Abstract
During the last 10 years the number of porcine models for human bacterial infectious diseases has increased. In the future, this tendency is expected to continue and, therefore, the aim of the present review is to describe guidelines for the development and reporting of these models. The guidelines are based on a review of 122 publications of porcine models for different bacterial infectious diseases in humans. The review demonstrates a substantial lack of information in most papers which hampers reproducibility and continuation of the work that was established in the models. The guidelines describe overall principles related to the inoculum, the animal, the infected animal and the post-mortem characterization that are of crucial importance when porcine models of infectious diseases are developed, validated and reported.
Introduction
The use of pigs as experimental animals is increasing.
1
In the EU 239,089 pigs have been used for experimental purposes during the last 5 years (Figure 1(a)).
2
The use of pigs as experimental animals has primarily been in the areas of toxicology, metabolism, cancer and xenotransplantation.
1
However, during the last 10 years the number of porcine models for human bacterial infectious diseases has increased, and it is reasonable to assume that many more models will follow in the future (Figure 1(b)).
3
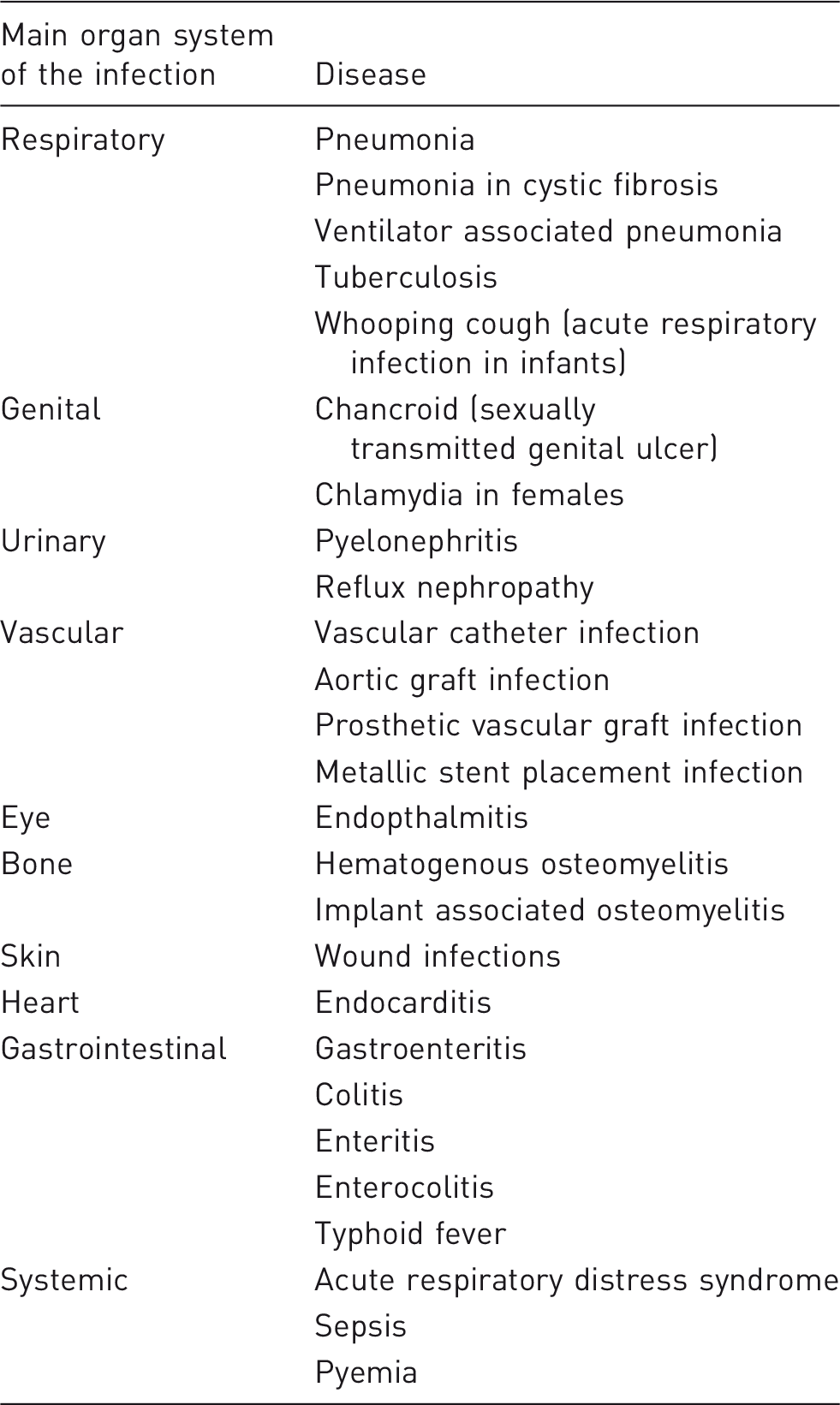
Pigs have been used to model the different human bacterial infectious diseases listed in Table 1. The aims of the different models were to study either the efficacy of novel and well-known antimicrobials, prevention and treatment of infection, specific virulence mechanisms of pathogens or the process of inflammation and disease development. The increased popularity of pigs as experimental animals is also reflected in several reviews about the advantages of porcine models in biomedical research.1,4–6
(a) Pigs used for biomedical research from 2012–2016 in the EU. Pigs are frequently used as experimental animals in most of the European countries.
