Abstract
Background
Lupus nephritis is one of the most serious complications of systemic lupus erythematosus. This study evaluates the prevalence and correlation between neutrophil gelatinase-associated lipocalin and other biomarkers associated with renal involvement in systemic lupus erythematosus.
Methods
Paired serum and urine specimens from 50 suspected systemic lupus erythematosus patients, characterized by antinuclear antibodies detected by indirect immunofluorescence assay and varying positive concentrations of anti-double stranded DNA antibodies by Crithidia luciliae immunofluorescence assay, were investigated. Of these 50 patients, 18 were identified with renal involvement based upon laboratory serology. Patients and healthy control serum samples (n = 50) were also evaluated for high avidity double stranded DNA IgG antibodies, anti-C1q IgG antibodies, and serum creatinine. The prevalence and relationship between biomarkers were evaluated using statistical methods.
Results
Serum and urine neutrophil gelatinase-associated lipocalin concentrations were significantly elevated in patients compared to controls, with a prevalence of 24% and 36%, respectively. These concentrations were also more markedly increased in systemic lupus erythematosus patients with renal involvement than those without. Spearman’s correlations between neutrophil gelatinase-associated lipocalin and other biomarkers tested ranged from 0.06 to 0.66 in all patients. Combined concordance as determined by Cronbach alpha coefficient between biomarkers was reduced from 0.71 to 0.58 (serum) and 0.62 (urine) when neutrophil gelatinase-associated lipocalin was removed.
Conclusions
Neutrophil gelatinase-associated lipocalin concentrations are elevated and demonstrate variable associated with other laboratory markers for renal involvement in systemic lupus erythematosus. Prospective longitudinal studies are needed to determine the optimal biomarker combinations for use in routine management of systemic lupus erythematosus patients at-risk for lupus nephritis.
Keywords
Introduction
Systemic lupus erythematosus (SLE) is a chronic autoimmune disease characterized by heterogeneity in disease manifestations as well as diversity in immunologic and therapeutic responses. Despite longstanding research efforts, precise diagnosis, variable disease presentation and course, and predicting response to treatment remains problematic, due to the lack of sensitive and specific biomarkers.1,2 A number of epigenetic, environmental, and hormonal factors have been reported to contribute to the immune abnormalities and clinical heterogeneity that occurs in SLE. It is likely that network interplay between these elements drives the production of a variety of autoantibodies, complement products, inflammatory markers, and other mediators identified as biomarkers to diagnose, monitor, stratify, and/or predict disease course, risk, or response to treatment.1–7 Those of laboratory and clinical interest are diverse autoantibodies and complement products, which in addition to clinical manifestations such as skin lesions, arthritis, renal disorder, haematologic changes, and neurologic disorders, are traditionally considered hallmarks of the disease.8,9
Of the different clinical manifestations associated with SLE, lupus nephritis (LN) is one of the most common and is associated with significant morbidity and mortality. In the US, approximately 35% of adults with SLE have clinical evidence of LN at the time of diagnosis, with an estimated 50–60% developing nephritis during the first 10 years of disease. 10 Among these patients, the progression to end-stage renal disease is estimated at a rate of 5–10%. 11 Improved methods for detecting LN or renal involvement would allow earlier treatment, thereby preventing irreversible impairment of renal function and damage. In place of invasive, subjective, and costly serial renal biopsies, laboratory testings such as creatinine clearance, urine protein, complement factors 3 and 4 (C3, C4), serum creatinine, and anti-dsDNA titres have been used for decades to follow onset, course, severity and/or renal involvement. It is, however, recognized that these analyses are inadequate as they are limited in responsiveness to change and therefore unsuitable for optimal patient care and management.6,10
In an effort to reliably predict LN, several candidate biomarkers have been identified.1,2,6 Among the autoantibodies, antichromatin/antinucleosome, high avidity (HA) dsDNA and anti-C1q antibodies have shown some promise as biomarkers of disease activity and/or renal involvement. Of the urine protein biomarkers, neutrophil gelatinase-associated lipocalin (NGAL), soluble vascular cell adhesion molecule 1 (sVCAM-1), monocyte chemotactic protein 1 (MCP-1), and tumour necrosis factor-like weak inducer of apoptosis (TWEAK) amongst others have received considerable attention in predicting renal involvement in SLE.1,2,6 Of these, NGAL, a small protein expressed in the neutrophils, certain epithelial cells, and the renal tubules in response to kidney injury, has been reported in a variety of acute and chronic diseases.12–15 In healthy physiologic conditions, NGAL concentrations are minimally expressed in urine and plasma, but rapidly increase in response to kidney damage to reach diagnostic thresholds within a limited period of time. This is in contrast to the more routine SLE kidney biomarkers that assess renal function such as creatinine, where increased concentrations may not be observed until 24 to 48 h after injury and often lack both diagnostic sensitivity and specificity.
