Abstract
The nnU-Net framework effectively automates hyperparameter selection; however, its fixed internal configurations—notably convolution kernel sizes—restrict its flexibility. This limitation is pronounced in 3D medical imaging, where anatomical structures undergo continuous spatial evolution along the Z-axis. In this study, we introduce a self-adaptive convolution module designed to dynamically tune the effective receptive field, matching the dynamic structural transformations of organs. By employing a differentiable soft-attention mechanism to aggregate candidate kernels, the network adaptively optimizes its scale sensitivity. This integration allows MSA2-Net to capture both global context and local nuances within feature maps. The module is strategically embedded into two core components: the multi-scale convolution bridge and the multi-scale amalgamation decoder. In the Bridge, it refines CSWin Transformer outputs by aligning features with the inherent spatial continuity of volumetric data, thereby mitigating redundancies that might otherwise hinder decoding. Simultaneously, the multi-scale amalgamation decoder leverages this module to precisely reconstruct organ details as their size and shape fluctuate across slices. This mechanism ensures the decoder preserves seamless topological intricacies within the feature maps, yielding superior segmentation accuracy. Leveraging this architecture, MSA2-Net achieves competitive Dice scores of 86.49%, 92.56%, 93.37%, and 92.98% on the Synapse, ACDC, Kvasir, and ISIC2017 datasets, respectively. Extensive experiments validate the model's robustness in handling complex spatial variations across diverse medical modalities.
Keywords
Introduction
Medical image segmentation plays a pivotal role in clinical diagnostics, empowering clinicians to rapidly identify and localize pathologies.1–3 This task entails the precise extraction and delineation of anatomical structures or lesions from diverse imaging modalities, such as computed tomography (CT), magnetic resonance imaging (MRI), and X-rays. At its core, segmentation aims to assign semantic labels to each pixel or voxel, effectively categorizing distinct organs, tissues, and pathological features to provide a granular understanding of the underlying anatomy.
Since their emergence, 4 convolutional neural networks (CNN) have become the dominant framework for medical image segmentation. By leveraging local receptive fields and parameter sharing, CNN efficiently extract essential features such as edges, textures, and organ structures, making them ideally suited for medical imaging scenarios characterized by strong spatial correlations. Among these architectures, U-Net stands as a quintessential model and remains the most widely adopted solution in the field. Its design comprises an encoder for hierarchical feature extraction and downsampling, a decoder for restoring spatial resolution via upsampling, and skip connections that bridge the two. These connections are critical for recovering the fine-grained spatial details often lost during the encoding process. Through this elegant symmetry, U-Net has significantly enhanced segmentation precision and generalization across various clinical applications.
Despite their success, CNN are inherently restricted by their focus on local modeling.5–7 The fixed dimensions of convolutional kernels often impede the effective capture of multiscale features, a challenge that is especially pronounced in complex multi-organ segmentation. This localized perspective hampers the integration of global context, ultimately compromising segmentation precision. To address this, various strategies have been advanced: MISSFormer 8 introduces an enhanced transformer context bridge to capture hierarchical representations; dilated convolution 9 expands the spatial coverage by increasing dilation rates without escalating computational costs; spatial transformer networks (STN) 10 employ learnable geometric modules for adaptive spatial alignment; and adaptive spatial feature fusion (ASFF) 11 utilizes spatial filtering to harmonize features across different resolutions. While these methods bolster performance to a degree, a fundamental bottleneck remains: kernel size is typically a predetermined hyperparameter. The inability to dynamically recalibrate the receptive field according to the specific characteristics of the training data significantly constrains the model's broader generalization.
nnU-Net
12
utilizes a heuristic approach based on “data fingerprints” and “pipeline fingerprints” to encapsulate essential dataset and network attributes, enabling autonomous hyperparameter tuning. However, these adjustments primarily target preprocessing configurations—such as voxel size, spacing, and batch size—while largely overlooking architectural hyperparameters, specifically convolution kernel sizes and padding. To bridge this gap, we present the multi-scale adaptive attention network (MSA2-Net). This architecture employs a novel differentiable dynamic kernel aggregation mechanism, allowing the network to adaptively synthesize kernels by analyzing the statistical characteristics of input features. The primary contributions of this work are summarized as follows.
Propose the self-adaptive convolution module (SACM), which dynamically senses the spatial scale of organs in each slice. By generating input-specific statistical fingerprints, the module automatically adjusts its receptive field. This mechanism allows the model to synchronize with the continuous spatial evolution of organs along the z-axis, ensuring optimal feature extraction whether the organ appears as a small tip or a full cross-section. An innovative multi-scale convolution bridge (MSConvBridge) is proposed based on self-adaptive convolution modules. This architecture utilizes a combination of multi-scale adaptive convolutions to delicately process the feature maps output by CSWin. Through this multi-layer approach, the MSConvBridge efficiently eliminates redundant striated textures between image features while maximally preserving the information within the feature maps, thereby significantly enhancing overall semantic consistency. This method not only optimizes feature representation but also strengthens the model's capability to handle complex scenes. Multi-scale amalgamation decoder (MSADecoder) employs recursive nested adaptive convolution groups, enabling the decoder to replenish information about small organs in the image while upscaling, and simultaneously preserving information about large organs in the image.
