Abstract
The horse is one of the few animals kept and bred for its athletic performance and is therefore an interesting model for human sports performance. The regulation of the development of equine locomotion in the first year of life, and the influence of early training on later performance, are largely unknown. The major structural protein in skeletal muscle, myosin heavy-chain (MyHC), is believed to be primarily transcriptionally controlled. To investigate the expression of the MyHC genes at the transcriptional level, we isolated cDNAs encoding the equine MyHC isoforms type 1 (slow), type 2a (fast oxidative), and type 2d/x (fast glycolytic). cDNAs encoding the 2b gene were not identified. The mRNA expression was compared to the protein expression on a fiber-to-fiber basis using in situ hybridization (non-radioactive) and immunohistochemistry. Marked differences were detected between the expression of MyHC transcripts and MyHC protein isoforms in adult equine gluteus medius muscle. Mismatches were primarily due to the presence of hybrid fibers expressing two fast (2ad) MyHC protein isoforms, but only one fast (mainly 2a) MyHC RNA isoform. This discrepancy was most likely not due to differential mRNA expression of myonuclei.
S
The MyHC proteins are believed to be primarily transcriptionally controlled. The time course and threshold stimulus needed to trigger changes at the mRNA level are therefore important aspects of gene regulation. (O'Neill et al. 1999). Changes in MyHC mRNA should precede changes in the corresponding protein products in fibers that undergo fiber-type transformation, thus leading to a mismatch in the distribution of transcripts compared to that of the corresponding protein products (Andersen and Schiaffino 1997; Jaschinski et al. 1998). By monitoring changes in expression of both MyHC mRNA and protein, a potential clue can be obtained for the direction of change in MyHC gene expression. To investigate the RNA expression of the MyHC genes, we first isolated part of the genes encoding the equine MyHC isoforms type 1, 2a, and 2d/x. The sequences of the clones were analyzed and their isoform specificity was determined. Second, our objective was to reassess myofiber typing in equine gluteus medius muscle by studying the correspondence between the MyHC proteins (IHC) and their mRNAs (in situ hybridization, ISH) to better understand the dynamic nature of fiber type determination and transition. For these analyses we used both transverse and longitudinal sections.
Materials and Methods
All chemicals were obtained from Merck (Amsterdam, The Netherlands) unless otherwise indicated.
Muscle Biopsies
Percutaneous muscle biopsies from the gluteus medius were taken according to the protocol of Lindholm and Piehl (1974). The biopsies were taken on an imaginary line drawn from the coxal tuber to the sacral tuber, at one third distance from the sacral tuber, perpendicular to the skin. They were taken as deep as possible (until resistance from the iliac wing was reached). The samples were rolled in talcum powder, mounted on cork blocks with the use of OCT embedding medium, and oriented such that the fibers could be sectioned transversely and frozen in liquid nitrogen. All samples were stored at − 80C. For the RNA isolation and subsequent cDNA isolation samples of six different horses (adult Dutch Warm Blood, age between 5 and 10 years, varying training status), five geldings and one mare were combined. For the ISH experiments, samples from an adult, untrained healthy horse (26-year-old Frisian stallion) were used. All procedures were reviewed and approved by the institutional animal care committee.
Oligonucleotides for 3′ RACE PCR
The oligonucleotides RoRi-dT17 and Ro were synthesized according to Harvey and Darlison (1991). RoRi-dT17 (5′-TCGATGGTCGACGCATGCGGATCCAAAGCTTGAATTCGAGCTTTTTTTTTTTTTTTT3′), which consists of 17 thymidine residues with an adapter sequence at the 5′ end, was used to prime first-strand cDNA synthesis. The Ro primer (5′ ATCGATGGTCGACGCATGCGGATCC3′) corresponds to the first 25 bases of RoRi-dT17 and was used in PCR together with either of the gene-specific primers FG2Exn40 (fast MyHC isoforms) (5′AGGAAGGTCCAGCACGAACTGGAGG3′) (Ennion et al. 1999), or RnExn40 (slow MyHC isoforms) (5′CGCAAGGTGCAGCACGAGCTGGAC3′) (McKoy et al. 1998) to amplify the 3′ ends of the myosin heavy chains. These gene-specific primers were designed from conserved sequences in exon 40 of the MyHC genes.
First-strand cDNA Synthesis
Total RNA was isolated using acid guanidinium thiocyanate-phenol-chloroform according to Chomczynski and Sacchi (1987), with minor modifications. RNA (4 μg) was dissolved in 7 μl of RNase-free water, heated at 70C for 10 min, and rapidly cooled on ice to eliminate secondary structure before cDNA synthesis. The cDNA synthesis reactions were assembled on ice in a 20-μl volume containing buffer (final concentration 10 mM Tris-HCl, pH 8.3, 15 mM KCl, 0.6 mM MgCl), 10 mM DTT, 0.5 mM of each nucleotide (Roche Molecular Biochemicals; East Sussex, UK), 66 μM RoRi-dT17 primer, 40 U RNase inhibitor (Promega; Southampton, UK), 4 μg total RNA, and 200 U M-MLV reverse transcriptase (GIBCO BRL; Paisley, UK) per μg RNA. Control reactions were set up as above but omitting reverse transcriptase. The reactions were incubated overnight at 37C and subsequently stored at −20C.
