoforms. It has also been occasionally noted by the authors that libraries may contain some outdated clones that encode proteins which do not match with the reference sequences, although it should be noted that many of the case studies reviewed here were from early versions of cell-based protein array libraries. Furthermore, while libraries are progressively improving, they can include some targets that are unlikely accessible to biologic candidates in vivo, such as cytoplasmic proteins, proteins located on intracellular membrane or plasma membrane proteins without any extracellular domain. Interactions with such proteins have been detected on those arrays 47 and may be secondary to compromised cellular membrane integrity, which could be an effect of the fixation process (Retrogenix Cell Microarray) or the fixation-free permeabilization process used to detach the adherent cells from the culture plate (MPA). Consequently, some of the detected interactions may not have any in vivo relevance. This also implies that interactions with surfaceome proteins might not occur at their final subcellular localization, notably because the high level of target expression may overwhelm the cellular transport machineries.
For all those reasons, CBPA studies are susceptible to false positives and negatives and should be regarded as screening assays. Any interactions (hits) require additional validation through orthogonal methods, and validation of any hits is important in any complete screening package to eliminate false positives.
Efforts are continuously made to improve and qualify CBPA assays. Notably, Integral Molecular’s Membrane Protein Array technology was recently accepted into the FDA ISTAND (Innovative Science and Technology Approaches for New Drugs) pilot program for evaluation as a drug development tool.22,46
Retrogenix Cell Microarray Platform
The Retrogenix Cell Microarray platform is an in vitro CBPA screening system that identifies binding partners for various therapeutic modalities. It uses a library of cDNAs covering around 6500 human proteins, including plasma membrane, secreted proteins and heterodimeric proteins. The library has no bias toward any protein classes and provides over 90% coverage of the cell surface and secreted proteome. It also includes over 98% of prenatal proteins present at 8 to 10 weeks of pregnancy, vital for data on potential off-target interactions in unrepresented patients in clinical trials. A cynomolgus monkey library is also available to evaluate off-target interactions in monkeys.
All test articles are initially assessed for background binding to untransfected HEK293 cells or cells transfected with positive and negative target controls. The CBPA is prepared by spotting each library cDNA vector onto microarray slides and allowing HEK293 cells to be transfected to express the protein encoded on each cDNA spot. TAs are added and binding to individual overexpressed proteins on cells is assessed using fluorescent detection reagents and a fluorescent slide scanner. Significant interactions can be further assessed using flow cytometry. 7
Integral Molecular—Membrane Protein Array
The Integral Molecular’s Membrane Protein Array platform is an in vitro CBPA screening system that identifies binding partners for various therapeutic modalities. It uses a library of cDNAs covering around 6000 human proteins, covering the majority of all potential membrane proteins, including intracellular; a secreted proteome library is under development. The library has no bias toward any protein classes and provides over 94% coverage of the full human membrane proteome, including developmental proteins.