2
(b) Number of publications where pigs have been used to model human bacterial infectious diseases. The colours indicate the target organ system infected in the models. References for all the models can be found in the supplementary material. Human bacterial infectious diseases modelled in pigs.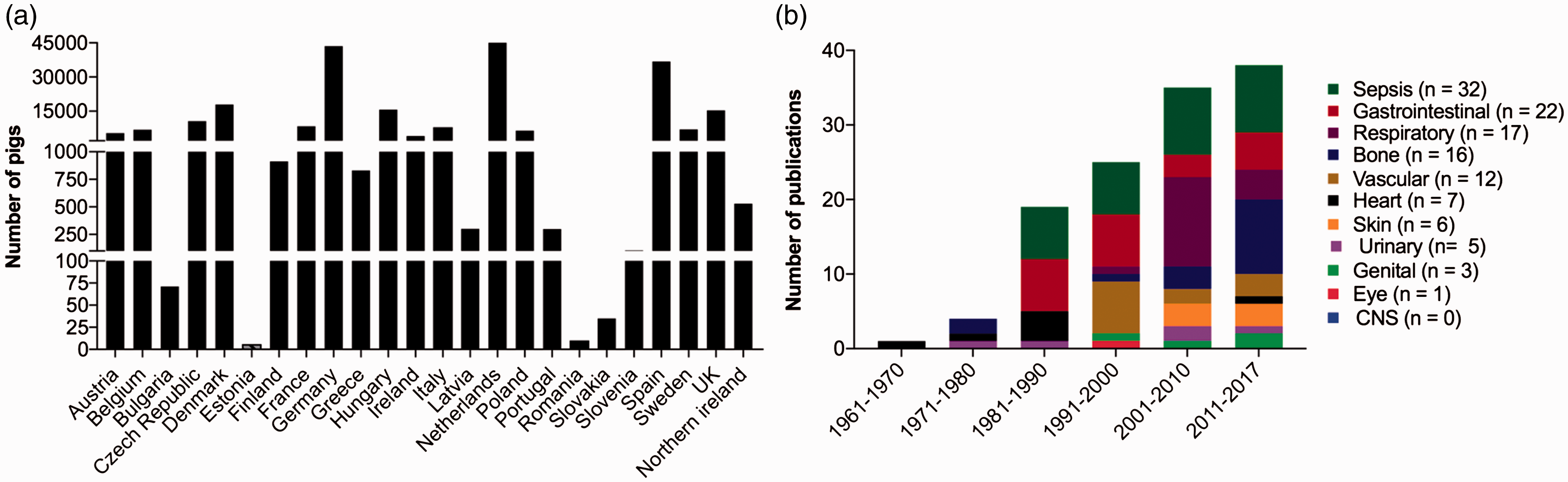
Animal models of infections are important tools to study different aspects of infectious diseases in humans because they provide all factors involved in a pathogen–host interaction. 7 The reason why pigs are preferred as experimental animals is due to their size, anatomy and physiology which are comparable to humans.4,6 Recently, the complete genome of the domesticated pig was sequenced 8 and more laboratory reagents specific for porcine tissue have become available. 9 However, two additional points make pigs especially favourable for modelling of infectious diseases in humans. Firstly, the immune response of pigs is comparable to that of humans as the porcine and human genome show similarity, both structurally and functionally, among immune-related proteins. 10 Furthermore, as in humans, pigs have a large percentage of neutrophilic granulocytes in the peripheral blood. 11 Second, porcine models are advantageous due to spontaneous development (naturally-occurring infections) of the same infectious diseases as in humans. 3 As an example, the pathogenesis and lesions of spontaneously occurring chronic pressure wounds, endocarditis, pyelonephritis and hematogenous osteomyelitis are comparable to those seen in humans and are also often caused by the same bacteria. 3 Homologous models (same aetiology, pathogenesis and symptoms) are described as the most ideal for capturing the complex nature of diseases. 12 Beside all the advantages of using porcine models in translational research, it is also important to consider the differences between pigs and humans. As an example, there are some major anatomical differences in the gastrointestinal tract. In pigs the caecum, which has a remarkably larger size, and colon are arranged in a series of centrifugal and centripetal coils differing from those in humans. 4 In a review by Swindle et al. the anatomical similarities and dissimilarities between pigs and humans are described in detail for all organ systems. 4 Another difference, which is very important for bacterial challenge trials is the body temperature. In normal pigs the rectal temperature can vary between 38–40℃ which can make it difficult to define the occurrence of pyrexia. 13
For scientific, ethical and economic reasons, animal experiments should be appropriately designed, correctly analysed and clearly described. However, it has been documented recently that there is a lack of detail in the way research using animal models is reported. 14 Furthermore, several authors have stated the need for more efficiency in translational medicine and called for international guidelines to set higher standards for the validity of animal models in different disciplines. 15 Therefore, the aim of this review was to investigate the details of the reported data from existing porcine models of bacterial infectious diseases in humans. Furthermore, the aim was also to describe guidelines of overall principles that are of crucial importance when porcine models are developed, validated and reported.
Material and methods
Data collection
Papers were collected by a comprehensive search of Medline, Web of Science and Google Scholar. The aim of the search was to find papers in which pigs were used to model bacterial infections in humans. Prior to the search, it was decided to allocate all papers into groups based on the target organ-system of the infection. The following target-organ systems were defined: systemic (sepsis models); respiratory; urinary; brain; gastrointestinal; genital; eye; heart; skin; bone; vascular (graft/catheter). Thereafter, Group A search words were combined with Group B search words in the databases:
Group A search words:
Porcine model OR pig model OR swine model NOT guinea, human OR man; Infection OR infectious OR bacteria OR micro-organism OR pathogen OR disease NOT virus.
Group B search words:
Lung OR pneumonia OR respiratory OR airway; Kidney OR nephritis OR urinary OR cystitis; Brain OR CNS OR neuro OR encephalitis OR meningitis; Gastro OR intestine OR enteritis OR colitis; Genital OR reproductive; Sepsis OR pyemia OR bacteraemia OR bacteremia; Graft OR catheter; Eye OR orbital OR endophthal OR retin OR uve OR kerat; Bone OR bone infection OR osteomyelitis; Heart OR endocarditis; Skin OR wound.
