Abstract
The mitochondrial matrix contains endogenously biotinylated proteins. These proteins can cause unexpected background signal when biotin–avidin- or biotin–streptavidin-based detection systems are used in immunocytochemistry. Here we show that this reactivity can be deliberately exploited, using a simple anti-biotin reagent, to obtain strong and highly specific labeling of mitochondria by both light and electron microscopy. The signal is substantially stronger than when either avidin or streptavidin is used to detect the endogenous biotin. These results confirm the accessibility of protein-bound endogenous biotin to exogenous probes, and localize the biotinylated enzymes to the mitochondrial matrix.
Keywords
The presence of these endogenously biotinylated proteins in mammalian cells can lead to unexpected background problems when the biotin–avidin system is used in immunocytochemistry, e.g., for detection of in situ hybridization of biotinylated probes (Varma et al. 1994) or for streptavidin–biotin immunodetection at the light microscopic level (Satoh et al. 1992).
Here we describe the optimization of a system to exploit the presence of high concentrations of biotindependent carboxylases within mitochondria to develop a simple, strong, and highly organelle-specific labeling protocol for these organelles, which is equally effective at the light or EM level.
Materials and Methods
Normal rat kidney (NRK) cells were grown as an adherent cell line on glass coverslips in tissue culture dishes in DMEM with 10% (v/v) fetal calf serum at 37C in a 5% carbon dioxide atmosphere.
For vital labeling of the mitochondria, coverslips bearing subconfluent NRK monolayers were incubated for 60 min at 37C in normal growth medium containing 5 μg/ml of freshly prepared Mitotracker Green FM (M7514; Molecular Probes, Eugene, OR). The Mitotracker Green was prepared as a 1 mg/ml stock in dimethylsulfoxide and was diluted 1:200 in growth medium immediately before use. Labeled cells were rinsed twice in PBS and fixed in methanol at −20C for 5 min. The fixed coverslips were rinsed again in PBS and mounted in Moviol (Longin et al. 1993).
For detection of biotin, NRK cells were grown, fixed, and washed as above. For biotin detection with avidin, the cover-slips were incubated in fluorescein-conjugated avidin (Vector Labs; Burlingame, CA) at a dilution of 1:100 of a 0.5 mg/ml stock in PBS at room temperature (RT) for 20 min. The coverslips were washed twice with PBS and mounted in Moviol. For streptavidin labeling the protocol was identical, but used 1:100 of a 0.5 mg/ml stock of Texas Red-conjugated streptavidin (Zymed Laboratories; San Francisco, CA).
For indirect immunodetection of biotin, coverslips were prepared as above and blocked in a solution of 0.2% (w/v) porcine gelatin in PBS for 5 min and then incubated with a monoclonal mouse anti-biotin antibody (200-002-096; Jackson ImmunoResearch Labs, West Grove, PA) diluted 1:100 in the blocking solution for 20 min at RT. After washing in PBS, the coverslips were incubated for 20 min in a 1:300 dilution of rhodamine-conjugated goat anti-mouse IgG (Cappell, distributed by Organon Teknika, Turnhout, Belgium) in blocking solution before washing in PBS and mounting in Moviol.
The tissue sections were obtained from normal rat kidney fixed in 4% (w/v) paraformaldehyde (PFA) in 250 mM HEPES buffer, pH 7.4, for 10 min on ice, followed by 50 min at RT in 8% PFA in the same buffer. After rinsing in PBS, the tissue blocks were cryoprotected in 2.1 M sucrose and frozen in liquid nitrogen. Sections were cut at approximately 100 nm thickness using a Reichert Ultracut S with the FCS attachment and collected onto gelatin-coated glass slides (for confocal microscopy) or Formvar-coated, glow-discharged EM grids (for electron microscopy). Tissue sections on glass slides were labeled using the monoclonal anti-biotin antibody exactly as described above.
All fluorescently labeled specimens were examined using a BioRad MRC 1024 confocal laser scanning microscope as described (Fricker et al. 1997). The data shown have been median-filtered with a 3 × 3 box and stretched to fill the entire grayscale range. No other image processing was used.
