Abstract
Although growth factors and cytokines play critical roles in skeletal muscle regeneration, intracellular signaling molecules that are activated by these factors in regenerating muscles have been not elucidated. Several lines of evidence suggest that leukemia inhibitory factor (LIF) is an important cytokine for the proliferation and survival of myoblasts in vitro and acceleration of skeletal muscle regeneration. To elucidate the role of LIF signaling in regenerative responses of skeletal muscles, we examined the spatial and temporal activation patterns of an LIF-associated signaling molecule, the signal transducer and activator transcription 3 (STAT3) proteins in regenerating rat skeletal muscles induced by crush injury. At the early stage of regeneration, activated STAT3 proteins were first detected in the nuclei of activated satellite cells and then continued to be activated in proliferating myoblasts expressing both PCNA and MyoD proteins. When muscle regeneration progressed, STAT3 signaling was no longer activated in differentiated myoblasts and myotubes. In addition, activation of STAT3 was also detected in myonuclei within intact sarcolemmas of surviving myofibers that did not show signs of necrosis. These findings suggest that activation of STAT3 signaling is an important molecular event that induces the successful regeneration of injured skeletal muscles.
A
The sequential but distinctive events of mpcs in regenerating muscles are regulated by growth factors and cytokines, such as the hepatocyte growth factor (HGF), the fibroblast growth factors (FGFs), the platelet-derived growth factor (PDGF), the insulin-like growth factors (IGFs), and the leukemia inhibitory factor (LIF) (Florini et al. 1991; Grounds 1999). Moreover, the significance of LIF in muscle regeneration has been demonstrated in vitro and in vivo. LIF belongs to the interleukin-6 (IL-6) family of cytokines composed of IL-6, ciliary neurotrophic factor (CNTF), oncostatin M (OM), cardiotrophin-1 (CT-1), and interleukin-11 (IL-11), which share gp130 as a signaling receptor in their functional receptor components (Taga 1996; Takeda and Akira 2000). LIF stimulates proliferation and survival of cultured myoblasts (Austin et al. 1992; Kurek et al. 1996b; White et al. 2001) and induces the formation of larger myotubes in vitro (Vakakis et al. 1995). Furthermore, administration of LIF to damaged muscles and the mdx mouse model of Duchenne muscular dystrophy accelerates muscle regeneration (Barnard et al. 1994; Kurek et al. 1996a, 1997). Signals for LIF mRNAs were observed in myonuclei and/or nuclei of mpcs located in the periphery of myofibers at the early stage of muscle regeneration, and these signals disappeared in newly formed myotubes (Kurek et al. 1997; Kami and Senba 1998).
The action of LIF on skeletal muscle regeneration is mediated by a functional LIF receptor composed of two signal-transducing proteins, LIF receptor-β (LIFR) and gp130. It has been reported that rapid upregulation of these two receptor mRNAs after muscle injury was detected in myonuclei and/or nuclei of mpcs in injured muscles but that these signals disappeared in newly formed myotubes (Kami et al. 1999, 2000). Signal transduction after LIF binding to the LIFR/gp130 complex is accomplished preferentially through the Janus kinase (JAK) signal transducer and activator of the transcription (STAT) pathway. Activation of the LIFR/gp130 complex induces tyrosine phosphorylation of STAT proteins (preferentially STAT3) by receptor-associated JAKs. Phosphorylated STAT proteins subsequently form hetero- or homodimers and translocate to the nucleus, where they exhibit transcriptional activity on target genes. Additional phosphorylation of STAT3 proteins at serine residues has been suggested to enhance transcriptional activity of STAT3 proteins (Taga 1996; Takeda and Akira 2000). Potential target genes of STAT3 have been known to include cyclin D1, c-fos, JunB, Bcl-2, and Bcl-xL (Coffer et al. 1995; Fukada et al. 1996; Jenab and Morris 1996; Bromberg 2001), which mediate several biological functions such as proliferation, differentiation, and survival of myoblasts (Lassar et al. 1989; Rahm et al. 1989; Li et al. 1992; Rao et al. 1994; Skapek et al. 1995; Dominov et al. 1998, 2001; Olive and Ferrer 1999). Therefore, monitoring STAT3 signaling in regenerating muscles may be important to further understand the regulatory mechanism of skeletal muscle regeneration.
We analyzed the activation of STAT3 signaling in the regeneration of skeletal muscles induced by crush injury, identifying mpcs and distinct regenerative stages of mpcs (i.e., activation, proliferation, and differentiation). In this study, satellite cells were identified by c-Met, dystrophin, and laminin immunolabeling, and nuclei expressing both proliferating cell nuclear antigen (PCNA) and MyoD proteins were identified as nuclei of proliferating myoblasts. Expressions of the cyclin-dependent kinase inhibitor p21, myogenin, and AchR in myoblasts determined that these myoblasts were in the post-mitotic stage. The present study shows that rapid phosphorylation and nuclear translocation of STAT3 proteins are exclusively induced in the activated satellite cells, proliferating myoblasts, and surviving myofibers during skeletal muscle regeneration. These results provided a molecular basis for further understanding of the muscle regeneration mechanism.
Materials and Methods
Muscle Crush Injury and Tissue Preparation
Adult Wistar rats (200–250 g) were used in this study. Muscle crush injury was achieved without a skin incision as described previously (Kami et al. 1993, 1995). Briefly, a cylinder (640 g) was dropped from a distance of 250 mm onto a plate (diameter 1.0 cm) placed on the belly of the medial gastrocnemius muscle of the right leg of a rat anesthetized with ethyl ether. Injured muscles and their uninjured contralateral counterparts were resected under pentobarbital anesthesia 3 hr, 1, 3, or 5 days after the muscle crush injury (three rats per day). Gastrocnemius muscles freshly dissected from anesthetized animals were frozen in liquid nitrogen and stored at −85C until use. All experiments were carried out with the approval of the Osaka University of Health and Sport Sciences Animal Ethics Committee.