Related work
nnU-Net
In medical image segmentation networks, there are a number of hyperparameters. Researchers often have to adjust these hyperparameters through repeated experiments during the network design process. This adjustment is typically reliant on the individual researcher's experience, making the process highly inefficient. Excessive manual adjustments to the network structure can lead to overfitting for specific datasets. The impact of non-structural aspects of the network might have a greater influence on the segmentation task. Therefore, nnU-Net was developed, focusing not on modifying the specific network architecture but on adjusting the hyperparameters in dataset preprocessing, training scripts, and post-processing. Although nnU-Net's approach has improved training efficiency, it overlooks adjustments of the hyperparameters within the network structure itself. The adaptive convolution module proposed in this article builds on the nnU-Net framework, capable of adapting the size of the convolutional receptive field according to the characteristics of the dataset.
U-shaped architectures for biomedical segmentation
In the realm of biomedical image analysis, the U-Net architecture has established itself as a fundamental benchmark due to its ability to function effectively with limited annotated samples—a common constraint in clinical datasets. Its symmetric topology, featuring a contracting path for context capture and an expanding path for precise localization, perfectly aligns with the requirement to delineate anatomical boundaries. While the encoder hierarchy captures deep semantic representations of tissues and organs, the skip connections are pivotal in recovering the high-frequency spatial information lost during downsampling, ensuring that fine-grained structures like tumor margins are preserved in the final segmentation map.
Building upon this foundation, recent variants have sought to address U-Net's limitations in global context modeling. For instance, UNet++ introduced nested and dense skip connections to bridge the semantic gap between encoder and decoder sub-networks. Attention U-Net integrated gating mechanisms to suppress irrelevant regions in input images while highlighting salient features useful for specific tasks. More recently, Transformer-based U-Net variants like Swin-UNet have attempted to leverage self-attention for long-range dependency modeling. However, these methods often replace the inductive bias of convolutions entirely or rely on static convolutional patches, missing the opportunity to dynamically adjust the feature extraction scale based on the specific anatomical characteristics of the input scan.
