Abstract
Background
Dengue is the most common arboviral illness worldwide. While most infected patients recover, a proportion of them develop severe complications or fatality. Nevertheless, the pathophysiological mechanisms which distinguish the disease severity and associated complications are not clearly understood. We studied blood profiles of dengue patients in order to identify microRNAs that could play a role in these pathophysiological mechanisms.
Methods
Blood samples from 26 dengue-infected patients were collected within 0–14 days of infection. Together with samples obtained from six healthy individuals, microRNA profiles were generated to identify significantly altered microRNAs upon dengue infection. Profiles of patients with influenza were also used to determine the disease specificity of these altered microRNAs. Their discriminative power to distinguish dengue from influenza was then tested statistically.
Results
Several significantly altered microRNAs were identified in patients with dengue. Twelve microRNAs were specifically altered upon acute dengue whereas 14 microRNAs exhibited similar expression between dengue and influenza. Seventeen microRNAs which could potentially distinguish dengue-related complications were also identified. Expression of miR-24-1-5p, miR-512-5p and miR-4640-3p distinguished mild dengue from those exhibiting liver complications whereas miR-383 was significantly upregulated in mild dengue compared to those diagnosed as severe dengue with fluid accumulation.
Conclusions
We identified two panels of microRNAs – one specific for dengue and the other common to dengue and influenza. We also report on the differentially expressed microRNAs in patients with mild versus severe dengue, which could be the basis for the complications seen in them.
Introduction
Arthropod-borne infections make up approximately 16% of the estimated global burden of infectious diseases with dengue being the most prevalent. 1 Annually about 50–100 million dengue virus (DENV) infections are estimated to occur on a global level and the annual economic cost of dengue management is projected to be at least US$587 million. 2 Yet, this is believed to be an exaggerated underestimation. 3 The DENV is a single-stranded positive strand RNA virus belonging to the flavivirus family and four serotypes with about 65% genetic homology are in circulation globally. Despite this, the variants are antigenically distinct serotypes and thus are classified as DENV1-4. Although the majority of DENV infections are asymptomatic, 4 they can also induce mild to severe clinical manifestations in individuals which can last up to 14 days. Upon infection, the DENV takes control of the host’s replicative machinery as the progression of the pathogenesis requires coordination of the molecular processes occurring within both the virus and the host. For instance, the DENV capsid interacts with the human death-domain associated protein (DAXX) and increases Fas-mediated and CD137-meditated apoptosis of liver cells.5,6 Thus, the response to DENV infection triggers a myriad of changes in the host transcriptome, which is subsequently reflected in the proteome expression. Yeast two-hybrid assays identified about 403 interactions between the DENV and host proteins. 7
Another level of regulation that is also mediated on the host transcriptome is by endogenously transcribed microRNAs (miRNAs). The human genome encodes for hundreds of miRNAs which are capable of regulating proteome expression via the manipulation of the host transcriptome.8,9 miRNAs are an abundant class of short oligonucleotides ranging from 20 to 25 nucleotides in length and are categorized as non-coding regulatory RNAs. They interact with their target mRNA via partial or perfect complementarity resulting in degradation and/or translational repression of the transcript and this eventually affects protein production. This mechanism, known as RNA interference (RNAi), is also an important antiviral defence process, as silencing the RNAi components such as Dicer, Drosha, Ago1 and Ago2 led to an increase in DENV replication. In fact, the DENV NS4B (dengue virus non-structural protein 4B) was implicated as a suppressor of the RNAi pathway causing downregulation of most miRNAs. 10 Though miRNAs are considered to be defence intermediaries against infection, several host miRNAs have also been reported to be crucial for propagation of some viruses. miR-29a in T-cells was found to strongly inhibit replication of HIV-1 via a highly conserved target site in the viral genome, 11 while elevated expression of miR-146a promoted Hendra virus replication by modulating NF-κB activity. 12
As the DENV mediates its pathological effects via its entry into the bloodstream, with consequences on the host’s immune system, liver and endothelial cells of blood vessels, 4 whole blood profiles will provide crucial information of the condition. Thus, we propose blood as an ideal candidate to study the miRNA changes occurring as a consequence of dengue infection. Nevertheless, the effect of DENV infection on the global host miRNA transcriptome is not well-characterized, particularly in whole blood. As miRNAs also possess regulatory potential and are often released into the blood stream in response to infection, in this study, by utilizing blood miRNA profiles, we aimed to elucidate crucial miRNAs regulating the complex host response during dengue infection. Identification of these miRNAs could aid in the understanding of the complexities involved in the regulation of host response and the complex interplay between the host responses and virus upon DENV infection.
