Abstract
Introduction
Glioblastoma is the most common malignant primary brain tumor in adults, which is associated with a poor prognosis, with median overall survival between 16 and 20 months despite maximum treatment. 1 Initial treatment for glioblastoma includes maximum safety surgical resection and adjuvant chemoradiation. 2 Monitoring of patients after completion of initial treatment relies heavily on follow-up magnetic resonance imaging (MRI) to detect recurrence of the disease. Therefore, accurate analysis of MRI changes and the true condition of the tumor is of great significance for clinical decision-making.
Conventional treatment regimens vary in the degree of benefit of treatment for each patient with glioblastoma because of differences in genetic and epigenetic composition, protein expression levels, metabolic or bioenergetic behavior, and biochemical and structural composition of the microenvironment, resulting in different histopathological and radiographic features of individual tumors in different patient.3–5 Therefore, personalized treatment plans for individual patient tumor characteristics are increasingly advocated, which can assess the real situation of each patient to maximize the survival rate after glioblastoma treatment. However, so far, the standardization of glioblastoma responses and imaging indicators to determine disease progression has proven to be quite difficult, for example, with the addition of temozolomide to treatment criteria, clinicians have found that these imaging criteria do not accurately distinguish between true progression (TP) and false progression (PSP).6,7 PsP is a subacute treatment-related effect that typically occurs within 3 months of the completion of chemotherapy, and its imaging features lack specificity for clinical manifestations of tumor recurrence, which leads to clinical difficulties in developing follow-up treatment options, delaying optimal treatment for patients. 8
The rapid and widespread application of artificial intelligence technology confirms that it has unlimited possibilities, such as early radiomics research combined with machine learning, through computer analysis of image features that cannot be detected by the eye and extract valuable imaging features, classification of high-grade gliomas and low-grade gliomas,9,10 which has made imaginations have a strong interest in these unique imaging features, and led to the continuous deepening of research, but since the WHO classification in 2016, the classification of glioblastoma is also more based on the genetic level,11,12 This also means that for more accurate diagnostic and prognostic assessments, it is necessary to consider the differences between different genotypes of glioblastoma.13–15 Radiomics research has also begun to combine machine learning methods for research,16,17 but such methods are often based on simple linear models, the calculation method is not complicated, but in clinical practice, the changes of each patient are often non-linear, which also means that more realistic algorithms are needed to be applied to the clinic as much as possible, and radiomics and machine learning require a lot of image annotation, wasting a lot of energy and time. Deep learning (DL) is a part of machine learning that has developed in recent years (Figure 1), the study of DL neural networks can be achieved quickly and efficiently. 18 The neural layers of these neural networks are hundreds to thousands of layers, and more deep information can be obtained. The robustness of the model constructed by combining this information with clinical data is more stable. After training a DL model, it must be determined whether the model fits the data and whether its fit is affected by any bias. Model fitness can mainly be determined by observing the loss curve obtained during training. We aim to summarize recent applications of DL in glioblastoma prognosis prediction.
The relationship among AI, ML, DL.
Applications of deep learning in glioblastoma diagnosis and classification of glioblastoma
In previous studies, it was found that different molecular phenotypes of glioblastoma determine different tumor heterogeneity, which indicates that the prognosis of patients with glioblastoma will also have different outcomes,
19
so it is important to determine the molecular typing of glioblastoma, and conventional surgery requires surgical removal to take tissue specimens, and then molecular typing is detected; For glioblastoma located deeper in the brain and adjacent to important functional areas, if its molecular typing can be known in advance in a non-invasive manner, then preoperative protocol development and postoperative treatment evaluation are of good significance.
20
Radiomics-derived imaging phenotypes are associated with molecular markers to create “radiogenomics” models.
21
It is a rapid and reproducible tool for assessing tumor subtypes, mutant states, and intratumoral heterogeneity; using these features, it is used to non-invasively predict tumor progression, survival, and response to targeted therapies. DL is significantly more efficient than conventional radiomics and shallow learning because it enables automatic feature extraction through a convolutional network (Figure 2). Islam et.al
22
combined gene expression data with radiological features such as shape, geometry, and clinical information. Utilizing the TCGA (Cancer Genome Atlas) dataset and synthesizing missing MRI patterns using a fully convolutional network in a conditional generative adversarial network, a survival prediction model is finally generated and it works well. While there is still a small gap between the accuracy of DL and shallow machine learning, this gap is likely to continue to improve as algorithms continue to change.