Immunostaining
Primary antibodies used in this study were as follows: a rabbit polyclonal anti-laminin (1:3000; Sigma, St Louis, MO), a rabbit polyclonal anti-phospho STAT3 (Tyr705) (P-STAT3) (1:100; Cell Signaling, Beverly, MA), a mouse monoclonal anti-MyoD (1:50; DAKO, Carpinteria, CA), a mouse monoclonal anti-dystrophin (1:100; Sigma), a mouse monoclonal anti-desmin (1:100; Zymed, So. San Francisco, CA), a mouse monoclonal anti-cyclin-dependent kinase inhibitor p21 (1:50; BD PharMingen, San Diego, CA), a mouse monoclonal anti-c-met (1:50; Santa Cruz Biotech., Santa Cruz, CA), and a mouse monoclonal anti-PCNA (1:50; DAKO).
Five-μm or 12-μm frozen cross-sections were mounted on 3-amino-propylethoxysilane-coated slides and fixed with 4% paraformaldehyde in a 0.1 M phosphate buffer (pH 7.4) for 15 min. The sections were then washed in 0.1 M PBS and incubated with 0.1 M PBS containing 10% normal serum and 0.3% Triton X-100 at room temperature (RT) to block nonspecific staining. For triple-immunofluorescence staining, the sections were incubated simultaneously with the primary antibodies diluted with 0.1 M PBS containing 5% normal donkey serum, 0.3% Triton X-100 from 16 hr to 48 hours at 4C. The sections were washed in 0.1 M PBS and incubated with the secondary antibodies diluted with 0.1 M PBS containing 5% normal donkey serum, 0.1% Triton X-100 overnight at 4C. Fluorescein-conjugated donkey anti-mouse IgG was used for the mouse monoclonal primary antibodies and rhodamine-conjugated donkey anti-rabbit IgG was used for the rabbit polyclonal antibodies (Chemicon; Temecula, CA). The sections were then washed in 0.1 M PBS, and mounted in Vectashield mounting medium with DAPI (Vector Labs; Burlingame, CA) to visualize the nuclei. Immunofluorescence-stained sections were viewed on an Olympus microscope with epifluorescence using a X20 or X40 objective.
For immunohistochemical staining, the sections were incubated with the primary antibody diluted with 0.1 M PBS containing 5% normal goat or horse serum, 0.3% Triton X-100 from 16 hr to 48 hr at 4C. The sections were washed in 0.1 M PBS and incubated with the biotin-conjugated secondary antibodies diluted with 0.1 M PBS containing 5% normal goat or horse serum, 0.1% Triton X-100 at RT. Biotin-conjugated horse anti-mouse IgG was used for the mouse monoclonal primary antibodies (Vector Labs) and biotin-conjugated goat anti-rabbit IgG was used for the rabbit polyclonal antibodies (Chemicon). The sections were then washed in 0.1 M PBS, and incubated with a Vectastain Elite ABC kit (Vector) and diaminobenzidine (DAB). Immunostained sections were stained with hematoxylin to visualize the nuclei.
Probes for In Situ Hybridization
The oligonucleotide probe for acetylcholine receptor-α (AchR) was complementary to nucleotides 1216–1263 of the published rat AchR sequences (Witzemann et al. 1990). The oligonucleotide probe for myogenin was complementary to nucleotides 56–100 of the published rat myogenin sequence (Wright et al. 1989). The specificity of these oligonucleotide probes was verified by a sequence comparison using computer analysis and a DNASIS DNA homology search (Hitachi; Tokyo, Japan). No significant homology with any other genes in primates or rodents was found. These oligonucleotide probes were labeled by the 3'-end-labeling method using [35]S-dATP and terminal deoxynucleotidyl transferase and had a specific activity of 1.0–3.0 × 105 cpm/μl. The labeled oligonucleotide probes were applied to sections at 0.5 × 106 cpm/slide.
In Situ Hybridization
Frozen cross-sections of gastrocnemius muscle 12 μm thick were thawed on 3-amino-propylethoxysilane-coated slides, then hybridized with radiolabeled probes as follows. Sections were fixed with 4% paraformaldehyde in a 0.1 M phosphate buffer (pH 7.4) for 15 min, then rinsed in 2 X SSC (0.3 M NaCl, 30 mM Na citrate). The sections were incubated with 1 μg/ml proteinase K (Sigma) at 37C for 5 min. After two rinses in 2 X SSC, they were dehydrated in 70%, 80%, 90%, 95%, and 100% ethanol, then air-dried. The sections were incubated with a hybridization buffer [50% formamide, 4 X SSC, 0.12 M phosphate buffer, pH 7.4, 1 X Denhardt's solution, 0.2% SDS, 0.25 mg/ml tRNA, 10% dextran sulfate, 100 mM dithiothreitol (DTT)] containing the radiolabeled oligonucleotide probes at 37C for 16 hr. After hybridization, the sections were washed with three changes of 1 X SSC at 55C for 20 min each, followed by dehydration in 80%, 90%, 95%, and 100% ethanol, and then air-dried. To visualize the signals for the specific mRNAs, the sections were dipped in Ilford K15 autoradiography emulsion diluted 2:3 with distilled water at 45C. After exposure at 4C for 4 weeks, they were developed in Kodak D19 for 4 min at 20C and then fixed in 24% sodium thiosulfate solution for 5 min. Lastly, the sections were stained with hematoxylin and eosin.
To ascertain the specificity of the nucleotide sequences designed as probes for AchR and myogenin mRNAs, we observed hybridization signals as follows. An AchR oligonucleotide probe revealed distinct signals in the neuromuscular junctions of the intact myofibers and myonuclei of denervated myofibers. The myogenin oligonucleotide probe has been characterized previously (Kami et al. 1995). Taken together, preliminary examinations and previous observations revealed that the probes used in this study could detect specific signals for each mRNA.
