Abstract
The biological actions of interleukin-6 (IL-6), leukemia inhibitory factor (LIF), and ciliary neurotrophic factor (CNTF) are mediated via respective functional receptor complexes consisting of a common signal-transducing component, gp130, and other specific receptor components, IL-6 receptor α (IL-6R), LIF receptor β (LIFR), and CNTF receptor α (CNTFR). IL-6, LIF, and CNTF are implicated in skeletal muscle regeneration. However, the cell populations that express these receptor components in regenerating muscles are unknown. Using in situ hybridization histochemistry, we examined spatiotemporal expression patterns of gp130, IL-6R, LIFR, and CNTFR mRNAs in regenerating muscles after muscle contusion. At the early stages of regeneration (from 3 hr to Day 2 post contusion), significant signals for gp130 and LIFR mRNAs were detected in myonuclei and/or nuclei of muscle precursor cells (mpcs) and in mononuclear cells located in extracellular spaces between myofibers after muscle contusion, but IL-6R mRNA was expressed only in mononuclear cells. At Day 7 post contusion, signals for gp130, LIFR, and IL-6R mRNAs were not detected in newly formed myotubes, whereas the CNTFR mRNA level was upregulated in myotubes. These findings suggest that the upregulation of receptor subunits in distinct cell populations plays an important role in the effective regeneration of both myofibers and motor neurons.
Keywords
S
A line of evidence suggests that interleukin-6 (IL-6), leukemia inhibitory factor (LIF), and ciliary neurotrophic factor (CNTF), which are members of the IL-6 family of cytokines consisting of IL-6, LIF, CNTF, interleukin-11 (IL-11), oncostatin M (OM), and cardiotrophin-1 (CT-1), are implicated in the degenerative and regenerative reactions of myofibers. Proteolysis of skeletal muscle proteins can be initiated by IL-6 in vitro and in vivo (Goodman 1994; Ebisu et al. 1995). Transgenic mice that overexpress IL-6 show enhanced muscle catabolism due to IL-6-induced activation of the lysosomal cathepsin (B and L) system (Tsujinaka et al. 1995, 1996). LIF enhances myoblast proliferation in vitro, but myotubes do not respond to LIF stimulation (Austin et al. 1992; Bower et al. 1995). Administration of LIF to regenerating muscles in vivo accelerates muscle regeneration (Barnard et al. 1994). Very low levels of IL-6 and LIF mRNAs are detected in normal adult muscles (Brown et al. 1994) and, in response to muscle injury, they are upregulated in both injured myofibers and mononuclear cells located at the site of the muscle injury (Kurek et al. 1996, 1997; Kami and Senba 1998). Protein and mRNA of CNTF are abundantly generated in Schwann cells of adult peripheral nerves, and nerve injury increases the release of CNTF from Schwann cells (Sendtner et al. 1992; Ip et al. 1993). Exogenous application of CNTF to regenerating muscles in vivo accelerates myotube differentiation (Marques and Neto 1997).
The biological actions of IL-6, LIF, and CNTF are mediated via their respective functional receptor complexes (Kishimoto et al. 1994). The receptor complexes for the IL-6 family of cytokines utilize gp130 protein as a common signal-transducing component. IL-6 interacts with cells through a receptor complex that consists of a ligand-binding component, the IL-6 receptor α (IL-6R), and gp130. LIF receptor β (LIFR) is a transmembrane signaling subunit, and LIF signaling is mediated by the heterodimerization of LIFR and gp130. The complex of gp130 and LIFR is also utilized by other members of the IL-6 family of cytokines, such as CNTF, OM, and CT-1. The action of CNTF is mediated via a tripartite complex consisting of a ligand-binding component, CNTF receptor α (CNTFR), gp130, and LIFR. IL-6R and CNTFR also exist in soluble forms (sIL-6R and sCNTFR). IL-6 bound to sIL-6R elicits an IL-6-specific response in cells that express gp130 on their surfaces, and the complex with CNTF and sCNTFR acts on the cells that express gp130 and LIFR. There are no data showing IL-6R expression in adult skeletal muscles. On the other hand, expression of gp130, LIFR, and CNTFR is detected in intact muscles and this receptor expression is increased in denervated muscles (Ip et al. 1993; Helgren et al. 1994). Although there are many reports demonstrating that LIF, IL-6, and CNTF induce degenerative and regenerative reactions of myofibers, the cell populations that express their receptor components in injured muscles have not yet been elucidated.
In the present study, to determine target cells for LIF, IL-6, and CNTF and to further clarify the roles of these cytokines in muscle regeneration, we examined the spatiotemporal expression patterns of gp130, IL-6R, LIFR, and CNTFR mRNAs in regenerating muscles using in situ hybridization (ISH) histochemistry. Muscle regeneration is induced by muscle contusion, a model of muscle crush injury that has been used to induce degenerative and regenerative reactions in both myofibers and intramuscular nerves (Kami et al. 1993, 1995, 1999; Kami and Senba 1998). The expression of LIFR mRNA in regenerating skeletal muscles after muscle contusion was briefly described in our previous study (Kami et al. 1999). In the present study, we show in detail the expression patterns of LIFR mRNA in regenerating muscle.
Materials and Methods
Muscle Contusion and Tissue Preparation
Adult (200–250 g bw) Wistar rats were housed in individual cages and fed commercial pellets and water ad libitum. Muscle contusion was achieved without skin incision as described previously (Kami et al. 1993, 1995). Briefly, a cylinder (640 g) was dropped from a distance of 250 mm onto a plate (diameter 1.0 cm) placed on the belly of the medial gastrocnemius muscle of the right leg of a rat anesthetized with ethyl ether. Injured muscles and their uninjured contralateral counterparts were resected under pentobarbital anesthesia at 3 hr, 6 hr, 1, 2, 3, 5, or 7 days after muscle contusion (three rats per day). For ISH analysis, gastrocnemius muscles freshly dissected from anesthetized animals were frozen in liquid nitrogen, and stored at −85C until use. All experiments were carried out with the approval of the Osaka University of Health and Sport Sciences Animal Ethics Committee.
