Abstract
Although gcm was first recognized for its role in specifying glial cell fate in Drosophila melanogaster, its mammalian counterparts are expressed predominantly in non-neural tissues. Here we demonstrate expression of the mouse and human GCM 1 proteins in placenta. We have prepared a highly specific antibody that recognizes the GCM 1 protein and have used it to assess the temporal and spatial expression profile of the protein. In both mouse and human placenta, the protein is associated with cells that are involved with exchange between maternal and fetal blood supplies: the labyrinthine cells of the mouse placenta and the syncytio- and cytotrophoblasts of the human placenta. Using the full-length hGcm 1 cDNA as a probe, we have mapped the gene on human chromosome 6p12 by fluorescent in situ hybridization.
E
First identified in D. melanogaster, the gcm gene (also called glide for
Surprisingly, the mammalian Gcm 1 mRNA is more highly expressed in non-neural tissue and is only weakly expressed in brain (Kim et al. 1998; Basyuk et al. 1999). The murine, rat, and human Gcm 1 genes are expressed in placenta at levels easily detectable by Northern blotting, whereas they are expressed in neural tissues at levels that can be revealed only by reverse transcription-PCR (Kim et al. 1998; Basyuk et al. 1999; Janatpour et al. 1999). The expression of the Gcm 1 gene in non-neural cells suggests that this putative transcription factor has retained many characteristics of the Drosophila protein but participates in non-neural developmental processes.
Our analysis has revealed that although Gcm 1 expression in mouse placenta intersects that of several other transcription factors, its precise pattern is spatially and temporally distinct (Basyuk et al. 1999). This specificity implies a unique function. In furthering our effort to define the function of the Gcm 1 gene, we describe the preparation of polyclonal anti-sera specific for mouse and human GCM 1, the cell types expressing the GCM 1 protein in placenta, and the chromosomal localization of human Gcm 1 gene.
Materials and Methods
Isolation of Human Gcm 1 cDNA
An anonymous human cDNA clone was identified in the expressed sequence tags database (dbEST, Genbank, R62635; Washington University, St Louis, MO and Merck, Darmstadt, Germany) that exhibited striking sequence similarity to coding regions of the Drosophila gcm gene. This EST clone was used as a probe to isolate a 2.8-kb cDNA from a human placental library (Stratagene; La Jolla, CA). Both strands of cDNA were sequenced by dideoxy method using 35S-labeled nucleotides. Our nucleotide sequence agrees perfectly with that deposited in GenBank by Yamada et al. (1999) in the coding region but has three single nucleotides additions in the 3′UTR.
Chromosomal Localization of hGcm 1 Gene by FISH
FISH experiments were done on lymphocytes isolated from human blood and cultured in α-minimal essential medium (α-MEM) supplemented with 10% fetal calf serum and phytohemagglutinin at 37C for 68–72 hr. The lymphocyte cultures were then treated with BrdU (0.18 mg/ml) (Sigma; St Louis, MO) to synchronize the cell population. Synchronized cells were washed three times with serum-free medium to release the block and cultured at 37C for 6 hr in α-MEM with thymidine (2.5 μg/ml; Sigma). Cells were harvested and slides were made by using standard procedures including hypotonic treatment, fixation, and air-drying. FISH experiments were performed using a biotinylated 2.8-kb Gcm 1 cDNA as probe. The cDNA probe was labeled with biotin–dATP using a Bio-Nick labeling kit at 15C for 1 hr (Gibco-BRL; Gaithersburg, MD) (Heng et al. 1992). The procedure for FISH detection was performed as previously described (Heng et al. 1992; Heng and Tsui 1993). Briefly, slides were baked at 55C for 1 hr. After RNase treatment, slides were denatured in 70% formamide in 2 × SSC for 2 min at 70C, followed by dehydration with ethanol. The probe was then denatured at 75C for 5 min in a hybridization buffer consisting of 50% formamide and 10% dextran sulfate and was loaded on the denatured chromosome slides. After overnight hybridization, slides were washed at 37C in 50% formamide, 2 × SSC and then in 2 × SSC. FISH signals and the DAPI banding pattern were recorded separately by taking photographs, and the assignment of the FISH mapping data with chromosomal bands was achieved by superimposing FISH signals with DAPI-banded chromosomes (Heng and Tsui 1993).