The following data were extracted from all included publications: bacterial strain used for inoculation (including origin); inoculation dose; inoculation volume; animal data (age, breed, sex, immune status, number); housing data (acclimatization period, diet, individual/group housing, isolation stable, specification of housing condition); inoculation route; anaesthesia during inoculation; surgery during inoculation; methods used to monitor local and systemic infection in alive animals; pain management; time frame; full-body necropsy; occurrence of secondary systemic bacterial (inoculum) spread; methods used to characterize local and systemic inflammation post mortem; methods used to identify the inoculated bacteria post mortem. Finally, the impact of each study was evaluated by registration of the number of citations (with and without self-citation) in Scopus. The extracted data and all references can be found as supplementary material.
Results and discussion
In total, 122 different papers were collected for the present review (Figure 1(b)). All the models followed the same structure: a pathogen was introduced into an animal, which then became infected for a fixed period of time followed by post-mortem analyses. Therefore, the results were organized in four sections dealing with the inoculum, the pig, the infected pig and post-mortem characterization (Figure 2).
Guidelines for development and reporting of porcine models for human bacterial infectious diseases. The inoculum, the pig, the infected pig and the post-mortem characterization must be well considered and reported to allow reproduction. The bullets listed in the figure should be mandatory in all papers reporting porcine models of human bacterial infections.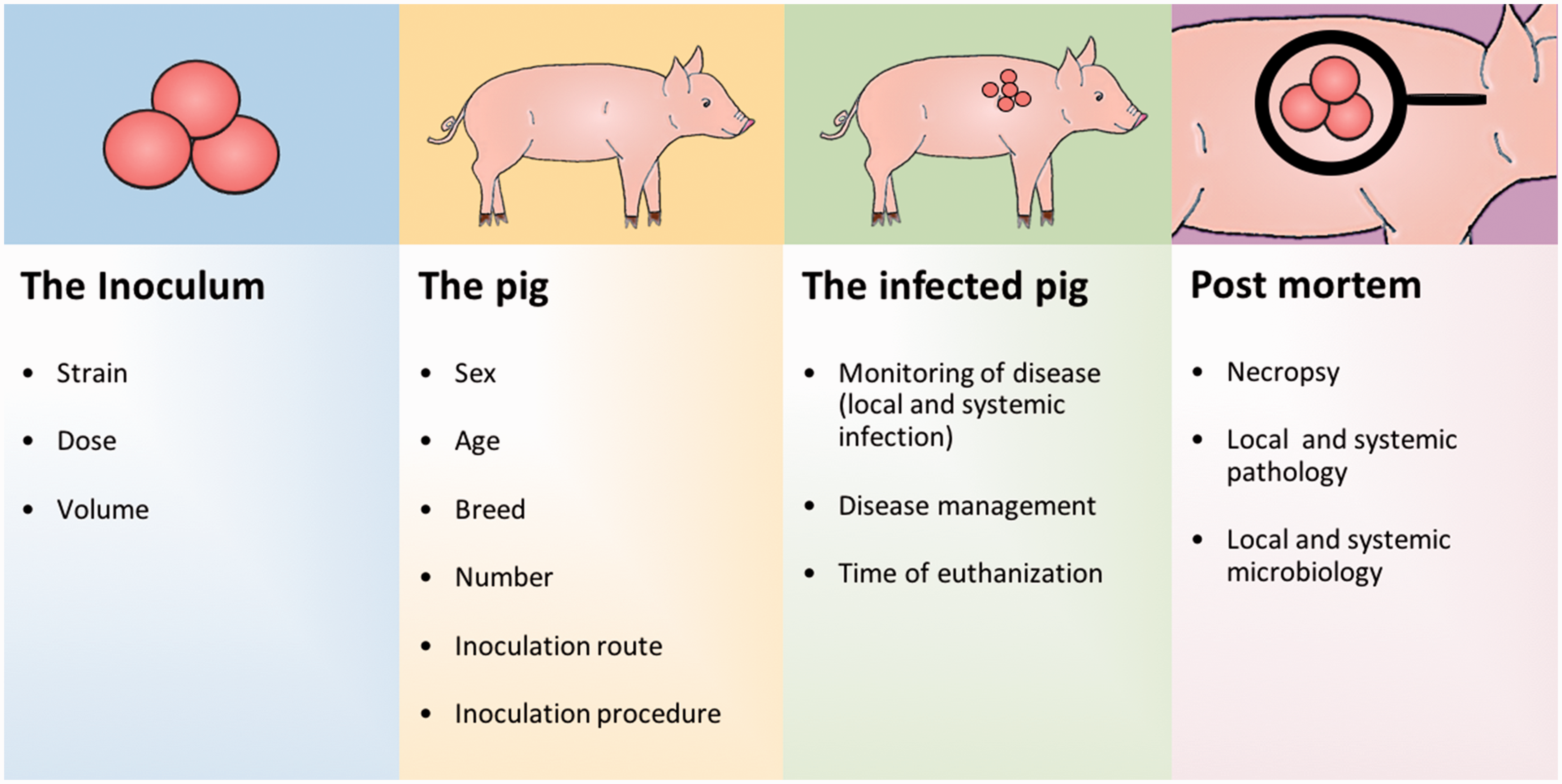
The inoculum
The inoculum consists of an exact dose of a specific bacterial strain suspended in a fixed volume (Figure 2).16–20 The different bacterial strains used for inoculation of pigs for human bacterial infections are shown in Figure 3(a). The most commonly used gram positive and gram negative bacterial species, regardless of target organ system, were (a) Bacterial species used in porcine models of human infections. (b) Inoculation doses (colony forming units; CFU) used in porcine models of human bacterial infections with respect to target organ system for the infection. Doses reported as an interval – for example, 102–4 – have been reported as 102, 103 and 104. In studies where multiple doses of the same 10 factor was used, only one dose was reported. (c) Animal number, (d) age and (e) time-frame reported in porcine models of human bacterial infectious diseases. The highest inoculation time for each study was chosen if multiple time points of euthanasia were described. The colours indicate the target organ system in the models.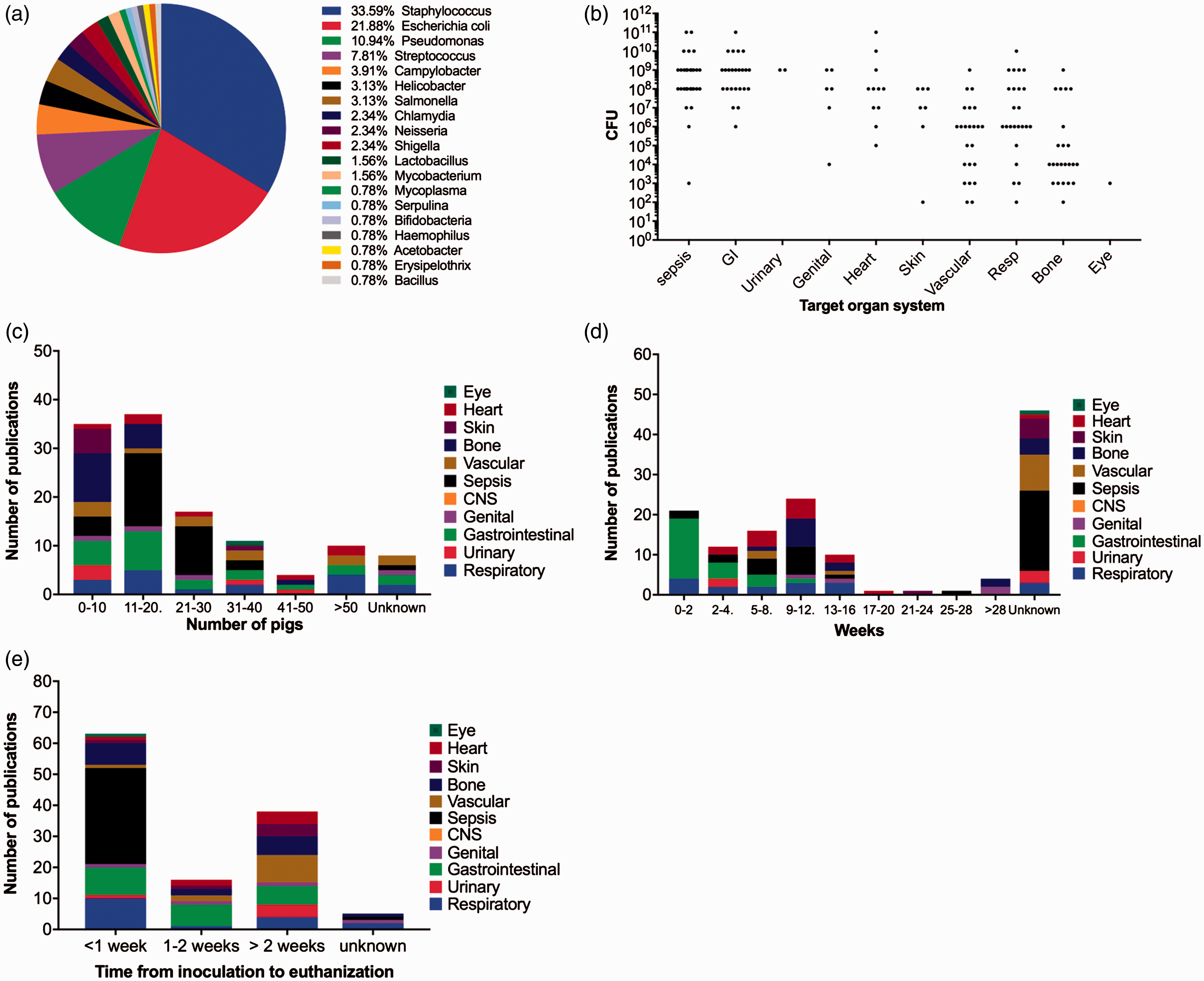