RACE-PCR
Double-stranded cDNA was amplified from the first-strand cDNA reaction in a PCR using either FG2Exn40 or RnExn40 primer. Reactions were performed in a 100-μl volume containing reaction buffer (final concentrations 10 mM Tris-HCl, pH 8.3, 50 mM KCl, 1.5 mM MgCl2), 0.2 mM each of dNTPs (Roche Molecular Biochemicals; East Sussex, UK), 25 μM each Ro and exon 40 primers, and 10 μl first-strand cDNA. Using a “hot start” 2.5 U of Taq polymerase (Roche Molecular Biochemicals) was added. Amplification was then carried out by 32 cycles of denaturation at 94C for 1 min followed by annealing at 55C for 1 min, followed by elongation at 72C for 2 min. A final elongation step at 72C for 10 min was included to ensure that all PCR products had 3′ A overhangs. PCR products were analyzed by gel electrophoresis on 1% agarose gels. PCR products were excised and purified using Wizard PCR purification kit (Promega) and subcloned into the pGEM-T vector (Promega) or the TpCRII vector (Invitrogen; Breda, The Netherlands) using the T-A cloning method. DNA sequencing was performed by the chain termination method (Sanger et al. 1977). Sequence analyses were performed using BLAST searches on the EMBL database. The alignment of sequences and the creation of the dendrogram in Figure 4 were created using the multiple alignment program DNAMAN (version 2.51, Lynnon BioSoft). The sequences are submitted to GenBank under the accession numbers AY187001, AY187002, and AY187003 for type 1, 2a, and 2d/x, respectively.
In Situ Hybridization
Transverse serial sections (10 μm) were made with a cryostat at −20C, placed on Superfrost Plus slides (Menzel; Merck, Amsterdam, The Netherlands), dried for 1 hr, fixed for 20 min with 4% paraformaldehyde in 1 × PBS, pH 7.4 (Zeller and Rogers 1989), washed in 1 × PBS, dehydrated, and stored at −80C. For the ISH experiments the slides were returned to room temperature, rehydrated, and rinsed in 1 × PBS. After digestion with 20 μg/ml of proteinase K at 37C for 3 min, acetylation with 0.25% acetic anhydride in 0.1 M triethanolamine for 10 min was done according to Zeller and Rogers (1989). After several rinses in 1 × PBS and refixation for 2 min in 4% paraformaldehyde followed by several washes in 1 × PBS, the sections were completely dehydrated before hybridization. Procedures for hybridization with digoxigenin-labeled riboprobes were adapted from Horton et al. (2001), with some modifications.
The cDNAs included MyHC 1 (276 bp) in pGEM-T, 2a (278 bp) in pGEM-T, and 2d/x (282 bp) in pCRII-TOPO. Riboprobes were synthesized with digoxigenin-labeled UTP in antisense direction by in vitro transcription from the linearized cDNA templates, according to the manufacturer's guidelines (Roche Molecular Biochemicals; Almere, The Netherlands) and purified by a Qiagen RNeasy kit (Westburg; Leusden, The Netherlands). The riboprobes (500 ng/ ml final concentration) were suspended in 40% (deionized) formamide, 1 × SSC, 10% dextran sulfate, 1 × Denhardt solution [0.02% Ficoll, 0.02% polyvynilpyrrolidine, 0.02% bovine serum albumin (BSA)], 0.67 M NaCl, 0.1 μg/μl yeast tRNA, and 0.1 μg/μl herring sperm DNA, heated at 80C for 5 min. Prehybridization (45 μl) was performed for 30 min. Approximately 30 μl of probe was used per slide, overlaid with a coverslip. Hybridization was performed overnight at 45C in a humidified In Slide Out incubator (Boekel Scientific; Merck, Amsterdam, The Netherlands). Coverslips were removed by rinsing in 6 × SSC at 60C, followed by two high-stringency steps at 60C for 20 min in 0.5 × SSC and 20% formamide and two rinses in 2 × SSC at room temperature. Unhybridized probe was digested with 2 μg/ml RNase A in 0.5 M NaCl, 10 mM Tris-HCl, pH 8.0, at 37C for 30 min, followed by five washes in 2 × SSC at room temperature and another high-stringency wash for 10 min. The sections were rinsed twice with 2 × SSC and maleic buffer (0.1 M maleic acid, 0.15 M NaCl, pH 7.5). Methods for the detection of hybridized probes were adapted from the manufacturer's protocols (Roche Molecular Biochemicals). Tissue sections were blocked twice for 15 min and once for 1 hr with 5% inactivated BSA in maleic buffer at RT. Blocking was replaced with sheep anti-digoxigenin Fab–alkaline phosphatase conjugate (1:2000 in 1% BSA in maleic buffer) overnight at RT. After several rinses with the same buffer, sections were washed twice with 0.1 M NaCl, 0.1 M Tris-HCl, pH 9.5, 50 mM MgCl2, 0.1% Tween-20 at RT. Alkaline phosphatase activity was visualized by incubation with 0.18 mg/ml BCIP, 0.34 mg/ml NBT, in the buffer described above. The staining was allowed to develop for approximately 16 hr at RT, rinsed with distilled water, and embedded in Aquamount. Control sense probes were negative (results not shown).