Materials and methods
Patient enrolment and sample collection
The study protocol was approved by the National Healthcare Group’s Domain Specific Institutional Review Board (NUS-IRB Ref Code: 09/641). Patients were identified as dengue patients through routine clinical laboratory diagnostic techniques involving primarily NS1 antigen detection, the polymerase chain reaction (PCR) and serology. 13 The clinical status of the patients was assessed by a single experienced infectious diseases physician (PAT) using the World Health Organization definitions of dengue fever (DF), DF with warning signs and severe dengue. Consenting participants contributed blood within a period of 14 days from the onset of the infection for the study, and these were collected in RNALater™ (Ambion, TX, USA). The total RNA was isolated using Ribopure™ Blood RNA isolation Kit (Ambion, TX, USA) according to manufacturer’s protocol. RNA concentration was determined using ND-1000 Spectrophotometer (Nanodrop™, Rockland, DE, USA) and its integrity was verified using Agilent 2100 Bioanalyzer (Agilent Technologies, USA) and denaturing agarose gel electrophoresis.
MicroRNA array and validation
Locked nucleic acid-modified oligonucleotide (Exiqon, Vedbaek, Denmark) probes (miRBase version 18.0) were used for the microarray. Total RNA (1 μg) was 3′-end-labelled with Hy3 dye using the miRCURY LNA™ Power Labeling Kit (Exiqon, Vedbaek, Denmark) and hybridized on miRCURY LNA™ Arrays (seventh generation chips) according to the manufacturer’s protocol, using the MAUI® hybridization system. Hybridization images were scanned, digitized using InnoScan 700 microarray scanner (Innopsys, Carbonne, France) and analysed using Partek® 6.6 Genomics Suite software (Copyright, Partek Inc., St Louis, MO).
Validation of miRNAs microarray in blood was carried out using TaqMan quantitative real-time PCR. 14 All reactions were carried out according to the manufacturer’s protocols and recommendations (Applied Biosystems, USA). The PCR reactions were carried out using the Applied Biosystems 7900 high throughput sequence detection system. Both randomly selected and miRNAs that are differentially regulated and found to be important in the diagnosis/prognosis of DF were selected for validation of the microarray data. All reactions were performed in triplicate and repeated four times, using individual patient samples. The fold change for the expression was then calculated based on 2−ΔΔCt. U6 snRNA was used as the endogenous control for blood miRNAs and the healthy controls (n = 6) were used as calibrator. The fold change for miRNA expression was calculated for respective day of infection (n = 3) and were then averaged for acute and recovery phases.
Statistical analysis
Analysis of profiling data was performed according to previously published report 15 that included background subtraction, normalization and hierarchical clustering. Briefly, background-subtracted median signal intensity of 100 was selected as a threshold value for inclusion of significantly detected miRNAs. Global sample variability was assessed by principal components analysis (PCA). First stage of normalization was carried out against a group of endogenous controls and the spike-in controls for each chip to avoid technical and experimental variations among the healthy and DENV-infected samples. The normalized signal intensity value was log2 transformed and the differentially regulated (dengue patients versus healthy controls) miRNAs with an absolute fold change ≥1.2 and ≤ −1.2 and P ≤ 0.05 were selected after Benjamini–Hochberg false discovery rate (FDR) correction following multiple comparisons. The FDR method was used to filter the differentially expressed miRNA. All statistical analyses were performed using the statistical tools provided by Partek® 6.6 Genomics Suite software. The data are presented as fold change; One-way ANOVA followed by Bonferroni correction was used for multiple comparisons. 15 The FDR method was used to filter the differentially expressed miRNA. Further selection of miRNAs was based on receiver-operator curve (ROC) analyses which were performed using the MedCalc (MedCalc Software, Belgium) statistical tools.