(1) Tumor localization and segmentation
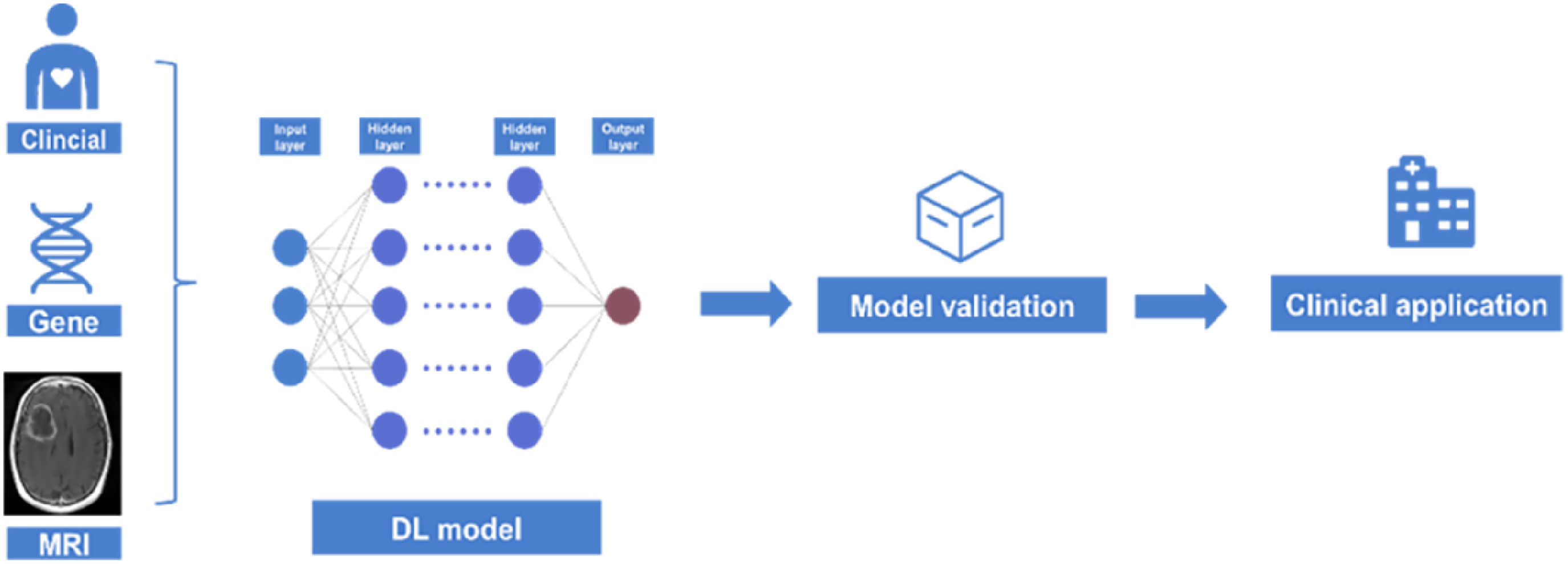
The training process of a multi-parameter deep learning model.
Defining the location of the tumor is clearly related to subsequent treatment, which directly affects the formulation of clinical decisions,
23
and a “probabilistic radiology map” of more than 500 patients with glioblastoma shows that stereotacticity is related to the incidence of tumors and age, resection range, gene expression and survival data.
24
After that, the DL model is used to automatically identify and locate the tumor. This facilitates rapid tumor discovery and precise localization in later work. When the localization is completed, it needs to be quantified, which requires the ability to accurately segment the tumor to obtain internal information, and the previous study quantified glioblastoma by manually or semi-automatically annotating training radiomics and machine learning models, which is time-consuming and the resulting features are often superficial and low-order image features. DL neural networks have many algorithms so far, such as the classical convolutional neural network (CNN, artificial neural network) and the later U-NET, 3D U-NET, V-net, by entering image data for model training to achieve automatic segmentation of tumors, and then obtain deep image features.,25–27 such a model can be combined with radiomics and machine learning models, and can also continue to be combined with neural networks to build clinical evaluation models.
(2) Tumor feature extraction and dimensionality reduction (3) Advanced MRI sequences and multimodal analyses
After the completion of tumor recognition and segmentation, how to process the obtained segmentation model into a form that the computer can understand has become a big problem, this place often has two forms, one is the segmentation of the image using machine learning methods to extract its features,
28
one is to continue to directly through the neural network feature extraction,
29
these two ways in the order of magnitude of the extraction of features have obvious differences, the latter can get more features than the first method, And through the use of DL to extract many features do not require a lot of manual sketching, which has an important application prospect in the efficiency of later practical use, in the face of so many features, we need to reduce their dimensionality, so that these high-latitude data volume through a series of methods into a basic one-dimensional vector, and then filter it to get meaningful feature values for us. Once we have meaningful features, we can combine them with a range of other data sources for integration and analysis.