The MPA screen is performed using high-throughput flow cytometry. Before specificity testing, optimal conditions for screening are determined in an assay setup experiment using primarily human HEK-293 cells or avian QT6 cells, as a nonhuman cell line alternative, expressing positive and negative (mock-transfected) binding controls. Once conditions are optimized, protein targets are overexpressed using plasmids individually transfected into cells present in 384-well plates. Cells from each well are then pooled in duplicate in differing groups of 48 to facilitate the speed of flow cytometry detection, followed by matrix deconvolution. Protein targets identified as potential binding interaction in the MPA screen are tested individually in a titration experiment to validate the interactions.6,23
Cell-based Protein Array Comparison
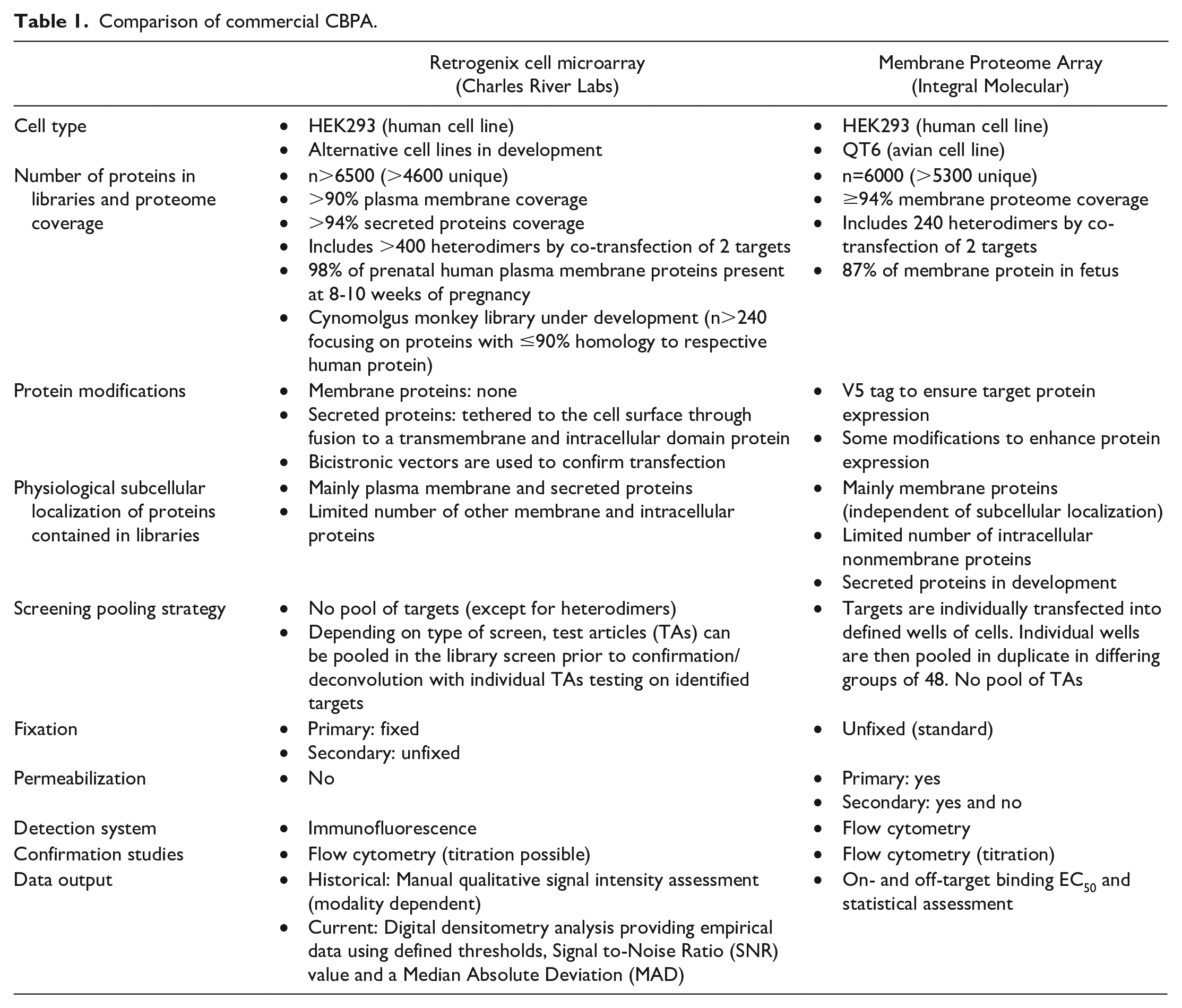
The Retrogenix and Integral Molecular CBPA technology are similar to the extent of enabling systematic and high-throughput analysis of therapeutic binding interactions with membrane proteins. Both assays are available as fee-for-service platforms. Distinctions between the two platforms include differences in technologies used to detect binding (immunofluorescence or flow cytometry), the coverage and quality of the protein libraries, use of live or fixed cells, screening pooling strategy (combining multiple targets or several TAs) and the timing of flow cytometry (screening or validation) potentially leading to variations in target and off-target binding due to inherent platform differences (Table 1).
Comparison of commercial CBPA.
General Use Cases
A variety of use cases have been employed by the contributors at various therapeutic development stages. Microarray can be used to support lead selection, candidate derisking, receptor deorphanization (identification of targets with unknown endogenous ligands or biologic functions), and format evaluation. Some companies may elect to evaluate the final candidate after full biophysical characterization, while others use the assay to evaluate several potential lead candidates earlier in preclinical development. CBPA can be used in conjunction with IHC-based methods for on- and off-target distribution and to compare specificity between commercial antibodies and lead antibody candidates. In addition, this may be employed when a suitable TCR method cannot be developed. Finally, microarrays may be employed in late development to de-risk clinical findings and further contextualize biodistribution data.
Cell-based Protein Array Case Examples
A variety of microarray case examples are presented below and as supplementary material. They represent a diversity of TAs (ADCs, bispecific antibodies, Fabs, IgG1, IgG4, scFv, VHH), at various stages of the biologic development and safety assessment pipeline– from lead selection to late development. The case examples below are intended for illustrative purposes and reflect the experiences of the contributors.
Case 1: Cell-based Protein Array for Early Selectivity Assessment and Candidate Selection