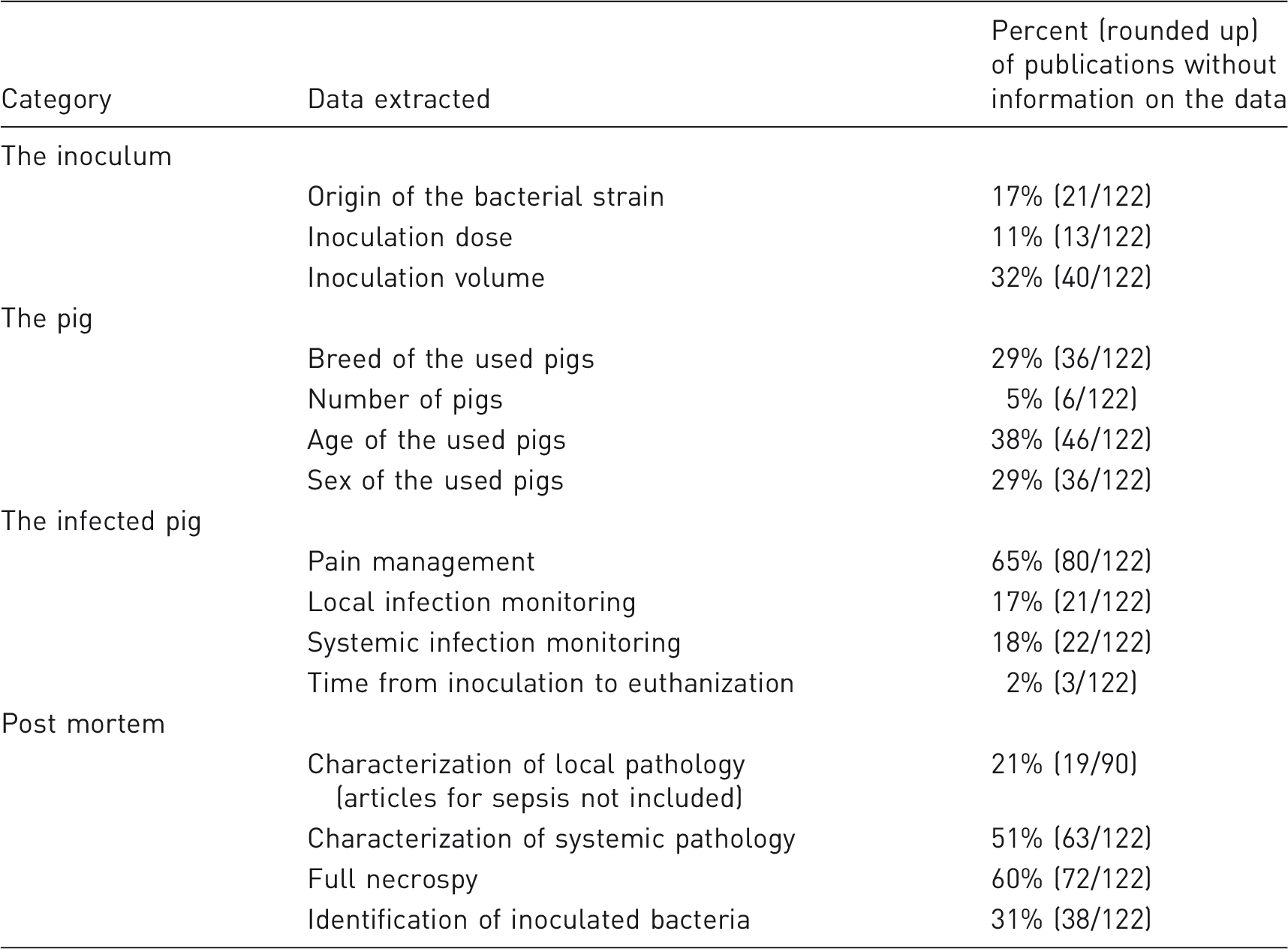
Overview of ‘not reported data’ in 122 publications describing porcine models of human bacterial infectious diseases.
The pig
Inoculation routes used in order to establish porcine models of human bacterial infectious diseases. It is registered if the inoculation routes demand anaesthesia and surgery (defined by cutting).
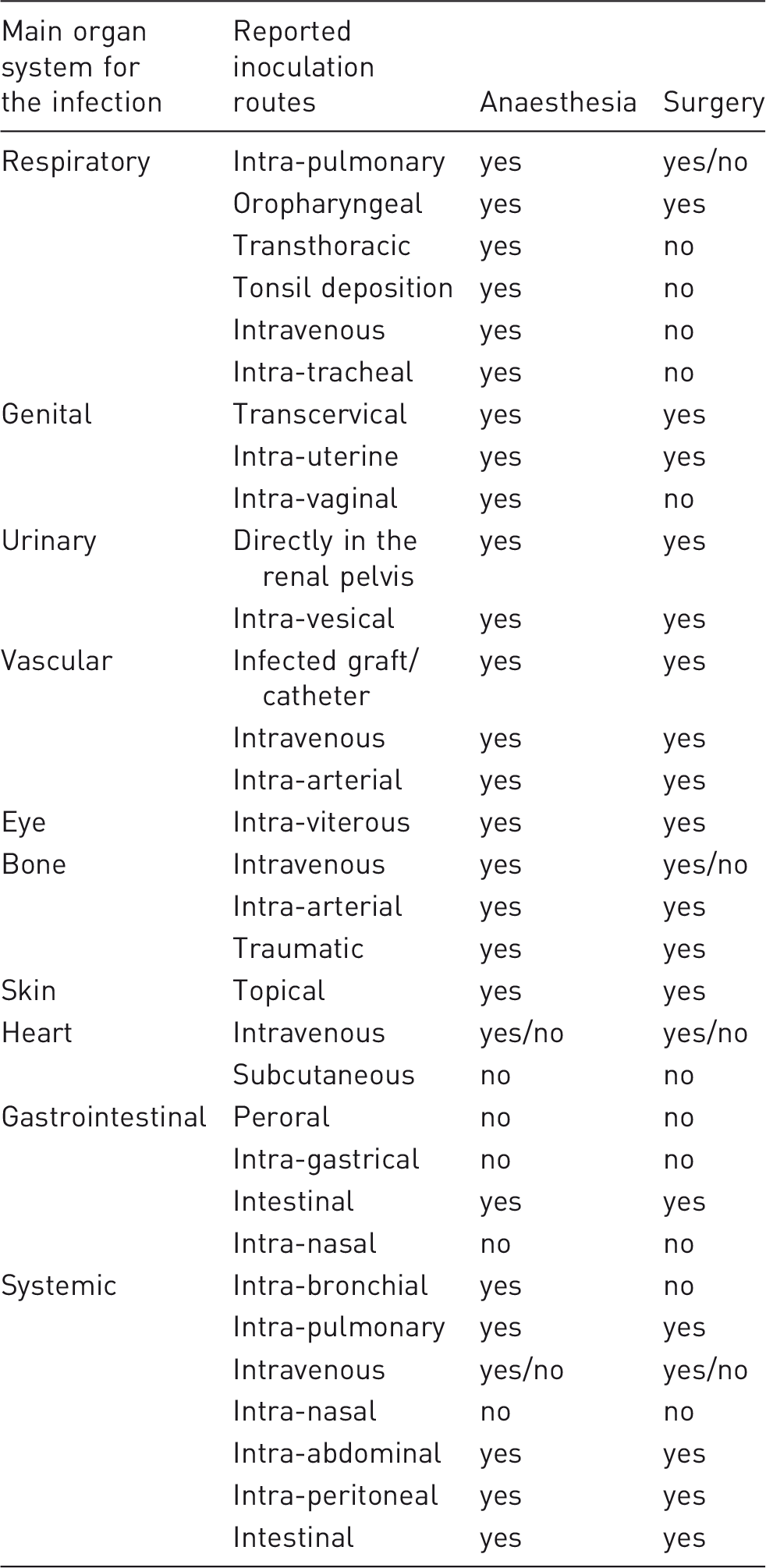
Reporting of housing details was in general found to be very limited. Only 22% of the studies report a period of acclimatization, even though a pre-study stabilization period is recommended for experimental pigs to be used in survival studies. 6 In 23 of the 122 papers it was mentioned if the animals were housed individually or in groups. Of these 23 studies, group housing was reported in 12 papers and the number of pigs housed together ranged from two to six. Swine are social animals and to reduce stress, it is recommended that they have the opportunity to interact with or see other members of their species. 6 The use of an isolation stable was reported in 13% of the papers included in the present review. Diet details were reported in 25% of the included papers and housing conditions – for example, pen size, flooring, bedding specification, humidity or temperature – were only reported in 11% of the papers.