Probes
A plasmid vector containing 3.0 kbp at the XhoI site of pSP72 (provided by Dr. Kishimoto, Osaka University) was digested with Pvu II and was self-ligated. The resultant plasmid containing 500 bp of the 3′ end of cytoplasmic domain was linearized at a SacI or XhoI site for the antisense and sense gp130 probes, respectively. A plasmid vector containing ~2.5 kbp at the EcoRI site of PUC18 (provided by Dr. Kishomoto, Osaka University) was digested with Hind III. The resultant cDNA fragment of 600 bp was inserted into pBS and linearized at the XhoI or XbaI site for the antisense and sense probes for IL-6 mRNA, respectively. Characterization of the LIFR probe has been described in a previous report (Yamakuni et al. 1996). For in situ hybridization, the [35S]-dUTP-labeled single-strand cRNA antisense and sense probes synthesized from the linearized template using the Riboprobe Gemini II system (Promega; Madison, WI) had a specific activity of 1.0–2.0 × 106 cpm/μl. The labeled cRNA probes were applied to the sections at 1.0 × 106 cpm/slide.
Three oligonucleotide probes for CNTFR were complementary to nucleotides 129–164, 470–500, and 1114–1500 of the published rat CNTFR sequence (Ip et al. 1993). The oligonucleotide probe for myogenin was complementary to nucleotides 56–100 of the published rat myogenin sequences (Wright et al. 1989). The specificity of these oligonucleotide probes was verified by sequence comparison using computer analysis and “DNASIS” DNA homology search (Hitachi; Tokyo, Japan). No significant homology with any other genes in primates or rodents was found. These oligonucleotide probes, labeled by the 3′-end-labeling method using [35S]-dATP and terminal deoxynucleotidyl transferase, had a specific activity of 1.0–3.0 × 105 cpm/μl. The labeled oligonucleotide probes were applied to sections at 0.5 × 106 cpm/slide.
In Situ Hybridization
Frozen cross-sections of gastrocnemius muscle 12 μm thick were thawed on 3-amino propylethoxysilane-coated slides, then hybridized with radiolabeled probes, as follows. Sections were fixed with 4% paraformaldehyde in 0.1 M phosphate buffer (pH 7.4) for 15 min, then rinsed in 2 × SSC (0.3 M NaCl in 30 mM Na citrate). The sections were incubated with 1 μg/ml proteinase K (Sigma Chemical; St Louis, MO) at 37C for 5 min. After two rinses in 2 × SSC, the sections were dehydrated in 70, 80, 90, 95, and 100% ethanol, then air-dried. The sections were then incubated with hybridization buffer [50% formamide, 4 × SSC, 0.12 M phosphate buffer, pH 7.4, 1 × Denhardt's solution, 0.2% SDS, 0.25 mg/ml tRNA, 10% dextran sulfate, 100 mM dithiothreitol (DTT)] containing the radiolabeled antisense oligonucleotide probes at 37C for 16 hr to detect the signals of the myogenin and CNTFR mRNAs. After hybridization, the sections were washed with three changes of 1 × SSC at 55C for 20 min each. Other series of sections were incubated with hybridization buffer containing the radiolabeled anti-sense or sense cRNA probes at 52C for 16 hr to detect the gp130, IL-6R, and LIFR mRNA signals. After hybridization, the sections were washed at 65C for 30 min with 50% formamide in 2 × SSC and 10 mM DTT. After incubation with RNase buffer (0.4 M NaCl, 10 mM Tris-HCl, pH 8.0, 50 mM EDTA-2Na) at 37C for 10 min, the sections were digested at 37C for 30 min with 25 μg/ml ribonuclease A (Sigma Chemical) in RNase buffer, then washed in RNase buffer for 10 min, 2 × SSC for 15 min, and 0.1 × SSC for 15 min at 37C, followed by dehydration in 80, 90, 95, and 100% ethanol, and then air-dried. To visualize the signals for the specific mRNAs, the sections were dipped in Ilford K15 autoradiography emulsion diluted 2:3 with distilled water at 45C. After exposure at 4C for 4 weeks, they were developed in Kodak D19 for 4 min at 20C, then fixed in 24% sodium thiosulfate solution for 5 min. Lastly, the sections were stained with hematoxylin and eosin.
Semiquantitative analysis was performed to make a general survey of the expression of gp130, IL-6R, LIFR, and CNTFR mRNAs in various regions of rat gastrocnemius muscle after muscle contusion. The relative signal level in each hybridized section was inspected and graded visually. Upregulated hybridization signal was designated “+” and background hybridization signal “–”.
Specificity of Hybridization Signals
To ascertain the specificity of the observed hybridization signals for gp130, IL-6R, and LIFR mRNAs, adjacent serial sections were hybridized with antisense or sense probes for each mRNA. Sense probes did not display specific hybridization signals in cells in regenerating muscles, whereas anti-sense probes detected abundant hybridization signals for gp130, IL-6R, and LIFR mRNAs (data not shown). In addition to the specificities of nucleotide sequences designed as probes for CNTFR and myogenin mRNAs, we observed hybridization signals as follows. The CNTFR oligonucleotide probe revealed significantly increased signals in myonuclei of denervated myofibers and axotomized spinal motorneurons. These cells were previously shown to upregulate CNTFR mRNA in the same situation (Helgren et al. 1994; McLennan et al. 1999). The myogenin oligonucleotide probe has been characterized previously and revealed prominently increased signals in desmin-positive mpcs and myotubes in regenerating muscles (Kami et al. 1995). Taken together, preliminary examinations and previous observations revealed that the probes used in this study can detect the specific signals for each mRNA.