Method
Self-adaptive convolution module
In volumetric medical datasets like Synapse, anatomical structures exhibit continuous spatial evolution along the z-axis. Consequently, the cross-sectional size of a single organ fluctuates significantly across different slices—appearing as a small, localized region in some frames while occupying the entire field of view in others. This variability exposes the fundamental limitation of standard convolutional operations: their fixed receptive fields.
As illustrated in Figure 1(b), a fixed kernel size faces a dilemma. Oversized kernels are prone to incorporating irrelevant background noise when processing slices where organ cross-sections are small or sparse, leading to the ‘background contamination’ shown in Figure 1(c). Conversely, undersized kernels fail to capture global contextual information when organs expand to larger scales. Given that multiple visceral organs often interweave with complex spatial dependencies, static kernels cannot optimally adapt to these dynamic changes.
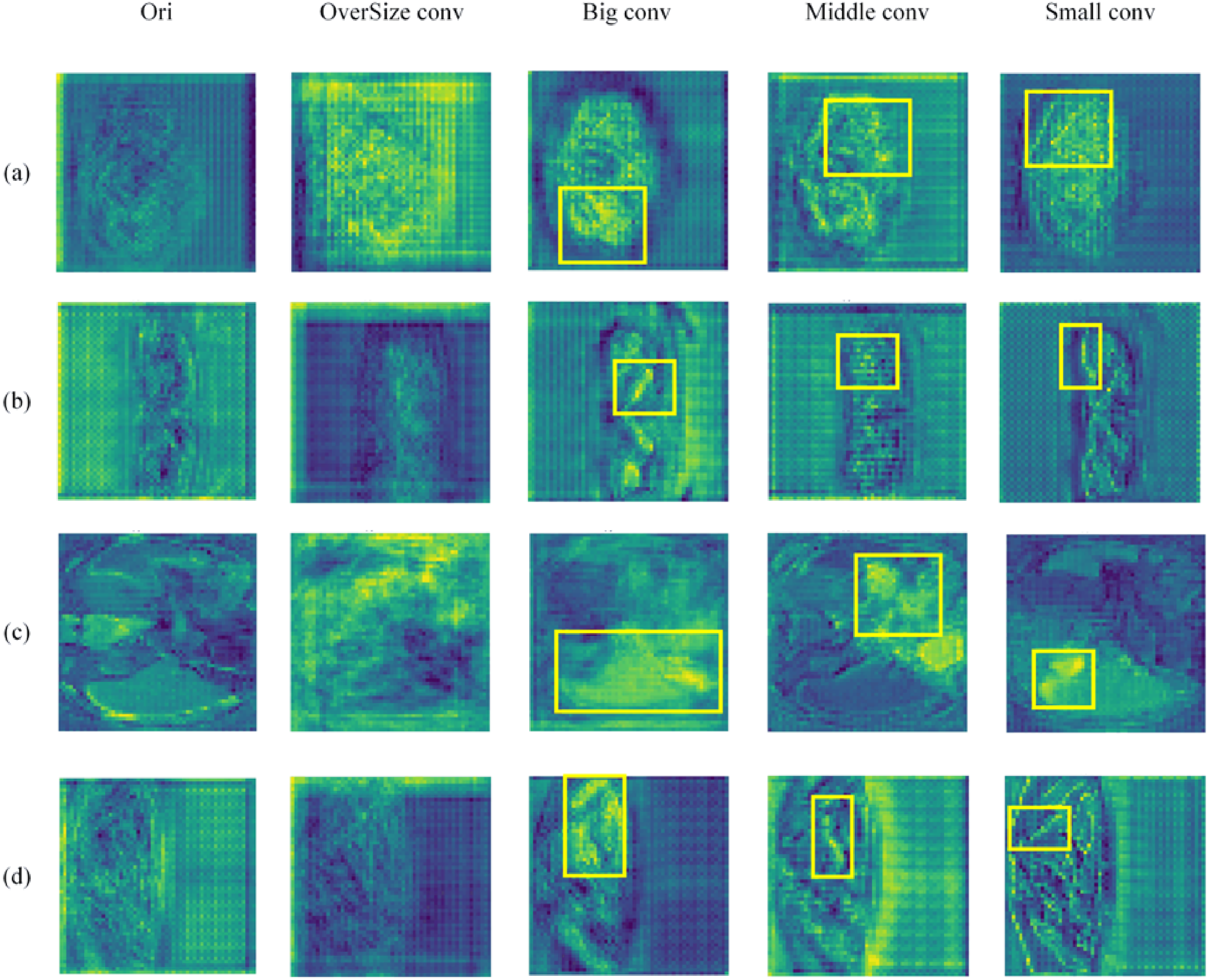

Impact of different kernel sizes on feature maps.
To resolve this, we propose the self-adaptive convolution module, which dynamically synthesizes kernels conditioned on the input feature statistics. Instead of a hard selection strategy, we formulate this as a soft attention mechanism, allowing the network to continuously adapt its receptive field to match the evolving organ dimensions slice-by-slice in an end-to-end manner.
Let 
Then, we employ a lightweight gating network (a two-layer MLP) to map the fingerprint 
Where 
Finally, the input 
Self-adaptive convolution module dynamically adjusts the convolutional receptive field according to dataset characteristics, ensuring the kernel size is neither too large nor too small—precisely covering most sub-features in the feature maps to achieve efficient multi-scale information extraction (Figure 2).
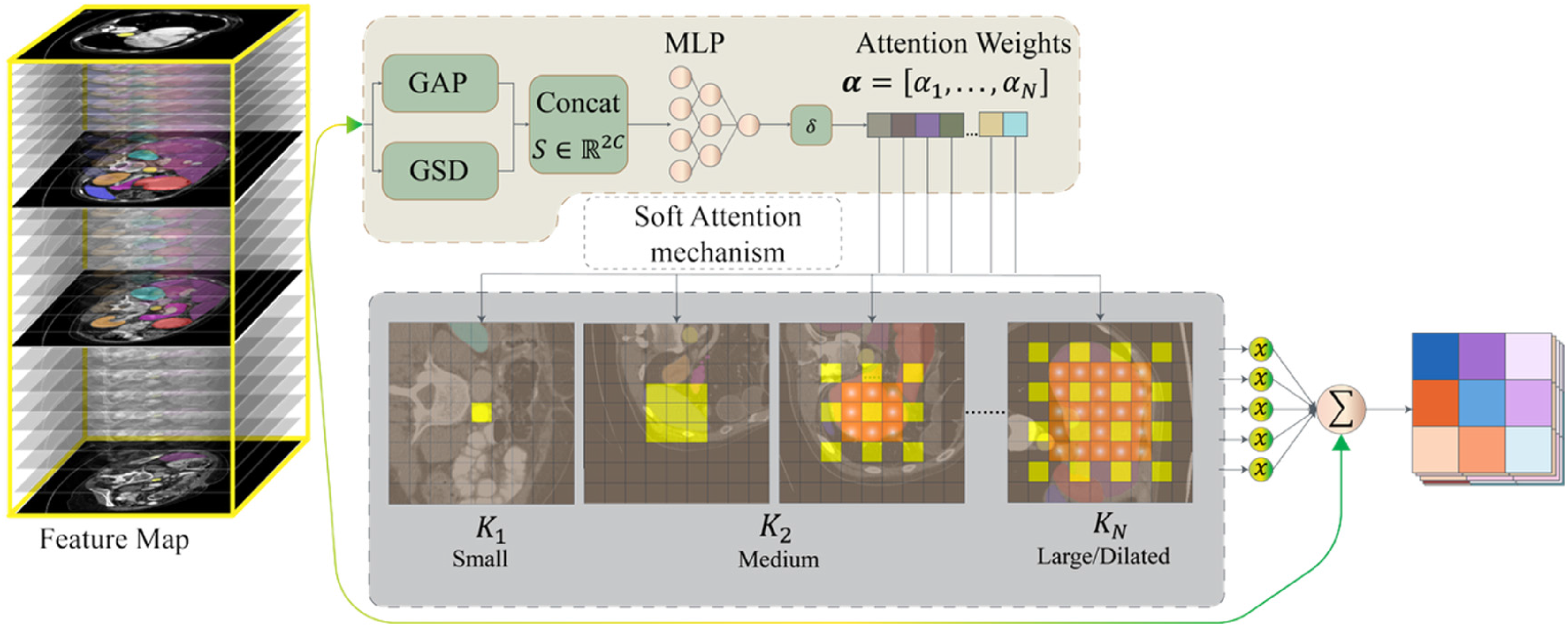
Architecture of self-adaptive convolution module.