The infected pig
It is necessary to consider how to monitor and handle the infected pig with regards to disease development. As the models replicate infectious diseases in humans, discomfort and signs of pain can be expected. In 34% of the papers reviewed analgesic treatment was reported during the study period. Analgesic treatment was primarily used in studies of sepsis, bone infections, infections of the respiratory system and catheter/graft associated infections.20,39,44,45 Both opioids and NSAIDs have been used as analgesic treatments in porcine models of human bacterial infections.20,45,46 A drawback of NSAIDs is their anti-inflammatory effect, however recently opioids have also been shown to have anti-inflammatory activity.
47
In a porcine model of osteomyelitis, macroscopic bone lesions with sequester formation have been established despite daily oral NSAID treatment.
39
Beside analgesic treatment, diseased pigs should have an increased number of daily inspections and special requirements to minimize their discomfort, such as fluid therapy or oxygen supply, should be established if possible.
25
In the 122 papers collected for the present review, the development of a local infection in the main target organ system was monitored by either clinical signs, imaging techniques (e.g
Post-mortem characterization
To characterize a porcine bacterial infectious disease model, it is important to focus on both local and systemic signs of infection post mortem.16,17,21,23,27,39,41,43,52,53 Local inflammatory lesions will occur in the main target organ and have been characterized by several methods (Figure 4(a)). However, most often macroscopic pathology and histology were applied. The inflammatory lesions should be described and quantified by semi-quantitative scoring systems or objective measurements.17,39 It is relevant, in all porcine models of bacterial infectious diseases, to include a definition of infection. Not all inoculated animals may develop an infection, or the same grade of infection. Therefore, it is important to have criteria that define if an animal was infected or not.22,39,54 From the main target organ the local infection can spread either locally into the adjacent tissue or systemically. A local spread will be registered in the description of the lesions pattern of distribution. However, an infection in the main target organ may result in an infective embolus (bacteria and particular material in the blood) which can settle in other organ systems and initiate secondary lesions. This will result in signs of infection, which can interfere with the signs caused by the primary infection of interest.18,30,52 Therefore, it is always important to look for and describe the occurrence of secondary, embolic infectious lesions. Macroscopic pathology and histology of internal organs (i.e. lungs, kidneys, spleen and liver) were the predominantly used methods to exclude secondary foci of infection (Fig. 4(a)).18,19,39,55,56 However, in half of the porcine models reviewed, occurrence of systemic inflammation was not described, and a full necropsy was only reported in 40% of the papers (Table 2). Microbiologically, it is important to look for the inoculated bacteria both locally within lesions and on inserted implants/devices and systemically.
57
In 28% of the publications (sepsis models excluded) the inoculated bacteria were recovered from blood or other organs than the local organ in question, or animals were reported dead due to sepsis. The techniques listed in Figure 4(b) have been used to localize and re-isolate the inoculum post mortem. The lungs of pigs trap blood-borne pathogens due to the presence of specialized pulmonary intravascular macrophages.
20
Therefore, lung samples for quantitative microbiology were commonly used to look for a hematogenous spread of the inoculated bacteria.18,31,56 Culture (by the use of swabs) was frequently used, however, the technique will not identify bacteria located inside the tissue.
58
Clinically, it has been demonstrated that swabs are less effective compared to biopsies for the diagnosis of infections.
58
The occurrence of bacterial biofilm formation should be considered in future porcine models of bacterial infections.
59
Biofilm can be formed both inside tissue and on inserted implants and devices, and biofilm is an important factor in sustaining infections in humans.
59
During the last decades, it has been realized that bacterial biofilm formation is involved in chronic infections.
59
Chronic infections are of increasing concern due to their high healthcare burden and, recently, diagnostic and therapeutic guidelines for biofilm infections were developed by ESCMID (European Society of Clinical Microbiology and Infectious Diseases).
59
The guidelines raised the following question: what research is urgently needed to improve diagnosis and treatment of bacterial biofilm infections? Among others, one of the answers was the development of better animal models that realistically reflect infectious diseases in humans. Biofilm formation can be confirmed by different types of microscopy and sonication of implants.