To support the selection of a bispecific antibody prior to administration in monkeys, three candidate molecules were profiled on the Retrogenix Human Cell Microarray to assess their specificity. Instead of the final bispecific candidates (targeting CD3 and an undisclosed target D), biotinylated Fabs targeting target D were used. As shown in Figure 1, the individual Fabs all recognized their primary target with medium to strong intensity. However, candidate 2 also recognized cells overexpressing two unrelated proteins: off-target A with weak to medium intensity and off-target B with weak intensity.
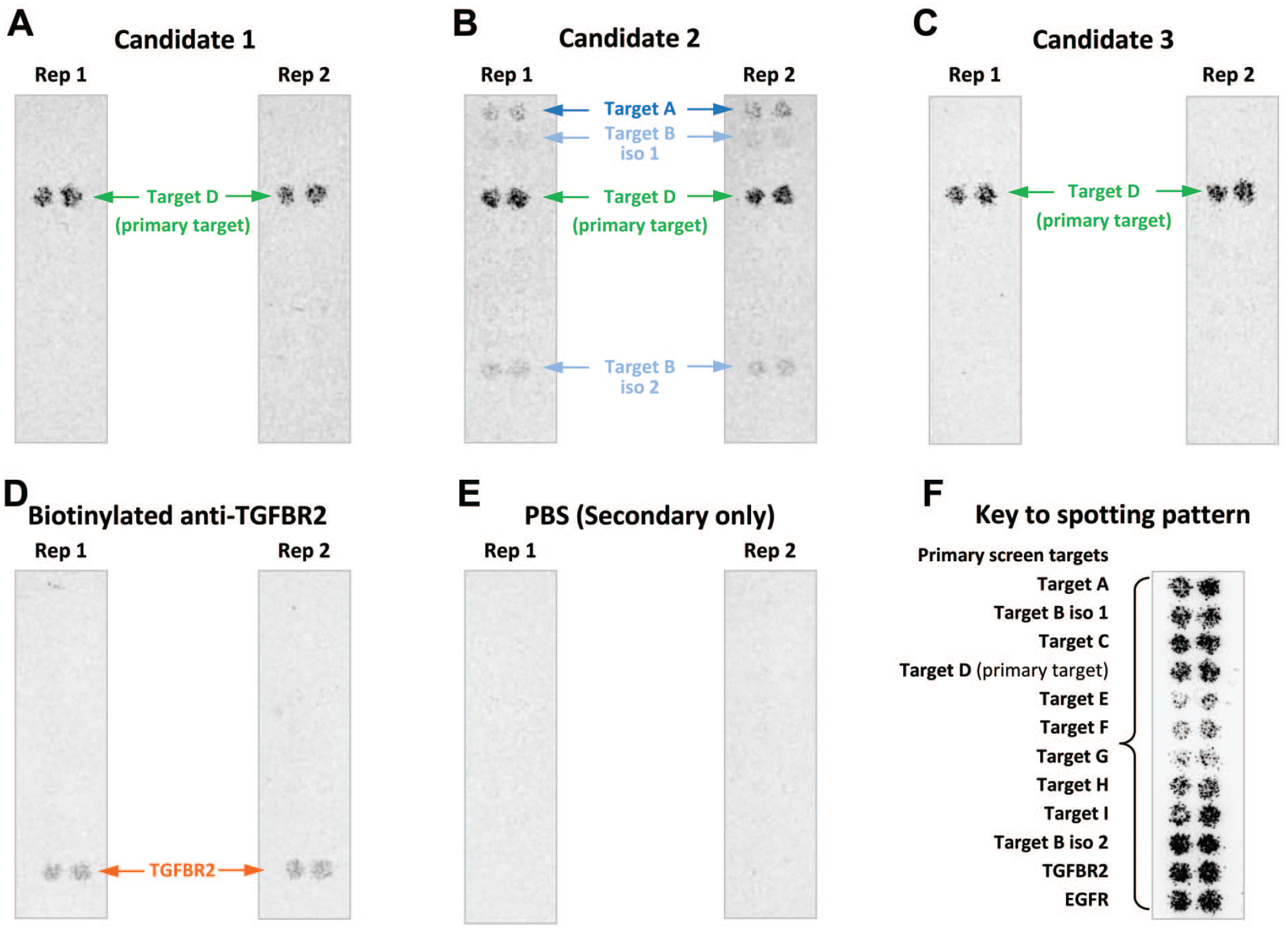
Early selectivity assessment of Fabs with a Retrogenix screen. Primary screen: 4575 bicistronic expression vectors, each encoding a full-length human plasma membrane protein and ZsGreen fluorescent protein, were arrayed in duplicate across 13 microarray slides. Human HEK293 cells were used for reverse transfection/expression. 3 biotinylated Fabs (Candidates 1, 2, and 3) were pooled, and the pool was added to each slide after cell fixation giving a final concentration of 10 µg/mL of each biotinylated Fab. Detection of binding was performed by using AlexaFluor647 streptavidin followed by fluorescence imaging. Two replicate slides were screened for each of the 13 slide-sets. A protein “hit” is defined as a spot showing a raised signal compared with background levels in the Alexa 647 channel. Ten primary hits were identified (data not shown).
To address the relevance of those off-targets a follow-up functional assessment was performed. The bispecific molecules were incubated with activated T cells and a cell line which express the off-target(s) but not the primary target D. Candidate 2 induced death of cells expressing the off-target “A” in a dose dependent manner (starting at single digit nanomolar concentrations), unlike candidates 1 and 3. Taking into account these data, the projected clinical Cmax, and tissue distribution of off-target A, candidate 2 was terminated. In contrast, the weak intensity off-target binding to target “B” did not lead to a functional consequence, as candidate 2 did not significantly kill cells expressing the off-target “B.”
Commentary: This case exemplifies how CBPA and followed by orthogonal hit validation can support early lead selection. Pivotal TCR studies are typically conducted on the final molecule, in parallel with investigational new drug-enabling toxicity studies. However, it is beneficial to eliminate nonspecific drug candidates earlier in development.
Case 2: Evaluation of Cell-based Protein Array Platform Consistency
A total of 5 molecules were profiled on both CBPA platforms (Table 2). The primary targets were consistently identified (4/4 for Retrogenix Human Cell Microarray, 3/4 for MPA). For the biologic product 2, the off-target “L” whose signal was clearly above assay thresholds was also identified by both platforms (Table 2, biologic product 2). Overlap was lower for weaker signal off-targets (Table 2, biologic products 4 and 5). No orthogonal validation was performed for the latter.
Testing of 5 biological products on commercial cellular array platforms.
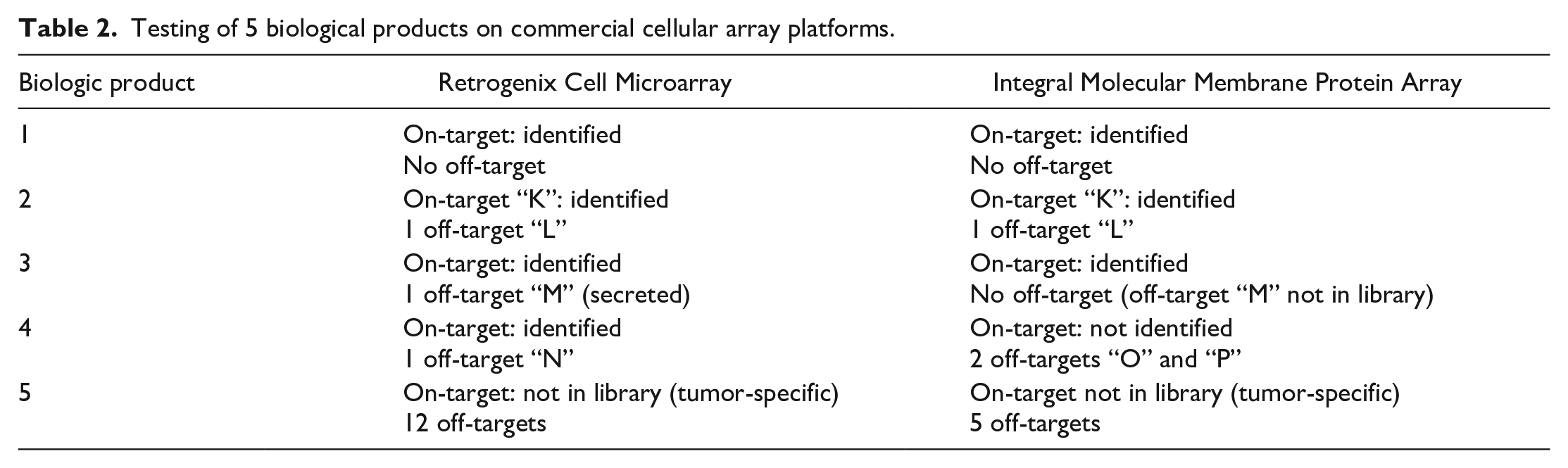
Those molecules were also profiled on a cell-free protein array but only 1 of the 4 primary targets were identified and the CBPA off-target “L” of biologic product 2 was not identified. One of the test molecules (biologic product 5) showed a high number of off-targets on all test platforms (12 for Retrogenix, 5 for Integral Molecular, 87 for the cell-free protein array) and overlap was limited to 6 hits between Retrogenix and the cell-free protein array.