Results
Discrimination of Cell Types Expressing Receptor mRNAs
Specific hybridization signals for gp130, LIFR, IL-6R, and CNTFR mRNAs were detected in nuclei of several cell populations in regenerating muscles. We defined cell types showing significant hybridization signals for these receptor mRNAs as follows. Myonuclei were easily distinguished when they were located in the center of the cytoplasm of myofibers at 3 hr post contusion. It was hard to distinguish myonuclei when they were located at the edge of myofibers. We observed the relationship between myofibers and nuclei at higher magnification. Nuclei located within myofibers were defined as myonuclei/nuclei of mpcs. Nuclei located in extracellular spaces between myofibers were defined as mononuclear cells. Newly formed myotubes were confirmed as small myofibers with centrally located myonuclei and signals for myogenin mRNA.
Effect of Muscle Contusion on gp130 Expression
Signals for gp130 mRNA were at background level in intact myofibers, but intense signals were detected in the endothelium-like cells of blood vessels (Figures 1A–1C). At 3 hr after muscle contusion, gp130 mRNA expression was upregulated in injured muscle tissues (Figure 1D). gp130 mRNA was expressed in mononuclear cells located in extracellular spaces between myofibers (Figure 1E). Signals for gp130 mRNA were also detected in a myonucleus located in the center of the cytoplasm of a myofiber (Figure 1F) and in myonuclei/nuclei of mpcs at 3 hr postcontusion (Figure 1G). On Day 1 postcontusion, myonuclei/nuclei of mpcs expressing gp130 mRNA were still detected (Figures 2A–2C), and then obvious signals for gp130 mRNA were observed in many mononuclear cells on Day 2 postcontusion (Figures 2D and 2E). Low levels of mononuclear cells expressing gp130 mRNA were observed in extracellular spaces between myotubes until Day 7 post contusion, although gp130 mRNA expression in myonuclei/nuclei of mpcs decreased after Day 2 post contusion (data not shown).
Myogenin, a member of the MyoD family, is first upregulated in myonuclei/nuclei of mpcs at 6 hr post-contusion (Grounds et al. 1992; Kami et al. 1995). Furthermore, at Days 5 and 7 post contusion, some of the newly formed myotubes with centrally located myonuclei expressed myogenin mRNA (Kami et al. 1995). Therefore, we used the expression of myogenin mRNA as a marker for newly formed myotubes and examined whether gp130 mRNA was expressed in myotubes using adjacent serial sections hybridized with probes for gp130 and myogenin mRNAs (Figures 2F–2I). On Day 7 post contusion, myogenin mRNA expression was detected in some myotubes (Figure 2I), but signals for gp130 mRNA were not detected in these myotubes (Figure 2G).
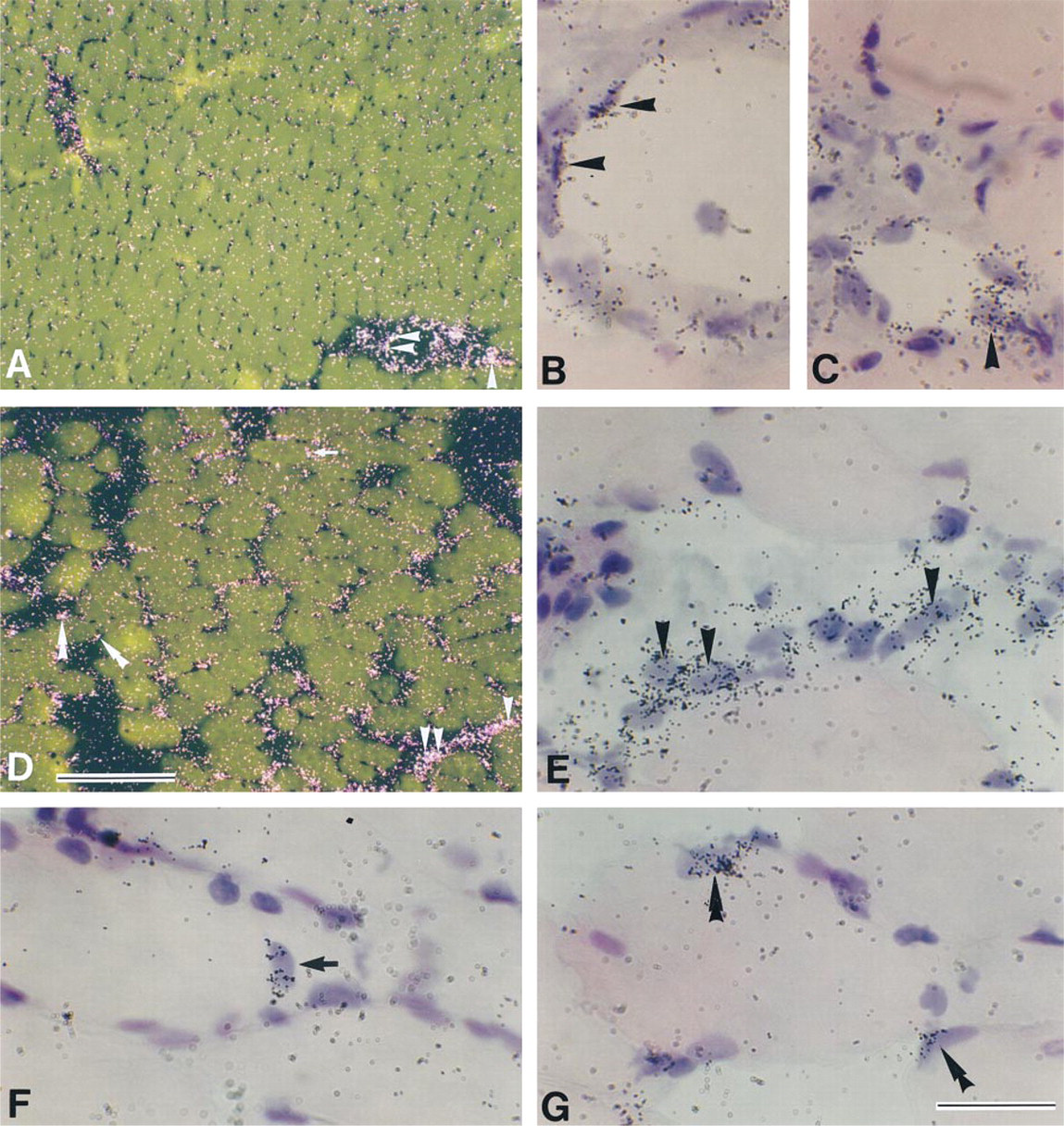
Darkfield (
Effect of muscle contusion on LIFR expression Signals for LIFR mRNA were at background level in intact myofibers (Figure 3A) but, as in the case of gp130, intense signals were detected in the endothelium-like cells of blood vessels (Figure 3B). At 3 hr after muscle contusion, upregulation of the LIFR mRNA level was observed in injured muscle tissues (Figure 3C). Signals for LIFR mRNA were detected in myonuclei/nuclei of mpcs (Figure 3D). At Day 1 post contusion, several mononuclear cells located in extracellular spaces between myofibers expressed LIFR mRNA (Figure 3F). LIFR mRNA expression in myonuclei/nuclei of mpcs decreased after Day 2 post contusion, but a few mononuclear cells expressing LIFR mRNA were observed at low levels until Day 7 post contusion (data not shown).