Preparation of GCM 1 Antiserum and Western Blot Verification of Its Specificity
The portion of the mouse Gcm 1 cDNA encoding the carboxyl-terminal region of the protein (amino acids 169 to 437; Altshuller et al. 1996) was subcloned into the fusion protein expression vector pGEX-2T (Amersham Pharmacia; Poole, UK) in frame with the vector's GST encoding sequence. The fusion protein, expressed in bacterial strain BL21, was purified according to the manufacturer's instructions. A rabbit polyclonal antibody was generated against the GST–GCM 1 fusion protein (Pocono Rabbit Farm and Laboratory; Canadensis, PA). Antisera having high titers were combined and purified by a two-step process. First, the antibodies recognizing the GST portion of the antigen were removed by repeated passage over matrix-bound GST (Pierce Chemical; Rockford, IL). Second, the treated antisera were then affinity-purified by adsorption to and elution from a column of matrix-bound Gcm 1 peptide (residues 169–437) (Harlow and Lane 1988). The specificity of these antisera were assessed by Western Blotting of GST–GCM 1 fusion protein and cell-free extracts prepared from mouse or human placenta.
On the basis of the sequence homology between the mGcm 1 and the hGcm 1 cDNA, the anti-mouse GCM 1 antibody should also crossreact with the human protein. To verify this conclusion, we used the antisera in Western blotting analyses of samples of placental extracts. Soluble proteins were extracted from third-trimester human and E 15.5 mouse placenta by desegregating and sonicating the tissue in extraction buffer (0.1 M PBS, 10 mM DTT, 1% Triton X-100, 1 mM PMSF, 10 μg/ml aprotin, 10 mM pepstatin). Particulate material was removed by centrifugation. Samples were adjusted to 1 × SDS loading buffer and separated on a 10% denaturing polyacrylamide gel as previously reported (Laemmli 1970). The resolved proteins were then electrolytically transferred to PVDF nitrocellulose membrane using a Bio-Rad transfer apparatus (Bio-Rad; Hercules, CA), extensively washed in water, and incubated overnight in blocking solution (5% dry milk in 0.1 M PBS, 0.2% Triton X-100). The blot was incubated for 1 hr with anti-mGCM 1 antiserum diluted 1:1000 in blocking solution. After extensive washing in PBS/0.2% Triton X-100 (three washes of 20 min each), it was incubated for 1 hr with alkaline phosphatase-conjugated anti-rabbit IgG (Jackson Immunoresearch; West Grove, PA) at 1:5000 dilution. After several washes in PBS/0.2% Triton X-100, the blot was developed using the chemiluminescent protein detection system (Immunstar; Bio-Rad).
Immunohistochemistry
Sections of human first- and third-trimester and mouse E 8.5 to E15.5 placentas were used to investigate the expression of GCM 1 protein by immunohistochemistry. Briefly, paraffin sections (10-μm thickness) were deparaffinized in xylene and rehydrated via a graded series of ethyl alcohol. Sections were then incubated for 20 min with 1% hydrogen peroxide in 10% methanol to inhibit endogenous peroxidase activity. After saturation with 10% normal goat serum in 0.1 M PBS/0.2% Triton, sections were incubated overnight at 4C with the affinity-purified anti-GCM 1 antibody diluted 1:100 in the case of mouse and 1:40 in the case of human tissue. After incubation with the primary antibody, sections were washed in PBS/0.2% Triton and the secondary antibody, peroxidase-conjugated goat anti-rabbit IgG (Jackson ImmunoResearch Laboratories; West Grove, PA) was then applied for 1 hr. After washing with PBS, the bound antibody was visualized by detection of the peroxidase activity using diaminobenzidine as substrate (Sigma). The reaction was monitored for 5 min and the sections were then rinsed with distilled water and mounted in xylene-based mounting medium. Sections were analyzed by brightfield microscopy (Axiophot; Zeiss, Oberkochen, Germany). For the above immunohistochemical procedures, controls were performed by replacing the primary antibody with preimmune serum. Further controls were also performed by omitting the primary antibody. Controls were always free of labeling.
