Abstract
Background
Hypertensive disorders of pregnancy are defined as syndromes characterized by high blood pressure that develop after 20 weeks of pregnancy. The pathogenesis of hypertensive disorders of pregnancy has not been fully elucidated, and the most effective treatment is the termination of pregnancy. Therefore, methods for an early predictive diagnosis should be developed to rescue both the mother and the child.
Methods
Previously, we explored the serum peptides whose concentration varied specifically between patients with hypertensive disorders of pregnancy and normal pregnant women using peptidomics analysis and identified seven candidate marker peptides. To quantify the marker peptides more reliably, we attempted to quantify these peptides simultaneously by multiple reaction monitoring using liquid chromatography/tandem mass spectrometry (LC-MRM/MS). Non-labelled and stable isotope-labelled forms of the seven peptides were synthesized as standards, and the multiple reaction monitoring transitions for their quantification were determined.
Results
As the retention of the peptides by the reversed-phase column was dependent on their hydrophobicity, two solvent compositions were required for the retention of all peptides. Under these conditions, we detected the peptides by LC-MRM/MS using a column-switching method. Further, we succeeded in quantifying the peptides in the serum of pregnant women using stable isotope dilution.
Conclusion
Our new peptidomics method has great value for peptides, particularly those that have no specific antibody as a detection tool. Using this system, the serum peptides in patients with hypertensive disorders of pregnancy can be validated as diagnostic markers of hypertensive disorders of pregnancy. Further, this method can potentially be applied to the general quantification of other serum peptides.
Keywords
Introduction
Hypertensive disorders of pregnancy (HDP) are syndromes associated with pregnancy that commonly affect maternal, fetal and neonatal morbidity/mortality.1–4 Despite various efforts to overcome these syndromes, especially the disease known as preeclampsia, they constitute a broad spectrum of life-threatening disorders that cannot be adequately managed. 5 From a biological perspective, this physiological phenomenon of pregnancy (or the fetal parasitic invasion into maternal body via the placenta) is a reproductive strategy restricted to mammals, and HDP may inevitably cause disasters in humans, who have relatively long pregnancy compared with other mammals. The pathophysiology of the syndromes, which were identified in the era of Hippocrates in fourth century BC, is complicated, and there is still no established therapy for the so-called toxaemia of pregnancy. 6
To understand the pathophysiology of the syndrome more precisely, we previously examined the serum from patients with HDP using peptidomics analysis (BLOTCHIP-mass spectrometry [MS] 7 ). 8 This method is a one-step direct-transfer technology for matrix-assisted laser desorption/ionization (MALDI)-time-of-flight (TOF)/MS analysis. Compared with conventional liquid chromatography/tandem mass spectrometry (LC-MS/MS), this technology can directly analyse almost all peptides in clinical samples that are capable of binding to abundant resident proteins.8,9
Based on data obtained from the peptidomics methods described above, we found changes in several peptides in the serum of HDP patients, and established simple immunological quantification methods (enzyme-linked immunosorbent assays [ELISAs]) for candidate disease biomarkers (DBMs) for further validation studies.8,10 However, some peptides are unsuitable for detection by ELISA because the production of specific antibodies (Abs) is not always technically easy in some cases. In fact, we encountered the difficulty in producing antibodies that specifically recognized the peptide P-2858. 10 Therefore, we developed an alternative quantification methodology for a candidate serum HDP marker peptide (a peptide, difficult to produce specific Abs) using multiple reaction monitoring (MRM) mass spectrometry. 11
To evaluate the correlation between the concentration of candidate DBM peptides and the pathophysiology of HDP more precisely, we attempted to develop a method for measuring seven candidate DBM peptides 8 simultaneously using an LC-MRM/MS system. As the BLOTCHIP analysis applied in our previous study is a MALDI-TOF/MS-based system, the ions of the peptides generated differed from those generated using LC-electrospray (ESI)/MS performed in this study. Therefore, we used standard synthetic peptides for each peptide. Based on experimental data obtained for each synthetic peptide, we established a quick, reliable MRM method for quantifying the peptides.
Materials and methods
Reagents
Acetonitrile, formic acid (FA), methanol, 2-propanol (all LC-MS grade) were obtained from Wako Pure Chemical Industries, Ltd (Osaka, Japan). Trifluoroacetic acid (TFA) and normal mouse serum were also from Wako. MonoSpin C18 spin column and GL-tip GC for peptide preparation were from GL Sciences, Inc. (Tokyo, Japan).
Peptides
Non-labelled (NL) and stable isotope-labelled (SI) forms of peptides selected as the DBM candidates 8 were synthesized (Scrum, Inc., Tokyo, Japan). SI peptides were designed enough to be separated from the NL peptides on mass spectrometry. The nomenclature and sequence of each peptide and its source proteins were shown (Supplementary Table S1). The nomenclature of each peptide was based on its molecular weight originally detected in the previous report. 8 The symbols of SI peptides and the position of SI amino acids were indicated by underbars (Supplementary Table S2). Each lyophilized peptide was dissolved at a concentration of 1 mg/mL in 0.1% FA and stored at –80°C until use. The solution for diluting each peptide was 10% acetonitrile, 0.1% FA containing the peptide fraction of the normal mouse serum.
Blood samples
The human serum sample was prepared according to the protocol approved by the Ethic Committee board members of Juntendo University and Yamagata Saisei Hospital (registration #19–017 and #144), respectively. The serum sample was stored at –80°C until use.