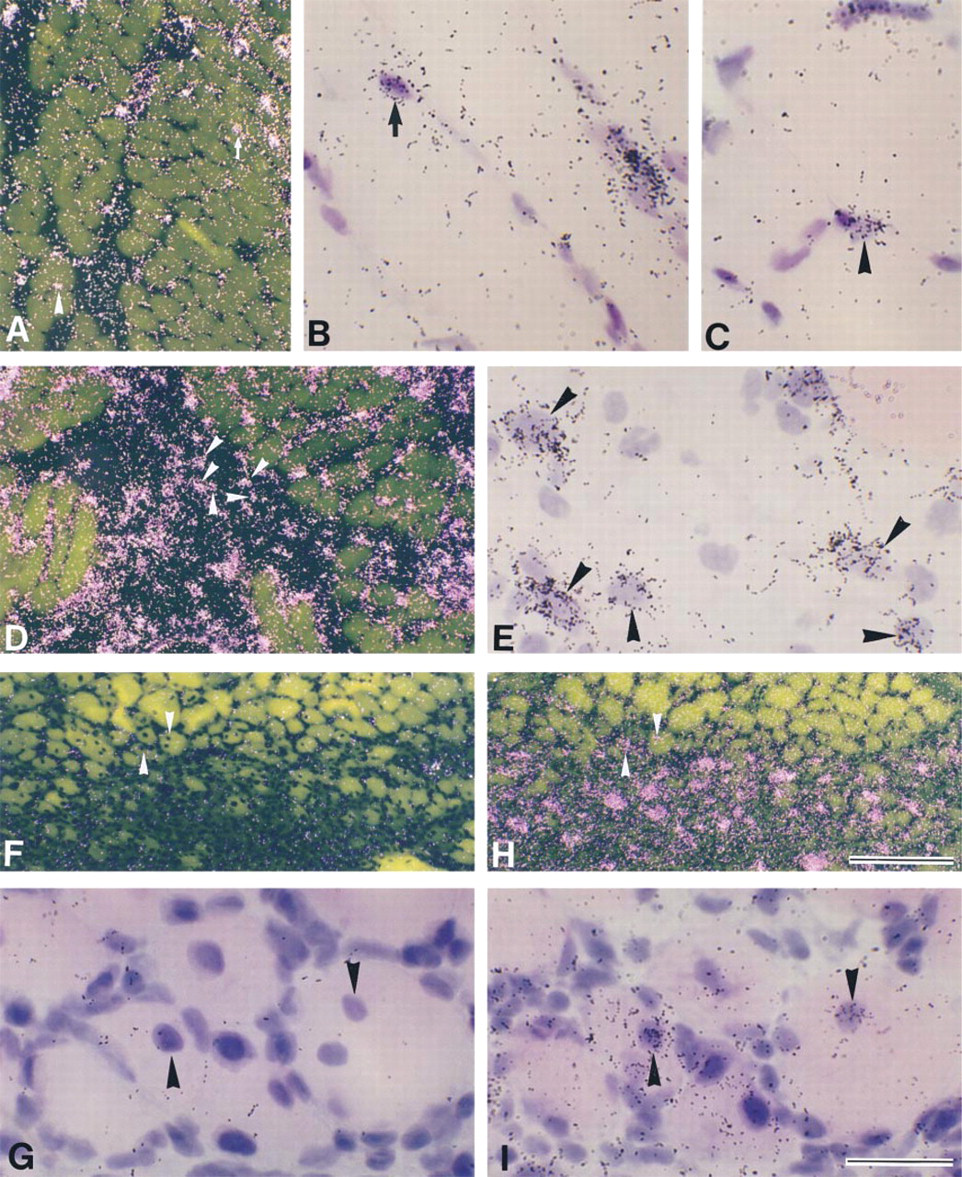
Darkfield (
Darkfield (
Darkfield (
To determine whether expression of LIFR mRNA can be detected in newly formed myotubes, adjacent serial sections were hybridized with probes for LIFR and myogenin mRNAs (Figures 3G–3J). At Day 7 postcontusion, some myotubes expressed myogenin mRNA (Figure 3J), but LIFR mRNA expression in these myotubes was not detected (Figure 3H).