Results
Sequence Homology Between mGcm 1 and hGcm 1
The sequence of the 2.8-kb cDNA clone that we isolated revealed a large open reading frame (nucleotides 199–1506) that encodes a 436-amino-acid protein identical to the hGCM 1/a protein sequence previously reported (Akiyama et al. 1996; Yamada et al. 1999). The hGcm 1 cDNA that we identified contains a modest 5′-UTR sequence of 198 nucleotides and a large 1.28-kb 3′-UTR sequence (not shown). The high degree of sequence similarity between the human and murine GCM 1 proteins has been reported previously (Akiyama et al. 1996) and is illustrated in Figure 1 because it is central to the crossreactivity of the polyclonal antisera. Not surprisingly, these proteins show the highest level of similarity in the DNA binding domain, the GCM motif. However, even the transactivation domain shows striking 61% amino acid identity and 71% sequence similarity. These observations, together with those of others (Akiyama et al. 1996; Altshuller et al. 1996; Yamada et al. 1999), suggest that the gcm genes are highly conserved in mammals, perhaps reflecting a conserved function for these genes. Nevertheless, the intensity of the signals from the human protein (Figures 2 and 3) is considerably weaker than that from the mouse. Although this difference could be due to differences in the concentration of the two antigens in their respective tissues, it appears equally possible that it is a result of the sequence divergence in the antigens (Figure 1).
Specificity of the GCM 1 Antiserum
The specificity of the crude and affinity-purified anti-mouse GCM 1 antibodies was analyzed by Western blotting, as illustrated in Figure 2. After removal of the GST-specific antibodies by repeated adsorption to matrix-bound GST, the rabbit antiserum recognized the GST–GCM 1 fusion protein used as positive control but not GST itself (Figure 2, Lanes 1 and 2). The antiserum recognized a protein of approximately 50 kD, the expected size of GCM 1 protein in mouse placenta (Figure 2, Lane 3). A faint band at approximately 90 kD was occasionally seen but only with the partially purified antibodies. The affinity-purified antibodies recognized a single antigen in extracts of mouse E10 and human third-trimester placenta (Figure 2, Lanes 4 and 5). The human protein has an apparent molecular weight that is slightly larger than predicted. Discrepancies between predicted size and that estimated from SDS gel electrophoresis are commonplace and may result from the residual secondary structure of the protein or from post-translational modifications. The immunochemically identified bands obtained with both mouse and human placental extracts were completely abolished after preabsorption of the GCM 1 antiserum on the matrix-bound GCM 1 peptide used in the affinity-purification step (data not shown).

Sequence homology of mouse and human GCM 1 proteins. The alignment of human (this report) and mouse (Altshuller et al. 1996) GCM 1 proteins reveals extensive regions of amino acid sequence identity (dark shaded regions) and similarity (light shaded regions). The mGCM 1 and hGCM 1 proteins show a high level of similarity in the DNA binding domain, and the transactivation domain shows 61% amino acid identity.
Cellular Localization of GCM 1 in Human and Mouse Placenta
Earlier attempts by in situ hybridization to determine the cell type expressing the mouse Gcm 1 gene identified the chorionic and, in later development, the labyrinthine trophoblast cells of the mouse placenta (Basyuk et al. 1999). The availability of well-characterized antisera allows the extension of this investigation to the protein level in mouse and human placenta. As anticipated from our previous in situ hybridization results (Basyuk et al. 1999), immunomicroscopy revealed intense staining of the chorionic cells and the ectoplacental cone of mouse E8.5 placenta (Figure 3A and 3B). The staining (Figure 3A) was remarkable in that neither the adjacent mesodermal cells nor the embryo proper had detectable staining at this development time. The GCM 1 protein was almost wholly confined to the nuclei of chorionic and trophoblast cells (Figure 3B). The variation of signal strength seen in our previous in situ hybridizaton (Basyuk et al. 1999) was not seen in the immunocytochemical analyses, suggesting that the cellular protein level is more stable than the mRNA level. The cognate structures and cells of the placental villi stained in the human first-trimester placenta (Figure 3C and 3D). Close inspection of the immunomicrographs reveals examples of staining in both the cytotrophoblastic (Figure 3C) and syncytiotrophoblastic layers (Figure 3D) of the villi. As in the mouse, the staining is principally nuclear, but with enough cytoplasmic staining to clearly define the outer two cell layers of the villi. Moreover, GCM 1 immunoreactivity is only weakly revealed on sections of third-trimester placenta (not shown), suggesting that GCM 1 is principally expressed at early stages of cytotrophoblast differentiation.
Chromosomal Location of the hGcm 1 Gene
The full-length hGcm 1 cDNA was used to map the Gcm 1 gene by FISH. We mapped the chromosomal position of Gcm 1 by FISH analyses on metaphase chromosome spreads. Under the conditions used, the FISH detection efficiency was approximately 53% for the hGcm 1 probe (among 100 checked mitotic figures, 53 showed signals). In all cases, the signal was on both chromatid pairs at the same position on the chromosome. The DAPI banding pattern was used to identify the specific chromosome and to estimate the position of the fluorescent signal. The FISH signals (Figure 4A) all mapped to the short arm of chromosome 6 (Figure 4B). Figure 4C shows a histogram summarizing the results of 10 spreads. We conclude from these data that the human Gcm 1 maps to human chromosome 6, region p12. This region of chromosome 6 is known to be syntenic to parts of mouse chromosome 9 (http://www.ncbi.nlm.nih.gov/Omim/Homology/human6.html) where we previously mapped the mouse gene (Basyuk et al. 1999).
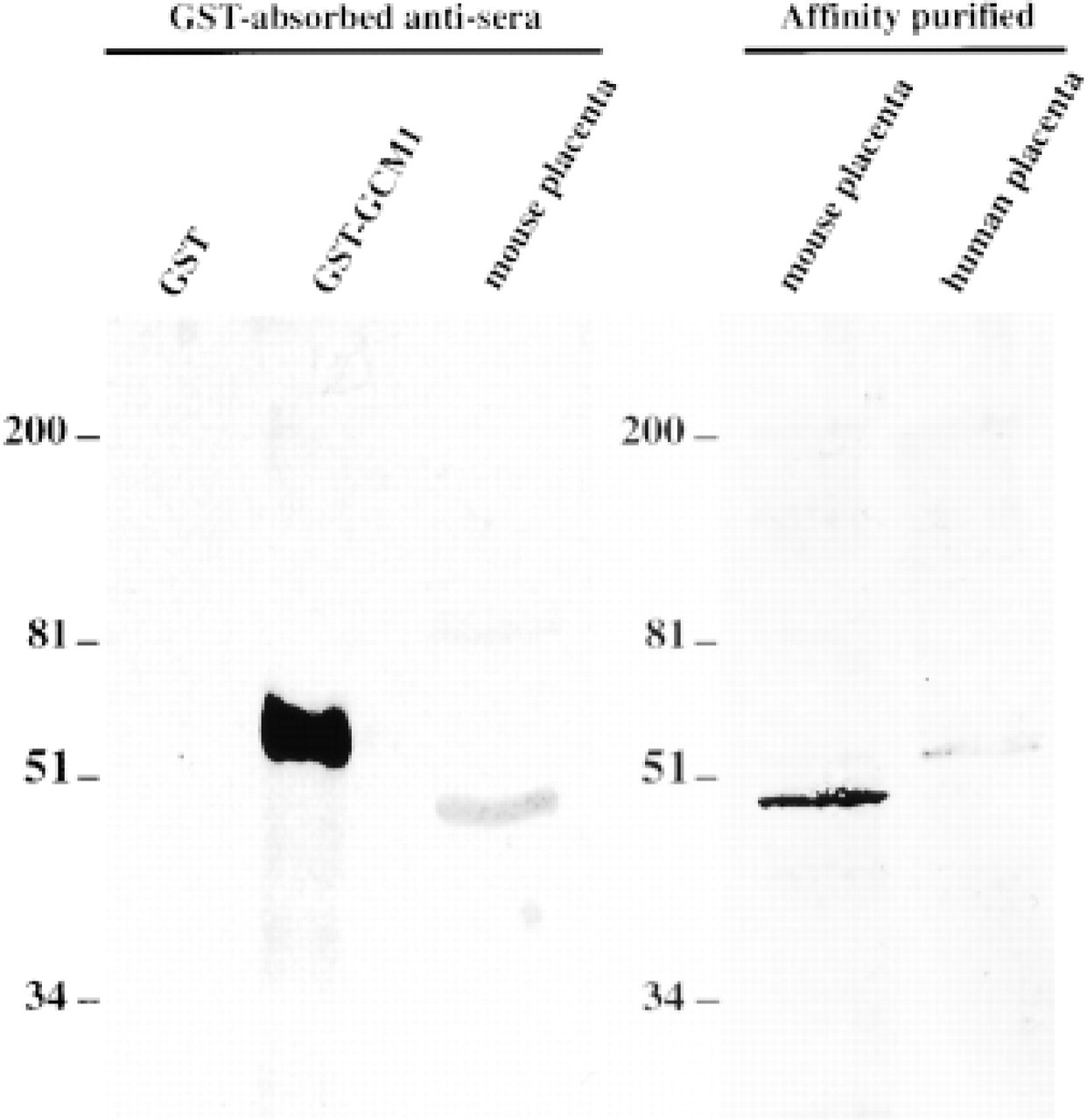
Immunoblot analyses and specificity of the anti-GCM 1 antibody. Western blotting analyses of the anti-GCM 1 (whole serum) preabsorbed over a GST-Sepharose column and after affinity purification. The anti-GCM 1 antibodies recognize a protein band of an apparent molecular weight of 50 kD in E15.5 mouse placenta (Lanes 3 and 4) and a slightly larger protein in third-trimester human placenta (Lane 5). The relative positions of the molecular weight standards run in parallel are indicated. For mouse and human placental extracts, 50 μg of proteins was used.
Discussion
Trophoblast invasion of the maternal uterus during human placentation is a highly complex process that involves cell proliferation, migration, and cell differentiation. Several known factors have been described in human syncytiotrophoblasts, including the Wnt genes (Nusse and Varmus 1992), the Hox gene (Oudejans et al. 1990; Taylor et al. 1997), and the integrin family (Yang et al. 1995; Aboagye-Mathiesen et al. 1996). In this study we have characterized a member of the human Gcm gene family and have demonstrated that the hGcm 1 gene, like its mouse homologue, is also expressed in placenta. These data therefore highlight the broader functions for Gcm genes in mammalian development and complement recent findings demonstrating that rat and mouse Gcm 1 mRNAs are mainly localized in trophoblastic cells (Kim et al. 1998; Basyuk et al. 1999). The in situ hybridization analyses, however, were limited by the low level of Gcm 1 mRNA and the resolution of this technique. To further these studies and circumvent these shortcomings, we have developed well-characterized, highly specific immunoreagents that are suitable for immunomicroscopy.