MRM condition
The values of mass-to-charge ratio (m/z) of precursor ions of each synthetic peptide were confirmed using QqTOF type LC-MS/MS system (LC: DiNa, KYA technologies Corporation, Tokyo, Japan, MS: QSTAR Elite, SCIEX, Framingham, MA, USA), and dominant fragment ions generated from the precursor ions by collision-induced dissociation (CID) were determined as described previously. 11 Based on the information of fragment ions, Q1 mass and Q3 mass for MRM transition were selected for each peptide. Other parameters, collision energy (CE), entrance potential (EP), declustering potential (DP), and collision cell exit potential (CXP), were optimized in QTRAP6500 LC-MS system (SCIEX, Framingham, MA, USA) by infusion analysis of each synthetic peptide.
Preparation of peptides from serum
One hundred microlitres of each serum sample were spiked with appropriate concentration of SI standard peptides and heated at 100°C for 10 min. After cooling on ice for 10 min, the sample was centrifuged at 20,400 × g at 4°C for 60 min. The supernatant was applied to a reverse-phase spin column MonoSpin C18, and the flow through fraction was pooled. Bound hydrophobic materials to the C18 spin column were eluted with 200 μL of 80% acetonitrile, 0.1% FA solution, and the eluted sample was evaporated. The sample was reconstituted with 2% acetonitrile, 0.1% TFA solution for LC-MS analysis. The four peptides (P-2127, P-2378, P-2858 and P-3156) were mainly recovered in this sample.
To the flow through fraction from the C18 spin column, which contained hydrophilic materials, one-tenth volume of 1% TFA was added. The sample was applied to a graphite carbon GL-tip GC device and bound materials were eluted with 60 μL of 80% acetonitrile, 0.1% TFA solution. After evaporation, the sample was dissolved in 2% acetonitrile and 1% TFA solution for LC-MS analysis. The three peptides (P-2081, P-2091 and P-2209) were mainly recovered in this sample.
Quantitative LC-MRM/MS analysis
We use nano-flow liquid chromatography (Eksigent 415, SCIEX, Framingham, MA, USA) coupled with a triple quadrupole type mass spectrometer with a linear ion trap, QTRAP6500. The peptide sample was separated with a reverse-phase C18 LC system (HiQ Sil C18W-3, the trap column; 0.5 mm I.D. × 1 mm length, the ESI analytical column; 150 μm I.D. × 10 cm length, KYA Technologies Corporation, Tokyo, Japan). A sample was loaded to a trap column, and the trap column was switched into the flow path of the analytical column after washing. The trap column was equilibrated with 0.1% FA for the conventional analysis or 0.1% TFA for the high retention at a flow rate of 2 μL/min. The ESI analytical column was equilibrated with 2% acetonitrile, 0.1% FA at a flow rate of 300 nL/min. Gradient elution was performed using 0.1% FA (solvent A) and acetonitrile with 0.1% FA (solvent B). The peptides were eluted from the columns by the linear gradient of 2–50% solvent B for 20 min at a flow rate of 300 nL/min. The columns were washed by solvent B and equilibrated under the initial condition for repeated use. Between each analysis, the washing program for cleaning columns was run to remove carry-over peptides with 5 μL of washing solution (acetonitrile/2-propanol/methanol/1% FA, 25% each).
The peptides were ionized in the positive ion mode using Nanospray III ionization source (SCIEX). Each peptide was detected by MRM scans. The MRM transitions (precursor/product) for detecting the peptides were established (Table 1).
MRM transition for detecting each peptide.
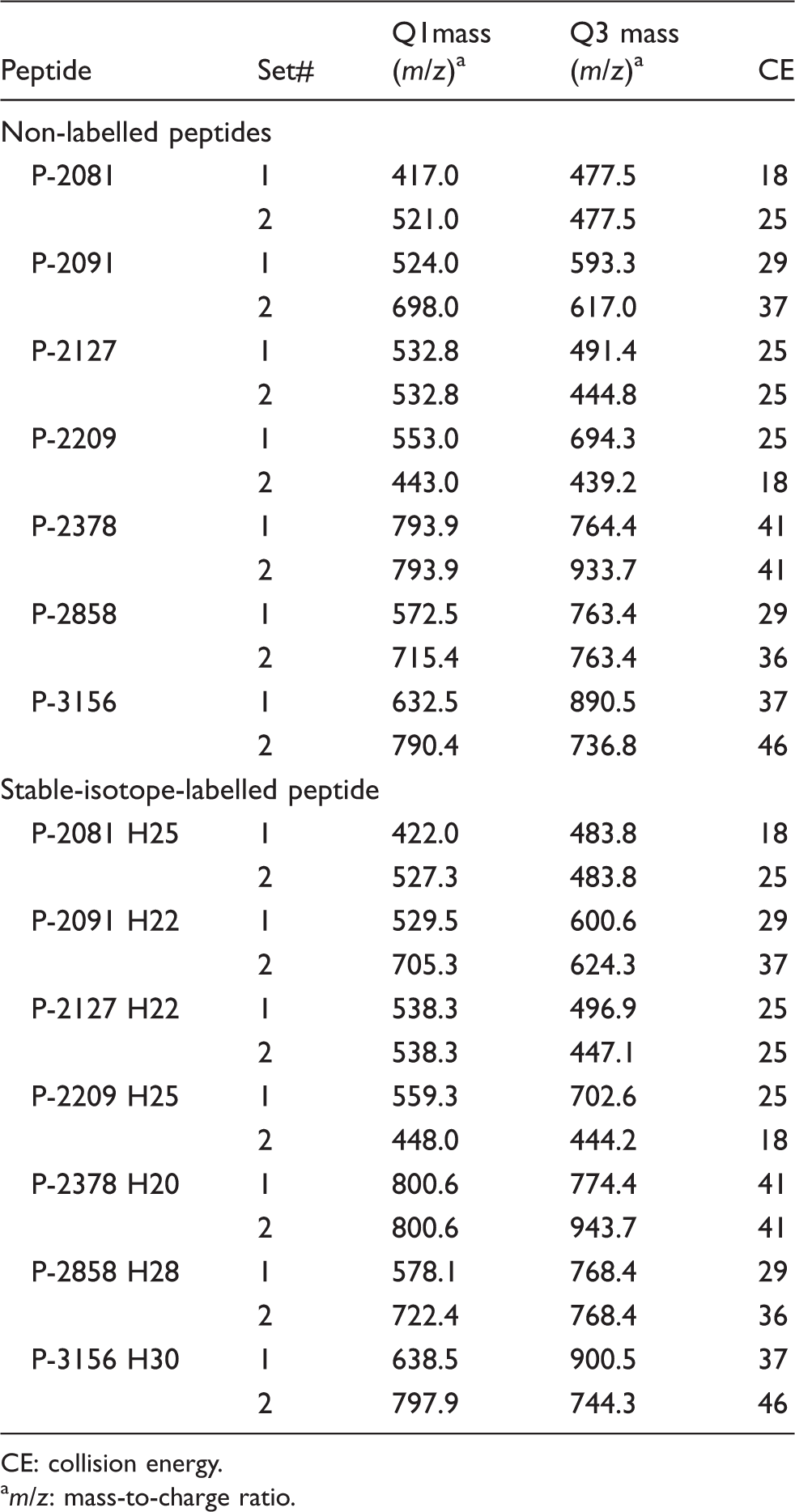
CE: collision energy.