Immunohistochemical Staining
The monoclonal antibodies (MAbs) used were previously shown to crossreact with horse myosins (Dingboom et al. 2002). MAb 219 reacts with type 1. MAb 333 reacts with type 2a, and MAb 412 reacts with type 1 and 2d/x (Bredman et al. 1991; Wessels et al. 1991). Transverse serial sections (10 μm) were obtained as described above. To perform the IHC the protocol was followed as described previously (Dingboom et al. 2002). Briefly, the slides were incubated in Pronase (0.1 mg/ml in PBS) for 30 min (omitted for MAb 412), rinsed in PBS, blocked in Teng-T (10 mM Tris, 5 mM EDTA, 0.15 M NaCl, 0.25% gelatin, and 0.05% Tween-20, pH 8.0) for at least 15 min, followed by rinsing in PBS and incubation overnight at RT with the MAbs at a dilution of 1:10 (333) or 1:25 (219, 412) in PBS. The slides were next rinsed with PBS and incubated for 90 min with a biotiny-lated horse anti-mouse polyclonal antibody (1:100 in PBS) (ABC peroxidase staining kit Elite (Vector Labs; Burlin-game, CA), rinsed in PBS, and incubated for 90 min with the components avidin (A; 1:100 in PBS) and biotin (B; 1:100 in PBS) of the ABC staining kit. After rinsing, the immunoreaction was visualized by incubation with 0.05% 3,3′-diaminobenzidine tetrachloride in 30 mM imidazole and 0.09% H2O2. The slides were then embedded in DePeX. This protocol was followed for the MAbs 219 and 333. For MAb 412, the slides were rinsed in 0.1 M acetate buffer (pH 6.0; 5-10 min) before visualization and di-ammonium nickel sulfate (2.5% in acetate buffer) instead of imidazole was used.
Analyses
A group of 296 contiguous fibers was used for fiber typing and calculation of fiber type composition. The muscle fibers were classified into type 1, type 2a, or type 2d/x, or type 2ad on the basis of their reactions with the different in situ probes and the MAbs.
Results
Isolation of Equine MyHC Isoforms
Sequence analysis of the isolated cDNA clones revealed that three different isoforms had been isolated. Identification of the isoforms was done by performing a BLAST search against all MyHC genes known in combination with the comparisons shown in this article. From these analyses, we could conclude that the isolated clones were the slow MyHC (type 1) and the two fast MyHC [type 2a and type 2d/x (called in the Figures 2d)] known in horse. We did not identify cDNAs encoding the 2b gene. The complete sequences of the different isoforms were aligned as depicted in Figure 1. The two fast isoforms isolated showed higher homology relative to each other (79%) than to type 1 (type 1 vs type 2a 61% homology; type 1 vs 2d/x 62% homology). Figure 2 depicts the alignment of the nucleotide and deduced amino acid sequence of the last 15 amino acids of the coding regions of the isolated horse clones. A comparison was performed with known sequences of human, mouse, rabbit, rat, bovine, and pig. From this comparison, it can be concluded that the type 2 isoforms show more homology to other type 2 isoforms than to the type 1 isoform. Of the 15 amino acids presented, six showed consistent differences between the type 1 and type 2 isoforms. These are the first A/V, the fifth D/E, the sixth I/V, the seventh G/H, the eleventh L/I, the twelfth N/S, resulting in a slight (2) charge difference. Lefaucheur et al. (1998) stated that in the pig the isoleucine at position 10 is characteristic for type 2d/x isoforms. These authors compared pig, rat, and human sequences. In Figure 2 we have included sequences from other species, showing variability in the identity of the tenth amino acid (valine or isoleucine) which therefore appears not to be isoform-specific. This clearly indicates the advantages of the availability of sequences of many species.

Nucleotide sequence alignment of the isolated horse MyHC genes. The stop codon is indicated in bold. The vertical line indicates the first nucleotide used in the comparison depicted in Figure 2. Identical nucleotide sequences relative to the horse type I (HORSE 1) sequence are indicated by dashes. Asterisks in the sequence of the type 2d/x (HORSE 2d) gene indicate identical nucleotide sequences relative to the type 2a (HORSE 2a) gene. Dots represent missing nucleotides.