Commentary: This case compares two CBPA platforms and a cell-free protein array technology. While more biologic products need to be compared in order to draw definitive conclusions, a key observation is that cellular assays show consistency for targets with a signal clearly above assay thresholds.
Case 3: Derisking Unanticipated Tissue Staining
A potential final clinical lead molecule, Candidate X (reformatted to a rabbit Fc) was evaluated for binding specificity using standard IHC methods on a human tissue microarray (TMA). Unexpectedly, intense staining was observed in tissues not known to express the target of interest, such as cardiac muscle, that was not observed with the isotype control. The same staining pattern was subsequently confirmed on full format tissue sections. Given the target expression profiling and known absence of the target antigen in the observed cell types/tissues, the IHC results were most likely an off-target interaction of the therapeutic binder.
To identify the off-target(s), human heart tissue homogenates were incubated with Candidate X and immunoprecipitation coupled with mass spectrometry analysis was used. This effort identified several peptides which by in silico analysis had homology to the intended target. The predicted off-targets had known expression in muscle, and therefore, the observed staining of these tissues with the Candidate X was consistent with the expected cellular location of these two targets.
A cell line, C2C12, was identified by transcriptomics and confirmed by proteomics to express the off-targets. Binding of Candidate X to cells was further confirmed by flow cytometry and immunofluorescence (IF) staining but was not observed with another antibody against the intended target or an isotype control antibody. In addition, surface plasmon resonance (SPR) studies further confirmed binding to the off-target proteins with high affinity.
At the time, it was not standard practice for the sponsor to use the CBPA technology to assess antibody specificity, but a Retrogenix Human Cell Microarray was eventually performed with Candidate X. The screen confirmed the off-targets previously identified, but also additional off-targets interactions. The team decided to deprioritize Candidate X and to screen another molecule as replacement, Candidate Y, with the Retrogenix Human Cell Microarray. Candidate Y demonstrated a better specificity profile and was selected as the new clinical lead.
Commentary: This case highlights the importance of considering early target derisking using CBPAs. It also illustrates the complementarity of IHC-based methods (unexpected tissue signal localization), CBPA and proteomics approaches in the identification of the off-target proteins. Results were consistent and confirmed by orthogonal approaches.
Case 4: Derisking Off-Target CBPA Interactions With Orthogonal In Vitro Assessments
In this example, the Retrogenix Human Cell Microarray was used to screen for off-target binding interactions of a bispecific human IgG1 antibody with primary targets A and B. Notably, this antibody was intended for a non-oncology indication. Route of administration was intravitreal. A proposal was made to two health authorities, the FDA and MHRA, to use CBPA data as an alternative to a TCR study, followed by assessment of potential off-targets, as needed. The proposal was considered to be “reasonable” by FDA and “acceptable” by MHRA.
The bispecific test article and isotype-matched controls (single target/isotype arms and isotype/isotype controls) were evaluated. Binding was screened against 5475 full-length human plasma membrane and secreted proteins and 371 heterodimers. The test article was bound to primary targets A and B (two isoforms each) in unfixed and fixed HEK293 cells. In addition, a very-weak to weak interaction was detected with off-target C in fixed and unfixed cells for both the TA and the single target B/isotype control, localizing the off-target interaction to the target B arm of the bispecific antibody.
The interaction with off-target C was further confirmed using flow cytometry, which showed strong intensity interactions between the test article and both primary targets and a medium intensity interaction to HEK293 cells transfected to overexpress off-target C. Quantitative dose-response flow cytometry provided an EC50 value that was ~77x higher for off-target C versus target B. Binding of the test article to transfected HEK293 cells expressing the cynomolgus monkey homolog of off-target C was also assessed by flow cytometry; a weak interaction was observed at higher concentrations of the test article, but no EC50 could be established.
Following these results, the sponsor developed a (off) target safety assessment and plan to better understand potential consequences resulting from an interaction with off-target C. Methods included a literature/in silico review, additional binding studies, and functional assessment. Interestingly, there was limited sequence homology between off-target C and target B (<50% between respective extracellular domains). Protein and/or RNA expression in human and cynomolgus monkey was reported in multiple sites and cell types, including immune cells. Flow cytometry on human whole blood using a commercial antibody to off-target C confirmed membrane expression on B cells, T cells, dendritic cells, and platelets, with the highest expression on B cells; however, no binding of the test article to primary B cells or other cell types was observed up to highest concentration tested. Similarly, no interaction with platelets was detected when using human-derived platelet-rich plasma. Surface plasmon resonance (SPR) also showed no binding to either human or cynomolgus monkey off-target C, but this result may have been due to potential differences due to avidity/affinity, or unsuitability of the test article for use in SPR. Finally, in vitro functional assessment was conducted using the PathHunter Checkpoint Receptor Signaling Assay, in which cells overexpressing off-target C were co-cultured with a reporter cell line expressing its receptor. The test article did not show any inhibitory effect on the off-target C pathway.