am/z: mass-to-charge ratio.
The MultiQuant 3.0 software (SCIEX) was used for quantifying peptide concentration. The chromatographic peak area of each peptide was calculated with the MultiQuant software MQ4 algorithm. The peptide concentration was measured by calculating the ratio of the peak areas of the NL and SI peptide.
Prediction of hydrophobicity of peptides
The hydrophobicity profile of each peptide was predicted from the amino acid sequence by the method of Kyte and Doolittle using GENETYX software (GENETYX Corporation, Tokyo, Japan). 12
Recovery test of peptides
The fixed amount of SI peptides and various amounts of NL peptides were added to the control normal mouse serum that did not contain the target peptides. The peptide fraction was extracted as described above and analysed using LC-MRM/MS. The calibration curves were built by linear regression analysis between the logarithmic value of the measured concentration and that of the expected concentration using the least-squares method.
Results
Determination of the MRM detection condition
The molecular ions of each NL peptide produced multiple protonated positive ions and the selected dominant precursor ions were fragmented by CID. Q1 mass, Q3 mass, and CE for the MRM transition were determined for each peptide (Table 1). The MRM transition of each SI peptide was determined based on the fragmentation pattern of the corresponding NL peptide. We confirmed that the transition worked well. Other machine-specific parameters (EP, DP and CXP) were determined by an automatic program (data not shown). Two sets of MRM transition parameters were obtained for each peptide (Table 1). The dominant set (#1) was used for quantification and the other set (#2) was used to confirm the peptide.
Simultaneous analysis of seven synthetic peptides
As an initial trial, we tried to separate the seven synthetic peptides (1 pmol each) using single-step reverse-phase C18 LC with 0.1% FA and acetonitrile as solvents and subjected the result online to MS to detect each peptide by MRM. Five peptides (P2091, P-2127, P-2378, P2858 and P-3156) were successfully detected, whereas P-2081 and P-2209 were not detected under our conventional nano LC conditions using acetonitrile and 0.1% FA as solvents (Figure 1(a)).
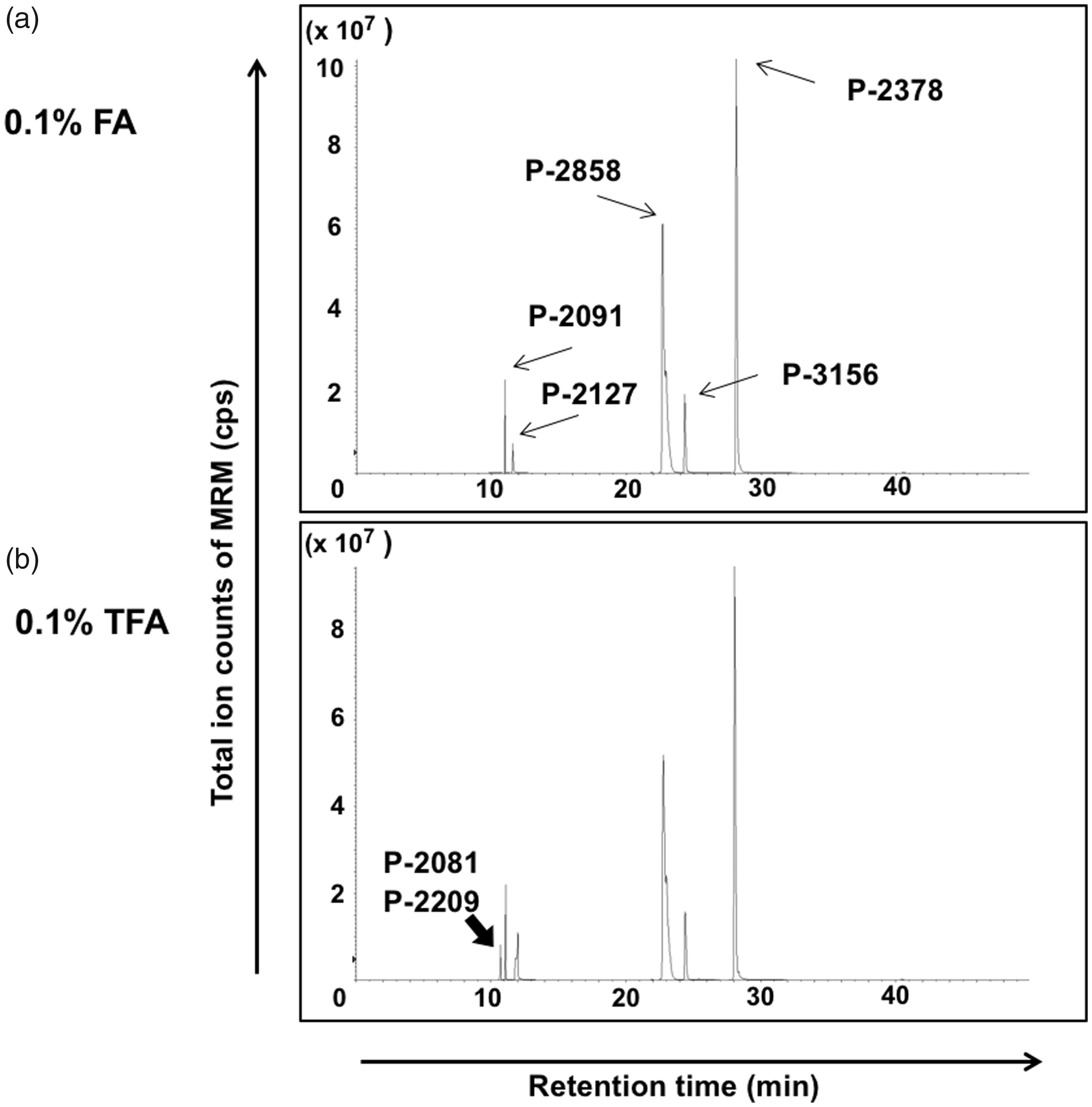