Results
Myonuclei Express Activated STAT3 Proteins
To investigate whether the STAT3 signaling pathway was activated in regenerating muscles after muscle crush injury, we performed immunohistochemical staining with a specific antibody for the Tyr705-phosphorylated form of STAT3 protein (P-STAT3). STAT3 proteins were detected as inactivated forms in various cells, including skeletal muscles and motor neurons (Megeney et al. 1996; Hass et al. 1999; Schwaiger et al. 1999; Zong et al. 2000). However, when tyrosine phosphorylation of STAT3 proteins occurs, the phosphorylated STAT3 is rapidly translocated into the nucleus and can never be detected in the cytoplasm (Taga 1996; Hass et al. 1999; Schwaiger et al. 1999; Takeda and Akira 2000). In intact and uninjured contralateral muscles, immunoreactivities for P-STAT3 could not be detected (data not shown). At 3 hr after injury, the earliest time point examined, the activated STAT3 proteins were observed in injured muscles (Figure 1A). Triple immunostaining using P-STAT3 and dystrophin antibodies and DAPI showed that immunoreactivites for the P-STAT3 were detected in nuclei located within dystrophin-positive sarcolemma of myofibers, indicating that activation of STAT3 proteins was induced in myonuclei (Figure 1D). Distinct expression of P-STAT3 in myonuclei could be detected until day 3 after injury. Myonuclei expressing the activated STAT3 proteins were detected exclusively in surviving myofibers without signs of disruption of the myofiber sarcolemma and necrosis of myofibers (Figures 1E-1H). Furthermore, P-STAT3-positive nuclei were also found outside the sarcolemma of myofibers (Figure 1D) and these were considered to be nuclei of satellite cells or inflammatory cells, which are distributed widely in injured muscles.
Activation of STAT3 Signaling in Activated Satellite Cells
To investigate if activation of STAT3 signaling is induced in satellite cells in regenerating muscles, satellite cells were identified using two strategies. Previous studies showed that c-Met, a receptor for hepatocyte growth factor (HGF), is expressed in both quiescent and activated satellite cells (Bottaro et al. 1991; Tatsumi et al. 1998; Anderson 2000). Therefore, we identified satellite cells by immunostaining with a c-Met monoclonal antibody. In our preliminary experiment, c-Met immunoreactivity was detected in small cells located in the periphery of myofibers in intact muscles, and more intense c-Met labeling was observed in injured muscles (data not shown). Although c-Met is a receptor in the plasma membrane, immunoreactivity was detected in the cytoplasm of satellite cells. Such an immunostaining pattern of c-Met in the satellite cells was previously reported by Tatsumi et al. (1998) and Anderson (2000), and may indicate vigorous production of c-Met proteins in ribosomes within satellite cells activated by muscle injury. At 3 hr after injury, the activated STAT3 proteins could be detected in nuclei of c-Met-positive satellite cells (Figures 2A-2D), indicating that activation of STAT3 signaling was induced in the activated satellite cells.
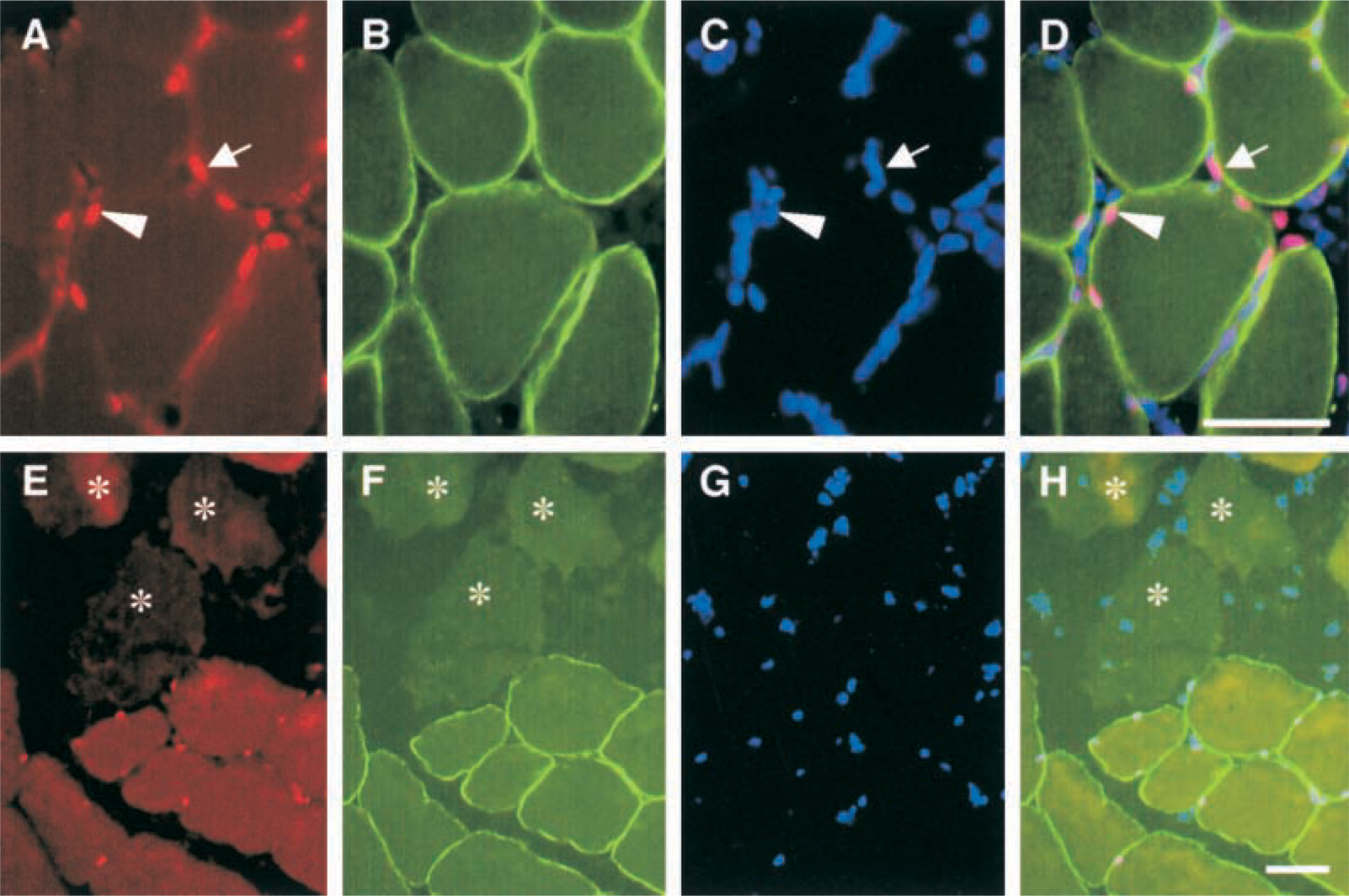
Photomicrographs showing localization of activated STAT3 proteins at 3 hr after injury. Triple immunostaining to view localization of (
Satellite cells are located between the sarcolemma and basement membrane of myofibers. On the basis of this feature, we identified satellite cells by triple and double immunostaining on 5-μm serial sections. Mammalian satellite cell dimensions are approximately 25 × 4 × 5 μm, and lengths of the rat satellite cell nucleus average 9–12 μm (Allbrook 1981; Watkins and Cullen 1988). Therefore, it is expected that a satellite cell nucleus can be divided into two serial sections. Triple immunostaining to visualize localization of P-STAT3, dystrophin, and nuclei was performed on a section (Figures 2E-2H), and an adjacent serial section was immunostained with laminin antibody and DAPI (Figures 2I-2K). These results showed that a nucleus located inside of laminin and outside of dystrophin was P-STAT3-positive at 3 hr after injury. Together with the present observation by c-Met immunolabeling, these results showed that activation of STAT3 signaling occurred in activated satellite cells.