In summary, an off-target interaction was identified using CBPA and confirmed by flow cytometry on transfected HEK293 cells. Lack of in vivo relevance for this interaction was supported by lack of TA binding in cells with endogenous off-target expression and lack of functional impairment of the off-target pathway by the TA. This discrepancy with CBPA results is likely due to higher expression levels in the CBPA system compared with endogenous levels of the off-target protein. Ultimately, this program moved forward into Phase 1 trials without a TCR study, despite the non-oncology indication.
Commentary: This case provides an example of early regulatory input on the alternative use of CBPAs in lieu of a TCR to support FIH studies for a nononcology indication. It also illustrates the importance of orthogonal validation to assess the biological relevance of hits.
Case 5: Investigating the Source of Unanticipated Clinical Adverse Events Using a CBPA
Following the observation of delayed hypersensitivity of unknown origin in patients treated with an ADC, a Retrogenix Human Cell Microarray screen was conducted. The primary target A of the ADC was identified, along with two additional potential off-targets (B and C). Those two interactions were also observed with an isotype ADC (targeting chicken lysozyme), suggesting they may be related to the drug payload and/or linker. Further validation was performed for off-target B. Notably, co-immunoprecipitation experiments showed that both ADCs (targeting either target A or chicken lysozyme) could trap a recombinant purified version of off-target B, unlike the corresponding naked antibodies (which do not carry the payload and linker). These results were further supported by a flow cytometry experiment, which showed that ADCs bind to monocytes, unlike naked antibodies (monocytes do not express the primary target but secrete off-target B, which can be recaptured at the monocyte cell surface by specific receptors). Although it has not been formally demonstrated, the contribution of this interaction to the clinical adverse events is plausible based on the known biology of off-target B and overlapping clinical findings with another ADC sharing the same payload.
In the GLP TCR study with the ADC, staining was observed in tissues where the primary target A of the ADC is expressed, but additional staining was detected in endothelium and pancreatic exocrine ducts. This staining pattern was consistent across cynomolgus monkey and human tissues. However, no ADC-related changes were noted in these tissues in the repeat doses toxicity studies. Contribution of those off-target tissue binding to clinical observations is unknown.
Commentary: This example highlights that TCR and CBPAs do not always correlate (here, TCR tissue staining does not match the known off-target B distribution). Target B being a secreted protein, it is possible that it was lost during TCR processing. Finally, it highlights the capability of CBPAs to identify some off-targets even when mediated through the chemical part of ADCs (linker or payload).
Case 6: Retrospective Evaluation by CBPA After TCR
This case involved a CBPA study that was conducted while the therapeutic candidate, a CD3 T cell engager against a myeloid target M was in Phase I trials for AML; the trial was eventually discontinued.
Both the CD3 bispecific antibody and the parental antibody against target M were evaluated in a GLP TCR study. Staining, both membranous and cytoplasmic, was observed with the parental antibody in the mononuclear leukocytes in blood smears, liver (Kupffer cells), lymph node, spleen, thymus as well as hematopoietic precursor cells in the bone marrow, consistent with the expression of target M on macrophages, monocytes and some lymphoid cells. Additional staining in mononuclear leukocytes in human lymphoid and selected nonlymphoid tissues was consistent with expression of CD3 for the bispecific antibody.
In the Retrogenix Human Cell Microarray, the parental antibody against target M was screened for binding against fixed human HEK293 cells, individually expressing 5861 full-length human plasma membrane proteins and cell surface-tethered human secreted proteins plus a further 371 human heterodimers and in a confirmation screen on both live and fixed cells. Binding to target M was observed as well as off-target binding to several related family members (F, and G) and another off-target (T) that is expressed on T cells. Binding was also confirmed by flow cytometry on live, unfixed, transfected HEK293 cells.
Following these findings, orthogonal methods were used to understand if the clinical safety findings were potentially related to the target or off-target. These included a literature/in silico review, additional binding studies (SPR, cell binding) and functional assessment. The interpretation was further complicated as target M and off-target F and G are expressed on similar cell types, including basophils, mast cells, and Kupffer cells that have contributed to the infusion-related reactions and liver findings, respectively.
Commentary: This example highlights a potential gap that may be missed in the TCR if the on-target and off-target distribution and expression profiles overlap on similar cell types.
Cases 7 and 8: Interpretation of Cell-based Protein Array Hits
Five antibodies were screened on the Retrogenix Human Cell Microarray and found to interact with CD209 and CLEC4M, raising concerns about their selectivity. However, those antibodies were not all targeting the same epitope (two targeted one epitope, while the other three targeted a nonoverlapping epitope), and contained completely distinct CDRs. This indicates that the interactions with CD209 and CLEC4M are not mediated by the antibody CDRs. This screen was part of the early selection process for candidates, and the antibodies were produced in yeast, not in their final mammalian cell line. It is known that antibodies produced in yeast have different glycosylation patterns, particularly containing mannose-rich glycans, 14 which both CD209 and CLEC4M can recognize.11,37,42 Therefore, it is possible that the observed interaction was due to yeast-specific glycosylation, which may not be relevant for the final product made in mammalian cells. However, the selectivity of the final product has not been assessed to confirm this. This case underscores the importance of considering post-translational modifications in addition to the candidate protein sequence when interpreting results of array screens.