Difference of peptide retention by solvent condition of the sample. Horizontal axis: Retention time on LC (min), Vertical axis: Total ion counts of MRM (cps; counts per second). Seven synthetic peptides were separated by a reverse-phase C18 LC with 0.1% formic acid (FA) and acetonitrile as solvents and introduced online to MS to detect each peptide by MRM. The acid composition in the injected sample solution was 0.1% FA (a) or 0.1% trifluoroacetic acid (TFA) added to 0.1% FA (b). (a) Arrows: the peaks of detected five peptides (P2091, P-2127, P-2378, P2858 and P-3156). (b) The peak indicated by the bold arrow was detected after addition of 0.1% TFA to the sample solution. This peak contained two peptides (P-2081 and P-2209).
Therefore, we hypothesized that these two peptides were not retained by the reverse-phase C18 column under these condition and examined the predicted hydrophobicity of the seven peptides. Compared with the five other peptides, the two undetected peptides lacked hydrophobic cluster that contributed to binding to the hydrophobic resin (Supplementary Figure S1).
To facilitate the retention of peptides in a reverse-phase column, TFA is generally used as an ion pair reagent. However, TFA is usually avoided in LC-MS because it suppresses the electrospray ionization. Therefore, we used TFA only for the injected samples and the trap column, and not in the analytical column, from which the eluate was introduced directly to the nanospray ionization source. As a pilot trial, a TFA concentration of 0.1% was used in both the sample and trap column. Under these conditions, peptides P-2081 and P-2209 were detected at almost the same retention time (Figure 1(b), bold arrow). Therefore, all of the synthetic standard peptides could be detected (Figure 1(b)). Although the detection intensity of the hydrophilic peptides was still lower than that of the hydrophobic peptides, TFA retained the two hydrophilic peptides in the reverse-phase column.
Two-step analysis of peptides
Next, we evaluated the influence of the TFA concentration in the sample on the detection intensity, varying the TFA concentration from 0.1% to 1.5%. The optimal TFA concentration was 1.0% for the detection of the four hydrophilic peptides (P-2081, P-2209, P-2091 and P-2127 (Figure 2(a)), whereas the detection of the three hydrophobic peptides (P-2378, P-2858 and P-3156) decreased as the TFA concentration in the sample increased (Figure 2(b)). Therefore, to improve the detection efficiency for each peptide, the solvent conditions should be based on the hydrophobicity of the peptides.