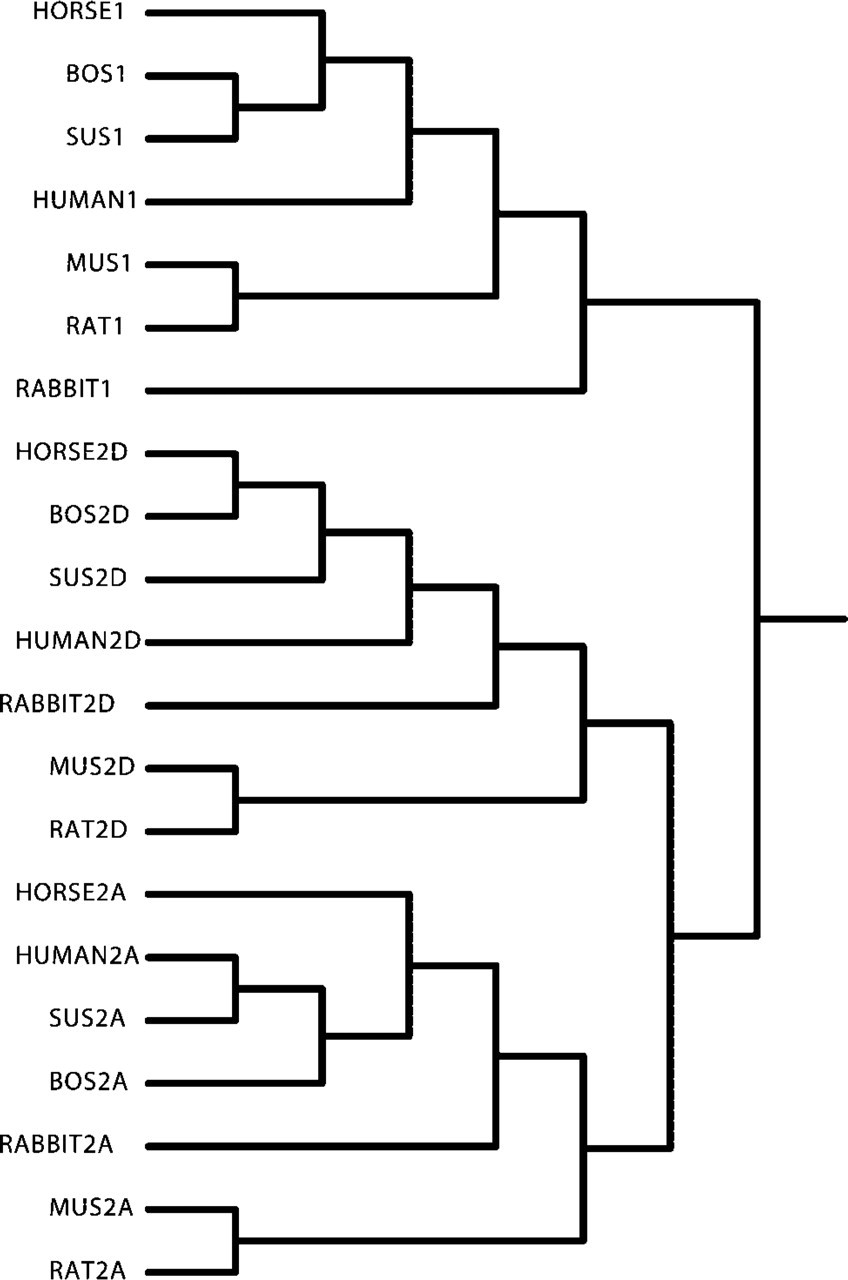
Figure 3 shows the nucleotide sequence alignment of the horse MyHC genes 3′ untranslated region (UTR) with sequences of the same species as in Figure 2. Interestingly, it can be seen that between species the sequence identity of the 3′UTR is much higher for the type 2 MyHC isoforms, suggesting that this region, as previously suggested, is either under stringent evolutionary constraints or that they have diverged much later than the type 1 isoform. There were no clear distinctions possible between a 5′ and 3′ part as previously suggested from the pig sequence (Lefaucheur et al. 1998). Figure 4 depicts the clustering relationship of the same genes as shown in Figures 2 and 3. The complete isolated sequences of the horse genes (Figure 1, coding and 3′UTR) and comparable regions of the other genes were used for construction of the dendrogram. Each MyHC isoform forms its own cluster, demonstrating again that, for each isoform, this part is more similar to the MyHC counterpart in the other mammalian species than to the other MyHC isoforms within a species. This indicates that this part of the genes is isoform-specific and confirms the validity of using this region for isoform-specific probes. Clearly, the type 1 and type 2 genes form separate clusters, again pointing at a later diversion of the type 2 isoforms compared to the type 1/type 2 diversion. As can be seen in Figure 2, the coding region is quite homologous, especially among the type 2 isoforms, emphasizing the need for 3′UTR sequences to produce probes that are specific enough to identify only a single MyHC isoform.
Identification of Myofiber Types by ISH and IHC
The complete equine MyHC sequences depicted in Figure 1 were used to generate isoform specific non-radioactive RNA probes. Because most ISH studies used radioactive probes and the equine muscle was to our knowledge not investigated by this type of analysis, we had to optimize the non-radioactive protocol for our experiments based on protocols for embryonic mouse and adult human muscle (Franco et al. 2000; Horton et al. 2001). Figure 5 (left panels) shows a panel of serial sections from the equine gluteus medius muscle processed for ISH with RNA probes specific for each MyHC isoform indicated. Fibers 1, 2, and 3 expressed exclusively MyHC type 1, 2a, and 2d/x, respectively, showing that the RNA probes were highly specific for each MyHC isoform (Figure 5). An analysis carried out on 296 fibers of the gluteus medius muscle showed that very few co-expressed two MyHC transcripts. The only co-expression detected was in 1.7% of the fibers, expressing mRNA of the 2a and 2d/x isoforms (type 2ad) (Table 1).