Effect of Muscle Contusion on IL-6R Expression
Signals for IL-6R mRNA were at background level in intact myofibers (data not shown). Upregulation of IL-6R mRNA was first observed at low levels in injured muscle tissues at 3 hr post contusion (data not shown), and then signals for IL-6R mRNA increased at Day 1 postcontusion (Figures 3A and 3B). Obvious myogenin mRNA expression has been observed in myonuclei/nuclei of mpcs at Day 1 post-contusion (Grounds et al. 1992; Kami et al. 1995).
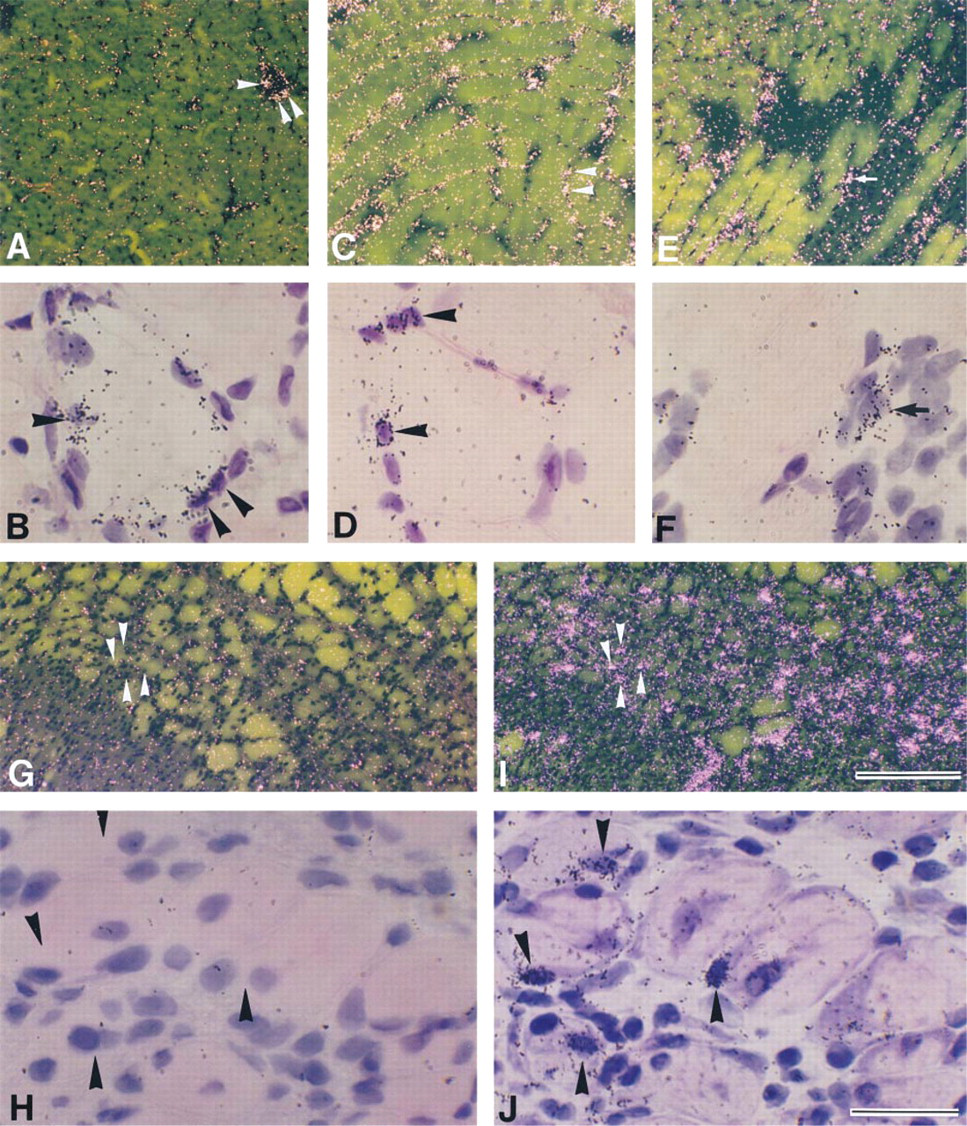
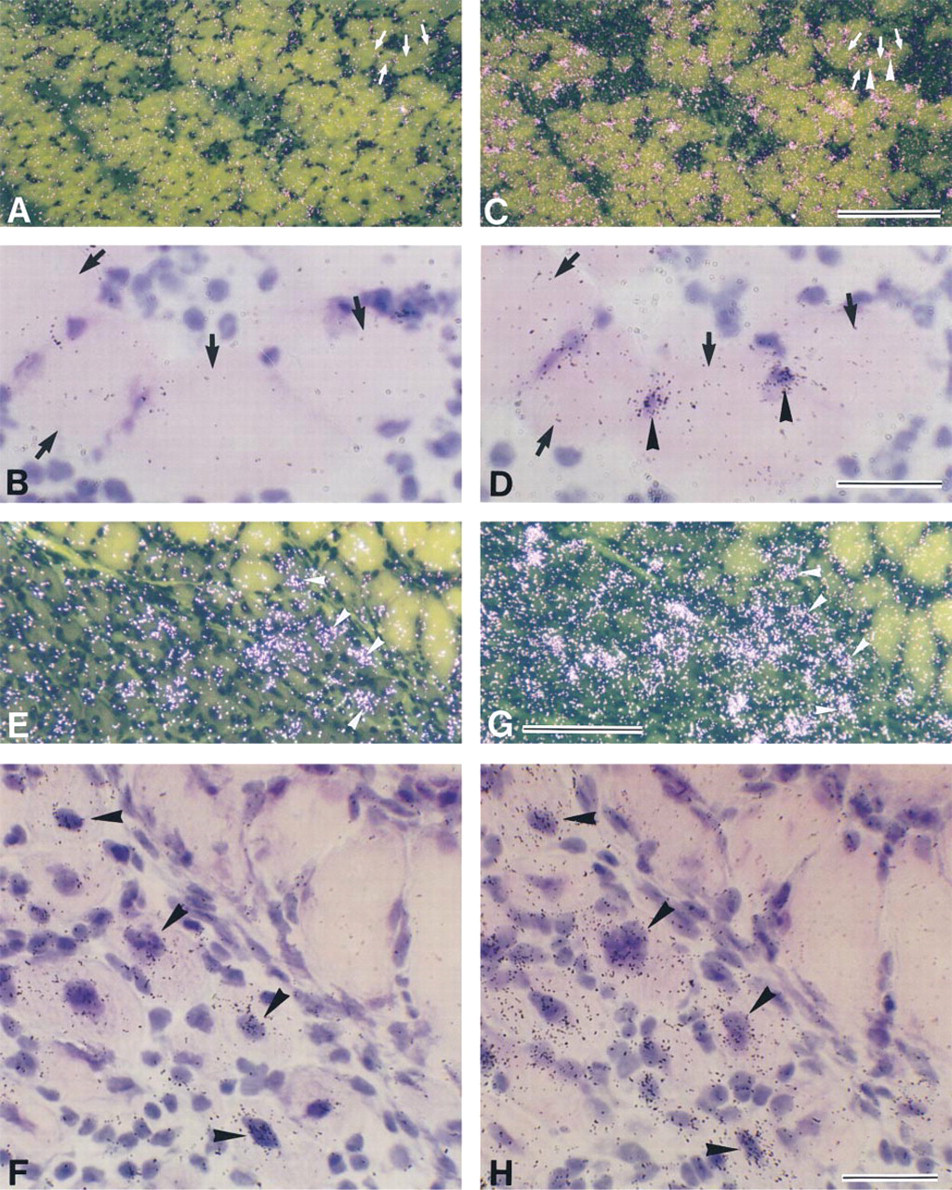
To determine whether IL-6R mRNA was expressed in myonuclei/nuclei of mpcs after contusion, adjacent serial sections were hybridized with probes for IL-6R and myogenin mRNAs (Figures 4A–4D). On Day 1 post contusion, some myonuclei/nuclei of mpcs expressed myogenin mRNA (Figures 4C and 4D). In contrast to myogenin expression, signals for IL-6R mRNA were not detected in myonuclei/nuclei of mpcs (Figures 4A and 4B), but significant signals for IL-6R mRNA were detected in mononuclear cells located in extracellular spaces between myofibers (Figure 4B). IL-6R mRNA expression throughout the regeneration process was detected only in mononuclear cells and was never observed in myonuclei/nuclei of mpcs. The