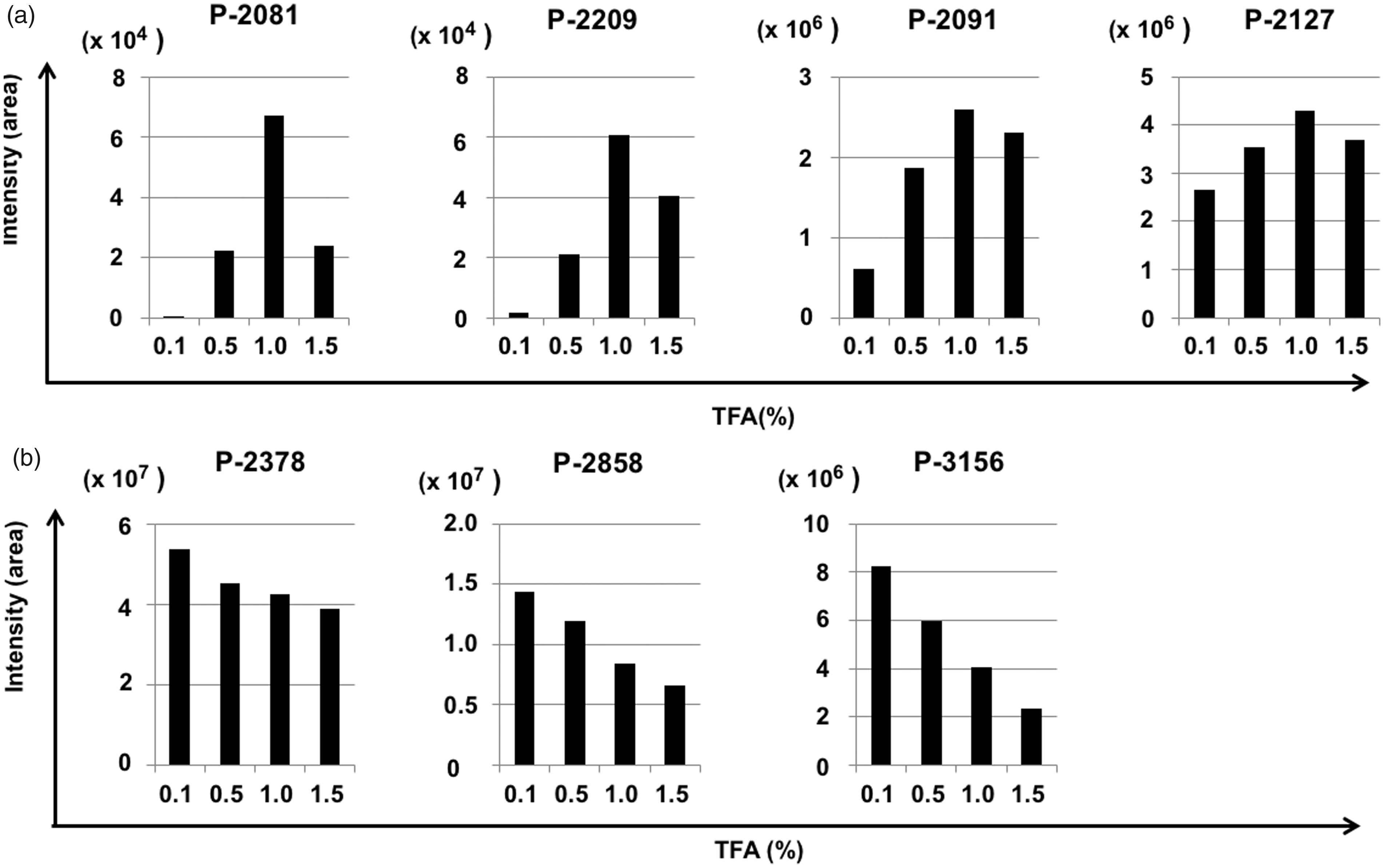
Influence of the TFA concentration in the sample on the detection intensity. The change of detection intensity by TFA concentration in the sample solution. (a) Four hydrophilic peptides (P-2081, P-2209, P2091 and P-2127), (b) three hydrophobic peptides (P-2378, P2858 and P-3156). Vertical axis: the detection peak area of each peptide. Horizontal axis: TFA concentration (%).
As the optimal TFA concentration in the sample for detecting peptides was dependent on their hydrophobicity, we applied a two-step analysis with different TFA concentrations. The peptides P-2127, P-2378, P-2858 and P-3156, were mainly recovered in MonoSpin C18-bound fraction, whereas P-2081, P-2091 and P-2209 were recovered from the flow through fraction of the C18 resin using graphite carbon resin (GC). For the C18-bound fraction, the optimal TFA concentration for peptides P-2378, P-2858 and P-3156 was 0.1%, whereas that of P-2127 differed. Nevertheless, a TFA concentration of 0.1% was sufficient for detecting peptide P-2127 (Figure 2(a)).
Accordingly, we used 0.1% TFA as an acid condition in the C18-bound peptide sample. For the hydrophilic peptides P-2081, P-2091 and P-2209 prepared with GC, 1% TFA was used as the acid condition in the sample. Further, to increase the sensitivity of detecting hydrophilic peptides, a concentrated sample was loaded in the analytical system. With this protocol, the peptides with a wide range of hydrophobicity could be analysed with one set of the LC-MS apparatus, and these technical improvements allowed us to quantify each peptide successfully.
Recovery test
To confirm the performance of the two-step analysis method described above, we performed a recovery test. Normal mouse serum spiked with each NL peptides at various concentrations was processed following the quantification protocol. Peptides P-2091, P-2209, P-2378 and P-2858 were recovered with good linearity at concentrations of 0.1 to 300 ng/mL, whereas peptides P-2081, 2127 and P-3156 were recovered with good linearity at concentrations of 1 to 300 ng/mL (Figure 3(a) and (b)).