The resulting screening features for data analysis and model establishment are not rigorous, because the clinical need is often a comprehensive model, clinical needs to consider the factors are multi-party, and accurate, therefore, the obtained features and effective clinical information to combine and get a usable value prediction model is the problem we need to study, at the same time, we also have to realize that MRI has many sequences, whether it is a conventional sequence (T1WI, T2WI) or a functional sequence (DWI, ADC, DKI, etc.), different sequences have their own unique advantages for the display of lesions, which also means that when we are conducting DL research, if we only analyze a single sequence, the information obtained is one-sided.30–32
Application of DL in the prognosis assessment of glioblastoma
Glioblastoma tends to have a poor prognosis, even if the tumor is completely removed by surgery, but patients often have a high risk of recurrence after 7 months of healthy storage,
33
glioblastoma has the characteristics of infiltrative growth, the actual invasion range is greater than the visible range, which means that even if the patient receives standardized treatment (including surgery and postoperative radiotherapy and chemotherapy), there is still a risk of recurrence, and because the perigma microenvironment of glioblastoma is complex,
34
based on simple imaging tests can not determine the actual depth of infiltration of GBM, This is detrimental to the clinical assessment of the prognosis of glioblastoma. Even in the process of postoperative chemoradiation, the pseudo-progression generated by chemoradiation and chemotherapy lacks the characteristics of imaging, and it is difficult to judge whether it is pseudo-progression or recurrence by the human eye alone, interfering with the subsequent treatment plan formulation. Therefore, in recent years, the prognosis assessment of glioblastoma using machine learning and DL has become an important research problem.
(1) DL-based assessment of the correlation between glioma peripheral range and prognosis (2) Application of DL prognostic prediction model combined with genotype and peritumoral glioblastoma
Glioblastoma progresses rapidly, and its depth of invasion often exceeds the scope of current MRI sequences, and the complex peritumoral microenvironment surrounding glioblastoma leads to effective surgical resection and postoperative chemoradiotherapy for glioblastoma patients. The risk of recurrence is often unknown, including the peritumoral microenvironment of glioblastoma, so it becomes a good way to try to construct a multivariate-based prognostic prediction model through machine learning. In previous studies, The most commonly used methods are radiomics, which can have good significance for prognosis prediction. However, it is still limited by the quality of manual segmentation and annotation and often consumes a lot of time. Therefore, the DL method is natural. also tried in this direction, Bani-Sadr A et al.
35
tried in this regard, obtained a model that can be automatically segmented through the training of a neural network, and extracted its features, and finally compared the obtained features with clinical information. Combined, a good performance evaluation was obtained. However, the difficulty of interpretability of the internal mechanism of DL becomes a big problem, which requires more experiments and methods to demonstrate.
Glioma is a general term for a group of neuroepithelial tumors with glial cell phenotype. The WHO Classification of Tumors of the Central Nervous System, released in 2021, integrates the histological features and molecular phenotypes of tumors, and proposes a new classification standard for tumors. This standard is currently an important basis for the diagnosis and grading of gliomas. The pathological diagnosis of gliomas should integrate histological types and related molecular markers. Histopathology can provide basic morphological diagnosis for glioma, and molecular pathology can provide more characteristics of tumor molecular genetic variation, which can directly affect clinical prognosis and the choice of treatment plan. Therefore, by incorporating molecular such as genotype and phenotype, It has also become a very challenging way to build DL prognosis prediction models based on information. Liu et al.
36
divided 266 patient images from hospitals and tcga-gbm into the training set, validation set and test set in a ratio of 6:1:1 by using a CNN, and then expanded the sample size based on data augmentation And verified the good performance of the DL model method in the prognosis of IDH wild-type and mutant patients.
Challenges in the clinical application of radiomics for glioblastoma:
Although DL has demonstrated potential in various aspects of glioblastoma treatment, these approaches have yet to be introduced into mainstream clinical practice. Barriers to translation include limited reproducibility of algorithms and less robust DL models. This requires that we follow the unified standard AI extension guidelines for clinical trial reporting in the process of artificial intelligence practice
(1) Image acquisition (2) Segmentation and feature extraction (3) Interpretability of DL Models
DL model training requires a large amount of data to support it, which also leads to obstacles in the construction of glioblastoma models. The increase in the amount of data can ensure the performance and robustness of the model, but due to glioblastoma, There are not many cases in each hospital. Therefore, there is an urgent need for a method to provide as many cases as possible for the construction of DL models. Therefore, the establishment of public datasets solves this problem very well. There are many good data sets for blastoma (eg, TCGA-GBM, IVY-GAP, etc.),37–39 and recently, the Pennsylvania Health System provided a large glioblastoma data set (UPENN-GBM),
40
the data set provides 501 cases and has complete clinical and molecular typing data, which makes the DL research of GBM can be better carried out. However, for public datasets, we still need to use some external datasets for external validation to prove the robustness of the model, which is necessary for DL.