Nucleotide and amino acid sequence alignment of the 3′ end of the coding region of the equine (horse) MyHC genes with known sequences of human (human), mouse (mus), rabbit (rabbit), rat (rat), bovine (bos), and pig (sus). Known sequences of the type 2b gene are included for completion. Stop codons are labeled with an asterisk. Type 2x/d is called type 2d in all the figures for clarity. The human type 1, type 2a, type 2d/x, and type 2b sequences were obtained from the EMBL under the accession numbers M58018, S73840, AF111785 and AF111783. The mouse type 1, type 2a, type 2d/x, and type 2b sequences were obtained from the EMBL under the accession numbers AJ223362, AJ002521, AJ002522 and AJ223361. The rabbit type 1, type 2a, type 2d/x, and type 2b sequences were obtained from the EMBL under the accession numbers Z34886, AF062351, U32574 and X05958. The rat type 1, type 2a, type 2d/x, and type 2b sequences were obtained from the EMBL under the accession numbers X15939, L13606, X72591 and X72590. The bovine type 1, type 2a and type 2d/x sequences were obtained from the EMBL under the accession numbers AB059400, AB059398 and AB059399. The pig type 1, type 2a, type 2d/x and type 2b sequences were obtained from the EMBL under the accession numbers U75316, AB025260, AB025262 (no 3′ trailer sequences included), and AB025261.
Using serial sections we compared the MyHC mRNA content of the fibers (Figure 5, left) with their MyHC protein composition, determined by IHC (Figure 5, right). Fibers 1, 2, and 3 exhibit specific staining for type 1, 2a, and 2d/x, MyHC protein, respectively. Clearly, the staining patterns of the ISH and IHC experiments show a high correlation. We analyzed the same 296 fibers used for the mRNA fiber typing again for the protein fiber typing. On the protein level more (21.4%) hybrid fibers were detected, and they all expressed the type 2a and type 2d/x isoform (type 2ad; Table 1). Of all fibers analyzed, 76.5% had the same RNA and protein expression; the rest showed a mismatch (Table 1). Most of these mismatches concerned fibers expressing type 2ad protein. An example is fiber no. 5 expressing type 2a RNA but type 2ad protein. Of the type 2ad fibers at the protein level, 19.7% had only type 2a RNA, 1.4% had only type 2d/x RNA, and 0.3% (one fiber) had type 1 RNA (Table 1). On the other hand, fiber no. 4 in Figure 5 expressed type 2ad at the RNA level but only type 2d/x protein. This is a typical result because type 2ad RNA expression was never matched by type 2ad protein expression (Table 1).
Discussion
We have isolated and characterized for the first time part of the genes encoding the equine MyHC type 1, 2a, and 2d/x. Isolation was performed based on conserved sequences in exon 40 of the fast and slow isoforms. We did not isolate any cDNA fragment encoding the 2b gene, making it unlikely that this gene is expressed in the gluteus medius of the horse. This finding is in line with previous publications on the expression of MyHC protein in equine skeletal muscle (Rivero et al. 1996,1999; Dingboom et al. 2002; Karlstrom and Essen–Gustavsson 2002), and equals the situation in humans, ruminants, and carnivores (Talmadge et al. 1996; Andersen and Schiaffino 1997; Tanabe et al. 1998; Arguello et al. 2001). In the past, it was believed that the expression of the 2b gene depended on body size. Larger animals were supposed to be unable to cope with the very fast 2b isoform (Rome et al. 1990; Smerdu et al. 1994; Tanabe et al. 1998; Rivero et al. 1999; Pette and Staron 2000). However, the 2b gene is expressed in the pig and the llama (Graziotti et al. 2001; da Costa et al. 2002; Lefaucheur et al. 2002). It has been suggested that the expression of 2b in pig skeletal muscle is due to rigorous selection for fast-growing muscle (Lefaucheur et al. 2002). The llama is also an animal that has been selected for meat quality (Graziotti et al. 2001). Horses have been domesticated for a long time but were mostly selected for different purposes (coordination, stamina). This has resulted in some differences in fiber type composition among breeds, but thus far not in the appearance of type 2b in equine skeletal muscle (Snow and Guy 1980; Rivero et al. 1995; Barrey et al. 1999). In humans, the MyHC 2b gene is transcribed in the masseter muscle and occasionally in the abdominal external oblique (Horton et al. 2001). However, no 2b protein was found in these muscles. Andersen et al. (2002) detected 2b RNA and protein in the human extra-ocular and laryngeal muscles, muscles known to be under extremely fine control. To get more insight into the expression and regulation of the MyHC 2b isoform in the horse we are planning to isolate and characterize this gene from the extra-ocular or laryngeal muscles.

Nucleotide sequence alignment of the horse MyHC genes 3′ UTR with comparable regions of the same genes as in Figure 2. The polyadenylation signals are underlined. Identical nucleotide sequences relative to the horse MyHC sequence are indicated by dashes. Dots represent missing nucleotides. The trailers of the type 2a and type 2d/x genes showed, on interspecies comparison, a higher level of homology than the trailer of the type 1 gene.
The isolated MyHC genes presented in this article cluster together with their expected counterparts when a relationship analysis is performed (Figure 2). Comparing the last 15 amino acids of the coding region reveals some characteristic differences between the type 1 and the type 2 isoform (Figure 2). In the type 2 isoforms, such differences do not appear to exist, although not many sequences are known, especially not of the 2b gene. Shoemaker et al. (1999) identified a potential localization signal in the 3′ trailer of the type 2 genes. The 2d/x localization sequence (TCTTTGTCACT) is present in all MyHC type 2d/x trailers here presented. However, not all of the MyHC type 2a trailers and only one type 1 (rat) trailer contained the suggested 1 and 2a localization sequence (TGTGAAACCTT), making it less likely to be a true localization signal. Obviously, the addition of more known sequences of other species is valuable when DNA sequences are analyzed in trying to identify potentially important sequences.