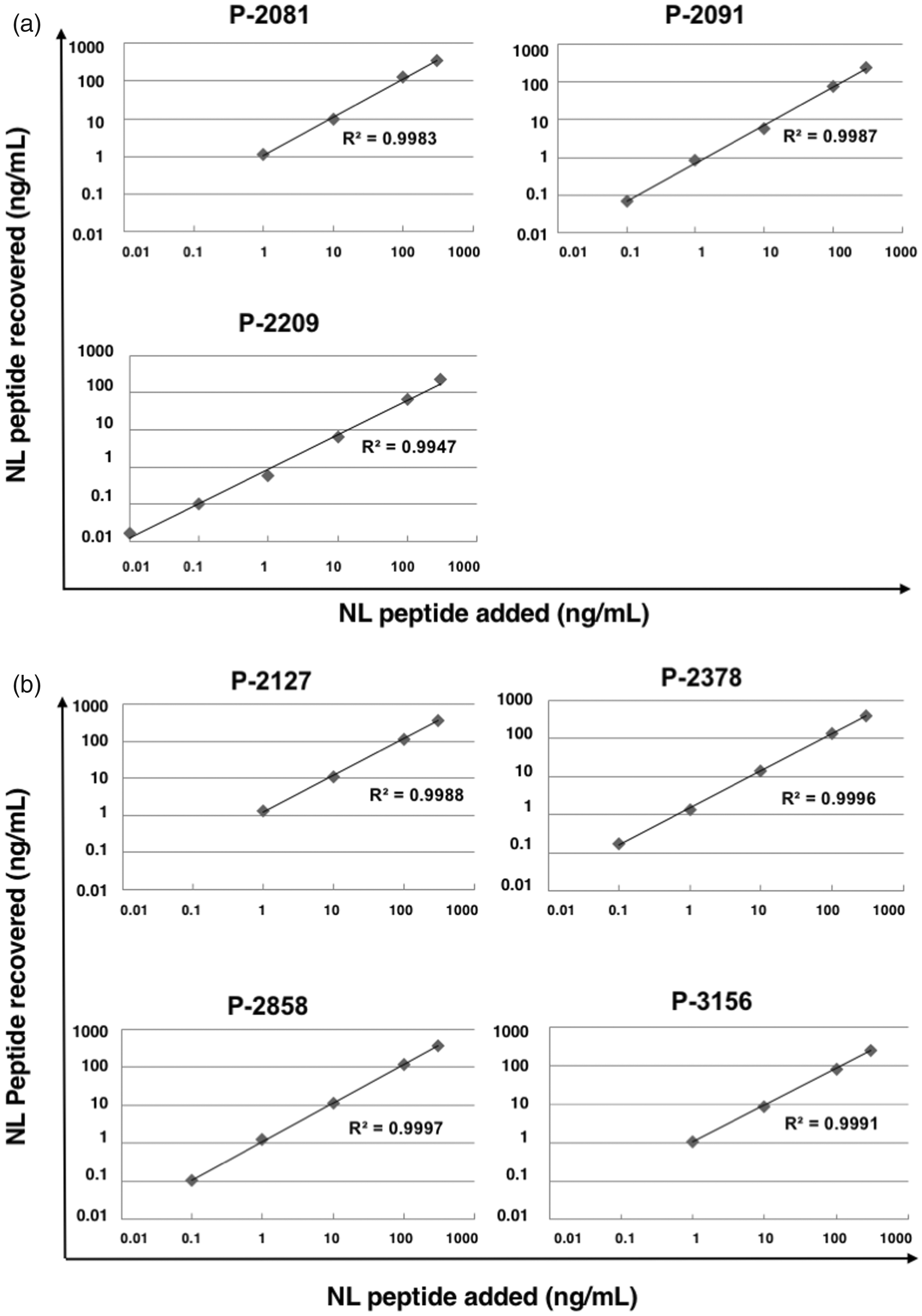
Recovery test of peptides. The fixed amount of SI peptides and various amounts of NL peptides were added to the control normal mouse serum. The peptide fractions prepared with a GC tip column (a) and with a C18 tip column (b) were analysed by LC-MRM/MS. Horizontal axis: the concentration of the NL peptide added, vertical axis: the concentration of the measured NL peptide calculated by the detection intensity of the SI peptide.
Quantification of seven peptides in the serum of pregnant women using SI peptides
A fixed amount of each SI peptide was added to serum from a healthy pregnant woman, and the extracted peptide fraction was analysed using LC-MRM/MS. Each internal standard SI peptide was detected and the corresponding native peptides extracted from the serum were detected at the same retention time on LC (Figure 4). We measured the concentration of the native peptide in the serum by calculating the ratio of the peak areas of the native and SI peptides. Of the peptide, P-2858 had the highest concentration and the P-2127 the lowest in this sample.
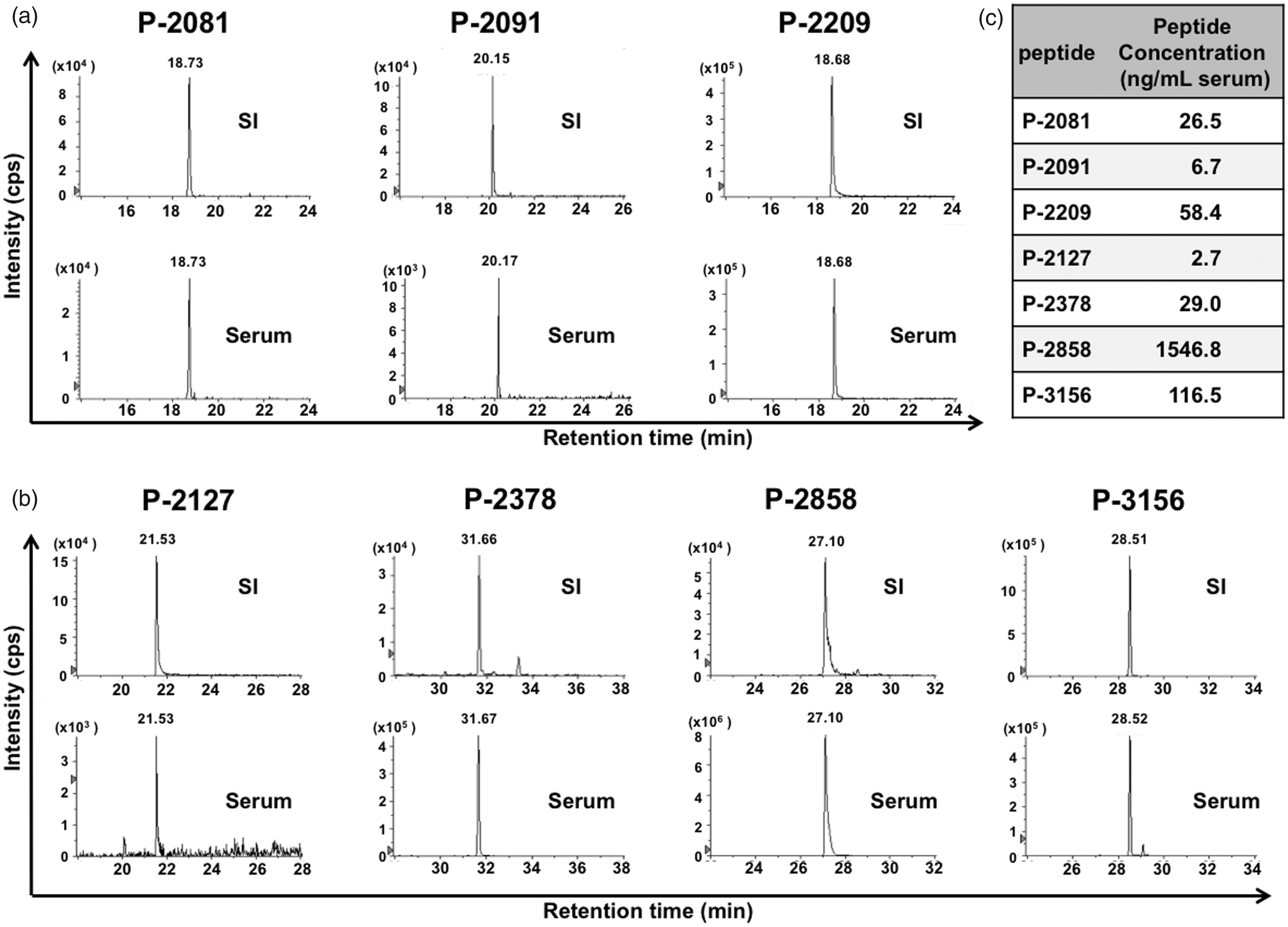
Quantification of seven peptides in the serum of a normal pregnant woman using SI peptides. The fixed amount of each SI peptides was added to the serum of a healthy pregnant woman. The peptide fractions were prepared with a GC tip column (a) and with a C18 tip column (b), and analysed using LC-MRM/MS. Horizontal axis: retention time, vertical axis: detection intensity. The concentration of each native peptide in the serum was calculated with the intensity of SI peptide (c).
Further, we confirmed that we could measure the concentration of each peptide in the serum of patients with HDP using LC-MRM/MS. Analytical data presented by box and whisker plots, obtained from 11 samples of HDP patients and 13 samples of normal pregnant women (these serum samples used were the same as those analysed previously by the peptidomic method using BLOTCHIP system 8 ) showed almost comparable results (Supplementary Figure S2) to our previous study. 8
Comparison between LC-MRM/MS and ELISA
To confirm the comparability between LC-MRM/MS and ELISA, we measured one of the candidate DBM peptides, P-2209, in the serum of patients with HDP both by LC-MRM/MS and by ELISA. Since we established the specific ELISA system to measure this peptide for clinical samples in our previous study, 10 P-2209 would be a good model peptide to compare two methods. Data obtained from these experiments showed a proportional relationship (R2= 0.77) between both methods (Supplementary Figure S3).
Discussion