Although considered the highest standard of segmentation, manual segmentation of images is labor-intensive and increases the risk of observer bias. In contrast, semi-automatic and fully automatic methods can improve robustness and reproducibility. The extracted features depend on the segmented regions and tumor margins (Figure 3), so segmentation is a critical step. Although automatic feature extraction has less variability in semantic feature scores, these methods still result in site-specific changes when imaging is obtained.
41

Differences remain between manual segmentation and algorithm-based automatic segmentation.
As DL models play an increasingly important role in many scenarios in people's daily lives, the “interpretability” of models becomes a key factor in determining whether users can “trust” these models (especially when we need to When machines give predictions and decision-making results for important tasks related to human life and health, property safety, etc.) 42 In the current wave of DL, many newly published works claim that they can achieve good performance on target tasks. Nevertheless, users still need to understand the reasons for drawing conclusions from a more detailed and concrete perspective in application scenarios such as medical, legal, and financial. Giving the model strong interpretability is also conducive to ensuring its fairness, 43 privacy protection performance, and robustness, explaining the causal relationship between the input and output states, and improving the user's trust in the product.
In order to make the DL model more “transparent” to users, researchers in recent years have started from the two perspectives of “interpretability” and “completeness,” and have studied the working principle of DL model prediction, decision-making results and DL. The internal structure and mathematical operations of the model itself are explained. So far, researchers in the field of interpretable DL have achieved gratifying achievements in three levels: “the processing of data by the network,” “the representation of the data by the network,” and “how to build a DL system that can generate self-explaining.” 44
As far as the network working process is concerned, the researchers explained the original model by designing a linear surrogate model, a decision tree model equal to the original model, but with stronger interpretability; in addition, the researchers also developed saliency maps, Methods such as CAM visualize 45 the most relevant raw data and computational processes for prediction and decision-making, giving a very intuitive explanation of how DL models work.
At data representation is concerned, the existing DL interpretation methods involve three research directions: “layer-based interpretation,” “neuron-based interpretation,” and “representation vector-based interpretation.” The important factors affecting the performance of DL models are explored in terms of scale, neuron function, etc. 46
As far as self-explaining DL systems are concerned, researchers have carried out research on attention mechanism, representation separation, explanation generation, etc., and realized the visualization of the model working mechanism in the “vision-language” multimodal task. Based on technologies such as InfoGAN and capsule network, the representations that have different effects on learning are separated, and fine-grained control of data representation is realized.
Conclusion
Compared with radiomics and shallow machine learning, DL can be a more powerful, non-invasive and effective method, but the development of this technology still requires continuous improvement in algorithms and clinical practice to provide clinical glioblastoma cells. The diagnosis and treatment of tumor patients and precise postoperative intervention provide more valuable information, in order to obtain a better prognosis.
Footnotes
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethical statement
Our study did not require ethical board approval because it did not contain human or animal trials
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work is supported by the National Natural Science Foundation of China (81871333, 82260340), Guizhou Province 7th Thousand Innovational and Enterprising Talents (GZQ202007086), 2020 Innovation group project of Guizhou Province Educational Commission (KY[2021]017), Guizhou Province Science & Technology Project ([2020]4Y159 and [2021]430), Guizhou Province Science & Technology Innovation Talent Team (CXTD[2022]006).
基金项目:
国家自然科学基金(81871333,82260340);贵州省第七批“千人创新创业人才”(GZQ202007086);贵州省精准影像与诊疗创新群体(黔教合KY[2021]017);贵州省科技支撑计划(黔科[2020]4Y159,[2021]430);贵州省脑科学科技创新人才团队(CXTD[2022]006)。
Author biographies
Pengyu Chen is a master in Brain Tumors and Artificial Intelligence. His area of research is Brain Tumors and the Application of Artificial Intelligence in Medicine.
Ping Wang is a master in Neuroimaging. Her area of research is Brain functional imaging.
Bo Gao is a professor in Neuroimaging. His area of research is Brain Molecular Imaging and Artificial Intelligence.