This article presents for the first time the application of a non-radioactive ISH procedure on skeletal muscle of a large animal. The technique allows comparison of the expression of MyHC isoform transcripts and protein on a fiber-to-fiber basis, using serial sections. Analyses of a large number of fibers (around 300) per biopsy are possible, an important advantage over single-fiber analysis.
The majority (76.5%) of the fibers analyzed expressed the same MyHC isoform on the RNA and protein levels. Mismatches were due to the occurrence of hybrid fibers. At the transcriptional level, hardly any hybrid fibers were detected, whereas at the translational level a substantial number of hybrid fibers were seen. Recently, the same type of result was presented in the skeletal muscle of the pig (Lefaucheur et al. 2002). Hybrid fibers are usually interpreted as transitory fibers (Pette and Staron 1997). However, because of the regular finding of substantial numbers of especially MyHC type 2ad protein hybrid fibers, these fibers are now more regarded as a “true” type, possibly as a functional intermediate between type 2a and type 2d/x fibers (Talmadge et al. 1996; Andersen and Schiaffino 1997; Lefaucheur et al. 1998; Andersen et al. 1999; Linnane et al. 1999; Rivero et al. 1999; Stephenson 2001; Dingboom et al. 2002). Because at the transcriptional level hardly any hybrid fibers are detected, no functional mRNA intermediate is apparently necessary between the type 2a and type 2d/x fibers. Fibers can change the expression of MyHC RNA in the course of days, instead of weeks for the protein [half-life RNA 2-3 days, protein 2-3 weeks (Andersen and Schiaffino 1997)], resulting in enough plasticity to respond to changing circumstances, possibly abolishing the need for mRNA hybrid fibers in normal skeletal muscles (Smerdu and Erzen 2001).
Fibers that contained two MyHCs at the protein level, but only one transcript, could be fibers that are converting to the type corresponding to the expressed mRNA (Serrano et al. 2001; Smerdu and Erzen 2001; Lefaucheur et al. 2002). An alternative hypothesis for the occasional appearance of a protein without its mRNA is that the entire fiber in question contains mRNA for more than one MyHC isoform but that each nucleus expresses only a single mRNA or expresses it in pulses as was demonstrated by Newlands et al. (1998). In this case, transverse sections would give a limited sampling of mRNA expression in a muscle fiber. However, in a partly longitudinal section stained for type 1 mRNA (Figure 6), no differences in RNA expression level were seen along a stretch of fiber (approximately 250 μm) containing multiple myonuclei (distance between myonuclei approximately 20 μm) (Murphy 1998) (Figure 6). The intensity of the signal seems to be higher at the periphery of the fiber, which is in accordance with previously described localization patterns (Russell et al. 1992). Also in human muscle, mismatch between MyHC mRNA and protein or MyHC-ATPase reactivity could not be attributed to variations in myosin composition along the length of the fiber because the staining of fibers did not change after extended analysis of serial sections (Andersen and Schiaffino 1997). Single-fiber analysis has shown that the MyHC expression can vary along the length of the fiber, but over distances in the millimeter range, much larger than the distances between the serial sections presented here (Peuker and Pette 1997; Conjard et al. 1998).
In conclusion, we have isolated and characterized part of the genes encoding the equine MyHC type 1, 2a, and 2d/x isoforms. The type 2b isoform was not identified. The isolated genes were used to generate isoform-specific probes for non-radioactive ISH experiments using a biopsy from the equine gluteus medius muscle. We compared the expression on the RNA level with the expression on the protein level on a fiber-to-fiber basis. Discrepancies were found that were not due to differences in expression between adjacent myonuclei. Co-expression of MyHCs was more common at the protein level than at the mRNA level and was mostly observed for 2a and 2d/x MyHCs, suggesting a fine tuning of these two genes and a strong influence of their expression on myofiber plasticity. The present study also shows that only the combination of IHC with ISH allows a clear understanding of the dynamic process involved in fiber type transitions by giving a potential clue to the direction of change in MyHC gene expression. Finally, because MyHC is the most abundant protein in muscle and because of the influence of its polymorphism on contractile, metabolic, and size properties of myofibers, further research is now possible to establish and understand the importance of MyHC transcriptional and translational polymorphism in growing and exercising horses.