Overall structure
The specific architecture of MSA2-Net is illustrated in Figure 3. The left portion of Figure 3 shows the encoder component of MSA2-Net, while the right portion displays the multi-scale adaptive decoder (MSADecoder). The middle section represents the multi-scale convolution bridge (MSConvBridge).

Architectural overview of MSA2-Net.
For a given image 
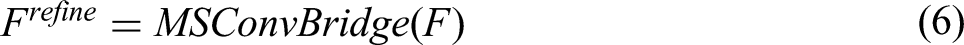
Finally, MSADecoder generates the segmentation map output by aggregating and upsampling these refined features: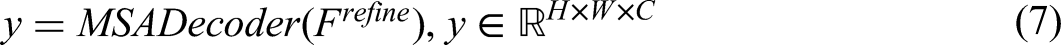
Encoder
To better adapt CSWin for medical image segmentation tasks, we modified the original encoder design of CSWin to preserve detailed information in feature maps while performing downsampling for feature extraction. MSA2-Net integrates the CSWin encoder with ResNet, where the CSWin encoder serves as the primary encoder and the ResNet equipped with adaptive convolution modules acts as the auxiliary encoder. The internal workflow at each encoding stage can be expressed as follows:

Here,
MSConvBridge
Although the encoder can effectively capture multi-scale information, directly feeding the original feature maps to the decoder may introduce redundant noise, thereby compromising segmentation accuracy. Convolutional operations function analogously to filters, capable of suppressing redundant information in feature maps while preserving critical features when detecting sub-structures. To enhance semantic consistency between the encoder and decoder, MSA2-Net incorporates a multi-scale convolutional bridge (MSConvBridge), which selectively refines and optimizes features within the skip connections. The operational mechanism of the MSConvBridge is detailed below.
For a given feature map 
Here,
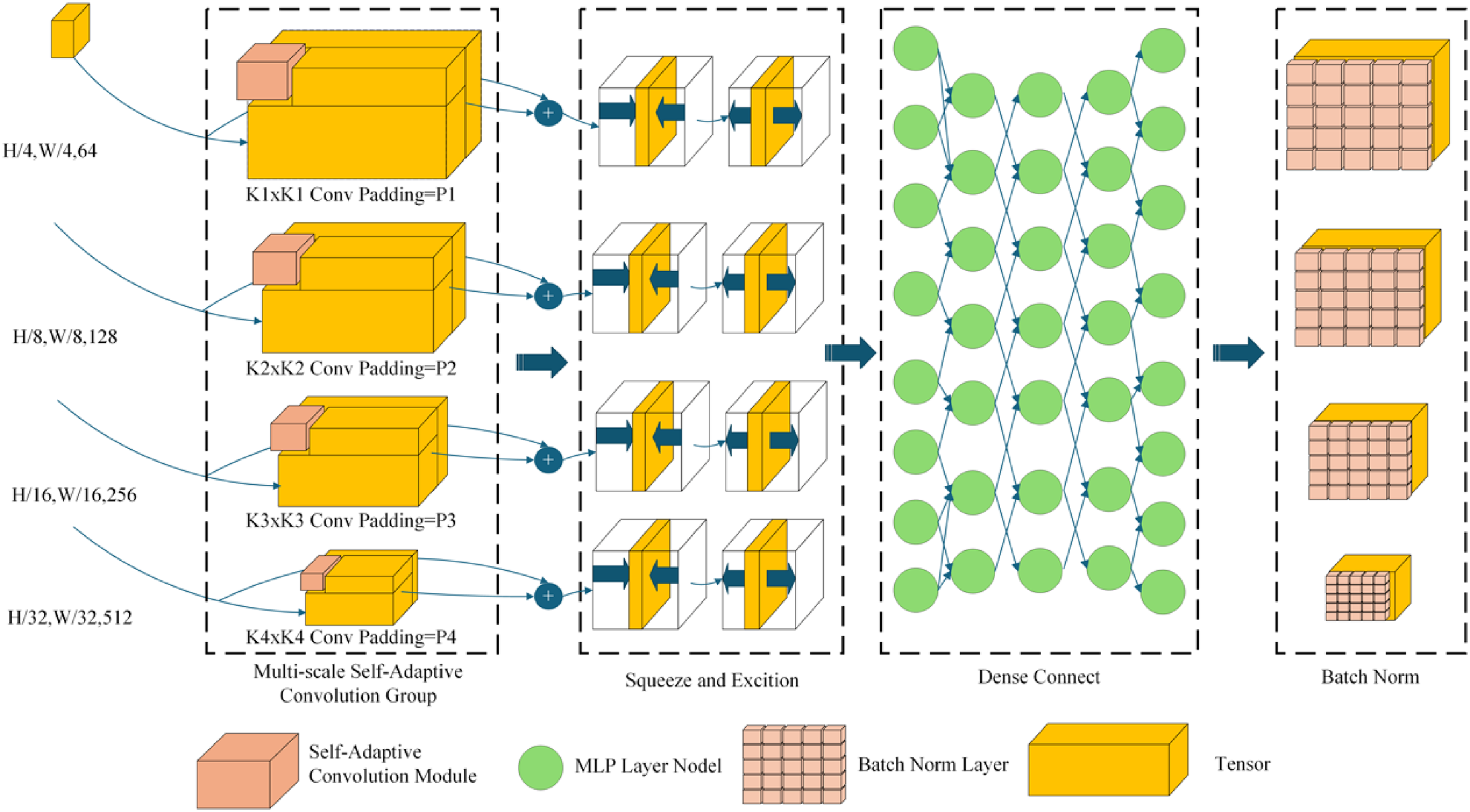
MSConvBridge architecture diagram.