The objective of this study was to investigate the prevalence and relationship between NGAL and other laboratory biomarkers associated with renal involvement in a cohort of patients under evaluation for SLE in a reference laboratory setting.
Materials and methods
Patient population and heathy controls
Fifty patient samples, with paired serum and urine specimens, received at ARUP Laboratories for antinuclear antibody (ANA) by indirect immunofluorescence assay (IFA) and anti-dsDNA IgG antibody by IFA titre (Crithidia luciliae indirect fluorescent test, CLIFT; both from INOVA Diagnostics, San Diego, CA) were investigated. Samples from these patients with suspected SLE (sSLE) were selected based on a positive ANA by IFA with an associated homogenous pattern and/or a positive CLIFT result. Of the 50 patients (40 females, 10 males; 13–60 years of age) fitting the above criteria, 18 patients (16 females, 2 males; 19–60 years of age) were receiving treatment from the University of Utah Hospital, Salt Lake City, Utah. Following testing, a comprehensive, retrospective chart review was performed for these 18 patients to incorporate clinical SLE diagnosis (cSLE) and other laboratory evaluations for renal involvement. In this investigation, renal involvement was defined by abnormal protein concentration, red blood cells, or cellular casts in the urine analysis, and abnormal total protein-to-creatinine ratio in random urine collection at the time of lupus evaluation and/or an episode of biopsy-confirmed nephritis within a month of laboratory analysis. Results were interpreted as abnormal based on reference ranges provided by the laboratory for each analysis.
Paired serum and urine samples from 50 local, self-reported healthy volunteers (22–64 years of age; 33 female, 17 male) served as controls for this study. All studies involving the use of human specimens were approved by the University of Utah Institutional Review Board (Salt Lake City, UT; protocols 7275 & 29507).
Detection of serologic and urine biomarkers
Serum samples were tested for HA anti-dsDNA IgG antibodies (ELISA, INOVA Diagnostics), anti-C1q IgG antibodies (ELISA, INOVA Diagnostics), and enzymatic serum creatinine (MODULAR P, Roche Diagnostics, Indianapolis, IN). The presence and concentration of NGAL in serum and urine were detected using the NGAL Test reagent kit (BioPorto Diagnostics, Hellerup, Denmark), a particle-enhanced turbidimetric immunoassay with application parameters for several automated methods; for this study the Roche cobas c501 was used. All testings were performed at ARUP Laboratories following manufacturers’ instructions. Manufacturers’ recommended cut-off values were utilized for HA dsDNA (>30 IU/mL) and anti-C1q (≥40 units) assays. Reference ranges for serum creatinine (35.4–106.1 μmol/L), serum NGAL (51–265 ng/mL), and urine NGAL (2–113 ng/mL) were established at ARUP Laboratories.
Statistical analysis
The prevalence of the specific biomarkers in each group was established using manufacturers’ recommended or laboratory established cut-off values, as indicated. The D’Agostino-Pearson omnibus K2 normality test was performed using GraphPad Prism (version 5.04, GraphPad Software, Inc., La Jolla, CA) to determine if the datasets were normal. Significance, as determined by P-values <0.05 from the Mann–Whitney test, was calculated using GraphPad Prism. All other analyses including Spearman correlation coefficient and Cronbach alpha coefficient were determined using SAS (version 9.3, SAS Institute Inc., Cary, NC).
Results and discussion
Along with the identification of novel serologic and urine biomarkers for identifying and managing SLE patients at-risk for renal involvement, the examination of the relationships between established and emerging tests to determine optimal strategies for patient care is crucial. Current biomarkers have been reported to have limited clinical significance in early prediction of renal involvement in SLE.1–3 In this preliminary cross-sectional laboratory investigation, the prevalence of serum and urine NGAL as well as routine tests for the evaluation of SLE was examined in a cohort of patients undergoing screening for SLE, along with a subset of patients with confirmed disease and renal involvement.
Baseline characteristics of study population.
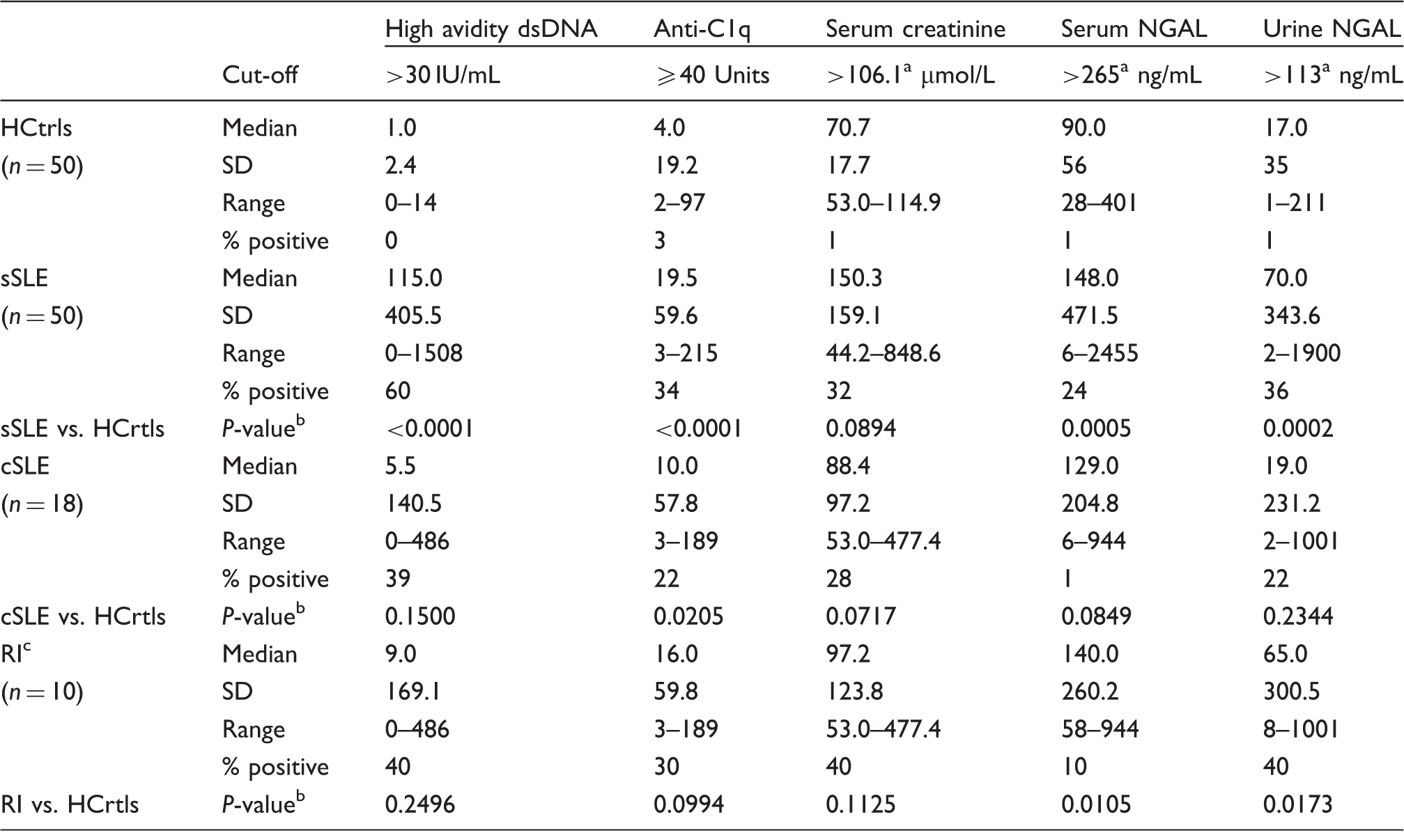
Cut-off values established at ARUP Laboratories.
P-values determined by the Mann–Whitney test.
Patients with RI came from the cSLE group.
HCtrls: Healthy controls; sSLE: suspected SLE; cSLE: confirmed SLE cases; RI: renal involvement; SD: standard deviations.
Demographics and laboratory characteristics of patients with confirmed SLE (cSLE).
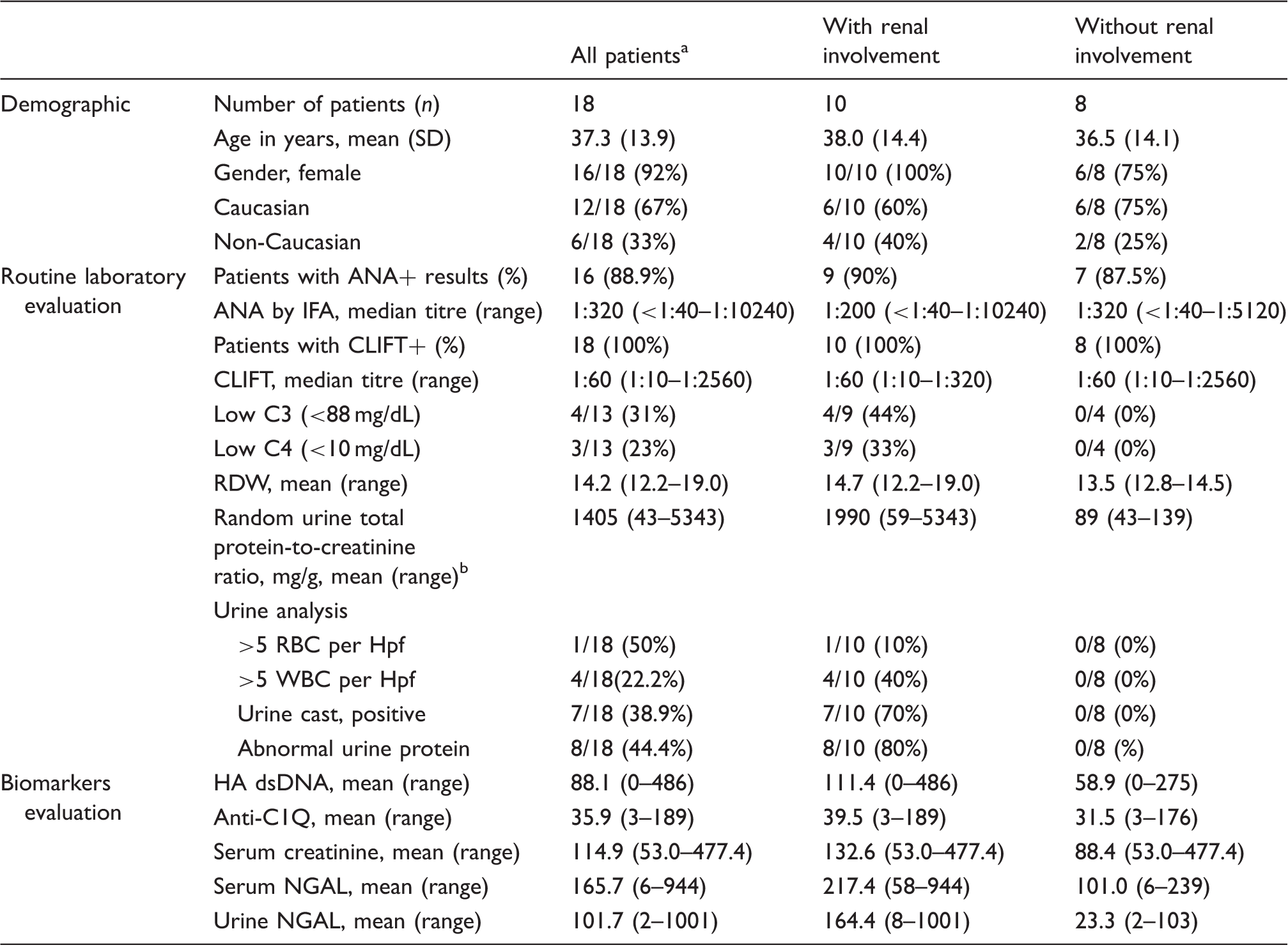
SD: standard deviation; C3: complement 3; C4: complement 4; RDW: red blood cell distribution width; Hpf: high power field.
Note: No significant differences were observed for any analyte between patients medians with or without renal involvement (P-value >0.05), except for urine protein (P-value 0.04).
All results for routine laboratory evaluation were not available for some patients. The total n was changed to account for this in order to calculate percentages.
Random urine total protein-to-creatinine ratio reference range: male 15–68 mg/g; female: 10–107 mg/g (18 years and older).
Of the five biomarkers tested, all except the anti-C1Q IgG test demonstrated significantly higher concentrations in patients with renal involvement compared to those without (Table 2). This observation may be limited by the cross-sectional nature of this investigation, the small sample size of the cohort as well as the lack of measures to define active LN and global disease activity. In a recent study by Bock et al., 16 anti-C1Q antibodies correlated not only with urine protein-to-creatinine ratio in patients with confirmed but also in those without confirmed renal involvement suggesting subclinical renal involvement. The observation that dsDNA IgG, serum creatinine, or urine NGAL concentrations are significantly elevated in patients with renal involvement than those without appears to be consistent with previous studies.13–21 Of the patients with renal involvement, only two patients were positive for a single biomarker (urine NGAL or HA dsDNA) of the five evaluated. All other patients tested positive for at least two or more biomarkers with urine NGAL and serum creatinine or HA dsDNA and anti-C1Q co-occurring in 60% of the cases.
Correlation between NGAL and other SLE biomarkers investigated.
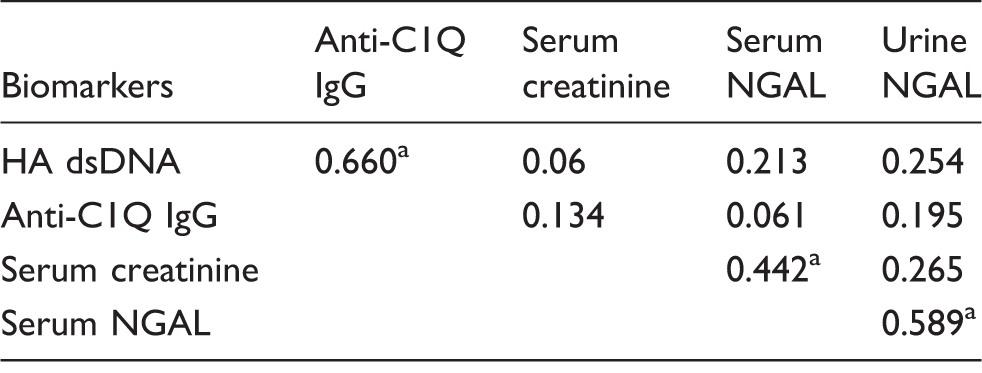
NGAL: neutrophil gelatinase-associated lipocalin.
Note: Comparisons between NGAL and indicated biomarkers were estimated by Spearman correlation coefficient in the suspected patient group (n = 50) and clinically defined cases with renal involvement. A value of 1.0 represents ideal correlation.
P-value <0.05
With the recent developments in biomarker discovery for predicting renal involvement in SLE, a number of investigations have examined the associations between specific urinary biomarkers as well as serum protein profiles and histological features of nephritis in an attempt to reliably predict disease.3,7,14,15,18–20 One of the seminal studies involving paediatric SLE patients, a population that is most adversely affected by LN, was reported by Brunner and colleagues.21 Based on the report from this investigation, the authors concluded that presence of defined urine proteins could predict specific LN signatures such as activity and chronicity. Although the current investigation is limited by the low number of cSLE patients with renal involvement, ethnic diversity, and disease activity score, there was a general trend for patients with renal involvement as defined by current laboratory tools to have higher levels and/or frequencies of all, if not most, biomarkers associated with LN. As previously reported by others and shown here, no single biomarker is sufficient in identifying and/or stratifying SLE patients at-risk for renal involvement.3,7,19 While SLE disease heterogeneity may partly be responsible for these observations, the prevalence of individual biomarkers lacks optimal sensitivity.
In conclusion, this study demonstrates that urine NGAL determination in addition to currently available laboratory tests for the evaluation of SLE may be useful in assessing renal involvement. Biomarker diversity and disease heterogeneity demands an informed approach geared towards the validation of biomarker ‘panels’ of renal involvement in SLE. Studies with well-defined endpoints that ensure a true measure of the intended clinical process in diverse cohorts coupled with robust analytical assays and high statistical power to confirm these panels are needed.
Footnotes
Acknowledgements
The authors thank Elisabeth Malmberg and Andrew Wilson for assistance with statistical analysis and Joshua Hunsaker for sample procurement and de-identification. The authors also acknowledge the provision of reagents from BioPorto Diagnostics and INOVA Diagnostics.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
Support for this study was provided by the ARUP Institute for Clinical and Experimental Pathology.
Ethical approval
The University of Utah Institutional Review Board approved this study (ref# 7275 & 29507).
Guarantor
AET.
Contributorship
SLL wrote the first draft of the manuscript, researched literature, performed testing, and data analysis. BBS was involved in protocol development, sample acquisition, and testing. KWD performed testing. JAS provided expertise in protocol development and data analysis. AET conceived the study, gained ethical approval, researched literature, performed data analysis and review. All authors reviewed and edited the manuscript and approved the final version of the manuscript.