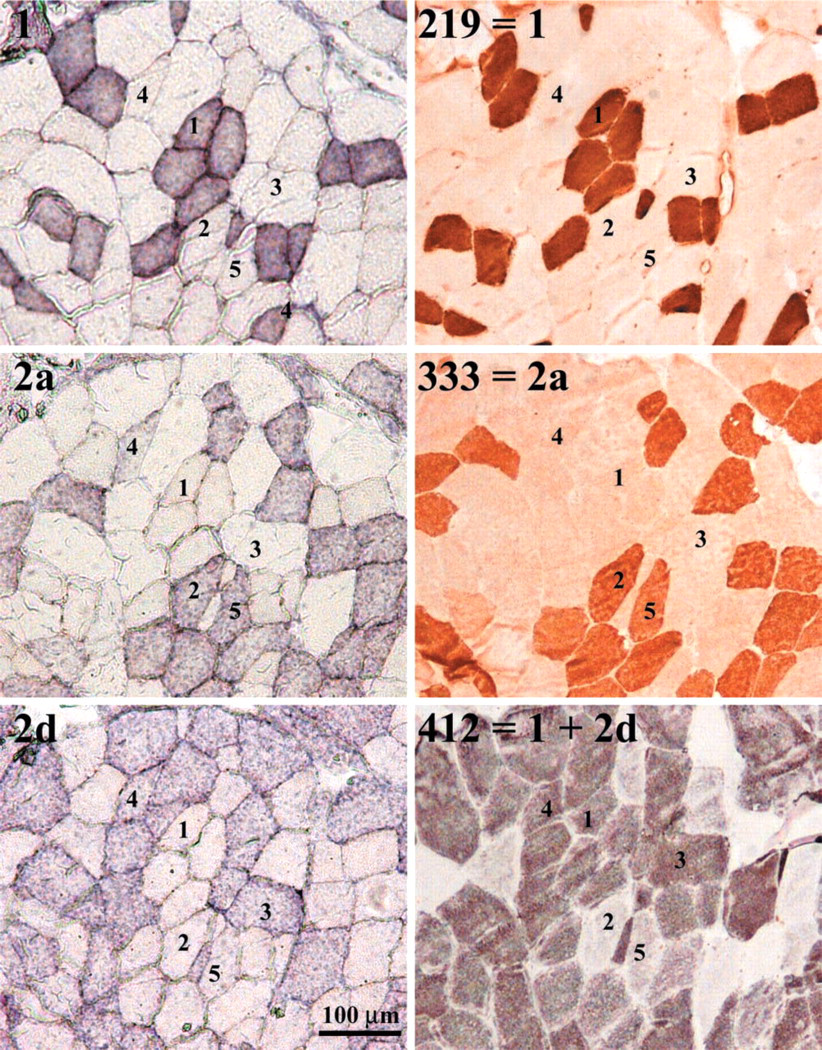
Identification of myofiber types by ISH using non-radioactive MyHC RNA probes (left panels) and immunohistochemistry (right panels) in serial sections. The specificity of the probes (left panels) and antibodies (right panels) are indicated. Numbers indicate fibers with typical expression patterns: (1) MyHC type 1 RNA and protein; (2) MyHC type 2a RNA and protein; (3) MyHC type 2d/x RNA and protein; (4) MyHC type 2ad RNA and 2d protein; (5) MyHC type 2a RNA and 2ad protein.
Identification of MyHC fiber types (%) in the equine gluteus medius muscle as assessed by ISH (RNA) and IHC (protein)a
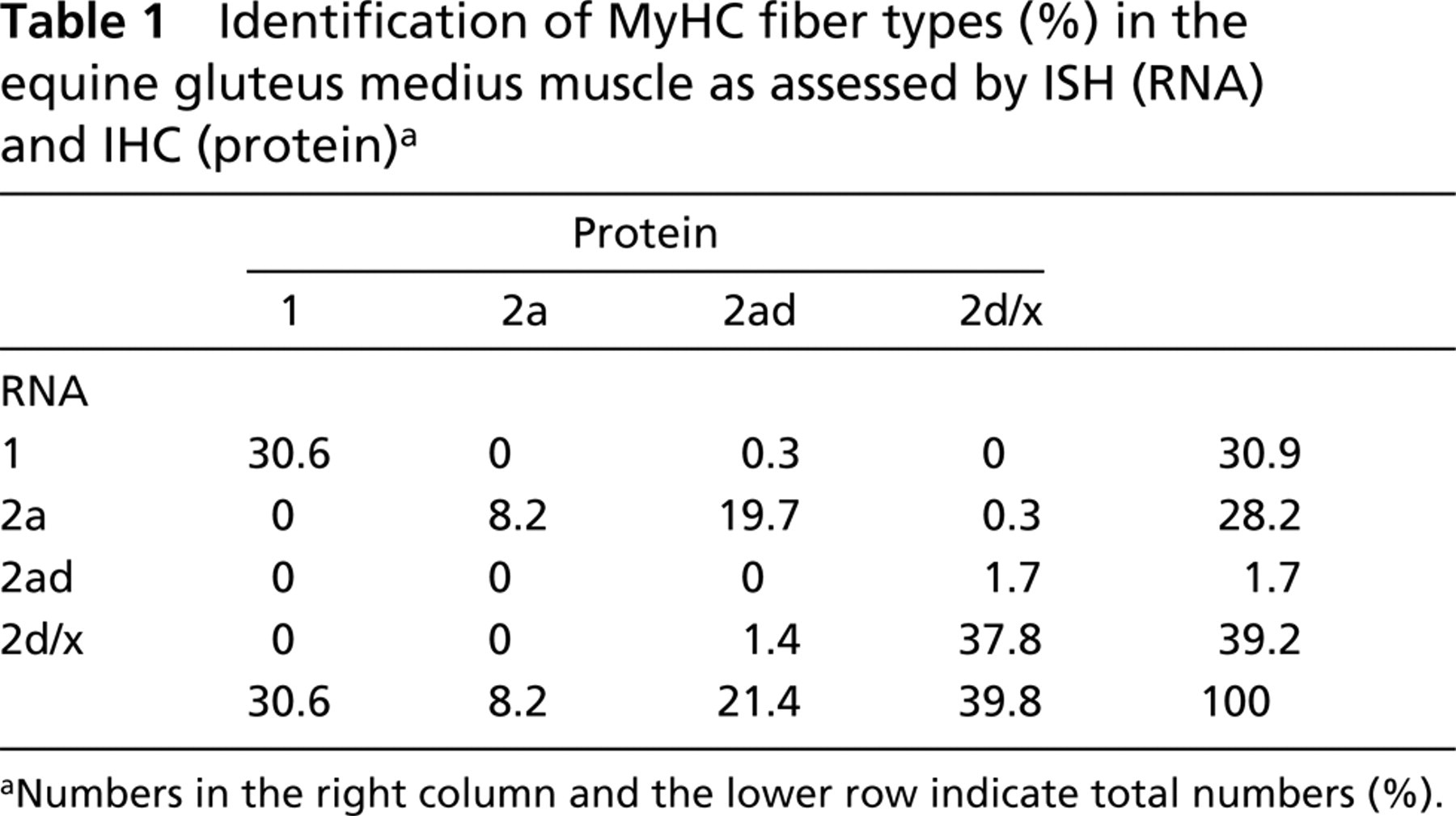
Numbers in the right column and the lower row indicate total numbers (%).