For electron microscopic immunocytochemistry, sections on grids were washed on droplets of PBS, blocked with 10% (v/v) fetal calf serum in PBS for 10 min, and incubated with the anti-biotin antibody at 1:10 dilution of the stock in 5% (v/v) fetal calf serum in PBS for 30 min. After washing the grids in PBS, they were floated for 20 min on drops of rabbit anti-mouse IgG (Cappell) diluted 1:50. The grids were then washed in PBS and incubated for 20 min with a 1:100 dilution of 9-nm protein A–gold stock (Slot and Geuze 1985). After washing in PBS and water, the grids were embedded in methocellulose-uranyl acetate (Tokuyasu 1980) and examined in a Leo EM 912 transmission electron microscope (Fricker et al. 1997). Images were collected at 80 keV using a Proscan 1024 × 1024 pixel cooled CCD camera.
Results and Discussion
Mitochondrial morphology in normal tissues and in tissue culture cell lines can be extremely complex. In many cell types the mitochondria form a dynamic branched network, constantly breaking and re-forming. The behavior of mitochondria in living cells can be demonstrated using vital dyes that are sequestered by active mitochondria in a membrane potential-dependent manner. These probes include rhodamine 123 (Grouselle et al. 1990) and the carbocyanine dye JC-1, which permits ratiometric estimation of mitochondrial membrane potential (Reers et al. 1991). These probes suffer from the disadvantage that their mitochondrial localization is an environment-sensitive equilibrium effect, so that staining is lost when the cells are washed or fixed. Mitotracker probes are cell permeant-specific mitochondrial probes that contain a mildly thiol-reactive chloromethyl moiety, which leads to retention of the dye by mitochondria after washing and aldehyde fixation (Poot et al. 1996).
Figure 1A shows that the Mitotracker probe Mitotracker Green FM decorates a complex, branched, reticular mitochondrial mass in methanol-fixed NRK cells. The dye gives a strong specific mitochondrial stain in living NRK cells (data not shown), and this staining pattern is not visibly altered when the cells are fixed in methanol at −20C.
Figure 1C illustrates the use of a fluorochrome-conjugated avidin to label NRK cells to which no biotinylated probe has been applied. Despite this lack of exogenous biotin, a strong labeling pattern is visible. The pattern is very reminiscent of the mitochondrial distribution in these cells (Figure 1A). Avidin is a 66-kD glycoprotein with a positive charge at physiological pH. Both its oligosaccharide moiety and its charge have been implicated in nonspecific background binding in indirect biotin-dependent detection systems (Wood and Warnke 1981).
To overcome the charge and oligosaccharide problems associated with avidin, it is possible to detect biotin with streptavidin, a nonglycosylated bacterial product of 60 kD with a near-neutral isolectric point and consequently very little net charge at physiological pH. Figure 1D illustrates the result when methanol-fixed NRK cells are incubated with a directly fluorochrome-conjugated streptavidin. Once again, no biotinylated probe has been used, and yet both the avidin and streptavidin probes are labeling similar widespread branched, tubular structures in the cytoplasm. In the case of the streptavidin there is an additional nuclear background.
The reason for this endogenous background reactivity with both avidin and streptavidin conjugates becomes apparent when an anti-biotin monoclonal antibody is used to display the endogenous biotin distribution in NRK cells (Figure 1B). A complex, branched tubular network of structures containing high levels of endogenous biotin is clearly labeled. Comparison of a specific mitochondrial probe (Figure 1A) with the anti-biotin labeling (Figure 1B) confirms that the high levels of biotinylated carboxylase enzymes within mitochondria make this organelle an excellent substrate for anti-biotin labeling. The anti-biotin labeling shown in Figure 1B is completely ablated if the monoclonal antibody is preincubated with excess free biotin, or if the specimen is preincubated with excess unconjugated streptavidin to block the endogenous biotin (data not shown), confirming that the mitochondrial staining by the monoclonal anti-biotin is indeed via endogenous biotin.
Figure 2A shows that the same anti-biotin labeling can be used to identify the mitochondria in sections of normal rat kidney that have been aldehyde-fixed and cryosectioned. Strong labeling of discrete cytoplasmic structures is seen in the cells lining the walls of the two tubules present in the section. The identification of these structures as mitochondria is confirmed by electron microscopic immunocytochemistry using immunogold labeling (Figure 2B). Prominent gold labeling is found within the mitochondria; the majority of the gold particles lie over the mitochondrial matrix. In some places an alignment of gold particles along the inner membrane and cristae is visible. These results are compatible with biochemical fractionation studies localizing these enzymes to the mitochondrial matrix compartment (Wolf 1995), and also with a report that methyl-crotonyl CoA carboxylase is associated with the inner mitochondrial membrane (Hector et al. 1980).