Activation of STAT3 Signaling in Proliferating Myoblasts
MyoD plays critical roles in development and regeneration of skeletal muscles, and rapid upregulation of MyoD mRNA and protein after muscle injury is detected in nuclei of all myogenic cells and myonuclei (Fuchtbauer and Westphal 1992; Grounds et al. 1992; Koishi et al. 1995; Cooper et al. 1999). In the present study, significant MyoD immunoreactivity was not detected in intact and 3-hr post-injured muscles (data not shown), but intense MyoD staining was observed in injured muscles at day 1 after injury (Figure 3A). Triple immunostaining using MyoD and P-STAT3 antibodies and DAPI detected many nuclei expressing both MyoD and P-STAT3 proteins (Figures 3A-3C). Because newly formed myotubes are never detected in regenerating muscles at day 1 after injury, these results suggest, that in addition to the myonuclei and nuclei of activated satellite cells shown in Figures 1 and 2, STAT3 signaling may also be activated in proliferating myoblasts.
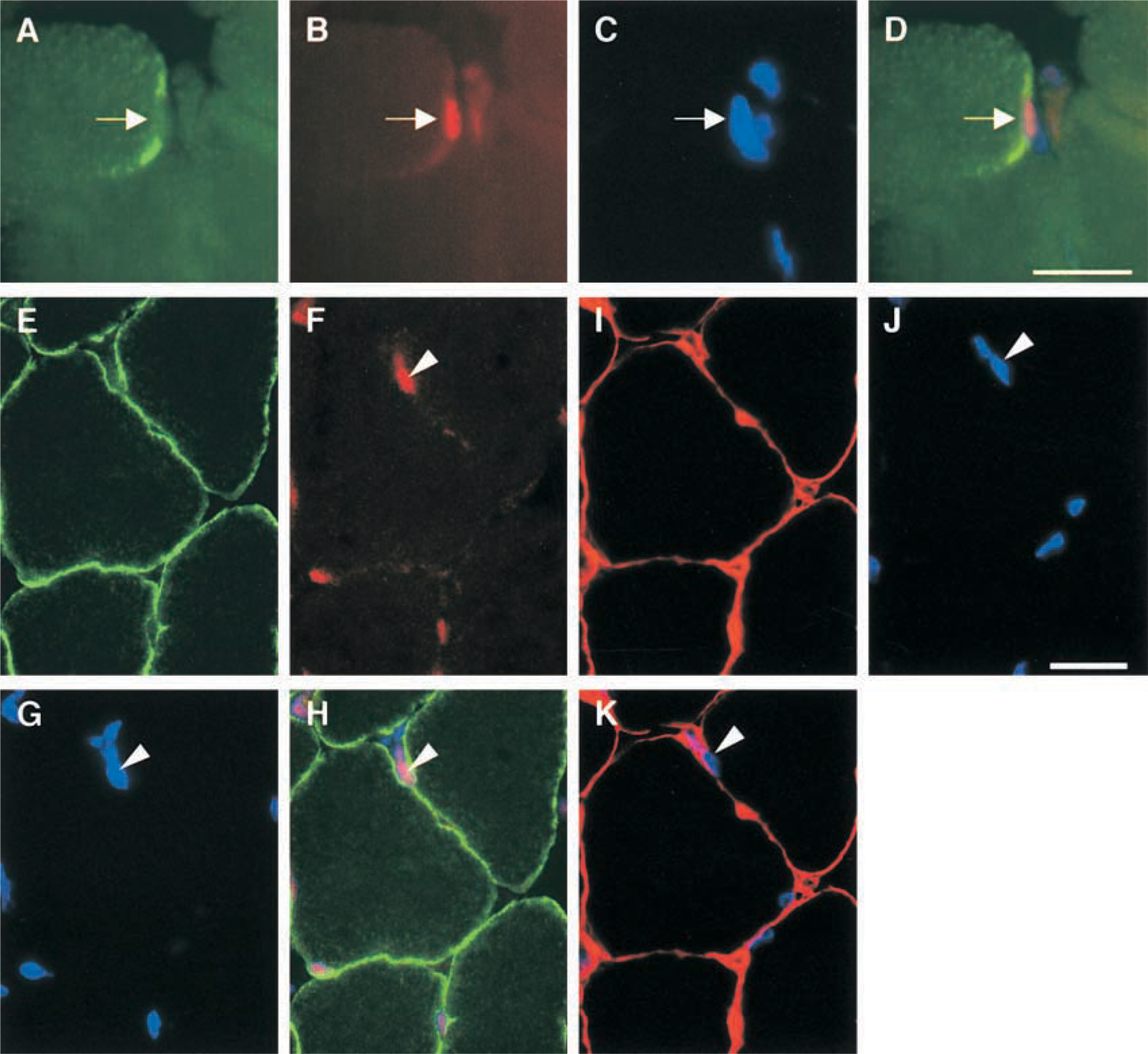
Photomicrographs showing activation of STAT3 signaling in satellite cells at 3 hr after injury. Triple immunostaining to view localization of (
To identify proliferating myoblasts in regenerating muscles, we performed triple and double immunostaining on adjacent serial sections prepared at a thickness of 5 μm as mentioned above. It has been known that proliferating cell nuclear antigen (PCNA) is an auxiliary protein of DNA polymerase δ, whose level correlates with DNA synthesis during the cell cycle, being maximal during the S-phase (Bravo et al. 1987; Baserga 1991). Yablonka-Reuveni and Rivera (1997) reported that the kinetics of the PCNA-positive nuclei correlated well with the kinetics of DNA-synthesizing satellite cells as determined by tracing [3H]-thymidine-labeled nuclei in vitro. In the present study, nuclei labeled with PCNA immunostaining were not detected in intact and 3-hr post-injury muscles, but significant PCNA-positive nuclei were observed in injured muscles at day 1 after injury (data not shown). Triple immunostaining using MyoD and P-STAT antibodies and DAPI showed nuclei expressing both MyoD and P-STAT3 proteins (Figures 3F-3H), and double immunostaining with PCNA antibody and DAPI on an adjacent serial section (Figures 3D and 3E) showed the existence of nuclei that display simultaneous immunoreactivities for PCNA, MyoD, and P-STAT3 (Figures 3D-3H). These results indicate that activation of STAT3 signaling was induced in proliferating myoblasts.