MSADecoder
To restore spatial resolution and accurately reconstruct fine structures such as organ boundaries and small organs, MSA2-Net employs a lightweight yet highly expressive decoding module—MSADecoder. As shown in Figure 5, the feature map 

MSADecoder architecture diagram.

Here,
Experiments
Environmental
MSA2-Net is implemented using PyTorch and trained on an NVIDIA A100 GPU platform. The CSWin backbone network incorporates pre-trained weights. Input images for the CSWin encoder are resized to
Dataset
In this section, we benchmark MSA2-Net against current state-of-the-art (SOTA) networks to demonstrate its superiority. Experiments are conducted on three datasets: the Synapse multi-organ segmentation dataset (Synapse), the automated cardiac diagnosis challenge dataset (ACDC), and Kvasir. Network performance is evaluated using the Dice coefficient and average Hausdorff distance (HD95). Results for comparative networks are obtained from previously published studies. The detailed data partitions for these three datasets are presented in Table 1.
Detail dataset partitions.
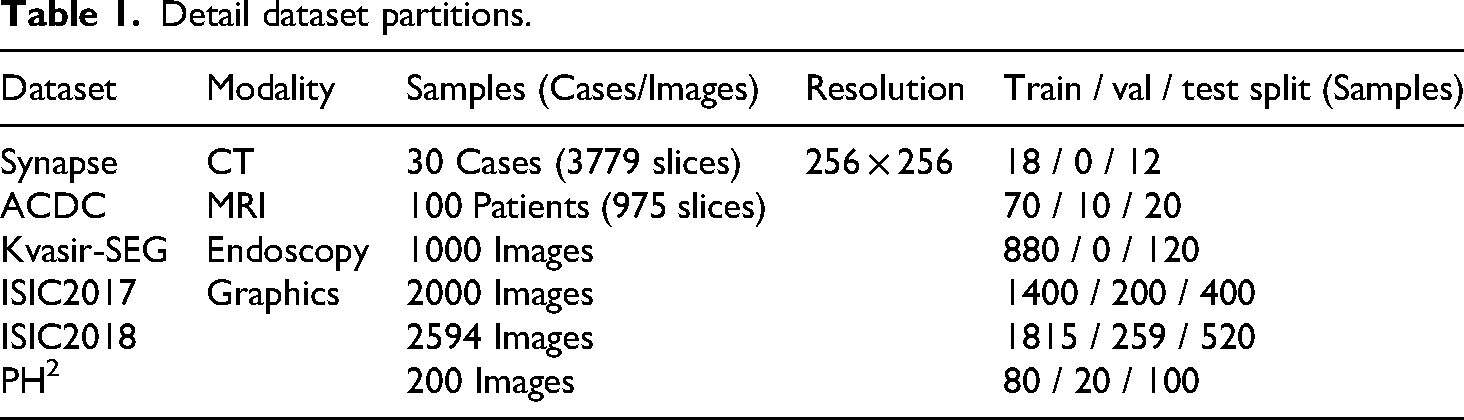

Experimental results and analysis
In this section, we will compare the performance of MSA2-Net with other state-of-the-art (SOTA) methods on the Synapse, ACDC, Skin Lesion Segmentation Datasets, and Kvasir-SEG datasets.
Performance of different networks on synapse
Table 2 presents the performance of MSA2-Net on the Synapse dataset, highlighting its superior capabilities. MSA2-Net achieved the highest average Dice coefficient of 86.49%, showing improvements of 5.52%, 9.30%, and 0.65% over MISSFormer, Swin-UNet, and AgileFormer, respectively. In terms of the HD95 metric, MSA2-Net recorded the second lowest distance (14.15), significantly outperforming MISSFormer (18.20), Swin-UNet (21.55), and TransCASCADE (17.34).
Performance of different networks on Synapse.

