Abstract
Background
False-positive cardiac troponin I results as a result of carryover have previously been reported on the Beckman Coulter AccuTnI assay. We sought to determine if the carryover problem had been resolved with the new AccuTnI + 3 assay.
Methods
Carryover experiments were performed in parallel on the Beckman Coulter Access2 analyser using the legacy AccuTnI and new AccuTnI + 3 assays. The same negative patient pool sample was analysed before and after a single analysis of an extremely elevated patient sample.
Results
Analysis of a single extremely high sample caused elevations above the 99th percentile cut-off, and thus false-positive cardiac troponin I results on both assays. Both assays demonstrated carryover and subsequent further elevations in negative pool results the following day.
Conclusions
Our study replicates our previously published findings of carryover and reagent pack contamination on the AccuTnI assay. Despite improvements on the new AccuTnI + 3 assay, carryover and reagent pack contamination are still present.
Introduction
Although cardiac troponin (cTn) measurement does not independently facilitate the diagnosis of acute coronary syndromes (ACS), the potential ramifications of falsely elevated cardiac troponin I (cTnI) results on patient care include prolonged emergency department stay, repeat laboratory testing and potentially invasive cardiac investigations. False-positive cTnI results have previously been reported on multiple analytical platforms. These false-positive results are random, not reproducible and unexplained despite intensive investigation.1,2 We have previously published our findings of carryover and reagent pack contamination on the Beckman Coulter AccuTnI assay. 3 Beckman Coulter have made significant improvements in the new AccuTnI + 3 assay, including an additional probe wash, and we sought to determine if the carryover has been resolved on the AccuTnI + 3 assay.
Methods
A negative patient serum pool with a cTnI concentration below the 99th percentile cut-off of 40 ng/L was prepared from remaining samples submitted to our laboratory for testing. The negative patient serum pool was prepared, divided into aliquots and stored frozen at −20℃ until use. Three levels of Bio-Rad® quality control (QC) material (Bio-Rad, Hercules CA, USA) were included at the beginning and end of each experiment to ensure acceptable analytical performance. One patient serum with an extremely elevated cTnI estimated at 1,800,000 ng/L was analysed in the run (final result obtained after 1 in 20 dilution with Beckman Coulter diluent as per manufacturer’s recommendations). Identical experiments using the AccuTnI and the AccuTnI + 3 assays were performed on the Beckman Coulter Access2, (Beckman Coulter Diagnostics, Brea, CA, USA). The sequence of testing was as follows: 3 levels of QC material, followed by 10 samples of the negative pool, the high sample analysed in singlicate, followed by another 10 samples of the negative pool. Another 10 negative pool samples were analysed the following day followed by 3 levels of QC. A modification of Broughton’s protocol was used to calculate carryover (a sequence of A1, A2, B1, B2, B3 where A = extremely elevated sample, B = negative pool) and calculation ([B1 − B3)]/[A2 − B3] × 100). 4 We only measured the extremely elevated sample in singlicate. Manufacturer stated performance characteristics of the assays are included in the supplemental file.
Results
The performance of the AccuTnI and AccuTnI + 3 assays for the 3 levels of QC material run and information pertaining to the LoD and LoB data from our method evaluation of the AccuTnI + 3 assay is included in the supplemental file.
Analysis of a single high sample (∼1,800,000.0 ng/L) followed by 10 negative pool samples demonstrated a significant increase in measured cTnI concentrations (see Figure 1). This increase was sufficient to exceed the 99th percentile cut-off (40.0 ng/L), causing false-positive results on both assays. The means (SD) of 10 negative pool samples prior to analysis of the high sample were 31.3 (10.3) ng/L on the AccuTnI, and 1.9 (0.7) ng/L on the AccuTnI + 3. The result immediately before the high sample, and three subsequent results were: 35.2; 84.4; 72.6; 39.0 ng/L on the AccuTnI and 2.6; 56.6; 25.1; 10.2 ng/L on the AccuTnI + 3 (an increase of 49.2 and 54.0 ng/L on the AccuTnI and AccuTnI + 3 respectively). Calculated percentage carryover is 0.00254% and 0.00257% on the AccuTnI and AccuTnI + 3 respectively.
Graphic representation of cTnI results on the same negative pool samples analysed on the Access2 by both the previous AccuTnI and new AccuTnI + 3 assay before and after analysis of the ∼1,800,000.0 ng/L sample (open points), as well as 24 h later (solid points). The arrow indicates where the high sample was analysed. The horizontal dashed line indicates the 99th percentile cut-off.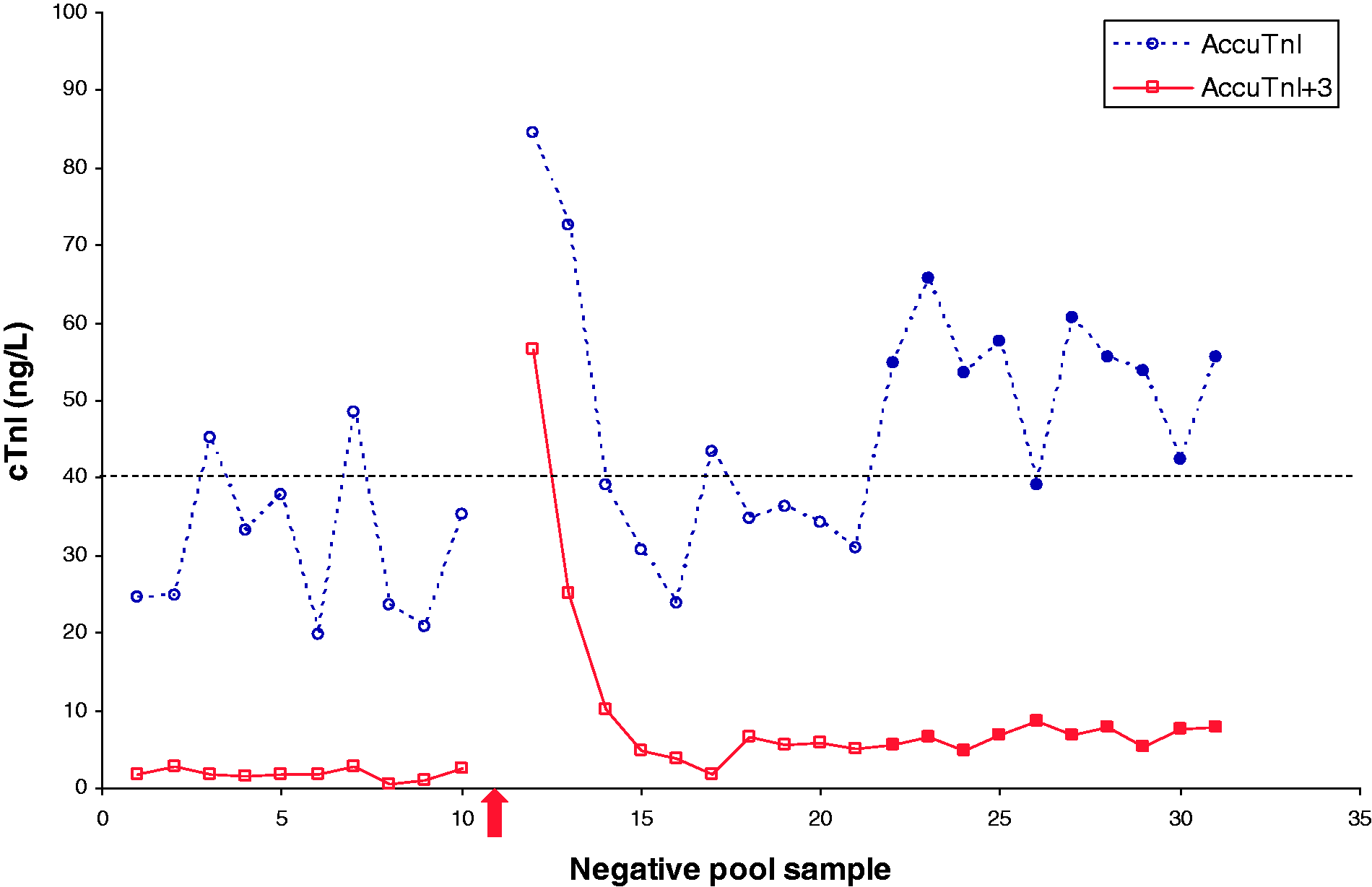
Both assays showed a statistically significant elevation in the mean of 10 replicates obtained on the negative pool 24 h later (AccuTnI: mean 53.9, SD 7.9 ng/L, P < 0.0001), AccuTnI + 3 (mean 6.8, SD 1.3 ng/L, P < 0.0001) when compared to the mean of 10 samples run before the high sample was analysed, using a paired t-test.
Discussion
The differences in results as well as precision obtained on the same negative pool between the AccuTnI and AccuTnI + 3 assays mirrors our method comparison study (unpublished data).
Other investigators have documented that the carryover problem with the AccuTnI assay has been resolved with the recently released AccuTnI + 3 assay5,6 but our evaluation does not support this finding. This study replicates our previously published findings on the legacy AccuTnI assay, and shows that although reagent pack contamination has been improved, carryover still occurs on the AccuTnI + 3 assay when extremely elevated samples are analysed. Despite improvements in the AccuTnI + 3 assay including the addition of an additional probe wash, the carryover is approximately equivalent with both assays. Calculated % carryover for the AccuTnI and AccuTnI + 3 was 0.00254 and 0.00257% respectively, which is not dissimilar to Lippi et al.’s findings of 0.003%. 6 However, the high sample they used was significantly less than ours (215,000.0 ng/L and 484,176.0 ng/L vs. 1,800,000.0 ng/L).
We are of the opinion that Broughton’s calculation of percentage carryover 4 offers false reassurance. Calculating the % carryover does not directly indicate to what extent a result may be influenced. A % carryover of 0.0025% may lead to clinically misleading results in one case, whereas a % carryover many times greater may be clinically irrelevant in another. This aspect is illustrated by cTn where it is important to accurately measure very low levels and where occasional samples with grossly elevated levels may be encountered. In our setting, with ∼256,000 cTnI performed annually (data for 2014) in our laboratories, we encountered 22 samples with cTnI concentrations exceeding 500,000 ng/L, or ∼2 per month.
Conclusions
Carryover from extremely elevated cTnI in patient samples, may result in false-positive results for patient samples analysed thereafter. Our study replicates our previous findings, and demonstrates that carryover still occurs on the AccuTnI+3 assay.
Footnotes
Acknowledgements
We gratefully acknowledge the assistance of Anna Bransden for collecting and analysing the samples at the RBWH.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Ethical approval
Not applicable.
Guarantor
UW.
Contributorship
UW, CJP, JPJU and MG developed the protocol. MG collected samples at TPCH. UW performed the data analysis and wrote the first draft of the manuscript. All authors reviewed and edited the manuscript and approved the final version.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