level of IL-6R mRNA peaked at Days 1 and 2 post contusion, and only a small number of IL-6R mRNA-positive mononuclear cells were observed at low levels until Day 7 post contusion (data not shown).
To confirm whether expression of IL-6R mRNA was present in newly formed myotubes, adjacent serial sections were hybridized with probes for IL-6R and myogenin mRNAs (Figures 4E–4H). Myogenin mRNA expression was observed in some newly formed myotubes (Figure 4H). However, signals for IL-6R mRNA were not detected in these myotubes (Figure 4F).
Effect of Muscle Contusion on CNTFR Expression
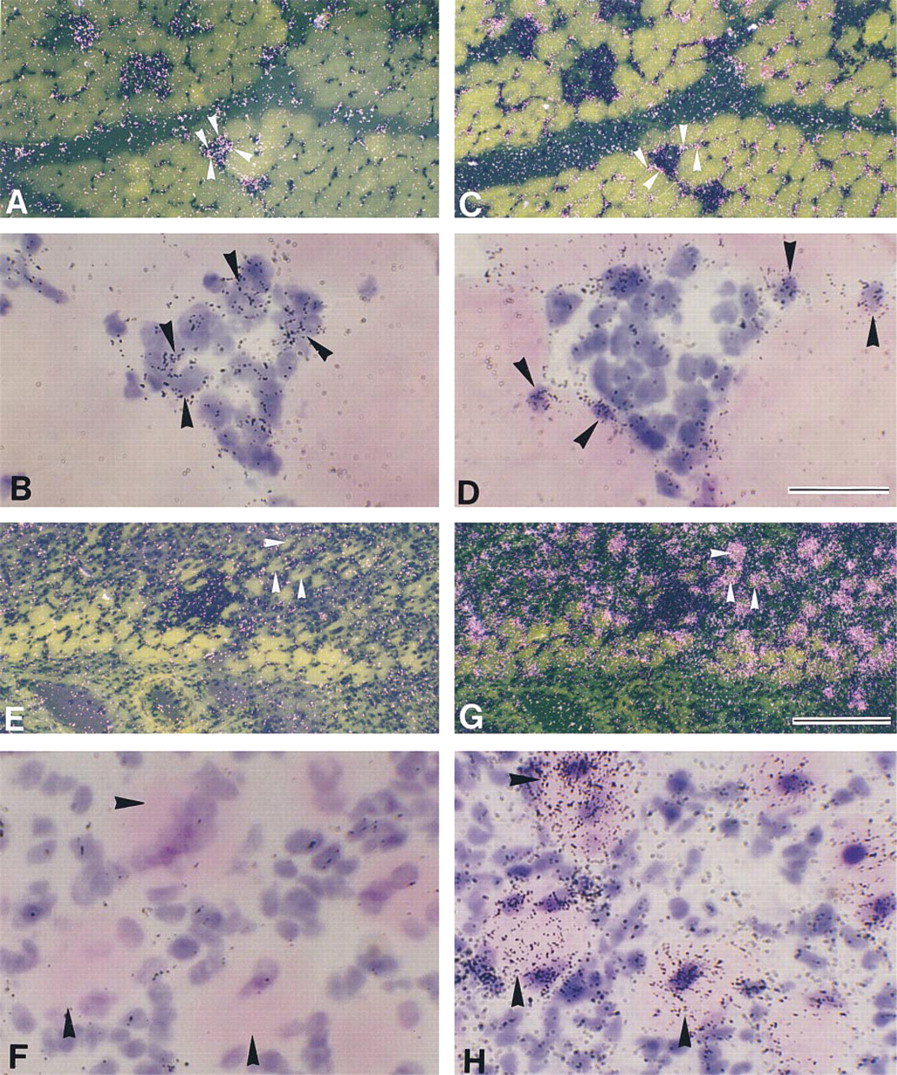
Signals for CNTFR mRNA were at background level in intact myofibers (data not shown). To determine whether CNTFR mRNA was expressed in myonuclei/nuclei of mpcs after contusion, adjacent serial sections were hybridized with probes for CNTFR and myogenin mRNAs (Figures 5A–5D). At Day 1 postcontusion, some myonuclei/nuclei of mpcs expressed myogenin mRNA (Figures 5C and 5D), but signals for CNTFR mRNA were not detected in myonuclei/nuclei of mpcs in injured myofibers that expressed myogenin mRNA (Figures 5A and 5B). Furthermore, CNTFR mRNA expression was not observed in mononuclear cells throughout the regeneration process (data not shown).