Results
Differential expression of miRNAs in dengue patients
A total of 32 individuals consisting of six healthy controls and 26 dengue patients between day 0 and day 14 of/after infection were recruited for the miRNA profiling phase of this study. The clinical characteristics of the patients recruited for the study are shown in Supplementary Table 1. The miRNA profiling of total RNA isolated from their blood samples detected 1250 miRNAs (Sanger mirBase version 18). A total of 348 miRNAs were found to be differentially and significantly regulated among the dengue patients (based on fold change ≥1.2 and ≤ −1.2 and P ≤ 0.05 were selected after Benjamini–Hochberg FDR correction). Hierarchical clustering of normalized data showed two independent clusters based on early acute infection (day 0–day 4) versus recovery (day 5 onwards) regardless of disease severity (Figure 1(a)). The miRNA profiles of healthy controls were grouped together with the profiles of patients in recovery phase. However, their independent segregation was clearly evident in the principal component analysis (Figure 1(b)). To understand the importance of these miRNAs in relation to DENV infection, the changes in their expression patterns throughout disease progression (acute as well as recovery phases) were analysed. Verification of miRNA profiling data was performed using quantitative real-time PCR on 10 randomly selected miRNAs (hsa-miR-15a-3p, -214-3p, -218-5p, -320a, -340-3p, -383, -544a, -576-5p, -615-5p and hsa-miR-1270). The quantitative PCR data were observed to be consistent with the miRNA microarray results (Supplementary Table 2).
Blood miRNA profiling of DENV-infected patients. (a) Hierarchical clustering of dengue miRNA profiles. Distinct clustering was clearly seen between acute (≤4 days) and recovered (> 4 days) dengue patients. The profiles recovered patients clustered together with healthy controls. The data presented were subjected quantile normalization. (b) Principal Component Analysis (PCA). PCA analysis reflected that although the recovered patients clustered together with healthy controls, there was segregation among them on a planar level. The red circles represent acute dengue while blue represents healthy controls and green represents recovered dengue patients.