By examining Table 2, it can be observed that MSA2-Net achieved the best Dice scores for 5 out of 8 organs. These organs are all small in size, demonstrating that the MSA2-Net efficiently preserves information for small organs during upsampling. Due to space constraints, and considering that MISSFormer is the primary inspiration for MSA2-Net and SwinUNet is a classic algorithm in medical image segmentation, Figure 6 displays the segmentation results of MSA2-Net, MISSFormer, and SwinUNet on the Synapse dataset.
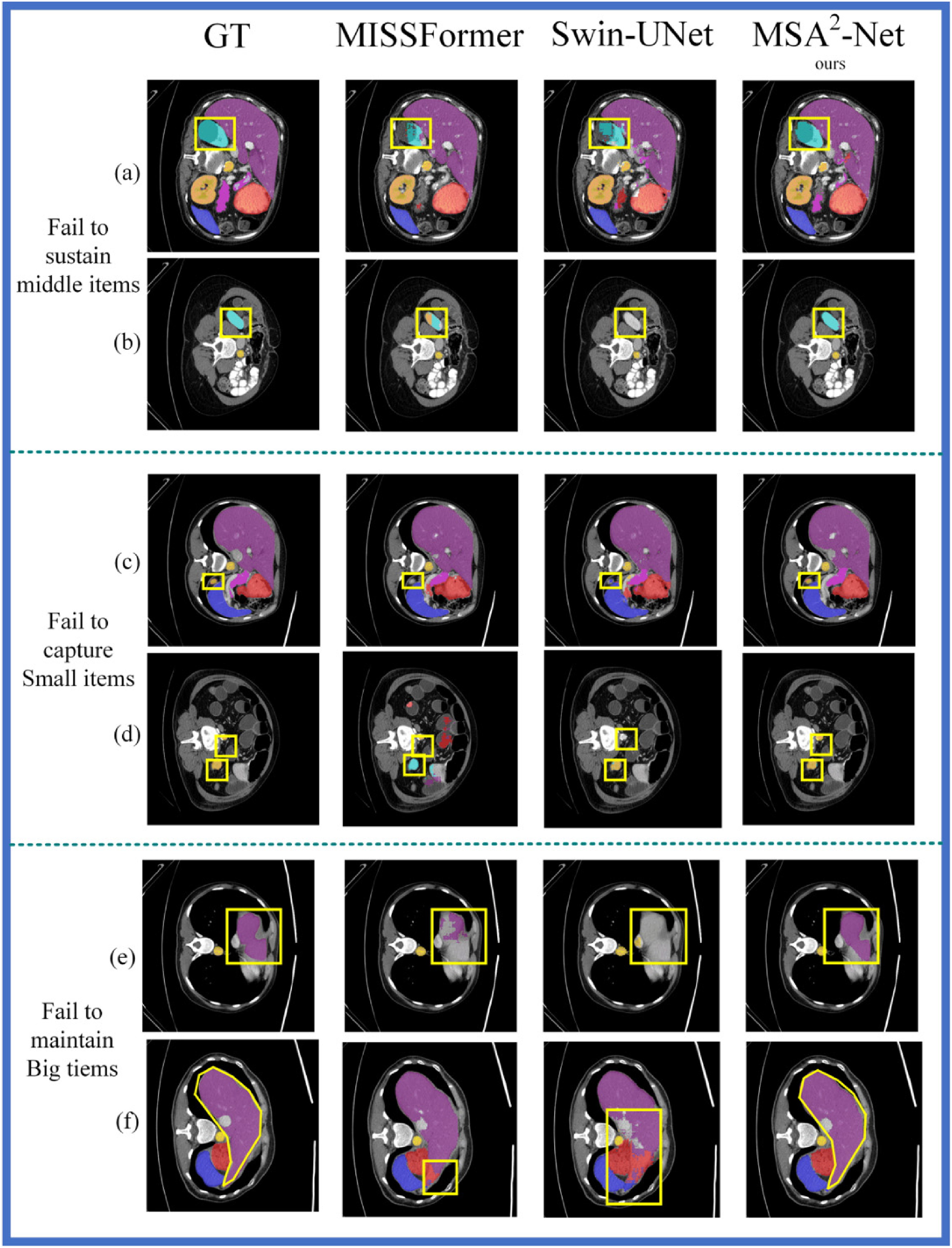
Segmentation effects of MSA2-Net, MISSFormer, and SwinUnet on the synapse.
Figure 6 provides a visualization of the segmentation task performed by MSA2-Net, MISSFormer, and Swin-UNet on the Synapse dataset. In Figure 6, the first column displays the Ground Truth images, the second column shows the segmentation images generated by MISSFormer, the third column presents the images produced by Swin-UNet, and the fourth column depicts the segmentation images created by MSA2-Net. The areas circled by the yellow line indicate the locations of sub-features where most methods failed to make accurate predictions.