To determine whether expression of CNTFR mRNA was detected in newly formed myotubes, adjacent serial sections were hybridized with probes for CNTFR and myogenin mRNAs (Figures 5E–5H). At Day 7 post contusion, signals for myogenin mRNA were observed in some newly formed myotubes (Figure 5H), and these myotubes also expressed CNTFR mRNA at significant levels (Figure 5F).
The cell populations that expressed gp130, LIFR, IL-6R, and CNTFR mRNAs in intact and injured muscles after muscle contusion are summarized in Table 1.
Discussion
Constitutive expression of gp130 and LIFR mRNAs in adult intact skeletal muscles has been detected by Northern blotting (Ip et al. 1993; Helgren et al. 1994). In the present study, we observed no signals for gp130 and LIFR mRNAs in intact myofibers, but prominent signals were detected in the endothelium-like cells of blood vessels, which have been shown to be responsive to LIF (Gendron et al. 1996). Therefore, the hybridization signals for gp130 and LIFR mRNAs in intact muscles detected by Northern blotting may be derived from the endothelium-like cells of blood vessels. At 3 hr post contusion, gp130 and LIFR expression began to be upregulated in myonuclei/nuclei of mpcs and increased levels were maintained until Day 2 post-contusion. However, both mRNAs were apparently downregulated in the newly formed myotubes. There are many data showing that LIF is upregulated in regenerating myofibers and that this contributes to the proliferation of myoblasts in vitro (Austin et al. 1992; Barnard et al. 1994; Bower et al. 1995; Kami and Senba 1998). The complex of gp130 and LIFR is also utilized as a signal transducer to mediate the biological actions of CNTF, OM, and CT-1 (Kishimoto et al. 1994), but there are no reports suggesting the synthesis of these cytokines in regenerating muscles or their roles in myogenesis in vivo or in vitro. Taken together, these findings suggest that one of the roles of LIFR and gp130 produced in injured myofibers and/or mpcs after muscle contusion is to turn on LIF signaling, which may induce proliferation of mpcs and transactivation of genes related to regenerative reactions in the early stage of muscle regeneration. Furthermore, LIF signaling may not be induced in myotubes in which both gp130 and LIFR are absent. It has been reported that cytokines of the IL-6 family activate the mitogen-activated protein kinase pathway or the Jak-STAT signal transduction pathway, both of which induce myoblast proliferation in in vitro experiments (Bennett and Tonks 1997; Megeney et al. 1996). Evidence demonstrating an activation of intracellular signal transduction pathways in regenerating myofibers is therefore required to further clarify the action of LIF on these cells.
Darkfield (
Several observations suggest the participation of IL-6 in myofiber degeneration (Goodman 1994; Ebisu et al. 1995; Tsujinaka et al. 1995, 1996). In this study, IL-6R mRNA expression was never observed in myonuclei/nuclei of mpcs and newly formed myotubes, but significant signals for IL-6R mRNA were detected in mononuclear cells after contusion. These results suggest that the IL-6 response is not directly induced in injured myofibers and mpcs after contusion. Investigations into the IL-6R system provided evidence that a complex of IL-6 and an sIL-6R might act on the cells that express gp130, but not IL-6R (Kishimoto et al. 1994). This suggests a possible role of IL-6R in regenerating muscles. If IL-6R is released from mononuclear cells and an IL-6/sIL-6R complex is formed, the IL-6 response may be induced in myofibers expressing gp130 after muscle contusion. Proteolysis of muscle proteins takes place in injured myofibers at early stage after injury, and IL-6 enhances myofiber catabolism by activating the lysosomal cathepsin system (Tsujinaka et al. 1995, 1996), which is activated in degenerating myofibers (Ishiura et al. 1983, 1986). Therefore, production and release of sIL-6R by mononuclear cells may be essential for the IL-6-induced activation of proteolysis in degenerating myofibers.
Mononuclear cells located in extracellular spaces between myofibers express gp130, LIFR, and IL-6R mRNAs. In this study, we did not determine the cell types of these mononuclear cells. However, macrophages, fibroblasts, neutrophils, and mpcs, all of which have been extensively observed in injured muscles, may be candidates for these mononuclear cells (Husmann et al. 1996). These cells produce and release growth factors and cytokines, such as fibroblast growth factors (FGFs), platelet-derived growth factor (PDGF), transforming growth factor-β (TGF-β), and LIF, all of which induce proliferation of mpcs, inhibition of myogenic differentiation, chemotaxis of mpcs and inflammatory cells, and synthesis of extracellular matrix components (Husmann et al. 1996). Therefore, to further elucidate the role of the IL-6 family in muscle regeneration, it is necessary to investigate the effects of the IL-6 family of cytokines on growth factor and cytokine production in these mononuclear cells.