In this study, we first established a reliable method for quantifying candidate DBM serum peptides simultaneously in HDP patients using LC-ESI/MS analysis. As these peptides were originally found using a MALDI-TOF/MS technique, 8 the binding property of the peptides to the reverse-phase LC column and their ionization efficiency on ESI were unknown at the beginning of this study. Indeed, two target peptides did not bind to the C18 column under conventional LC-ESI/MS conditions, and the ionization efficiency of these peptides was very low. By using TFA as a counter ion, the binding of the peptides to the column was improved. In addition, we concentrated these hydrophilic peptides using a GC tip column and succeeded in quantifying them. In this process, we observed that the predicted hydrophobicity of the peptides was useful for confirming the peptides.
Although LC-MRM/MS is useful for simultaneous analysis of the multiple peptides in each sample, this method also includes potential disadvantage as follows; (1) the sample should be prepared by prefractionation and desalting to suit the system. (2) The measurement of many samples should perform one by one, not in parallel. Consequently, the LC-MRM/MS possibly shows lower throughput in measurement of a great number of samples compared to other detection system such as ELISA.
ELISA is typically used to quantify proteins in serum, and there are many commercial ELISA kits. However, as the native peptides in body fluids have specific structures, commercial ELISA kits are often unsuitable for the exact measurement of native peptides. Further, it is sometimes difficult to make specific antibodies to the native peptides. 11 To overcome these problems, MS analysis using the MRM method described here is effective for molecular quantification. Once a peptide fragment is ionized and detected by MS, in principle, the peptide can be quantified using the MRM method. 13 Using the m/z values of the precursor and the fragment ions coupled with the retention time on LC, the MRM signal showed high specificity for detecting the targeted peptide. This is a strong advantage of the MRM method for measuring native peptides compared with conventional methods such as ELISA. 14 In addition, we confirmed the comparability between LC-MRM/MS and ELISA methods (Supplementary Figure S3).
Some peaks might be detected incorrectly in the same transition with the MRM method, as there are a number of peptide species in the serum-derived samples. Considering this possibility, we confirmed the actual multiple MRM transitions and retention time in LC to prevent peaks from being misassigned by using synthetic peptides as standards. Because the retention times of the native peptides exactly matched those of SI peptides added as internal standards, misassignment of the peaks was reduced.