Rows (a) and (b) in Figure 6 illustrate how other methods struggle to effectively capture medium-sized objects during the segmentation task on the Synapse dataset. In row (a), the area circled by the yellow box represents the left kidney. MSA2-Net segments the left kidney most completely, while MISSFormer and Swin-UNet fail to segment it as effectively due to their inability to preserve multi-scale information. In row (b), the yellow box again highlights the left kidney, but in a different channel dimension, resulting in a smaller ground truth value for the left kidney. MISSFormer retains some features of the left kidney because of its fixed-size convolution operation. In contrast, Swin-UNet does not employ convolution operations to maintain sub-features during the up-sampling process, which prevents it from detecting the left kidney. MSA2-Net successfully captures the multi-scale spatial information of the left kidney through the use of an MSADecoder, enabling it to correctly recognize the left kidney across different channel dimensions.
In rows (c) and (d), the area circled in yellow represents the gallbladder, a small organ. MISSFormer fails to capture the gallbladder because it uses a fixed-size convolution that filters out small features, while Swin-UNet misidentifies the gallbladder as other organs due to the lack of convolution operations during up-sampling. Only MSA2-Net successfully captures the gallbladder.
In rows (e) and (f), the difficulties in maintaining large objects are observed. In row (f), the yellow box highlights the liver. Although MISSFormer partially restores the liver segmentation due to its fixed-size convolution, it cannot fully capture the liver because its convolution kernel is too small, limiting its receptive field. Swin-UNet shows a significant discrepancy between the reduced liver and the original labeling because it lacks multi-scale convolution capabilities. In contrast, MSA2-Net, with its MSADecoder, completely restores the liver segmentation map, effectively handling the large object.
Performance of different networks on the ACDC and Kvasir-SEG
Table 3 presents the experimental results on the ACDC and Kvasir-SEG datasets. On the ACDC dataset, MSA2-Net outperformed SOTA competitors, achieving Dice score improvements of 1.08% and 1.88% over MISSFormer and Swin-UNet, respectively, with a notable peak of 90.95% in the RV subtask. This success on ACDC—which shares the volumetric nature of Synapse—validates that our model excels in capturing anatomical structures with spatial continuity.
Analysis of the results of the ACDC and Kvasir-SEG.

However, performance on the Kvasir-SEG dataset was less dominant. We attribute this to the fundamental difference in data characteristics. The design of our self-adaptive convolution module (SACM) relies on synthesizing kernels based on evolving statistical fingerprints, a strategy that thrives on the continuous z-axis variation found in CT/MRI scans. In contrast, Kvasir-SEG consists of discrete, independent endoscopic snapshots lacking inter-slice spatial correlations. Without the context of continuous scale evolution, the adaptive mechanism struggles to establish stable kernel adjustments for the erratic variations in independent samples, thereby limiting the performance gain compared to volumetric datasets.
Experimental results on skin lesion segmentation datasets
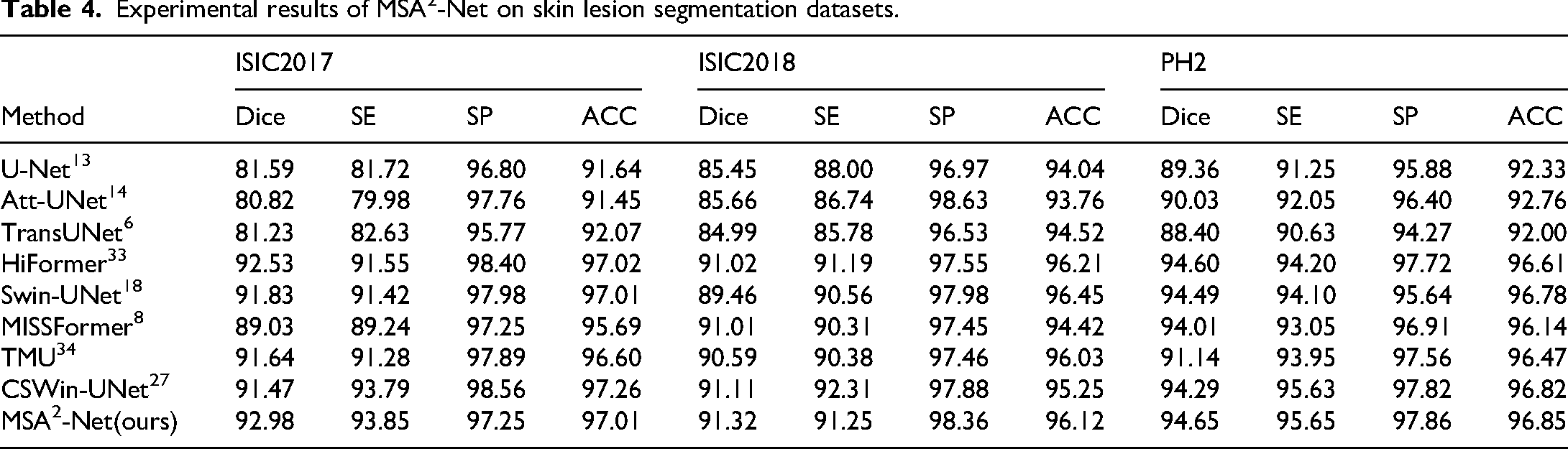
Table 4 shows the experimental results of MSA2-Net on the ISIC dataset. In all versions of the ISIC dataset, MSA2-Net achieved the best Dice scores, reflecting its strong segmentation performance and ability to accurately delineate skin lesion areas. Figure 7 presents the visual comparison of lesion segmentation by MSA2-Net and other advanced models on the ISIC2017 dataset. Rows (a), (b), and (c) contain lesions with complex backgrounds, testing the network's ability to filter redundant information. Rows (d) and (e) have lesions with simple backgrounds, testing the network's ability to capture detailed information. In row (a), MSA2-Net successfully identified most of the lesion area without being misled by background hair. In row (d), although all networks identified the main lesion area, MSA2-Net captured the most detailed lesion boundaries.