Cell types detected by ISH with probes for gp130, a LIFR, IL-6R, and CNTFR mRNAs in intact and injured muscles
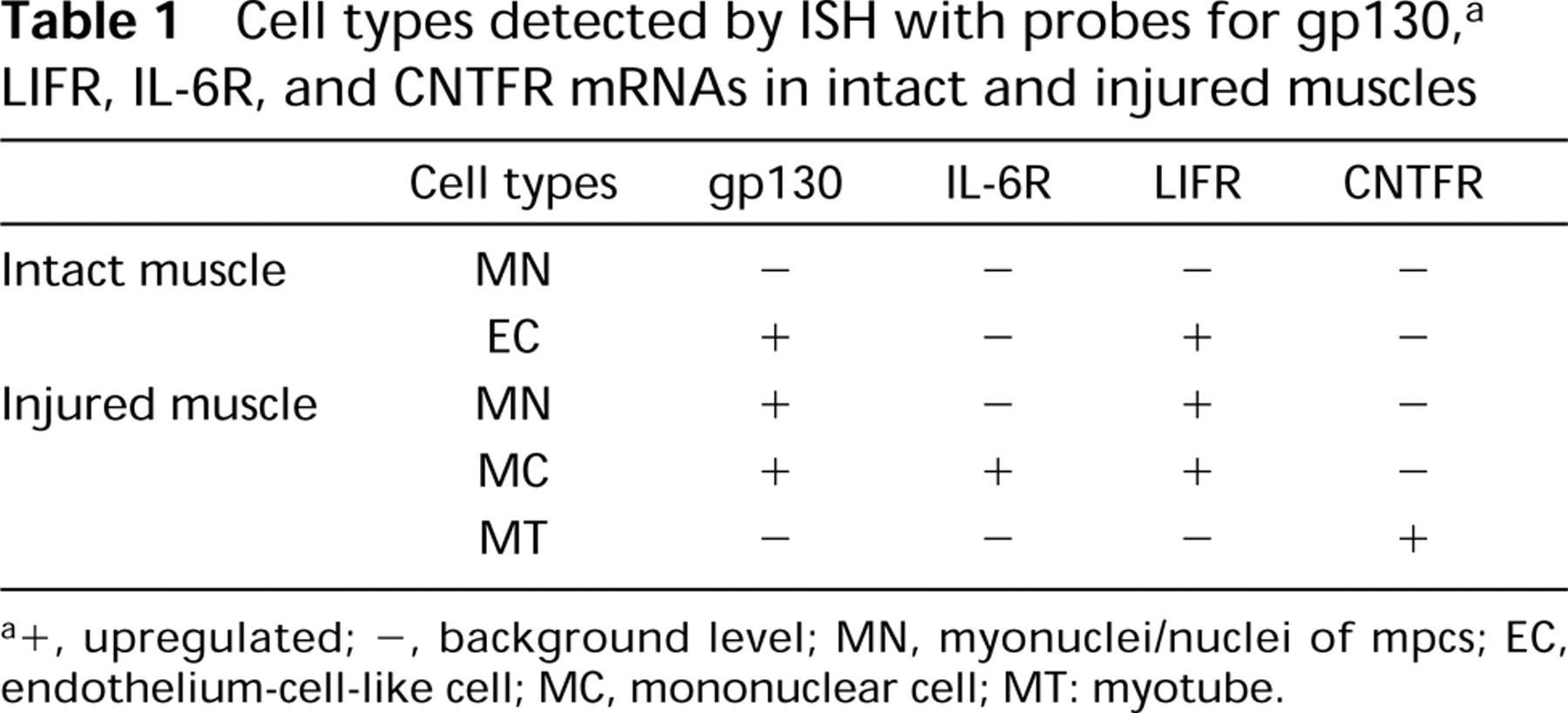

a +, upregulated; –, background level; MN, myonuclei/nuclei of mpcs; EC, endothelium-cell-like cell; MC, mononuclear cell; MT: myotube.
CNTFR expression was not detected at significant levels in myonuclei/nuclei of mpcs in intact and injured myofibers nor in mononuclear cells throughout the process of muscle regeneration. However, the level of CNTFR mRNA was significantly upregulated in some of the myotubes, whereas both gp130 and LIFR, the other components of functional CNTF receptors, diminished in myotubes. These findings suggest that myotubes are not targets for CNTF action. The effect of CNTF on myotube differentiation (Marques and Neto 1997) may be indirect. What is the role of CNTFR induced in myotubes? It is known that sCNTFR is released from myofibers in response to peripheral nerve injury (Davis et al. 1993), and CNTF is released from Schwann cells after nerve injury (Sendtner et al. 1992). We previously showed that the intramuscular nerves are injured by muscle contusion (Kami et al. 1999). These findings suggest that sCNTFR released from myotubes may form a complex with CNTF released from Schwann cells in injured intramuscular nerves (sCNTFR/CNTF complex). These complexes may be retrogradely transported in motorneurons in a receptor-mediated manner and may promote motor nerve sprouting and enhance myofiber reinnervation, as shown in previous studies (Curtis et al. 1993; Ulenkate et al. 1994).
In conclusion, this study has demonstrated that expression of gp130, IL-6R, LIFR, and CNTFR mRNAs in regenerating muscles is induced in distinct cell populations, such as myonuclei/nuclei of mpcs, mononuclear cells, and myotubes. These receptor subunits probably mediate actions of LIF, IL-6, and CNTF to facilitate the effective degenerative and regenerative reactions of myofibers and reinnervation on myotubes in regenerating muscles.
Footnotes
Acknowledgements
cDNAs for gp130 and IL-6R were generous gifts from Dr T. Kishimoto (Osaka University). Rat LIFR cDNA was a generous gift from Dr M. Minami and Dr M. Sato (Kyoto University).