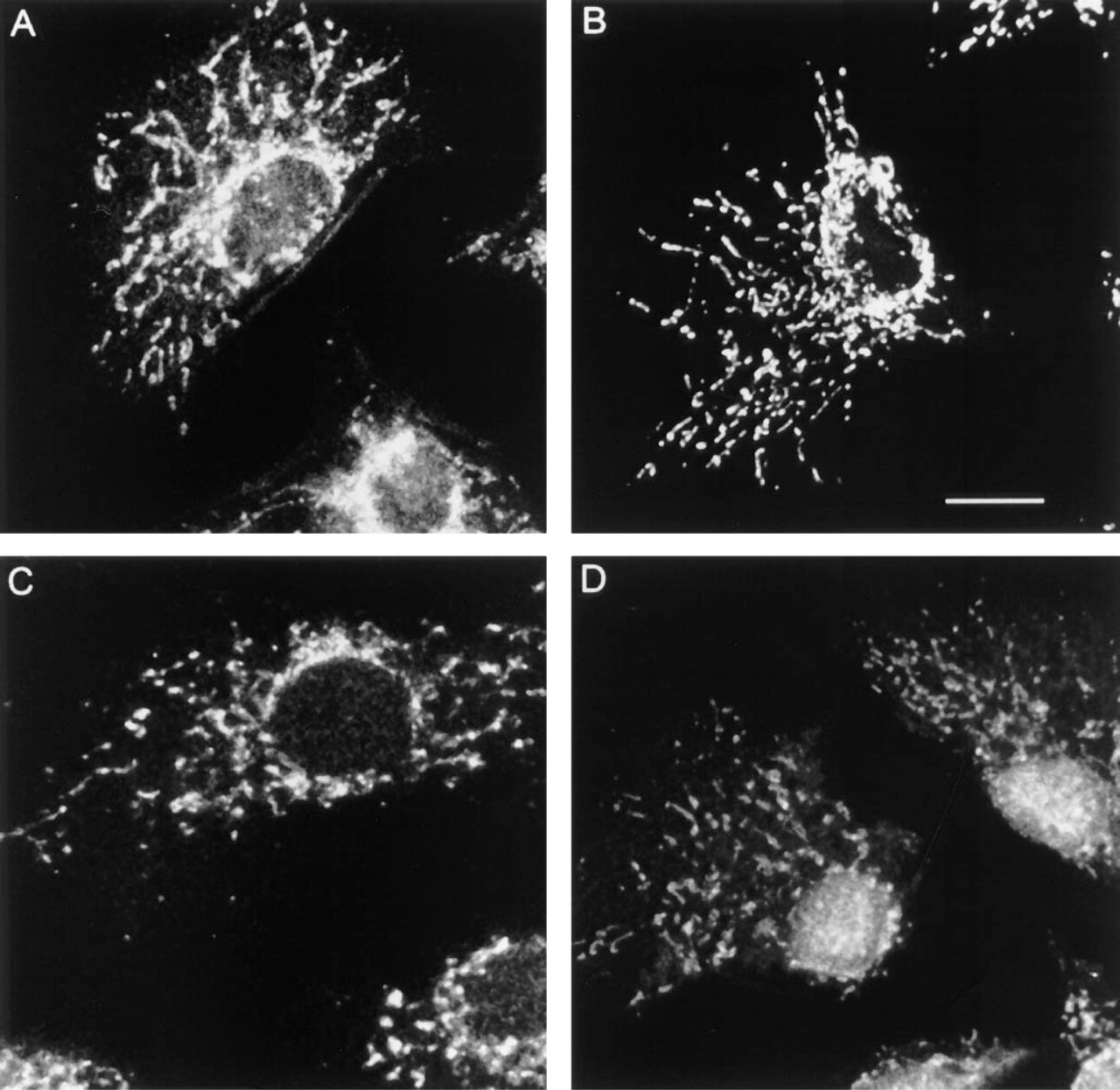
Mitochondrial labeling in NRK cells. Single confocal planes of methanol fixed/permeabilized NRK cells labeled with (
In this report we have demonstrated that avidin and streptavidin share the ability to recognize the endogenously biotinylated enzymes of the mitochondrion. This may give rise to a serious background problem and to false interpretation of the labeling produced by exogenous biotinylated probes if it is not taken into account. The problem has been recognized in nucleic acid hybridization experiments, and digoxigenin probes may be preferred over biotinylated probes for this reason (Yi et al. 1995). In an immunocytochemical study of manganese superoxide dismutase in rat stomach, Satoh et al. (1992) detected endogenous streptavidin binding reactivity only after Triton X-100 permeabilization. These authors ascribed the false-positive signal to endogenous biotin and surmised that the signal was mitochondrial, but added that EM localization of biotin would be needed to clarify this. The present study provides the EM immunocytochemical confirmation that endogenous biotin is detectable within the matrix of mitochondria, and additionally demonstrates that detection of endogneous biotin can occur under sample preparation conditions not involving Triton X-100 treatment.