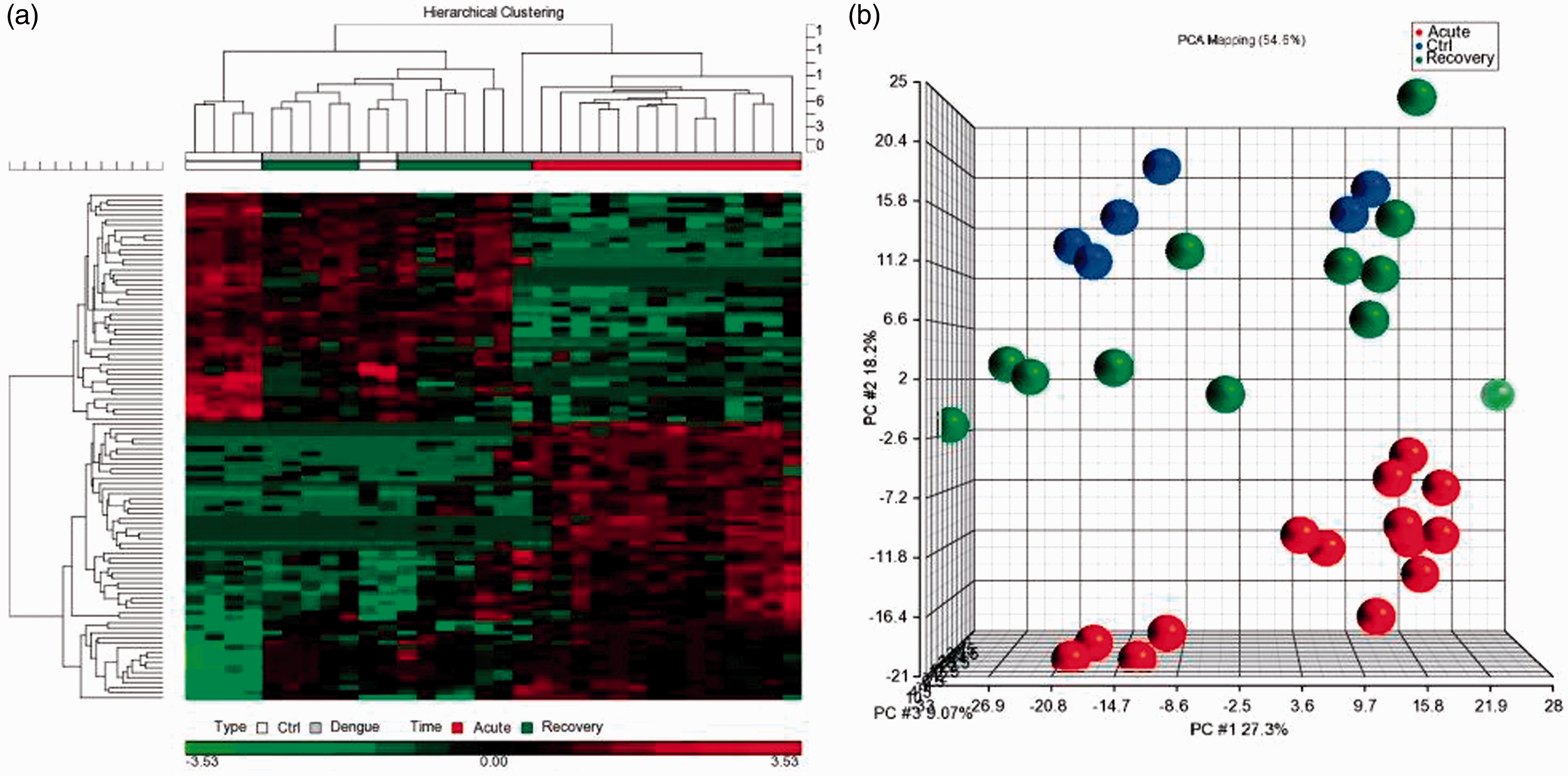
Regulation of miRNA expression during DENV infection
The miRNA profiles of all the dengue patients were compared to healthy controls. Applying inclusion criteria of fold-change ≥2 or ≤ −2 and a stringent FDR adjusted significance cut-off (P ≤ 0.01), 201 and 23 miRNAs were identified to be differentially expressed in acute and recovered dengue patients, respectively (Figure 2(a)). On the other hand, comparison of miRNA profiles of acute versus recovered patients showed differential expression for 177 miRNAs (Figure 2(a)). Overlapping these three datasets, 19 miRNAs (miR-19a-3p,-19b-3p, -33b-5p, -145-3p, -299-3p,-320a, -374a-5p, -423-3p, -548e, -576-5p, -584-5p, -614, -634, -652-3p -4451, -4503, -4529-3p, -4788 and -5583-5p) were seen to be differentially expressed in all dengue patients within two weeks of infection when compared to controls (Figure 2(b) and (c), Supplementary Table 3). Among these, miR-299-3p was distinctly upregulated and miR-576-5p was downregulated in all patients (Figure 2(d)).
miRNAs differentially expressed in DENV infection. (a) Venn diagram of differentially expressed miRNAs. Number of differentially regulated miRNAs in acute dengue and recovery with respect to healthy controls as well as acute dengue versus recovery are shown here. Number of miRNAs that could potentially differentiate one condition from the other two is represented in shaded regions. (b) Cluster of miRNAs differentially expressed in all dengue patients. Heatmap shows the 19 miRNAs which were found to be differentially expressed in all dengue patients within 0–14 days of infection. (c) Segregation of individuals according to disease versus