At present, HDP is a major leading cause of maternal, fetal and neonatal morbidity/mortality worldwide. The syndrome, as the most common and serious complication of pregnancy, is considered to be a multifactorial disorder, and the major pathophysiological disorder involves vascular endothelial damage leading to blood vessel spasms. 15 To date, it is widely accepted that numerous pathological abnormalities observed in the placentas of HDP patients have been suggested to be responsible for the syndrome. In some cases, such as inadequate differentiation and invasion of the trophoblast, dysfunction of the placental endothelium, immune maladaptation to paternal antigens and exaggerated systemic inflammatory response, are causes of eclampsia, a severe form of HDP. 16 The reduction of placental perfusion that causes the release of stress factors into the mother’s circulatory system and generalized maternal endothelial dysfunction likely results in multiple pathological changes as well as clinical manifestations of HDP.17,18 This hypothesis was supported by a study, indicating that hypoxia stimulates soluble Fms-like tyrosine kinase-1 (sFlt-1) production by placental cytotrophoblasts, 19 and that exogenous administration of sFlt-1 to pregnant rats leads to the development of hypertension and proteinuria. 20 However, the precise mechanism, especially concerning the onset of HDP, remains unknown.
Because the definition of the syndrome is based on blood pressure, the pathological disorder is already present at the time of diagnosis. An early diagnosis before the onset of hypertension is essential for preventing HDP. Many researchers have searched for DBMs that predict HDP. Although several DBMs of HDP have been proposed such as sFlt-1 20 and placental growth factor, 21 these molecules are still insufficient to predict the disease onset. Using the methods described here, we could potentially evaluate the native peptides in the serum of pregnant women with and without HDP to confirm the HDP DBMs. A follow-up study during pregnancy is necessary to establish DBMs that predict HDP.
Our method could be expanded to large-scale peptidomics analyses of other diseases. In fact, a similar approach was used to profile the plasma peptidome in cancer patients. 22 Our novel approach should enable the efficient evaluation of candidate DBM peptides.
Conclusion
Using LC-MRM/MS analysis, we established a method for simultaneous quantification of the seven peptides reported previously as candidate DBM for HDP. Depending on the LC conditions, two-step analysis may be applicable to a wide range of native peptides in body fluids of different hydrophobicities with differing retention in conventional reverse-phase columns. Our new peptidomics analysis method overcomes the inefficiency of establishing ELISA methods for evaluating candidate DBM peptides. This approach should contribute to the identification of essential DBM.
Supplemental Material
Supplemental Material1 - Supplemental material for The simultaneous quantification of candidate serum biomarker peptides for hypertensive disorders of pregnancy
Supplemental material, Supplemental Material1 for The simultaneous quantification of candidate serum biomarker peptides for hypertensive disorders of pregnancy by Mitsuaki Yanagida, Kensuke Hamamura, Kenji Takamori and Yoshihiko Araki in Annals of Clinical Biochemistry
Supplemental Material
Supplemental Material2 - Supplemental material for The simultaneous quantification of candidate serum biomarker peptides for hypertensive disorders of pregnancy
Supplemental material, Supplemental Material2 for The simultaneous quantification of candidate serum biomarker peptides for hypertensive disorders of pregnancy by Mitsuaki Yanagida, Kensuke Hamamura, Kenji Takamori and Yoshihiko Araki in Annals of Clinical Biochemistry
Supplemental Material
Supplemental Material3 - Supplemental material for The simultaneous quantification of candidate serum biomarker peptides for hypertensive disorders of pregnancy
Supplemental material, Supplemental Material3 for The simultaneous quantification of candidate serum biomarker peptides for hypertensive disorders of pregnancy by Mitsuaki Yanagida, Kensuke Hamamura, Kenji Takamori and Yoshihiko Araki in Annals of Clinical Biochemistry
Supplemental Material
Supplemental Material4 - Supplemental material for The simultaneous quantification of candidate serum biomarker peptides for hypertensive disorders of pregnancy
Supplemental material, Supplemental Material4 for The simultaneous quantification of candidate serum biomarker peptides for hypertensive disorders of pregnancy by Mitsuaki Yanagida, Kensuke Hamamura, Kenji Takamori and Yoshihiko Araki in Annals of Clinical Biochemistry
Supplemental Material
Supplemental Material5 - Supplemental material for The simultaneous quantification of candidate serum biomarker peptides for hypertensive disorders of pregnancy
Supplemental material, Supplemental Material5 for The simultaneous quantification of candidate serum biomarker peptides for hypertensive disorders of pregnancy by Mitsuaki Yanagida, Kensuke Hamamura, Kenji Takamori and Yoshihiko Araki in Annals of Clinical Biochemistry
Footnotes
Acknowledgements
We thank the clinical staff of Yamagata Saisei Hospital and Juntendo Urayasu Hospital for serum collection used in this study. We gratefully acknowledge Ms. Masako Miura and Ms. Chihiro Kobayashi for their excellent technical assistance.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This research was supported in part by Japan Society for the Promotion of Science Grants-in-aid #15K10686, 17K19719 and 17H04337, and in part by Matching Fund Subsidy for Private Universities from the Ministry of Education, Culture, Sports, Science and Technology, Japan.
Ethical approval
Ethic Committee board members of Juntendo University and Yamagata Saisei Hospital (registration #19–017 and #144).
Guarantor
YA.
Contributorship
MY conceived and designed the study, collected data, performed data analysis and interpretation, provided financial support and wrote the article. KH conceived and designed the study. KT provided financial support and directed the study. YA conceived, designed and supervised the study, provided financial support and wrote the article. All authors have given approval to the final version of the article.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