Visualization of MSA2-Net segmentation results on the ISIC2017 dataset.
Experimental results of MSA2-Net on skin lesion segmentation datasets.
Robust test
To strictly evaluate the stability and reliability of MSA2-Net, we conducted robustness tests on the Synapse datasets. While maintaining the original dataset partitioning schemes, we performed independent training runs for each model using different random seeds for weight initialization. We report the results in the format of “mean ± standard deviation”. This approach investigates whether the model's superiority is due to specific lucky initializations or inherent architectural robustness. To ensure a fair comparison, we retrained the representative baseline model under the exact same experimental settings as MSA2-Net.
As shown in Table 5, MSA2-Net demonstrates superior robustness compared to other architectures. MSA2-Net achieves the lowest standard deviation in average dice (1.05%), significantly more stable than TransUNet (1.65%) and Swin-UNet (1.55%). Even in the worst-case scenario (Mean - Std), MSA2-Net (85.05%) remains competitive with the average performance of AgileFormer (85.15%), and far surpasses the best average results of MISSFormer (81.20%). The robustness test confirms that MSA2-Net provides a statistically stable solution that reliably outperforms existing SOTA methods across repeated experiments.
Robustness analysis on synapse dataset (metric: dice %).

Ablation study
In this section, we performed ablation experiments on MSA2-Net using the Synapse dataset, as shown in Table 6. The results indicate that MSA2-Net achieves optimal performance with both the MSADecoder and the MSConvBridge, obtaining a Dice score of 86.49% and an HD95 of 14.13. When MSA2-Net is equipped with only the MSADecoder, the Dice score decreases by 9.16% and HD95 increases by 8.28. When equipped with only the MSConvBridge, the Dice score decreases by 9.93% and HD95 increases by 5.5. Without both the MSConvBridge and the MSADecoder, the Dice score decreases by 10.10% and HD95 increases by 7.49. Furthermore, when the model is not equipped with the SACM, performance declines even if the MSADecoder and MSConvBridge are retained. The primary purpose of the auxiliary encoder is to assist the encoder in preserving feature map details; therefore, the absence of this module leads to an increase in HD95.
Ablation study results.

Conclusion
In this work, we presented the MSA2−Net, a novel architecture designed to tackle the challenge of dynamic scale variation in medical segmentation. The core innovation, the self-adaptive convolution module (SACM), allows the network to dynamically synthesize kernels based on the statistical fingerprint of the input, thereby synchronizing the receptive field with the organ's spatial evolution. Additionally, we introduced the MSConvBridge to ensure semantic consistency by filtering redundant texture artifacts, and the MSADecoder to effectively reconstruct fine-grained details of small organs while preserving the global coherence of large structures.
MSA2−Net demonstrated superior performance across four prominent datasets (Synapse, ACDC, ISIC, and Kvasir-SEG). However, the performance discrepancy on Kvasir-SEG reveals an important insight: our adaptive mechanism thrives on the continuous spatial correlations typical of volumetric data (like Synapse and ACDC). In contrast, handling discrete, independent snapshots (as in Kvasir-SEG) remains a challenge, as the lack of inter-slice continuity limits the efficacy of our dynamic kernel adjustments. Future work will focus on decoupling the adaptive mechanism from spatial continuity to enhance robustness on snapshot-based 2D medical imagery.
Footnotes
Acknowledgements
Special thanks to Yuanyuan Li, whose profound insights and meticulous suggestions were instrumental in refining this work. I am truly grateful for her patience and perspective, which were a constant source of motivation during the most challenging phases of this project.
Author contributions
Conceptualization, Xiao Qin,Chao Deng.; methodology, Xiao Qin,Chao Deng.; software, Xiao Qin,Chao Deng.; validation, Xiaosen Li.; formal analysis, Xiaosen Li, Zhengyou Qin,Yuanxu Gong.; investigation, Xiaosen Li.; resources, Xiao Qin.; data curation, Xiaosen Li.; writing—original draft preparation, Chao Deng.; writing—review and editing, Xiao Qin.; visualization, Xiaosen Li.; supervision, Xiaosen Li.; project administration, Zhengyou Qin.; funding acquisition, Xiao Qin. All authors have read and agreed to the published version of the manuscript.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Ministry of Science and Technology of the People’s Republic of China, the Key Project of Science and Technology of Guangxi, (grant number STI2030-Major Projects 2021ZD0201900, AB25069247).
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Data availability statement
The datasets generated and/or analyzed during the current study are available in the following public repositories:
