Abstract
Background
The clinical utility of serum thyroglobulin in the follow-up of patients with differentiated thyroid carcinoma may be compromised by the presence of endogenous antithyroglobulin antibodies. To prevent interference by antithyroglobulin antibodies several groups have developed real-time PCR-based assays for quantification of blood thyroglobulin mRNA levels. For accurate quantification of thyroglobulin mRNA in blood preanalytical factors must be recognized and controlled. In this study, we evaluate the effect of different blood RNA stabilizing systems – the Tempus Blood RNA system and the PAXgene Blood RNA system – and storage time on RNA yield and quality, and thyroglobulin mRNA stability.
Methods
Blood samples from 11 patients previously treated for differentiated thyroid carcinoma were collected in K2-EDTA, Tempus and PAXgene tubes and maintained at room temperature. RNA was isolated following storage for 0 and 72 h, and RNA yield, integrity and purity was determined. Thyroglobulin, GAPDH and ACTB mRNA levels were quantified by semi-quantitative real-time PCR.
Results
The RNA yield was significantly higher for blood collected in Tempus tubes compared with PAXgene tubes following storage for 72 h at room temperature (P = 0.0011). High-quality RNA could be extracted from blood collected in PAXgene and Tempus tubes. Blood collected in K2-EDTA tubes, but not in PAXgene and Tempus tubes, showed significant changes in thyroglobulin mRNA levels following storage for 72 h at room temperature (P = 0.0263).
Conclusions
Stabilization of blood in PAXgene and Tempus tubes enables storage at room temperature for up to 72 h, without compromising thyroglobulin mRNA levels.
Introduction
Measurement of serum thyroglobulin (TG) is generally recommended in the follow-up of patients with differentiated thyroid carcinoma (DTC) after total or near total thyroidectomy and ablative radioiodine therapy. 1 The clinical utility of serum TG measurements may be compromised by the presence of endogenous antithyroglobulin antibodies (TGAb) and during thyroid hormone suppression therapy.2,3
Currently, all established methods for measurement of serum TG are affected to some extent by the presence of TGAb. 1 This is an important issue, since TGAb are detected in approximately 25% of patients with DTC 4 and 10% of the general population. 5 New methods are under development to prevent interference by TGAb and promising methods consist of TG assays based on liquid chromatography-tandem mass spectrometry (LC-MS/MS)6–8 and TG assays based on real-time PCR (RT-PCR).9–18 In contrast to other TG assays, the RT-PCR-based TG assays allow the detection of TG mRNA levels in blood. Despite promising, the outcome of studies involving TG assays based on semi-quantitative RT-PCR has been variable. Some find a correlation between TG mRNA levels and the presence of disease9,11 and others do not.10,12–17 Additionally, Boldarine et al. were able to differentiate DTC patients free of disease from those with metastases using quantitative RT-PCR for measurement of blood TG mRNA levels. 18 As previously noticed by several groups, this discrepancy in the outcome of RT-PCR-based TG assays could be related to methodological aspects such as different mRNA extraction protocols, different normalization strategies and the use of different primers.17–20 Moreover, preanalytical factors as type of collection tube, storage time and temperature, if not properly recognized and controlled, may also contribute to this discrepancy since they may influence on TG mRNA quality and stability and lead to induction or down-regulation of TG mRNA expression. Interestingly, a number of studies show that in blood, alteration of gene expression occurs following short ex vivo incubation as a result of induction, down-regulation or RNA degradation.21–24 Immediate processing of blood samples, although possible in explorative studies, is hardly realistic when collecting samples for clinical application. This emphasizes the importance of recognizing and controlling preanalytical variables when quantifying mRNA levels in blood. To avoid immediate processing of blood samples, collection tubes capable of stabilizing RNA at the time of blood collection has been developed. Such collection tubes include the PAXgene and Tempus Blood RNA tubes, which appear to stabilize gene expression in blood when compared with common collection tubes such as EDTA tubes.24–27
Apart from being a prerequisite for clinical application of RT-PCR-based TG assays, control of preanalytical variables is of utmost importance to obtain reliable analytical results, when testing the ability of RT-PCR-based TG assays to predict residual, recurrence or metastatic disease in follow-up of DTC patients after treatment. Although important only Eszlinger et al. have examined the influence of preanalytical variables on blood TG mRNA levels. 12 Briefly, the effect of different sample preparation methods and different storage time periods on RNA yield and GAPDH and TG mRNA concentrations was examined. Blood collected in EDTA or citrate tubes was used and citrate blood in combination with a subsequent separation of mononuclear cells resulted in the best RNA yield and the highest TG mRNA levels. Also Eszlinger et al. observed that RNA should be extracted as fast as possible from whole blood to detect low copy transcripts in a reliable manner. 12 Currently, it is not known how fast TG expression levels change following collection of blood in common collection systems. In the present study, we used blood from DTC patients to monitor the effect of different types of blood collection tubes, K2-EDTA tubes, PAXgene and Tempus Blood RNA Tubes, and storage time on TG mRNA quality and stability in order to establish a robust and standardized procedure for sampling and storage of blood TG mRNA for use in clinical practice.
Materials and methods
Patients
The study included 11 patients previously treated for DTC with total or near-total thyroidectomy and radioiodine ablation. Blood samples were collected during follow-up of patients in the Department of Oncology, Odense University Hospital, Denmark. The study was conducted in accordance with the Declaration of Helsinki of 1975, revised in 2000. All participants gave their informed consent, and samples were anonymized prior to analysis.
Blood collection and study design
Six blood samples were obtained from each patient; venous blood was collected in two K2-EDTA tubes (space for 4 mL blood) (BD Biosciences), two Tempus Blood RNA Tubes (space for 3 mL blood) (Applied Biosystems, Foster City, California, USA) and two PAXgene Blood RNA Tubes (space for 2.5 mL blood) (PreAnalytiX, QIAGEN/BD, Hilden, Germany). For each type of collection tube, one tube was immediately used for RNA isolation and served as reference (0 h) sample. Note that 0 h samples for PAXgene tubes were incubated for 2 h at room temperature before RNA isolation as recommended by the manufacturer’s. The second tube was placed at room temperature for 72 h prior to RNA isolation.
RNA preparation
Total RNA was extracted from 1 mL whole blood collected in K2-EDTA tubes using the TriReagent protocol (Molecular Research Center, Inc. Cincinnati, USA) following the manufacturer’s recommendations. DNA was removed from samples using the TURBO DNA-free Kit (Ambion, Austin, TX, USA) and RNA was further purified using the NucleoSpin® RNA Clean-up XS kit (Macherey-Nagel, Bethlehem, PA, USA). Total RNA from whole blood collected in PAXgene Blood RNA Tubes was extracted and treated with DNase I using the PAXgene Blood RNA Kit (PreAnalytiX, QIAGEN/BD, Hilden, Germany) according to the manufacturer’s instructions. Total RNA from whole blood collected in Tempus™ Blood RNA Tubes was extracted using the Tempus™ Spin RNA Isolation Kit (Applied Biosystems, Foster City, California, USA) according to the manufacturer’s instructions, and DNA was removed using the TURBO DNA-free Kit (Ambion, Austin, TX, USA).
RNA concentrations were measured using the NanoDrop ND-8000 Spectrophotometer (Thermo Scientific, Wilmington, DE, USA). Purity and integrity were assessed using the Agilent 2100 Bioanalyzer (Agilent Technologies, Mississauga, Canada) with the Eukaryote total RNA 6000 Nano Kit according to the manufacturer’s instructions. RNA integrity numbers (RIN) were calculated using the Agilent 2100 Expert Software.
cDNA synthesis and semi-quantitative real-time PCR
Blood TG mRNA levels and mRNA levels of reference genes (GAPDH and ACTB) were quantified using SYBR green-based semi-quantitative RT-PCR. Equal amounts of RNA (500 ng) were reverse transcribed using the High Capacity cDNA Reverse Transcription Kit (Applied Biosystems, Foster City, California, USA) according to the manufacturer’s protocol. For each sample, reverse transcription was performed in duplicate and cDNA was pooled prior to use. RT-PCR was performed on a StepOne Plus real-time instrument (Applied Biosystems, Foster City, CA, USA) using TG, GAPDH and ACTB oligonucleotide primers. Primer sequences and amplification efficiencies (AE) were: TG primers (116 bp amplicon) from Boldarine et al., 18 AE: 2.026; ACTB primers (87 bp amplicon) from Gresner et al., 28 AE: 1.998; GAPDH forward: 5′-GCCACATCGCTCAGACACCATGG-3′, reverse: 5′-TCCCGTTCTCAGCCTTGACGGT-3′ (215 bp amplicon), AE: 2.024. RT-PCR was performed using 2 µL cDNA (diluted 10-fold) in a 20 µL reaction volume containing 375 nM of each primer and 10 µL 2 × SYBR Green PCR Master Mix (Life Technologies, Grand Island, NY, USA). RNA extracted from blood samples collected in the same type of collection tube (Tempus, PAXgene, or K2-EDTA tubes) was analysed for a given gene in the same RT-PCR run, and reactions were prepared in triplicate. The following cycling conditions were used: 10 min at 95℃, followed by 40 cycles of 15 s at 95℃, 45 s at 55℃ and 45 s at 72℃. To test primer specificity, melt analysis was performed for each analysed gene after amplification by heating the reaction mixture from 60 to 95℃ at a rate of 0.3℃/s. Amplification reactions were also routinely checked for the presence of non-specific products by agarose gel electrophoresis.
Data analysis
Quantification cycle (Cq)-values were recorded with the StepOne Plus software v.2.3 (Applied Biosystems, Foster City, CA, USA), and raw Cq-values were exported into Excel-files and analysed using the qBasePLUS software (Biogazelle, NV, Belgium). Cq-values were converted to relative quantities using the method of qBasePLUS, which is a modification of the classic delta-delta-Ct method. 29 The modification takes multiple reference genes and gene-specific amplification efficiencies into account, as well as the error on the estimated amplification efficiency. 29 Briefly, Cq-values are first converted into relative quantities based on the gene-specific amplification efficiency. Normalization is then performed by dividing the relative quantities by a sample specific normalization factor, which is calculated by taking the geometric mean of the relative quantities of the reference genes. 29 Thus, normalized TG expression levels were determined using sample-specific normalization factors based on the geometric mean of the reference genes GAPDH and ACTB (information applied in respective figure legends).
Statistical analysis
Statistical analyses were performed using the GraphPad Prism (5.0 version) software (La Jolla, CA, USA). All analyses comprised independent experiments and differences between groups were analysed by two-tailed Student’s t-tests for paired and unpaired data. When analysing relative TG mRNA levels, 0 h samples were used as reference samples. Thus, to test these data for level of significance, the mean of 72 h samples was tested against a hypothetical value of zero using a two-tailed one sample t-test. The data are presented as means ± SD. Differences were considered to be significant at P < 0.05.
Results
RNA yield and quality
The Tempus Blood RNA system and the PAXgene Blood RNA system were systematically tested for their ability to stabilize RNA in blood following incubation at room temperature for 72 h. For comparison, blood was also collected in standard K2-EDTA tubes. First, the ability of the RNA stabilizing systems to preserve RNA yield and RNA quality was tested. Total RNA yield (∼3 µg/mL) was not significantly changed following incubation in PAXgene tubes for 72 h at room temperature (P = 0.646) (Figure 1). However, a significant change in RNA yield was observed following incubation in Tempus tubes (from ∼1.8 µg/mL to ∼5.1 µg/mL, P = 0.0143) and K2-EDTA tubes (from ∼3.3 µg/mL to ∼2.1 µg/mL, P = 0.0296) for 72 h at room temperature. After 72 h of incubation at room temperature the total RNA yield per mL blood was significantly higher for samples collected in Tempus tubes compared with samples collected in PAXgene tubes (P = 0.0011) (Figure 1).
Total RNA yield. Blood was collected in Tempus (grey bars), PAXgene (white bars), and K2-EDTA (black bars) tubes and stored at room temperature (RT) for 0 h (n = 11) or 72 h (n = 9), followed by total RNA isolation. Each bar represents the mean RNA yield per mL blood ± SD. *P < 0.05 and ** P < 0.01.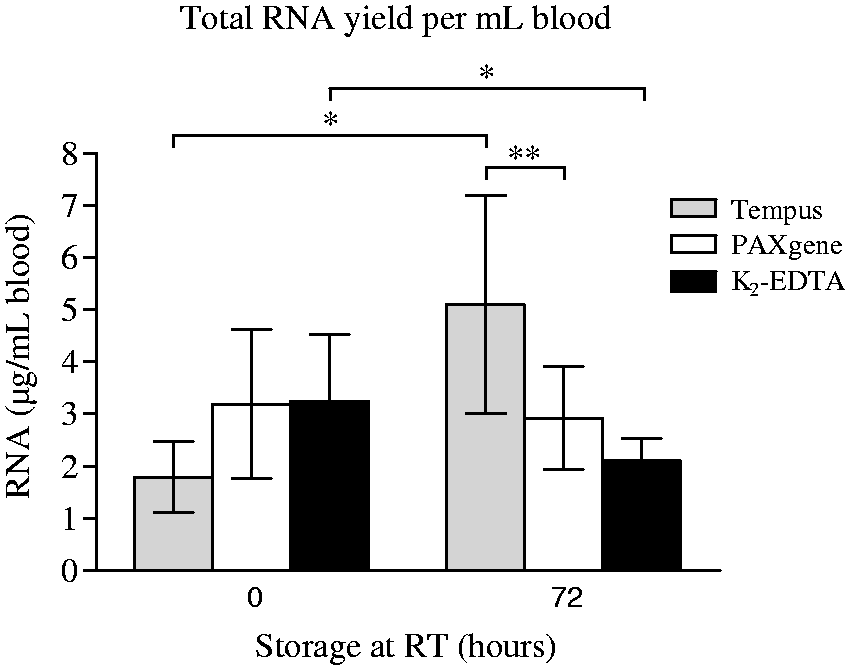
Quality of total RNA purified from blood collected in Tempus, PAXgene and K2-EDTA tubes.
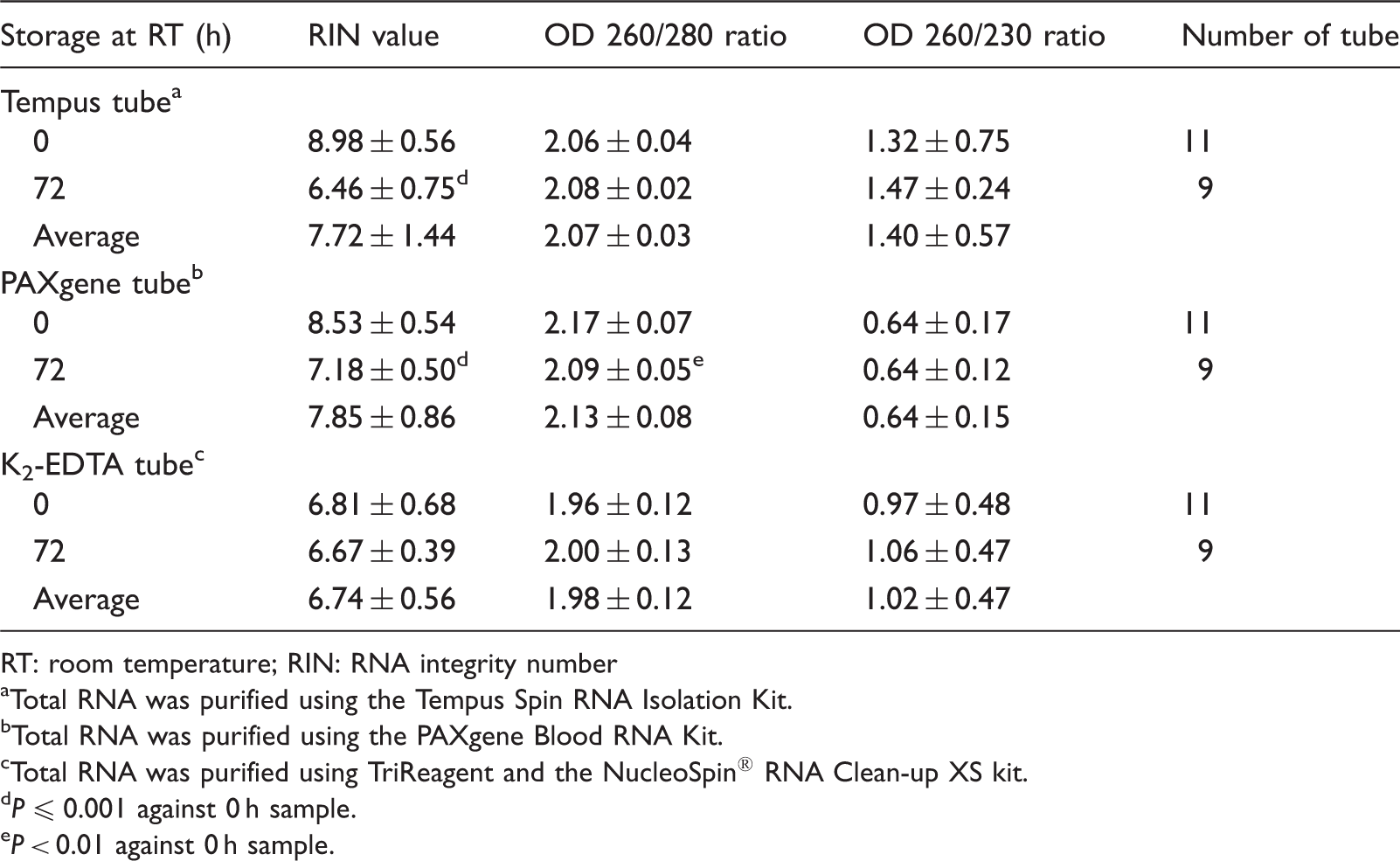
RT: room temperature; RIN: RNA integrity number
Total RNA was purified using the Tempus Spin RNA Isolation Kit.
Total RNA was purified using the PAXgene Blood RNA Kit.
Total RNA was purified using TriReagent and the NucleoSpin® RNA Clean-up XS kit.
P ≤ 0.001 against 0 h sample.
P < 0.01 against 0 h sample.
Average OD 260/280 ratios for RNA isolated from samples collected in Tempus, PAXgene, and K2-EDTA tubes were close to 2 (Table 1). Notably, the average OD 260/230 ratio for samples collected in PAXgene tubes was significantly lower than for samples collected in Tempus tubes (∼0.6 vs. ∼1.4, P < 0.0001) and K2-EDTA tubes (∼0.6 vs. ∼1.0, P = 0.0015). Incubation of PAXgene, Tempus and K2-EDTA samples at room temperature for 72 h had little effect on OD 260/280 and OD 260/230 ratios.
TG transcript stability
Incubation of blood samples in Tempus, PAXgene or K2-EDTA tubes at room temperature for 72 h leads to changes in relative TG mRNA levels when compared with 0 h samples (Figure 2(a)). For all samples collected in PAXgene and K2-EDTA tubes, relative TG mRNA levels decreased following incubation at room temperature for 72 h. In contrast, relative TG mRNA level increased for all samples (except two) collected and stored in Tempus tubes (Figure 2(a)). For samples collected in Tempus and PAXgene tubes, mean relative TG mRNA level were increased (∼1.4-fold) and decreased (∼1.4-fold), respectively (Figure 2(b)). In contrast to these changes that are within ±1.5-fold, the mean relative TG mRNA level for samples collected in K2-EDTA tubes decreased strongly (∼15-fold). Raw Cq-values for TG (and the reference genes ACTB and GAPDH) show similar trends as the observed changes in relative TG mRNA levels (Figure S1).
Storage effects on relative TG transcript levels. Relative TG transcript levels were evaluated by semi-quantitative RT-PCR using RNA extracted from blood collected in Tempus, PAXgene, and K2-EDTA tubes stored at room temperature (RT) for 0 h (n = 11) or 72 h (n = 9). Nine pairs of samples (consisting of one 0 h and one 72 h sample) were included together with two 0 h samples for which no 72 h samples were obtained. For each pair, the 0 h sample was used as a reference sample. (a) Each line represents the difference in relative TG transcript level of 0 h and 72 h samples for a given patient. The black line represents the mean log10-transformed values and the error bar indicates ± SD. The dashed lines indicate a change in the relative TG transcript level of ± 1.5-fold. (b) Each bar represents the mean log10-transformed relative TG transcript level and the error bar indicates ± SD. The dashed lines indicate a change in the relative TG transcript level of ± 1.5-fold.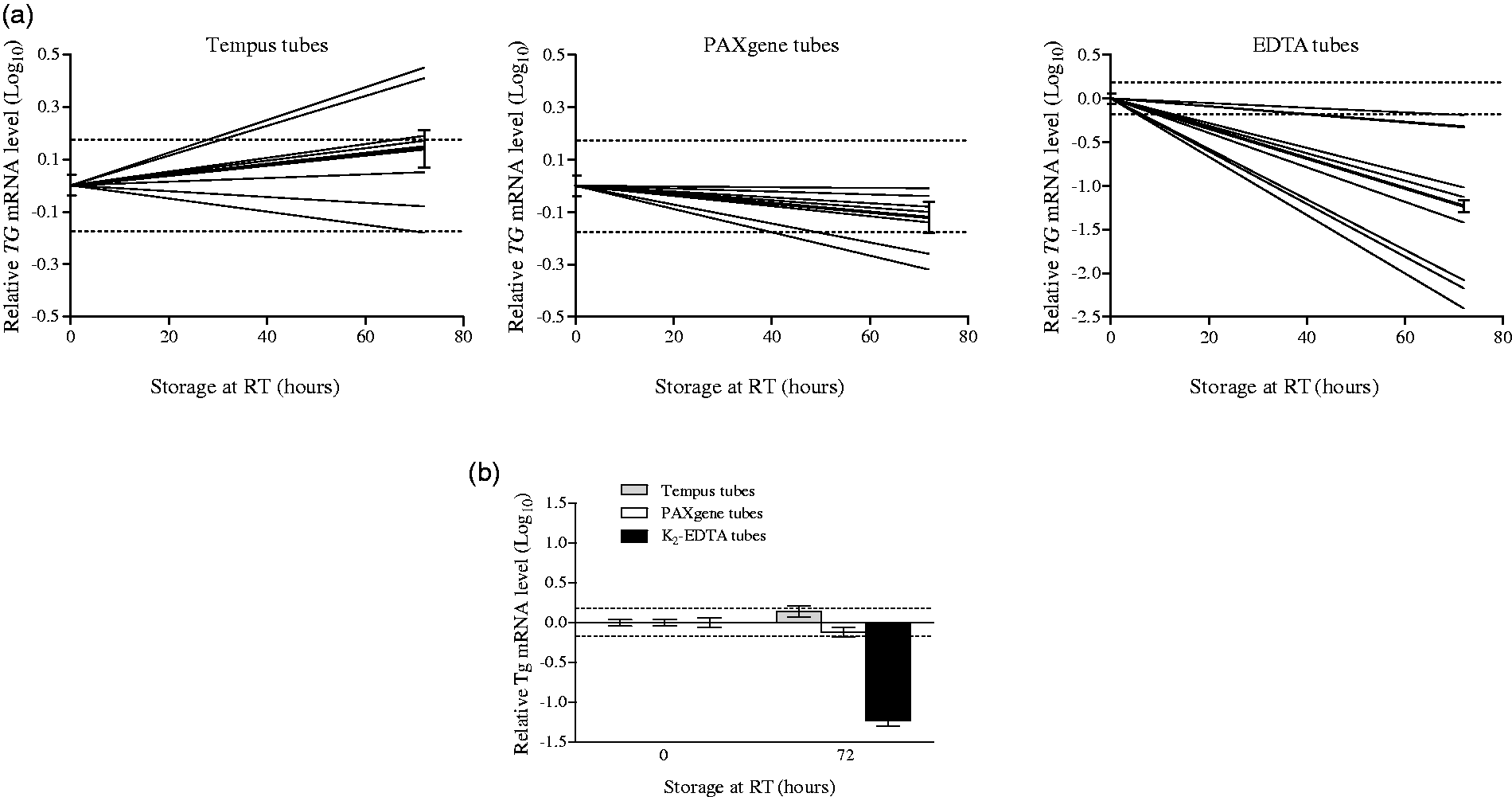
TG expression
TG expression levels were determined by normalizing relative TG mRNA levels to the expression of ACTB and GAPDH. TG expression changed for most pair of samples after incubation for 72 h at room temperature independently on type of collection tube (Figure 3(a)). However, mean TG expression levels were not significantly changed following incubation in PAXgene or Tempus tubes for 72 h (P = 0.9785 and P = 0.8508) (Figure 3(b)). In contrast, mean TG expression levels for samples collected in K2-EDTA tubes differed significantly following incubation at room temperature for 72 h (P = 0.0263) (Figure 3(b)).
Storage effects on TG expression levels. TG expression levels in blood were evaluated by semi-quantitative RT-PCR and normalized against ACTB and GAPDH levels. Blood was collected in Tempus, PAXgene, and K2-EDTA tubes and stored at room temperature (RT) for 0 (n = 11) or 72 h (n = 9) followed by total RNA isolation. Nine pairs of samples (consisting of one 0 and one 72 h sample) were included together with two 0 h samples for which no 72 h samples were obtained. (a) Each line represents the difference in normalized TG expression levels of 0 h and 72 h blood samples for a given patient. The black line represents the mean log10-transformed values and the error bar indicates ± SD. The dashed lines indicate a change in the normalized TG expression level of ± 1.5-fold. (b) Each bar represents the mean log10-transformed normalized TG expression level and the error bar indicates ± SD. The dashed lines indicate a change in the normalized TG expression level of ± 1.5-fold. *P < 0.05.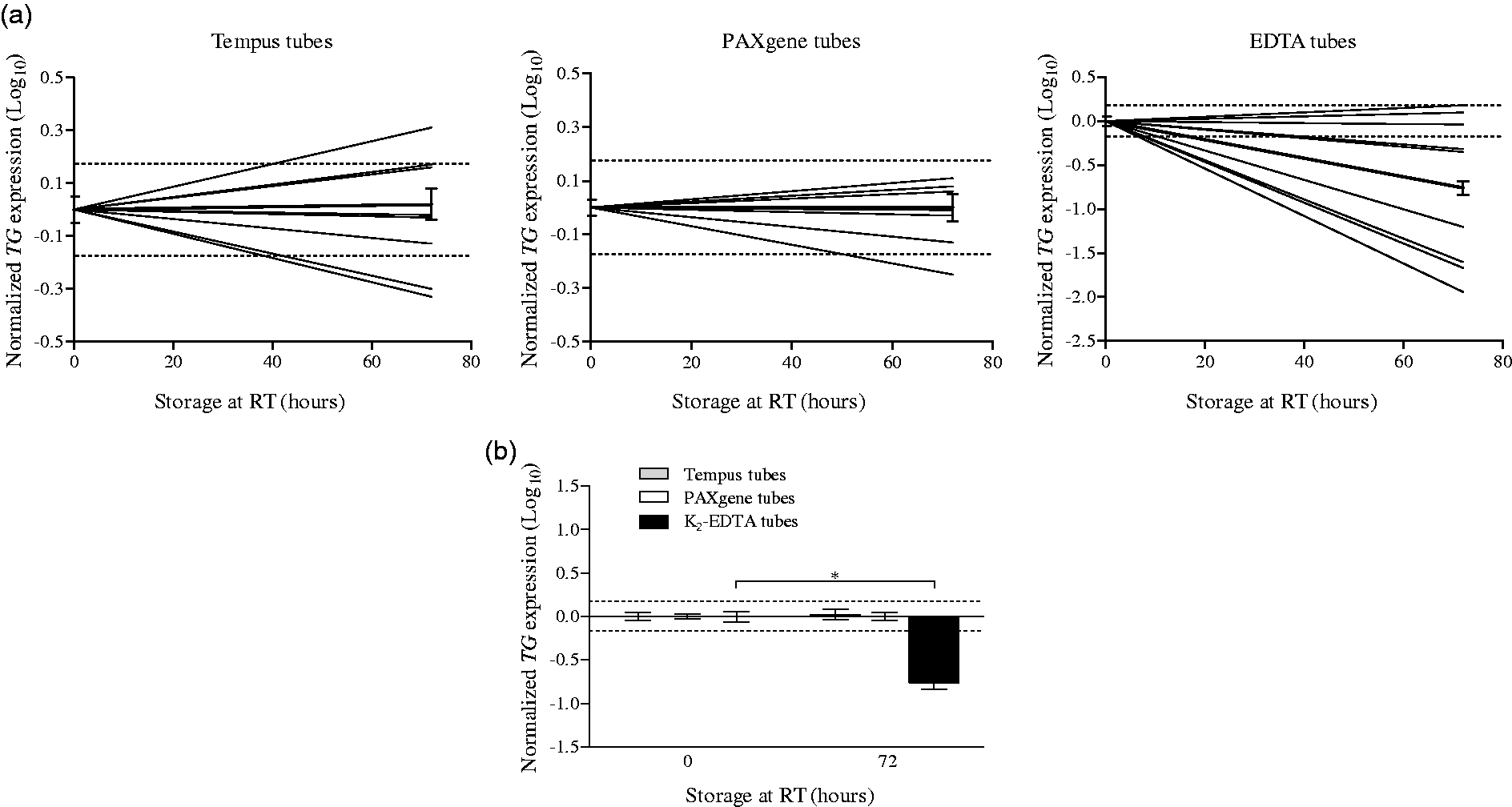
Discussion
Over the years, several RT-PCR-based assays have been developed to measure circulating TG mRNA in the follow-up of patients with DTC.9–18 However, only one of these studies 12 has focused on preanalytical issues, even though this may have a great impact on TG mRNA quality and stability. Preanalytical variables such as type of collection tube, storage temperature and time must be examined to establish robust and standardized TG mRNA RT-PCR protocols for clinical application. In the present study, we examined the ability of two commercially available RNA stabilizing systems to stabilize TG mRNA transcripts in blood of patients treated for DTC. TG mRNA stability in blood was tested at room temperature as this is preferable for easy incorporation into existing routine sampling methods. A storage time of 72 h was chosen, because this is the maximum period recommended for PAXgene tubes (for Tempus tubes the maximum period is 120 h).
The ability of the two systems to preserve RNA yield was examined. Zero h samples collected in Tempus and K2-EDTA tubes were processed immediately in contrast to 0 h samples collected in PAXgene tubes, which were incubated for 2 h at room temperature before RNA isolation as recommended by the manufacturer’s. Consistent with previous findings the RNA yield for samples stored in K2-EDTA tubes decreased significantly following incubation at room temperature for 72 h (Figure 1). 12 Surprisingly, a significant increase in RNA yield was observed when comparing 0 h and 72 h samples collected in Tempus tubes (Figure 1). The reason for this difference is currently unknown; however, a significant difference in RNA yield following storage of blood in Tempus tubes was also observed by Matheson et al. 30 In this study, blood from healthy adults were collected in Tempus tubes. From each individual, blood was collected in two Tempus tubes, which were incubated at room temperature for 2 h, followed by storage at either −80℃ or 4℃ for five days. Samples stored at 4℃ were shipped overnight at ambient temperature, and samples stored at −80℃ were shipped overnight on dry ice before RNA isolation. Surprisingly, they found that the RNA yield was significant higher in the samples that had been shipped overnight at ambient temperature prior to RNA isolation. Since all Tempus tubes were incubated at room temperature for 2 h before storage at −80℃ or 4℃ for five days, the observed difference in RNA yield most likely do not result from incomplete cell lysis. Similarly, Thach et al. 31 find a difference in RNA yield for blood stored at different time periods and at different temperatures in PAXgene tubes following incubation at room temperature for a minimum of 2 h. Hence, storage of blood in Tempus and PAXgene tubes for different time periods may affect RNA yield even when cell lysis is assumed to be complete. According to this and since blood sampling and extraction of RNA from Tempus tubes was performed as recommended by the manufacturer’s, the difference in RNA yield observed in this study is most likely not due to incomplete cell lysis.
A significant difference in yield per mL blood was observed between 72 h samples collected in Tempus and PAXgene tubes (Figure 1). We found higher RNA yields for Tempus tubes compared with PAXgene tubes, which is consistent with results previously reported by others.30,32,33
RNA integrity and purity were used to evaluate sample handling and processing. Mean RIN values for RNA isolated from 0 h samples indicate that better RNA integrity is obtained using PAXgene and Tempus tubes compared with K2-EDTA tubes (Table 1). Notable RIN values for samples collected in Tempus and PAXgene tubes decreased significantly following incubation at room temperature for 72 h (Table 1). Although RIN values for these samples were above 6 (RIN = 1; low RNA quality to RIN = 10; highest RNA quality 34 ), a decrease was unexpected since both Tempus and PAXgene tubes contain RNA stabilizing reagents, which are supposed to strongly reduce in vitro RNA degradation. Moreover, others do not see this kind of decrease in RIN values following incubation of blood at room temperature in PAXgene or Tempus tubes. 33 RNA purity was examined by measuring OD 260/280 and OD 260/230 ratios. For all blood collection systems, OD 260/280 ratios indicated high RNA purity. However, OD 260/230 ratios for samples collected in PAXgene tubes were significantly lower when compared with samples collected in Tempus and K2-EDTA tubes (Table 1). This is consistent with previous studies that also report lower OD 260/230 ratios for samples collected in PAXgene tubes compared with samples collected in Tempus tubes.33,35 As suggested by others, low OD 260/230 ratios may be the result of high salt content of the PAXgene elution buffer.33,35
To further validate the quality of RNA extracted from blood collected in PAXgene, Tempus and K2-EDTA tubes, we used semi-quantitative RT-PCR. Storage of PAXgene and Tempus tubes at room temperature for 72 h had some effect on relative TG mRNA levels (Figure 2(a)), however, mean levels were within ±1.5-fold (Figure 2(b)). In contrast, a pronounced storage-related change in mean relative TG mRNA level (∼15-fold) was observed for samples collected in K2-EDTA tubes (Figure 2(b)). Similar trends were observed for the non-normalized raw mean Cq-values since differences between 0 h and 72 h samples for TG were small for PAXgene (0.53) and Tempus (0.96) tubes but relatively large for K2-EDTA tubes (3.34) (Figure S1). Although pronounced storage-related changes in relative TG mRNA levels were observed for samples collected in K2-EDTA tubes only a significant change in RNA yield was observed and neither RIN values, OD 260/280 ratios nor OD 260/230 ratios changed significantly following storage for 72 h at room temperature (Table 1). Total RNA is primarily composed of ribosomal RNA (rRNA) with the majority of that comprising 28 S and 18 S rRNA species in mammalian cells. We used total RNA kits for measurement of RIN-values, thus measured RIN-values primarily reflect integrity of 28 S and 18 S rRNA species in our samples. Although it is generally believed that the integrity of the 28 S and 18 S rRNA species reflect the mRNA integrity, this is still unclear.36–38 We can therefore not exclude that the integrity of mRNA was compromised following storage for 72 h in K2-EDTA tubes, although RIN values do not indicate that.
For the three types of collecting tubes tested normalization of TG expression levels to GAPDH and ACTB expression levels decreased differences in mean TG expression levels between 0 h and 72 h samples (compare Figures 2(b) with Figure 3(b)). However, significant storage-related changes in TG mRNA levels were still observed for samples collected in K2-EDTA tubes (Figure 3(a) and (b)). Collectively, this indicates that for blood stored in K2-EDTA tubes, the utility of extracted RNA for measurement of TG expression levels decreases following storage at room temperature for 72 h. Several of the studies involving development of TG assays based on semi-quantitative RT-PCR for measurement of TG mRNA levels in blood use K2-EDTA tubes for collection of blood.14,16,17 Although blood samples were processed immediately in most of these studies, the study by Eszlinger et al. and our findings suggest that precaution should be taken using K2-EDTA tubes for collection of blood when measuring TG mRNA levels. Eszlinger et al. 12 did not use K2-EDTA tubes for collection of blood when quantifying TG mRNA since RNA yield and quality was relatively low. Hence, we cannot exclude that preanalytical factors may contribute to the variability previously observed in the outcome of TG assays based on semi-quantitative RT-PCR.9–17
One important limitation of this study is that all samples were left on the table for 72 h with a minimum of transport. Hence, we have not investigated the effect of transport on TG, GAPDH and ACTB mRNA quality and stability which may be an important issue.
In conclusion, our results suggest that high-quality RNA useable for measurement of TG expression levels could be extracted from blood stabilized in Tempus and PAXgene tubes for 72 h at room temperature. In contrast, RNA isolated from blood specimens collected and stored in a common blood collection system (K2-EDTA tubes) revealed significant changes in TG expression levels following incubation for 72 h at room temperature. Thus, K2-EDTA collection tubes can critically impact TG expression and are not recommended for collection of blood when measuring circulating TG mRNA levels in patients with DTC. We recommend the use of RNA stabilizing systems, e.g. Tempus or PAXgene tubes as part of a robust and standardized protocol for measurement of TG mRNA levels by RT-PCR.
Footnotes
Acknowledgements
The authors wish to thank laboratory technician Susanne Hillbrandt for invaluable technical assistance.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethical approval
This study was approved as a service evaluation project by The Regional Committee on Health Research Ethics for Southern Denmark.
Funding
This research was funded by Odense University Hospital.
Guarantor
SF.
Contributorship
SF, LB and SMP designed the study. SF performed the experiments and the data analysis. All authors contributed to data interpretation. SF wrote the first draft of the manuscript. All authors reviewed and edited the manuscript and approved the final version of the manuscript.