3
(a) Common methods used to characterize the local (bottom) and systemic (top) inflammatory response post mortem in porcine models of bacterial infectious diseases in humans. (b) Common methods used to isolate the inoculum post mortem in porcine models of bacterial infectious diseases in humans.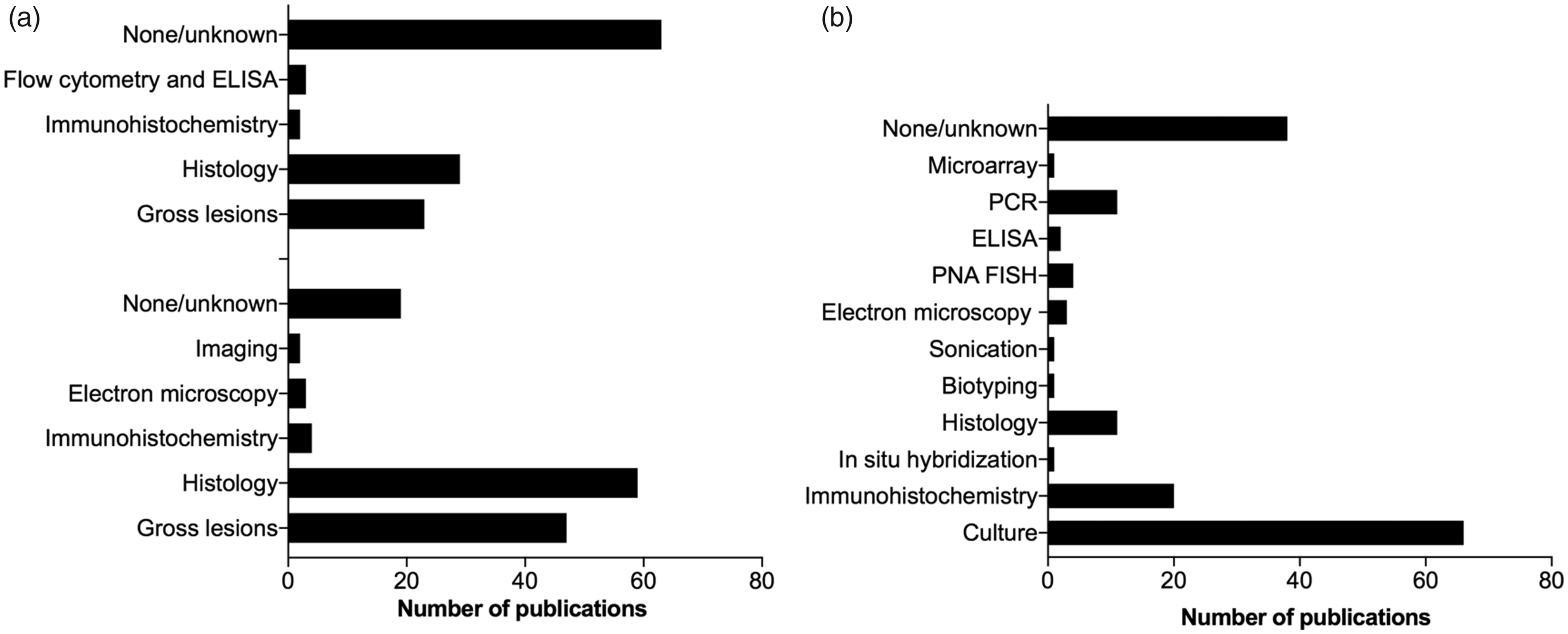
Citation impact
The analysed porcine models of bacterial infectious diseases in humans showed citation impact. In total, 88 of the 122 papers included in the present review were registered in Scopus. The mean number of citations for the 88 papers were 29.6 ± SEM 4.48 with self-citation and 22.73 ± SEM 3.617 without self-citation. If the porcine models were grouped by organ system the number of citations, without self-citations, from the highest to the lowest was as follows: urinary 85.5 ± SEM 83.5; respiratory 39.7 ± SEM 15.9; gastrointestinal 31.6 ± SEM 10.72; skin 21.4 ± SEM 5.9; sepsis 20.24 ± SEM 4.723; vascular 17.1 ± SEM 4.478; genital 11 ± SEM 6; heart 6.25 ± SEM 1.1 and bone 4.6 ± SEM 1.7. The 20 most cited porcine models of human bacterial infections were models of sepsis, pneumonia, wounds, graft infections, pyelonephritis and gastroenteritis (Figure 5). These most cited models were developed between 1975 and 2013 and they were cited regularly since publication – that is, the mean number of citations per year since publication = 7 ± SEM 2.6 (6 ± SEM 2.3 without self-citation). The most cited porcine model for human bacterial infections was published in Nature in 2013
60
and demonstrated pathophysiological aspects of cystic fibrosis and pneumonia: since publication this model has been cited 250 times by other authors.