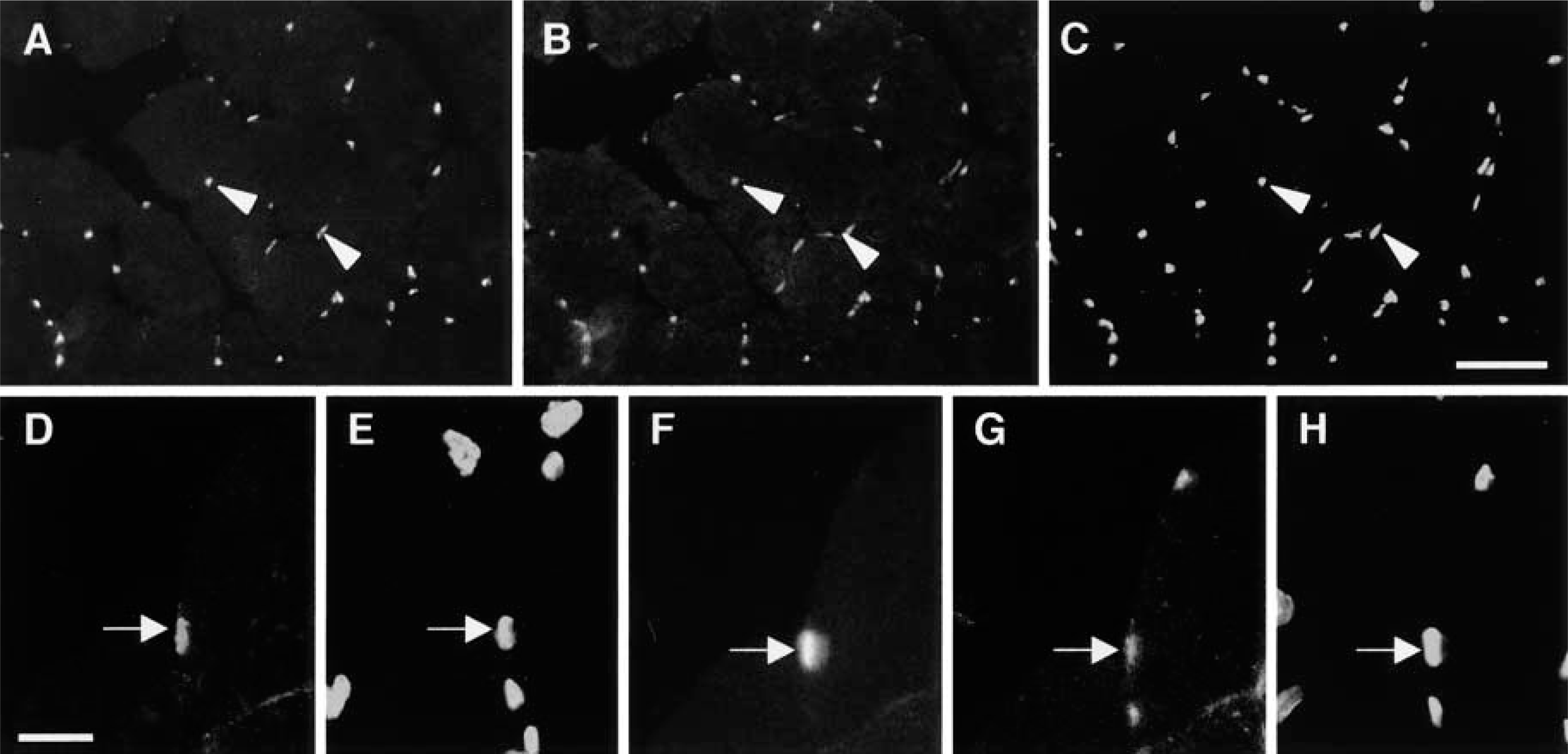
Photomicrographs showing activation of STAT3 signaling in proliferating myoblasts at day 1 after injury. Triple immunostaining to view localization of (
Activation of STAT3 Signaling Disappears in Differentiated Myoblasts and Myotubes
Myoblasts that stop proliferating are induced into the differentiation stage. Myotube formations in regenerating muscles progress within basement membranes as a scaffold for myoblast differentiation and then fuse, so that the myoblasts are frequently detected as a ring-like structure (Hurme and Kalimo 1991; Koishi et al. 1995; White et al. 2000). At day 3 after injury, many MyoD-positive myoblasts were observed as a ring-like structure within the remaining basement membranes that were not completely destroyed by muscle crush injury (Figures 4A-4D). It is well known that mRNAs and proteins for cyclin-dependent kinase inhibitor p21 (p21), myogenin, and AchR are upregulated in differentiated myoblasts and myotubes during myogenesis in vitro and in vivo (Halevy et al. 1995; Walsh 1997; Liu et al. 2000). Therefore, we analyzed the regenerative stage of myoblasts arranged as a ring-like shape using immunostaining or in situ hybridization to detect expression of p21, myogenin, and AchR. No significant p21 immunoreactivity was detected in intact and 3-hr after injury muscles (data not shown). At day 1 after injury, p21 immunoreactivity was detected in injured muscles, and increased and more intense p21-positive cells were detected at day 3 after injury. These cells formed a ring-like structure within laminin-positive basement membranes like that of MyoD (Figures 4E-4H). We investigated expression patterns of myogenin and AchR mRNAs in regenerating muscles using serial sections at day 3 after injury. Like MyoD and p21 proteins, desmin protein, another marker for activated satellite cells and myoblasts, was detected in cells arranged in a ring-like structure within the remaining laminin-positive basement membranes (Figures 4I and 4J). Serial sections showed that signals for both myogenin and AchR mRNAs could be detected in these myoblasts (Figures 4K and 4L). These results showed that myoblasts arranged as a ring in regenerating muscles were in the post-mitotic stage. However, because it is difficult to clearly distinguish between the differentiated mononuclear myoblasts and the young myotubes at the light microscopic level, the possibility that these cells are very young myotubes cannot be completely ruled out.
We assayed to ascertain whether activation of STAT3 signaling was also induced in differentiated myoblasts by immunostaining with MyoD and P-STAT3 antibodies. At day 3 after injury, MyoD-positive myoblasts were observed as a ring-like structure inside the remaining basement membranes (Figures 5A-5C), and analysis using serial sections showed that P-STAT3 could not be detected in these myoblasts (Figures 5D and 5E). Furthermore, at day 5 after injury, many myotubes that had centrally located p21- and MyoD-positive nuclei in their cytoplasm were detected, but activation of STAT3 signaling was not observed in myotubes (data not shown). The expression patterns for c-Met, MyoD, PCNA, p21, and P-STAT3 proteins in regenerating muscles are summarized in Table 1.
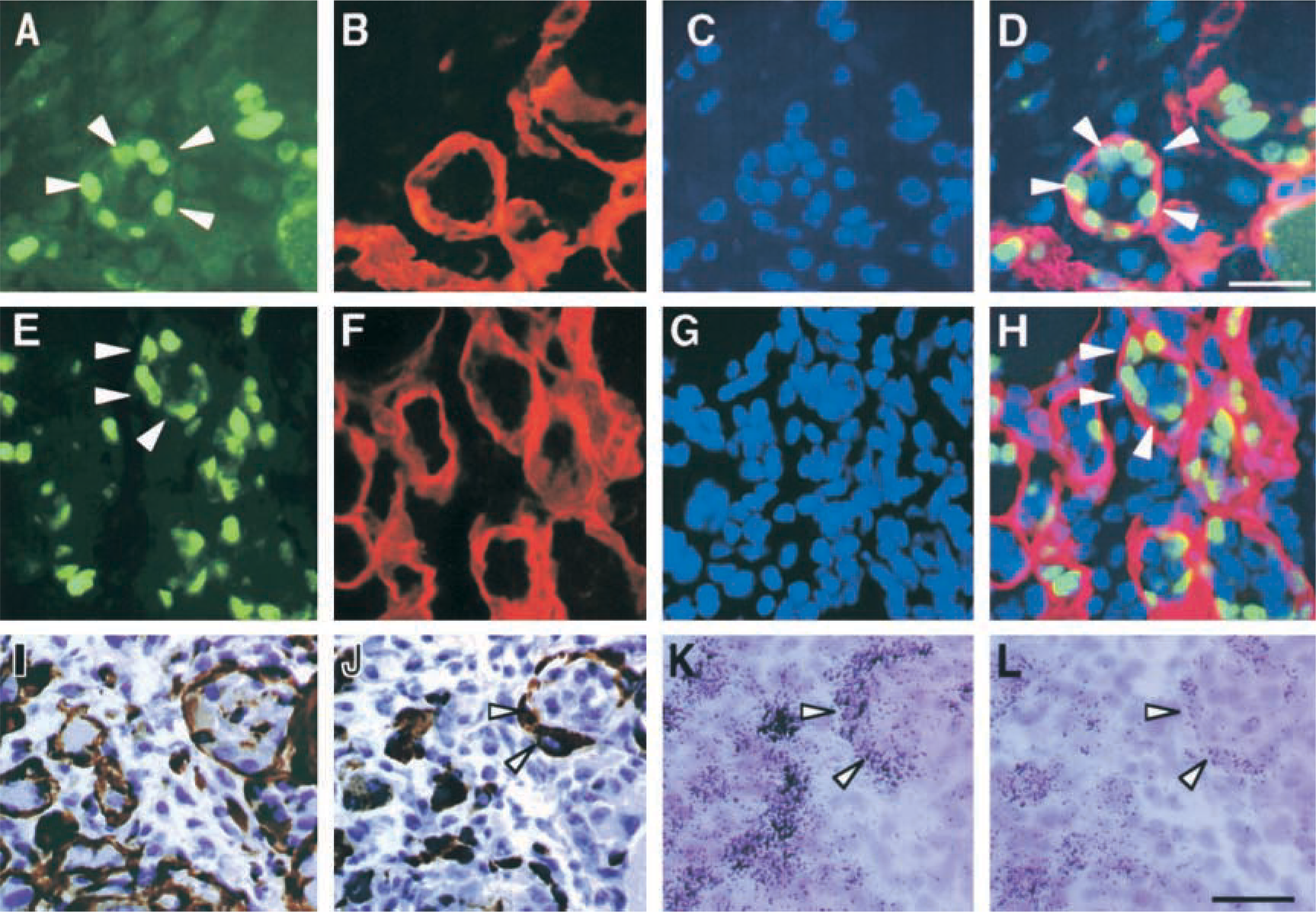
Photomicrographs showing localization of MyoD and p21 proteins and signals for myogenin and AchR mRNAs at day 3 after injury. Triple immunostaining to view localization of (
Discussion
Muscle regeneration progresses through sequential events consisting of activation, proliferation, differentiation, and survival of mpcs. Therefore, to determine the relationship between activated signaling molecules and the stages of mpcs, it is important to understand the molecular mechanisms of muscle regeneration. It is well known that the entire process of muscle regeneration is regulated by growth factors and cytokines (Florini et al. 1991; Grounds 1999). There is a line of evidence showing beneficial effects of LIF on muscle regeneration (Barnard et al. 1994; Kurek et al. 1996a, 1997). Therefore, we aimed to elucidate whether or not an LIF-associated signaling molecule, STAT3, is activated in mpcs in regenerating skeletal muscles. The present study showed that STAT3 signaling was activated in the activated satellite cells, proliferating myoblasts, and surviving myofibers, whereas it was no longer active in differentiated myoblasts and myotubes. The present results suggest that activated STAT3 protein may mediate important molecular events occurring at the early regenerative stage.