Similarly, a dual Fab construct with one arm targeting hyaluronic acid was profiled on the Retrogenix Human Cell Microarray platform. Binding was observed in cells overexpressing hyaluronan synthases (HAS2/HAS3) and to chondroitin sulfate proteoglycans (BCAN, NCAN and CD44). Hyaluronan synthases are essential to hyaluronic acid synthesis, while the chondroitin sulfate proteoglycans are known to interact with hyaluronic acid and in the case of BCAN to stabilize it. This suggests that those hits are unlikely unintended targets of the candidate but are exacerbating the binding to the Fab primary target through increased synthesis or stabilization of hyaluronic acid in over-expressing cells.
Commentary: Both examples highlight the need for careful interpretation of hits, since some detected interactions, while not artifactual, do not necessarily indicate a lack of selectivity of the test candidates. Case 7 also underscores the need for consideration between the test articles used in the screen and the final clinical drug product.
Discussion
Biologics have become mainstay pharmaceutical modalities and the perception of their high selectivity has contributed to their widespread use. However, increasing evidence suggests that their selectivity for the intended target is not absolute, with issues such as polyreactivity, polyspecificity, and on-target binding in unintended tissues, potentially leading to toxicity or suboptimal pharmacokinetic profiles. Literature-based examples below highlight some preclinical and clinical consequences of off-target interactions and demonstrate biologics selectivity is critical for patient safety, relevant translation of preclinical animal safety data, and reduced animal use.
An anti-FGFR4 antibody demonstrated rapid clearance, poor target tissue distribution and limited efficacy in a human xenograft mouse model. The antibody was shown to bind to complement C3 protein, a highly abundant serum protein. 4
The Biosafe Industry Survey of mAb and mAb-like molecules provides some examples where IHC staining correlates with toxicity. 31
Camrelizumab caused capillary hemangioma in the majority of treated patients, in contrast to patients treated with other anti-PD1 antibodies. Through the use of a CBPA, VEGFR2 was identified as a camrelizumab off-target. Follow-up work demonstrated that the antibody was a potent agonist of VEGFR2 that could drive vascular neogenesis potentially explaining the capillary hemangioma. 12
Administration of an antibody targeting amyloid beta protein oligomer resulted in severe toxicity in cynomolgus monkeys, including thrombocytopenia. 29 Monkey and human platelet factor 4 (PF-4), a protein known to be involved in heparin-induced thrombocytopenia, were identified as an off-target of this antibody.
An antibody targeting EGFRvIII, (an antigen not found in normal tissues) induced thrombocytopenia and platelet activation in cynomolgus monkeys. 48 It was demonstrated that binding to integrin αIIbβ3 contributed to this off-target toxicity. Interestingly, platelet activation could be recapitulated in vitro using macaque platelets but not with human platelets. This demonstrates that off-target might be species-specific. Consequently, it is crucial to recognize that some human-specific toxicities might not be observed in nonhuman primates.
Off-target mediated toxicity is not specific to antibodies. For example, engineered T cells expressing an affinity-enhanced TCR against HLA-A*01–restricted MAGE-A3 induced fatal cardiogenic shock in patients that was not predicted from preclinical studies. 27 Toxicity has been associated with the recognition of an unrelated peptide derived from the striated muscle-specific protein titin.
It is, however, important to note that a lack of selectivity in either TCR studies or CBPAs is not necessarily translating to notable toxicity:
Over 80% of Biosafe Industry members (21 out of 26 respondents) indicated that binding observed in the TCR did not translate to notable toxicity in nonclinical or clinical studies. 31 Similarly, while a retrospective analysis of data provided by Charles River Labs showed that >50% of tested molecules (modality agnostic) on the Retrogenix platform have at least one off-target identified (Figure 2) while one third of tested molecules exhibit polyspecificity on the Integral Molecular platform, 33 some CBPA case examples presented here indicate that they may not necessarily cause toxicity in vivo.
This lack of translation to toxicity could be attributed to assay artifacts, to true interactions that are not off-target driven (case 6), or because the actual interaction may not hold biological significance (case 4), which could be due to factors like low affinity, minimal native expression levels, or binding that does not result in functional consequences. Therefore, it is critical to perform follow-up studies to investigate the relevance of those unexpected binding. This includes alternative binding assays as well as functional assays. In our experience, stronger hits were more frequently validated than weaker hits by orthogonal methods.
It is important to note that biologics carrying highly potent cell-killing mechanisms such as ADCs, CD3-targeting bispecific antibodies or CAR-T are theoretically at a higher risk for safety concerns compared with effector null biologics if they are not selective.
In addition, it is possible that these methods might not detect actual off-targets that are concerning. Limited data are available to illustrate this, as molecules are often discontinued without further investigation upon identifying severe findings. A simplified, yet biased, method to estimate the false negative rate of the cell-based protein arrays is to evaluate their ability to detect the known primary targets of the tested biological products. Data from one sponsor indicated that out of the 59 primary targets present on the Retrogenix array against which molecules were tested, 57 were correctly identified.