The 20 most cited porcine models of bacterial infectious diseases in humans. The models have been regularly cited since publication. The last name of the first author and the year of publication is mentioned on the 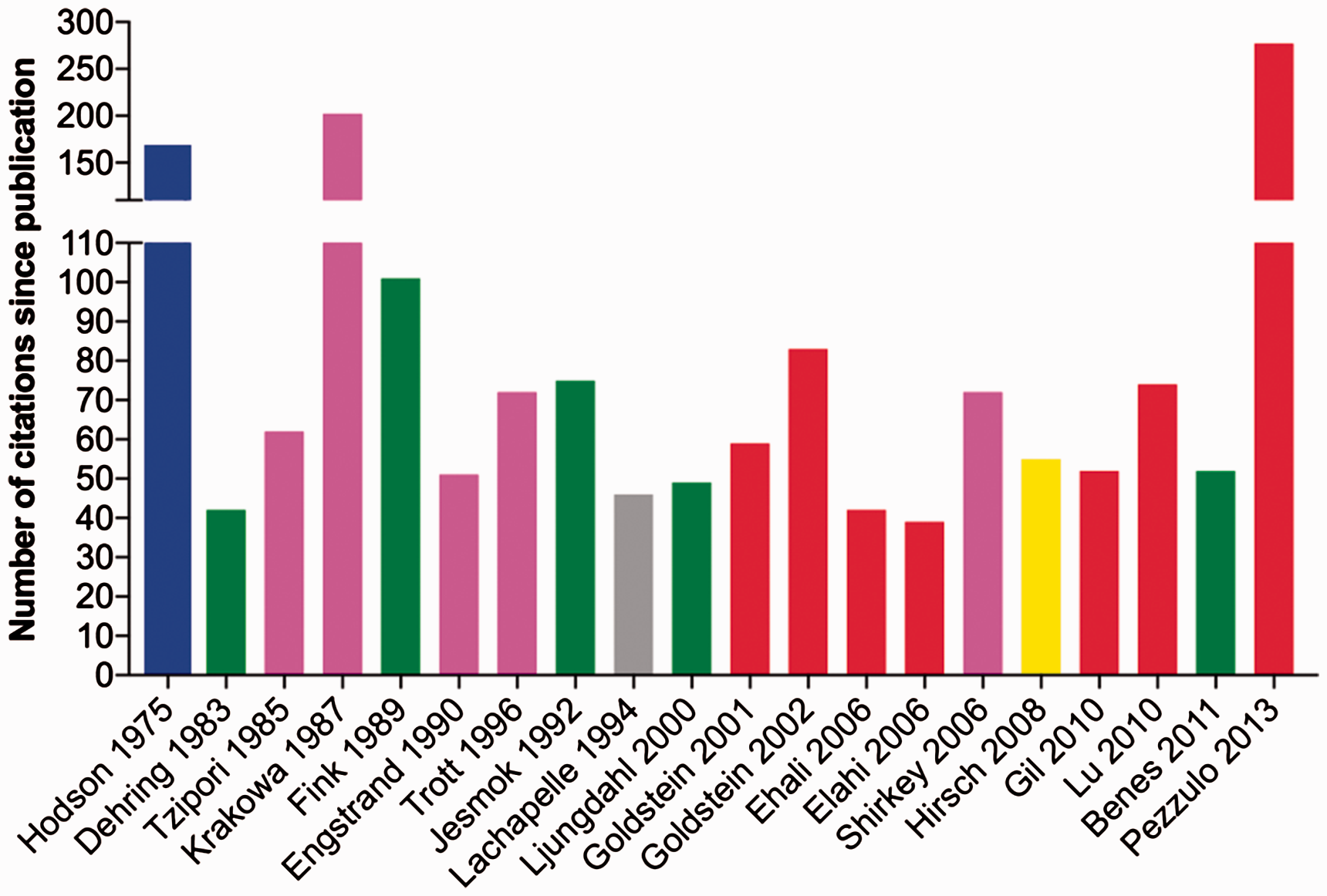
Guidelines for reporting of porcine models of bacterial infectious diseases in humans
The inoculum
The inoculum must be strictly reported and consist of the bacterial strain, the dose and the volume. The bacterial strain defines the virulence of the inoculum. Therefore, it is recommended to report as many details as possible for the used bacterial strain; for example, human or porcine origin, diseases/outbreaks caused by the strain, severity of disease development, expression of known virulence factors including antimicrobial resistance.
The pig
The breed, number, age and sex of the pigs should always be reported. Consider using mini-pigs if the growth of the pigs is a major drawback for the study or if mature pigs are essential. Report housing conditions. The inoculation route/procedure should reflect the natural pathogenesis which might demand anaesthesia and often also surgery.
The infected pig
Clinical signs of both local and systemic infection/response should be monitored, and it is important to report how the diseased animal is handled; for example, analgesic treatment, humane endpoints, any special requirements relevant for the disease. The time frame of the infection should reflect that of human infections.
Post-mortem characterization
A complete necropsy should be performed in order to disclose both local and systemic infectious lesions. Use blinded semi-quantitative or objective methods to evaluate the lesions. Include a definition of infection. Identify the inoculum and consider the formation of biofilm.
Conclusion
We have demonstrated an increase in the number of porcine models for bacterial infections in humans. Furthermore, we have also demonstrated that the models are regularly cited – that is, they have scientific impact. However, there is a substantial lack of information in most of the papers describing the models and, therefore, it was highly important to develop the present guidelines. It is important to emphasize that reporting of methods and their negative results such as ‘no signs of systemic inflammation were found post mortem’ is also valuable. Only then can one rely on, for example, that a high C-reactive protein response is caused by the local infection and not by a systemic spread of the inoculum. The reported guidelines should hopefully help authors to include all essential information in future publications as this will encourage reproducible, clear, comprehensive, accurate, concise and well-written manuscripts of porcine models for human bacterial infections. The guidelines will also help journal editors to facilitate improvement in the quality and validity of porcine bacterial infectious disease models. Recently, the ARRIVE (Animal Research: Reporting of In-Vivo Experiments) guidelines were developed to improve general design, analysis and reporting of research using animals. These guidelines are now endorsed by many scientific journals. However, the ARRIVE guidelines do not fulfil all the special requirements needed for optimal development and reporting of animal models for bacterial infectious diseases in humans. Application of the present guidelines for porcine models will build on the ARRIVE guidelines and comply with the principles of the 3Rs (replacement, reduction, refinement), as maximum information published for each model will decrease the number of unnecessary animal studies (reduction). Furthermore, reproducible, well-described and characterized animal models will have the least possible negative impact on animal welfare (refinement).
Porcine models of bacterial infections must be discriminative for the disease seen in humans. The optimal discriminative model reflects the aetiology, pathogenesis, distribution and appearance of lesions. Pathology (i.e
Supplemental Material
Supplemental material for Guidelines for porcine models of human bacterial infections
Supplemental material for Guidelines for porcine models of human bacterial infections by Louise K Jensen, Nicole L Henriksen and Henrik E Jensen in Laboratory Animals
Footnotes
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by a grant from the Danish Research Council (grant number 4005-00035B).
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