Possible Factors that Activate STAT3 Signaling
As mentioned above, rapid phosphorylation and translocation into nuclei of STAT3 proteins were induced at a considerably early time point (3 hr to day 1 after injury) after muscle injury. These results imply that an upstream factor(s) and their specific receptors connected to STAT3 signaling must also be induced in injured muscles at this time point. STAT3 is activated preferentially by the members of the IL-6 family of cytokines (Taga 1996; Takeda and Akira 2000). Our previous in situ hybridization data showed that, at 3 hr to day 2 after injury, transcripts for LIF, LIFR, and gp130 were expressed in myonuclei and/or nuclei of mpc located in the periphery of myofibers (Kami and Senba 1998; Kami et al. 1999, 2000). The appearance of LIF/LIFR/gp130 mRNA expression in the regenerating muscles coincides with the period of activation of STAT3 signaling, suggesting a direct causal relationship between these two events. On the other hand, expression of IL-6, another important signaling molecule for STAT3 activation, was also detected in injured muscles (Kurek et al. 1996b; Kami and Senba 1998). However, IL-6 receptor-α (IL-6R), a ligand-binding receptor in a functional IL-6 receptor complex, was exclusively expressed in interstitial mononuclear cells but not in myofibers or mpcs during muscle regeneration (Kami et al. 2000). In accordance with these in situ hybridization data, no beneficial effects of IL-6 on skeletal muscles are found in vivo or in vitro experiments (Goodman 1994; Ebisui et al. 1995). Therefore, IL-6 may not be directly involved in STAT3 phosphorylation. Likewise, any beneficial effects of other members of the IL-6 family of cytokines (i.e., CNTF, CT-1, OM, and IL-11) on muscle regeneration are not reported. Therefore, these findings and the present observations suggest that LIF is a strong candidate to induce activation of STAT3 signaling in regenerating muscles, although the involvement of other factors (e.g., granulocyte CSF, leptin, insulin, epidermal growth factor, platelet-derived growth factor) than the IL-6 family of cytokine cannot be completely ruled out (Bromberg 2001).
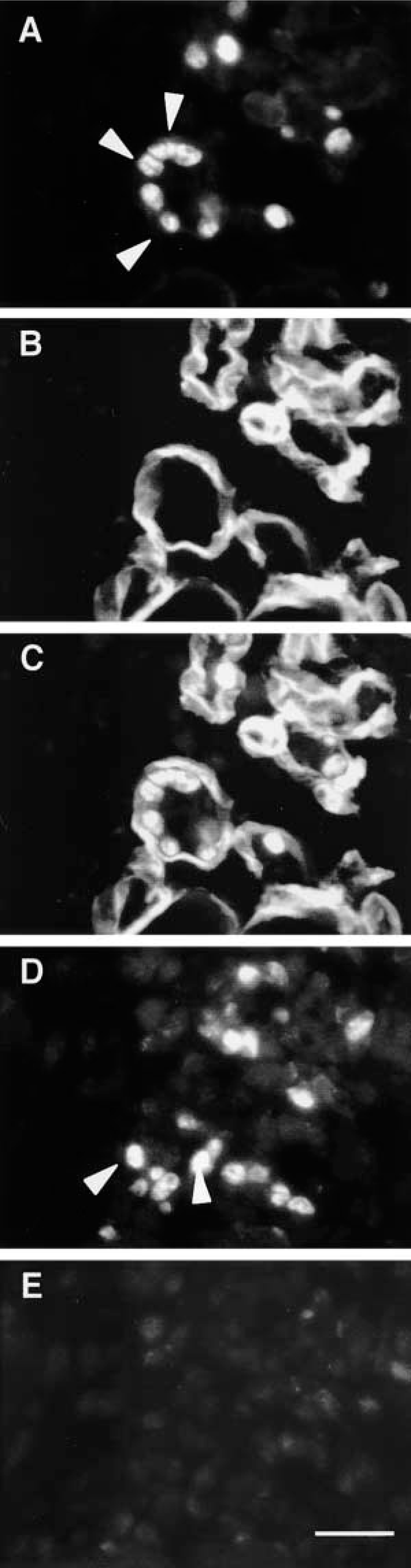
Photomicro-graphs showing inactivation of STAT3 signaling in MyoD-positive myoblasts at day 3 after injury. Double immunostaining to view localization of (
LIF also contributes to the formation of larger myotubes in vitro (Vakakis et al. 1995), suggesting that STAT3 signaling leads to myotube hypertrophy. However, present data showed that activation of STAT3 signaling was not detected in the differentiated myoblasts and myotubes. LIF is able to activate phosphatidylinositol 3-kinase (PI 3-kinase), an upstream effector for Akt (protein kinase B) activation (Oh et al. 1998). Activation of Akt protein has been shown to induce myotube hypertrophy (Rommel et al. 2001). However, the expression of LIFR and gp130 mRNAs was not found in myotubes in regenerating muscles (Kami et al. 2000), which implies that the LIF-induced larger myotube formation could not be a direct effect of LIF. It has been reported that LIF enhances the proliferation and survival of myoblasts in vitro and that the phosphorylated forms of STAT3 are detected in the proliferating myoblasts in vitro (Megeney et al. 1996; White et al. 2001). Consequently, such an increase of myoblast numbers induced by LIF stimuli may contribute to the formation of larger myotubes.
Possible Target Genes and Functions of Activated STAT3 Protein
Although phosphorylated STAT3 is rapidly translocated into nuclei to function as a transcription factor, the genes that are directly transactivated by STAT3 in skeletal muscles are not yet elucidated. In other cells, potential STAT3 target genes have been shown to include Bcl-2 and Bcl-xL, which act as anti-apoptosis factors (Fukada et al. 1996; Bromberg 2001). Bcl-2 is expressed in skeletal muscle cells at the early stage of myogenesis and inhibits apoptosis in vitro. Furthermore, Bcl-2-positive cells co-express markers of early stages of myogenesis such as desmin, MyoD, and Myf5, whereas Bcl-2 was not expressed in differentiated myotubes (Dominov et al. 1998). In developing skeletal muscle, Bcl-2 expression was limited to a small group of mononucleate, desmin-positive, and myogenin-negative muscle cells (Dominov et al. 2001). Examination of Bcl-2 expression in human myopathies showed that Bcl-2 immunoreactivity was detected not only in regenerating myofibers but also in apparently normal myofibers (Olive and Ferrer 1999). This distribution pattern resembles that of P-STAT3 in regenerating muscles shown in the present study.