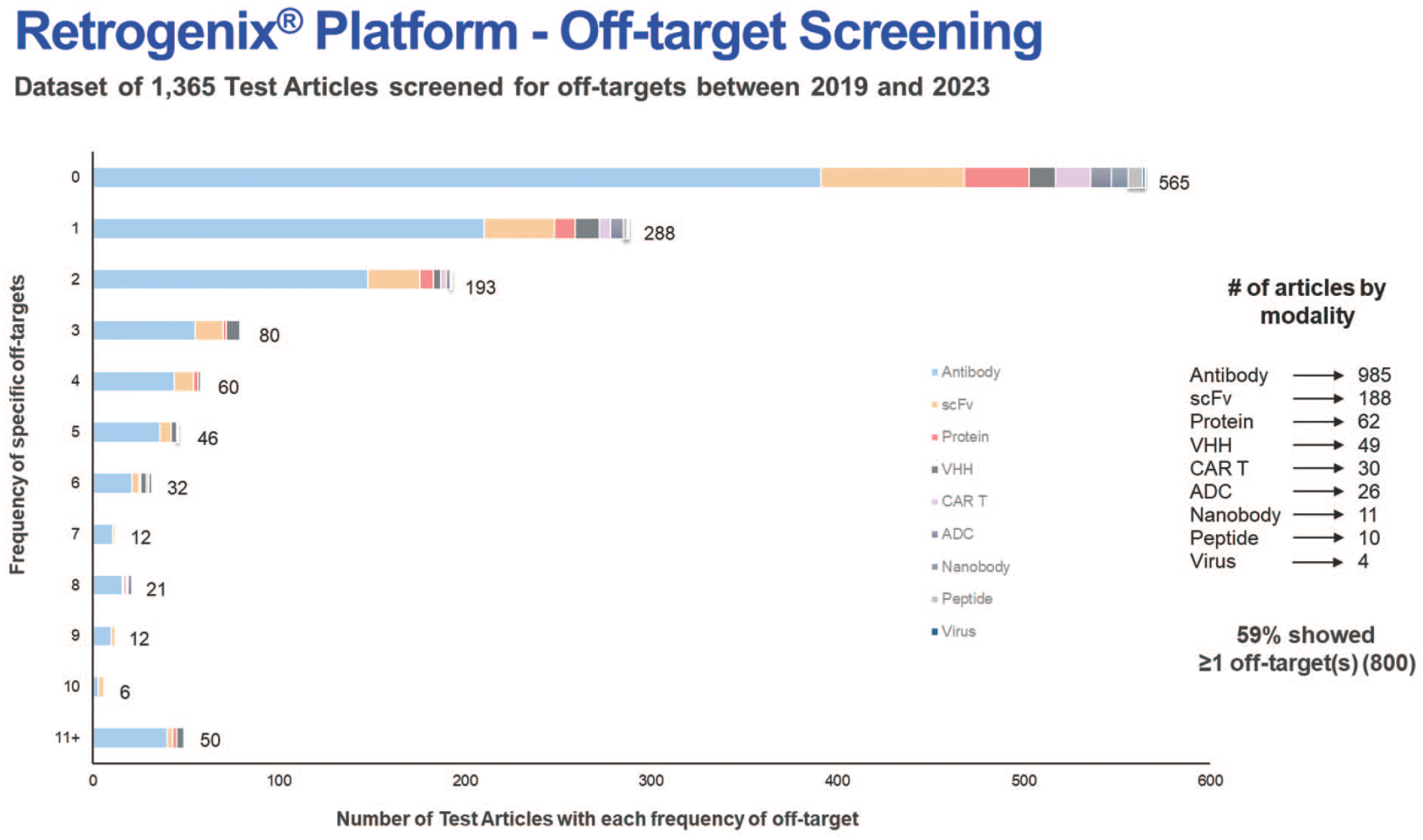
Summary of off-target interaction frequency for a variety of new molecular modalities. Retrogenix Cell Microarray specificity assay evaluated 1365 test articles represented from 2019 to 2023; 59% of all test articles included demonstrated >1 specific off-target interaction.
TCRs have been employed for over 3 decades to evaluate the on- and off-target binding of biologics, and the experience with CBPAs have extensively grown in the last 15+ years, with many screens being submitted to health authorities. Microarray data has been successfully used for IND and CTA submissions without TCR data for regulatory submissions.15,26,34 With regard to time and cost, it can be expected that a TCR may cost 2×-4× more and take several months longer to complete the characterization for one molecule compared with a microarray study. It is the author’s experience that this strategy is more commonly employed for oncology indications or when a suitable TCR method could not be developed. In addition, it is important to note that some companies have successfully moved their products to the clinic without either the TCR or CBPA data.
FDA and ICH guidelines indicate that best methodology(ies) should be used and justified in regulatory filings and with growing experience with CBPAs, a logical next step would be that health authorities, sponsors and assay providers discuss and align on how to improve biologics selectivity assessment. Various strategies have been proposed 31 and are recapped here with slight updates, including their (potential) context of use:
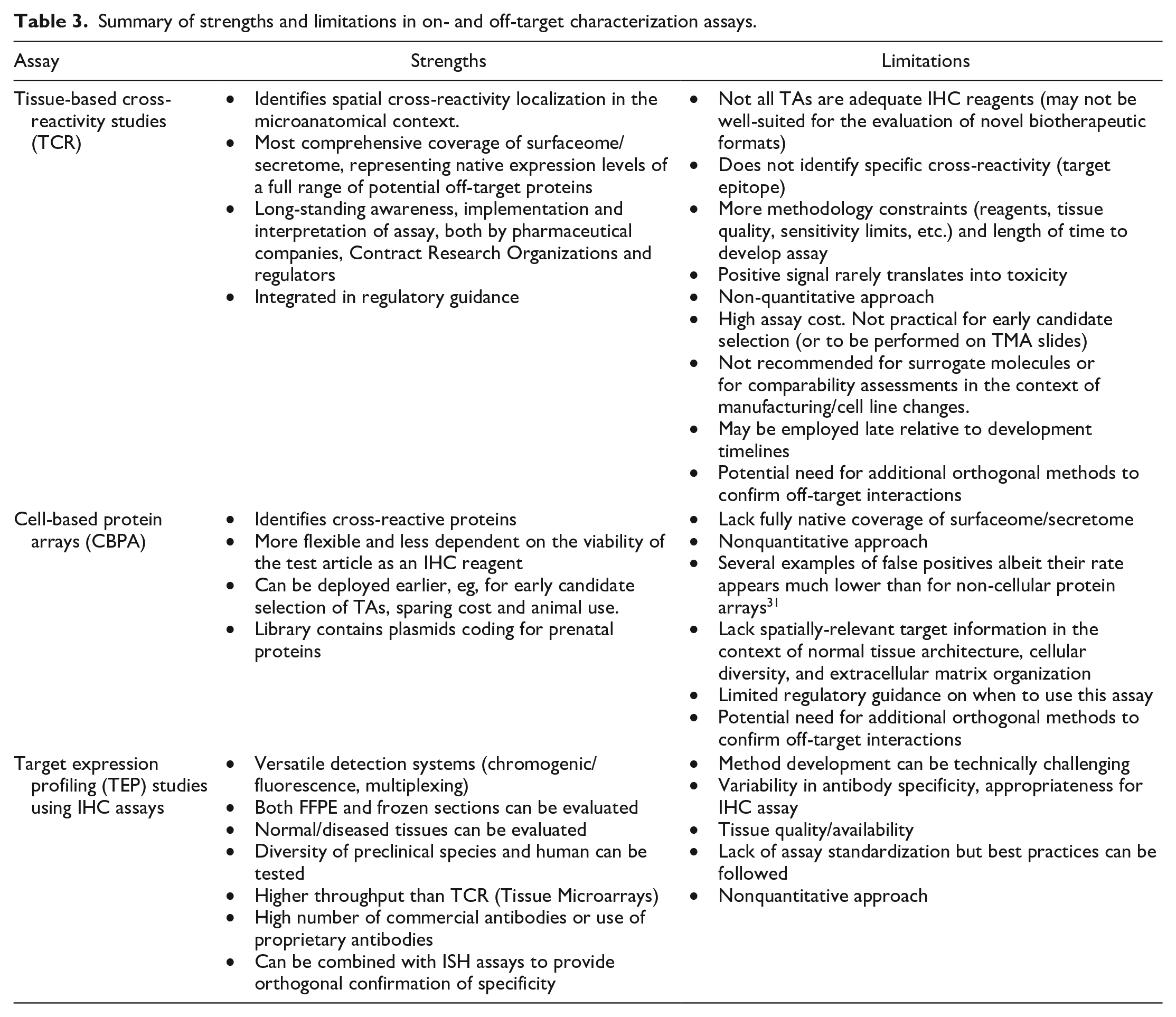
Conduct a standard TCR and employ a secondary array screen only in the instance of clear or possible cross-reactivity upon evaluation of the TCR data. This is the current standard practice for nononcology indications. Strengths and limitations of this approach are listed in Table 3.
Combine a standard TCR assay with a CBPA screen. This comprehensive approach is likely to increase the likelihood of identifying concerning off-targets, mitigating respective limitations of both approaches (Table 3). However, this could also increase the identification of spurious findings, potentially slowing down the development of new biologics and escalating their development costs. This approach is accepted for both oncology and nononcology indications but is not a standard practice.
Combine on-target localization data using an IHC-optimized surrogate antibody against the primary target with a CBPA screen. This would provide proper tissue expression data for the intended target and identity of potential protein off-target(s). Should such off-targets be confirmed, a subsequent off-target TEP assessment could be conducted. Additional considerations on strengths and limitations of assays used in this strategy are detailed in Table 3. This approach is appropriate for oncology indications and offer a solution for non-oncology biologics where TCR is not feasible. An on-target IHC approach may serve as a potential bridging study for biologics with array-only data when moving to a nononcology space, although to our knowledge, this method has never been endorsed. In the long term, subject to further qualification and consensus among assay providers, health authorities, and sponsors, this approach may be a suitable substitute for TCR.