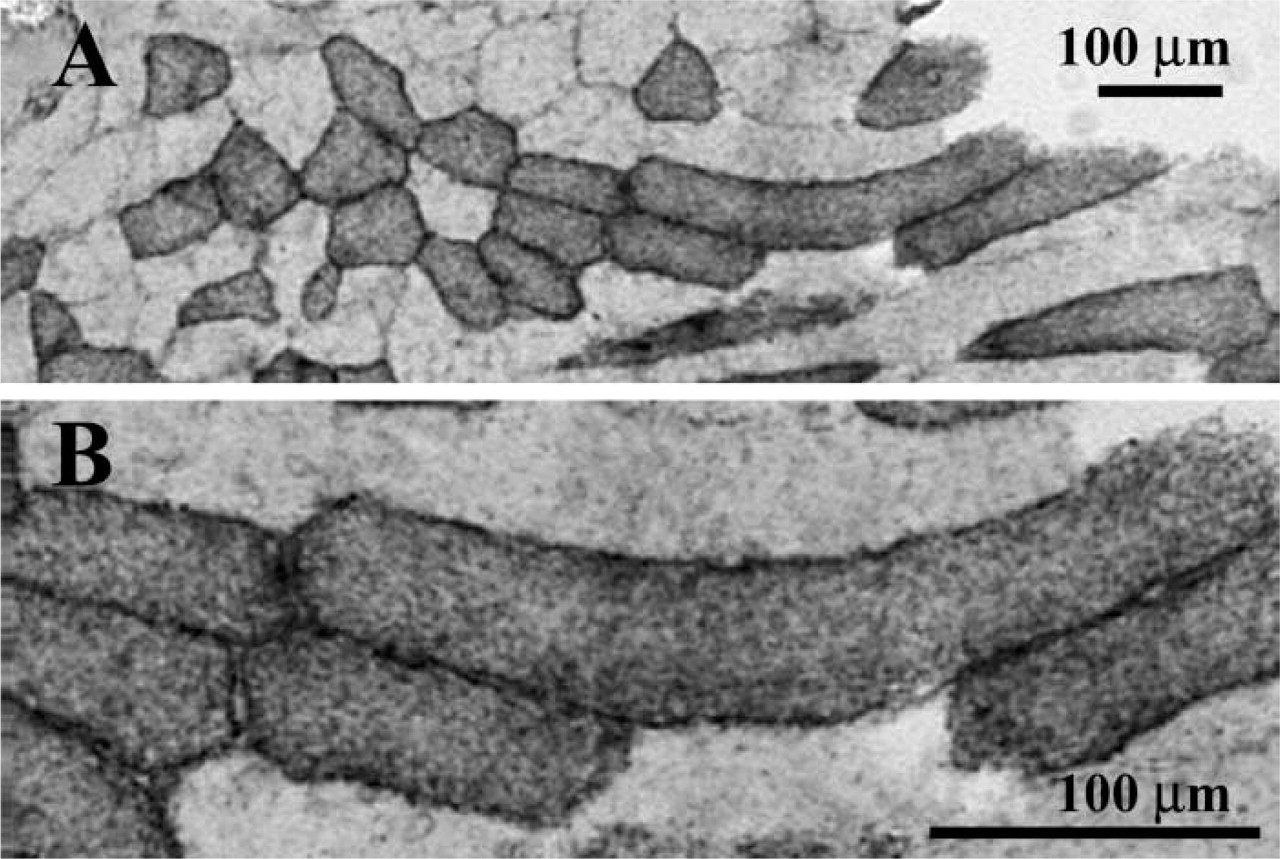
Part of a section stained with an MyHC type 1 RNA probe. (
Footnotes
Acknowledgements
We thank Dr A.F.M. Moorman and J.A.M. Korfage (University of Amsterdam) for the generous gift of the monoclonal antibodies. We are grateful for the skillful technical assistance of Miriam van der Belt and Ellen van der Wiel. Dr M. Horton (University of Pittsburgh) was invaluable for the development of the in situ hybridization protocol.