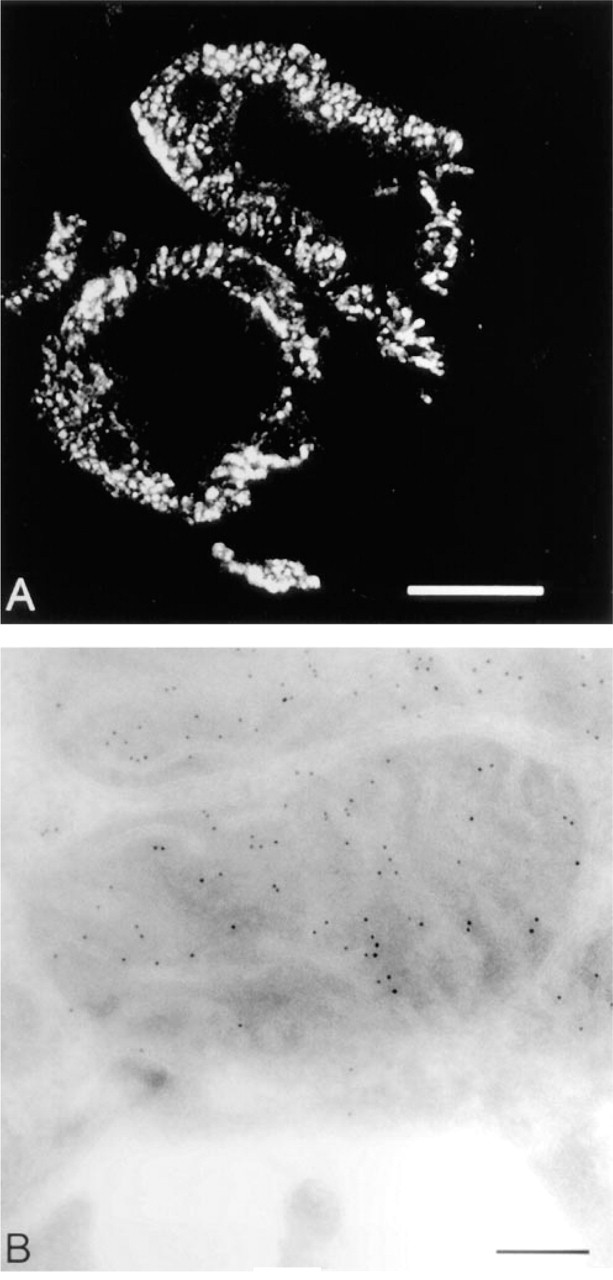
Mitochondrial labeling in tissue sections. Ultrathin cryo-sections from a single block of fixed rat kidney are shown. (
Finally, we have shown that the background caused by endogenous biotin can easily be exploited to achieve highly specific mitochondrial labeling at both the light and the electron microscopic level. In this case, optimal labeling is achieved not with avidin or streptavidin but by using a monoclonal anti-biotin antibody to detect the biotinylated mitochondrial enzymes.
Footnotes
Acknowledgements
Supported by a project grant from the Medical Research Council to DJV.
We are grateful to members of the laboratory for useful discussions and to Lance Tomlinson for excellent photographic assistance.