The mammalian GCM 1 and GCM 2 proteins show striking sequence similarities at their amino termini but no detectable similarities at their carboxyl termini (Akiyama et al. 1996; Kim et al. 1998). Consequently, to investigate the immunolocalization of GCM proteins in situ, we chose to raise antisera to these unique carboxy terminal domains by creating a separate GST fusion protein with each and using them as immunogens. As anticipated, the antibodies were specific and each recognized only the fusion protein used in its creation (data not shown). The specificity of these reagents was further ensured by affinity-purifying each against the GCM peptide portion of the fusion protein used for immunization.
In this communication we present the characterization of the GCM 1 immunoreagent and show that it crossreacts with the cognate human protein. Moreover, we show that the population of cells expressing this protein in placental sections appears larger than that expressing the Gcm 1 mRNA, as revealed by previous in situ hybridization in mouse and rat (Kim et al. 1998; Basyuk et al. 1999) and recently by Northern blotting in human placenta (Janatpour et al. 1999). This apparent discrepancy, which may be attributed to differential metabolic instability of the Gcm 1 mRNA compared to that of the GCM 1 protein and the differences in the sensitivity of the methods, underscores the usefulness of these immunoreagents. The assessment of Gcm 1 expression by in situ hybridation techniques appears to be limited by the instability of the mRNA. Both human and mouse Gcm 1 mRNA are characterized by a large 3′-UTR (Altshuller et al. 1996; Yamada et al. 1999) that contains purine-rich sequence that may explain the rapid decay or instability of Gcm 1 message (Chen et al. 1992). Using the anti-GCM 1 antibody, we have visualized the GCM 1 protein in the chorionic and labyrinthine trophoblasts of mouse placenta. The GCM 1 protein also can be identified in human placenta in both the cytotrophoblasts and the differentiated syncytiotrophoblasts of the first-trimester placental villi. In both animal species, the protein is largely confined to the nucleus but with some cytoplasmic expression as well.