Conduct an CBPA screen alone. This approach could be adequate for biologics targeting nonhuman targets or those not expressed in healthy tissue (such as tumor-specific targets). In addition, this strategy might be suitable for molecules aimed at a well-characterized target, provided there are no additional risks (such as increased affinity for the primary target or a more potent modality).
Summary of strengths and limitations in on- and off-target characterization assays.
Conclusions
Off-target binding assessment should be employed as part of a comprehensive approach in the development and safety assessment of biologic molecules. TCR studies have been the standard for numerous years, but cell-based protein array technology provides a complementary, and potentially alternative technology for binding assessments. The use of these assays will improve off-target assessment through the identification of specific cross-reactive proteins and support the assessment of diverse test article formats that are not amenable to TCR. Increasing regulatory experience with the inclusion of cell-based protein array data in submissions will enhance familiarity with this assay and potentially lead to expanded regulatory acceptance ultimately facilitating the development of novel therapeutic products for which animal models may not be relevant, and support 3R principles of reduced animal use.
Supplemental Material
sj-docx-1-tpx-10.1177_01926233241311259 – Supplemental material for Applications of Cell-Based Protein Array Technology to Preclinical Safety Assessment of Biological Products
Supplemental material, sj-docx-1-tpx-10.1177_01926233241311259 for Applications of Cell-Based Protein Array Technology to Preclinical Safety Assessment of Biological Products by Axel Vicart, Cam Holland, Kathryn Fraser, Frederic Gervais, Mark Aspinall-O’Dea, Nick Brown, Kirk Siddals, Géraldine Greiner, Vinicius Carreira, Elizabeth Galbreath, Maggie Willer, Saravanan Kaliyaperumal, Charles Wood, Tim MacLachlan and Elizabeth Clark in Toxicologic Pathology
Footnotes
Acknowledgements
The authors thank Ben Doranz and Rachel Fong from Integral Molecular and Diann Blanset for their critical review of this paper as well as Fionnuala McAleese Eser and Mathias Gehrmann from Bayer for their case report contribution.
Author Contributions
The authors are responsible for the contents and crafting of this paper with additional contributions noted in the Acknowledgements section.
Declaration of Conflicting Interests
The author(s) declared the following potential conflicts of interest with respect to the research, authorship, and/or publication of this article: At the time of manuscript drafting, MA-O’D, NB, FG, and KS are all employees of Charles River Laboratories.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