Activation patterns of the P-STAT3 protein in regenerating rat skeletal muscles a
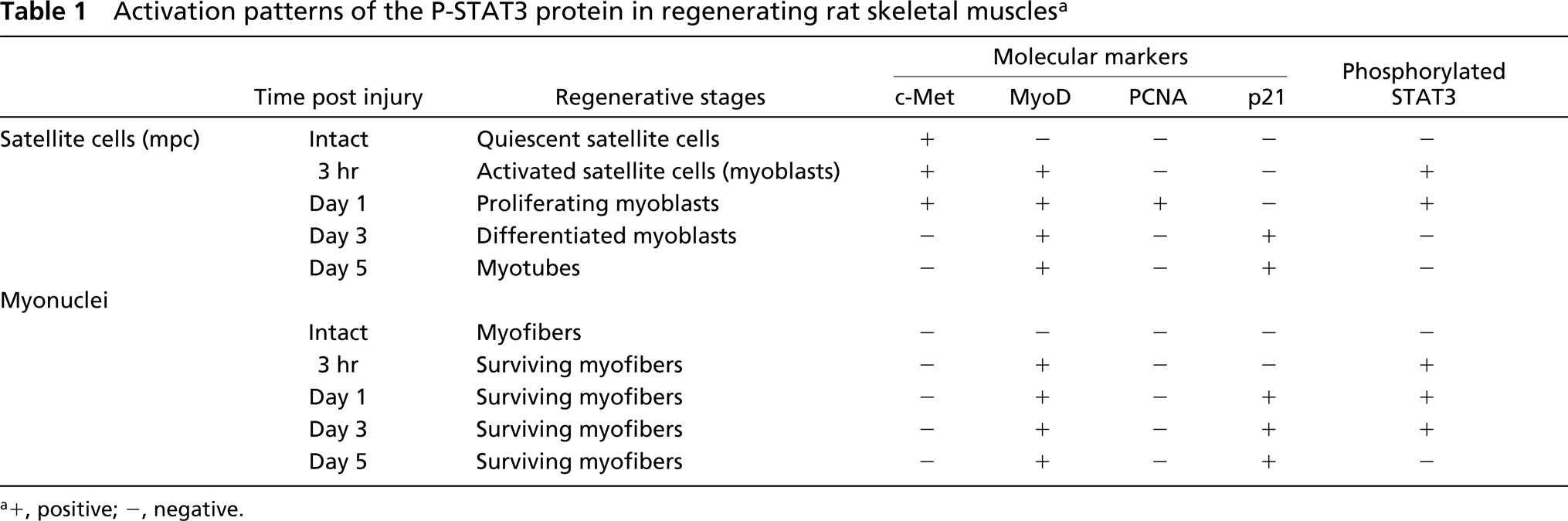

a+, positive; -, negative.
In an attempt to investigate regulation of cell death, neonatal muscles of Bcl-2-null mice revealed a reduced myofiber number compared with wild-type mice (Dominov et al. 2001), and direct evidence that survival of cultured myoblasts was enhanced by LIF was recently reported (White et al. 2001). These observations and the present results, taken together, suggest that one of the target genes of STAT3 in regenerating muscles may be Bcl-2 and that STAT3-induced Bcl-2 may serve to protect against apoptotic cell death of the activated satellite cells, proliferating myoblasts, and surviving myofibers.
Other potential STAT3 target genes are c-fos, junB, and cyclin D1 (Coffer et al. 1995; Jenab and Morris 1996; Bromberg 2001), which have been shown to be involved in myoblast proliferation and differentiation. Our previous study showed that expression of c-fos mRNA was induced in myonuclei and/or nuclei of mpcs located in the periphery of myofibers at 3 hr after muscle crush injury but that c-fos mRNA was not detected in newly formed myotubes (Kami et al. 1995). Therefore, the expression patterns of c-fos mRNA are also compatible with the activation patterns of STAT3 observed in the present study. c-Fos inhibits the formation of myotubes and expression of muscle-specific genes in vitro (Lassar et al. 1989; Rahm et al. 1989). Localization of JunB and cyclin D1 in regenerating muscles has not been reported. However, there is evidence to show that JunB suppresses transactivation of the muscle creatine kinase enhancer by MyoD and myogenin (Li et al. 1992). Proliferation of activated satellite cells is characterized by progression through successive steps of the cell cycle (Chambers and McDermott 1996), and forced expression of cyclin D1, which is important for the G1–S-phase transition of the cell cycle, inhibited the ability of MyoD to transactivate muscle-specific genes (Rao et al. 1994; Skapek et al. 1995). Therefore, if activated STAT3 transactivates c-fos, JunB, and/or cyclin D1 in the activated satellite cells and proliferating myoblasts, these proteins induced by STAT3 may inhibit myogenic differentiation. Inhibition of myoblast differentiation at an early regenerative stage may play an important role to ensure appropriate myoblast number through the progression of the cell cycle. Determination of the exact downstream targets of activated STAT3 in the activated satellite cells, proliferating myoblasts, and surviving myofibers is essential to elucidate the precise roles of this signaling molecule in regenerating muscles, and we are now attempting to confirm the hypothesis described above.
In summary, we have shown that activation of STAT3 signaling was exclusively induced in the activated satellite cells, proliferating myoblasts, and surviving myofibers in regenerating muscles, and LIF may be one of the strong upstream factors to activate this molecule. Protection of activated satellite cells, proliferating myoblasts and surviving myofibers from apoptotic cell death and inhibition of myoblast differentiation at the early regenerative stage are essential for complete muscle regeneration. Therefore, we propose that activated STAT3 may be an important signaling molecule that mediates these functions at the early stage of skeletal muscle regeneration.
Footnotes
Acknowledgements
Supported in part by a grant-in-aid (13670030) for Scientific Research from the Ministry of Education, Science, and Culture of Japan.