Successful development of the human embryo is largely dependent on the proper specialization of the extraembryonic tissue and the formation of a functional placenta. The individual roles of transcription factors such as Gcm 1 in placental development are largely unknown, and only recently have efforts been made to identify the key players in the transcriptional cascades that control trophoblastic differentiation (Janatpour et al. 1999). For example, Janatpour and colleagues (2000) have recently demonstrated that cytotrophoblast cells that constitutively express the
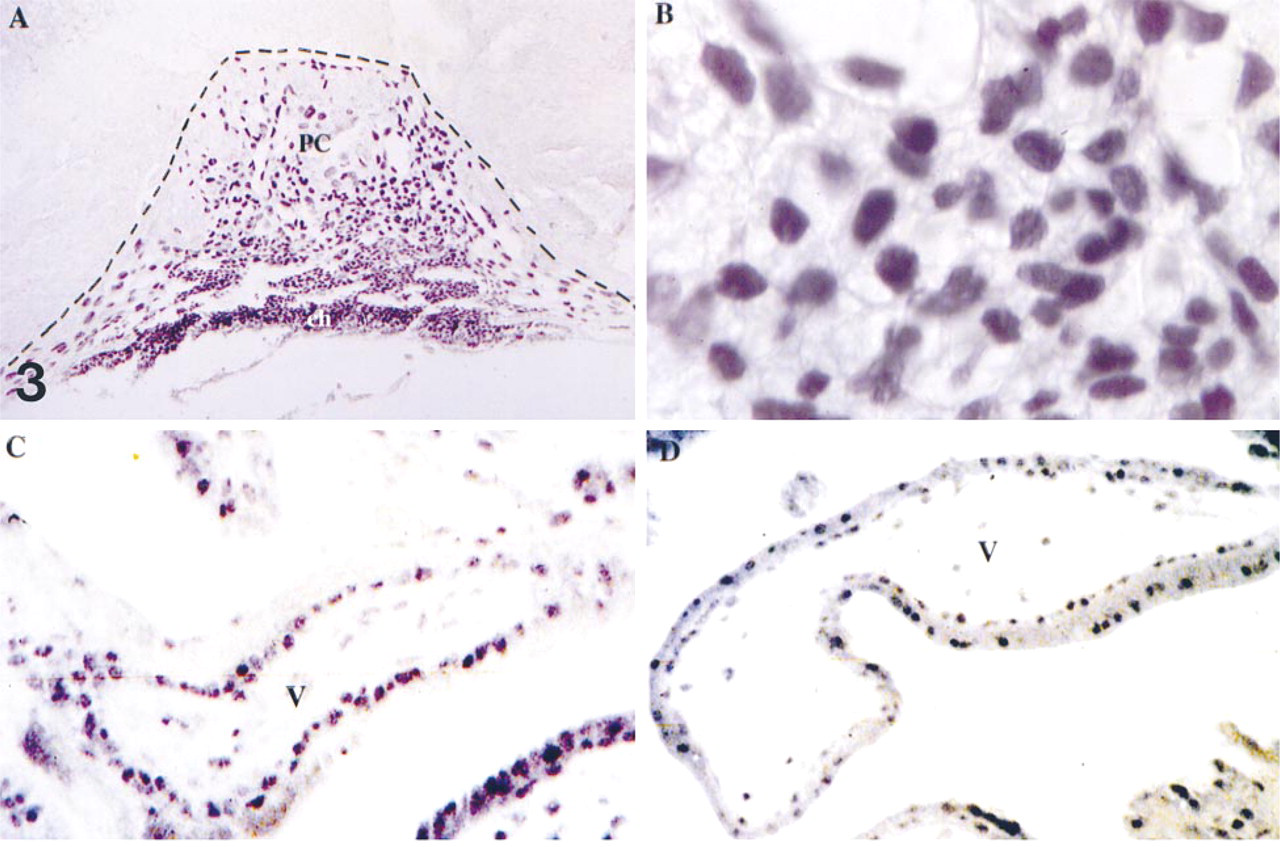
GCM 1 is expressed in mouse and human trophoblast cells. Immunolocalization of GCM 1 in trophoblast cells of E8.5 mouse placenta (
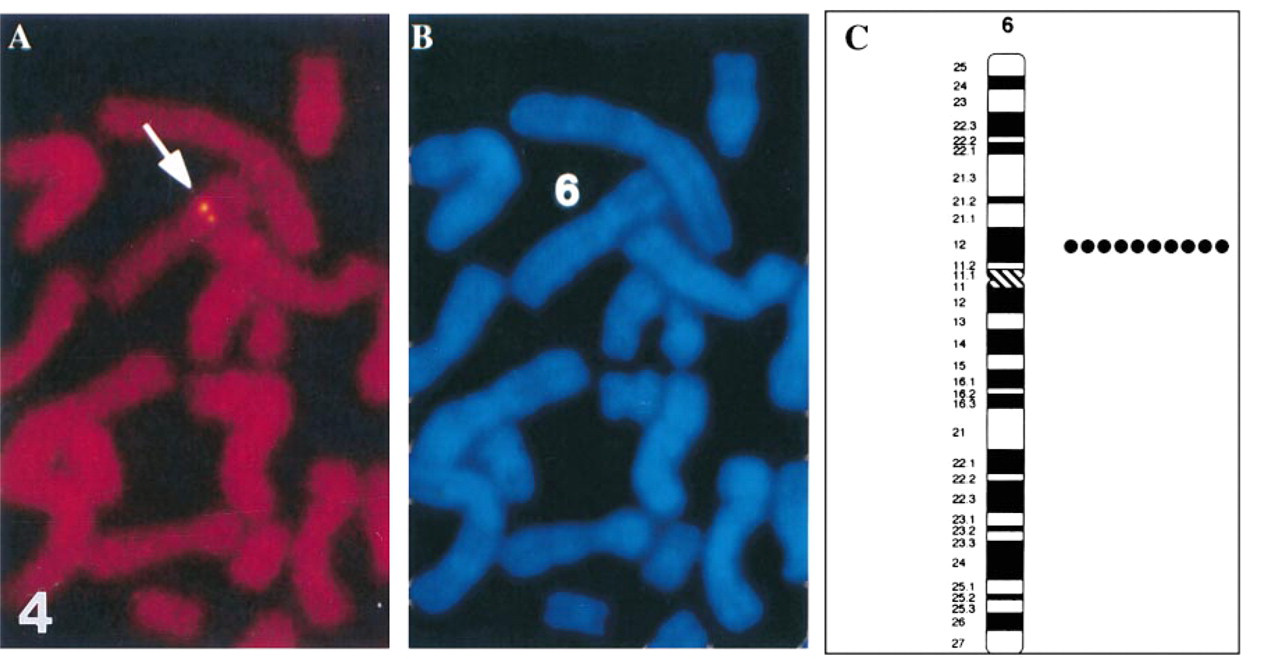
FISH mapping of the Gcm 1 on human chromosome 6. (
In this study we have mapped the Gcm 1 gene to human chromosome 6p12 but have not identified any known genetic diseases of placentation that map to this region. The hGcm 1 was previously mapped to chromosome 6 by physical mapping of ESTs to the interval D6S427–D6S294. This broad interval of chromosome 6 appears to include the band 6p12, the position to which we mapped the gene. Therefore, the FISH mapping data reported here are a higher resolution and refine the chromosomal localization of Gcm 1. These data also are consistent with the mapping of the mouse gene to mouse chromosome 9, band E1–E3 (Basyuk et al. 1999) because this region is syntenic to part of human chromosome (http://www.ncbi.nlm.nih.gov/omim/Homology/human6.html). Interestingly, the human Gcm 2 gene also maps to chromosome 6 but at 6p23, a position at some distance from the site of Gcm 1. Nevertheless, the proximity of the two genes suggests that some ancestral intrachromosomal duplication may account for this proximity.
Footnotes
Acknowledgements
Acknowledgments
We gratefully acknowledge the participation of Eugenia Basyuk and Christerfer Demarco in the early phases of the work. We also thank Nancy Amalbert for assistance in assembling and editing this manuscript.