controls. PCA plots show distinct segregation between healthy individuals and dengue patients based on the expression patterns of the 19 miRNAs. (d) Dot plots of miRNAs differentially expressed in dengue versus controls. Among the 19 miRNAs, miR-299-3p and miR-576-5p showed extensive segregation between dengue patients and healthy controls.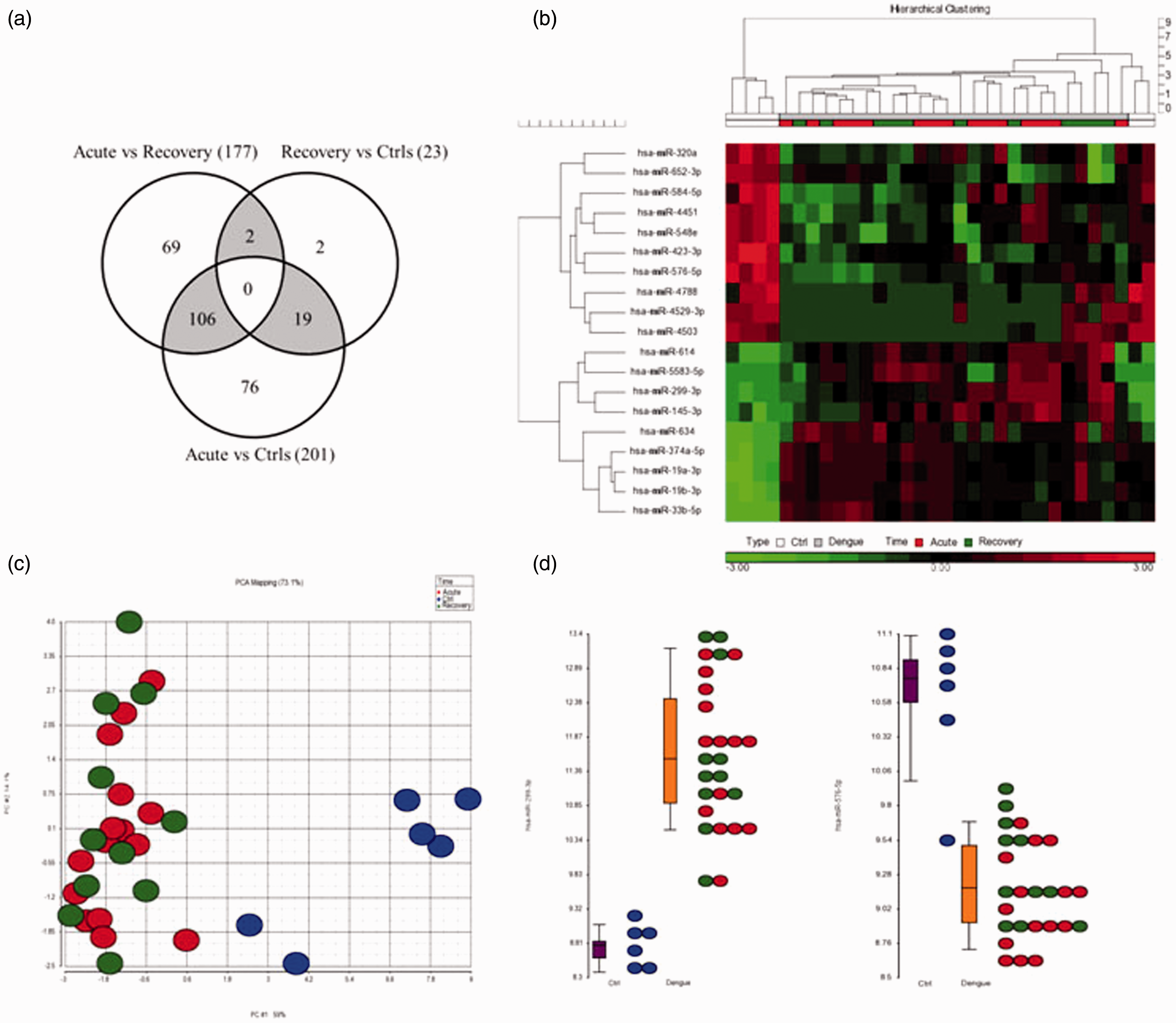
We also observed 106 miRNAs to be specifically altered during acute phase of dengue (Supplementary Table 3). Thus, this cluster of miRNAs was able to distinguish acute dengue patients from healthy as well as recovered individuals (Figure 3(a) and (b)). Within this cluster, miR-218-5p, -296-5p, -450b-5p, -499a-5p, -512-3p, -520d-3p, -520g, -524-3p, -544a, -548h-5p, -615-5p, -624-5p, -625-3p, -649, -892b -1181, -1204, -1225-5p, -1277-3p, -3937, -4259, -4327, -4446-5p, -4525 and -4640-3p were highly expressed during one to four days of dengue infection (Supplementary Figure 1). They were either undetectable or lowly expressed in healthy individuals and recovered dengue patients. On the contrary, miR-99b-5p, -491-5p, -499a-3p, -1273c, -3121-3p, -3591-3p, -4423-3p,-4434, -4444, -4490, -4501, -4694-3p, -4695-5p, -5000-5p, -5004-3p and -5004-5p were undetectable in early dengue, but expression was high in majority of the healthy controls and recovered dengue patients suggesting a suppression of these miRNAs' expression during the early phases of DENV infection (Supplementary Figure 2). Incidentally, miR-1270 and miR-1307-5p were the only miRNAs to be differentially expressed in recovered patients when compared to controls and acute dengue patients. Both miRNAs were seen to be downregulated in recovered dengue patients only (Figure 3(c)).
miRNAs differentially expressed in acute DENV infection. (a) Cluster of miRNAs differentially expressed in acute dengue patients. Heatmap shows the 106 miRNAs which were found to be differentially expressed in acute dengue patients within ≤ 4 days of infection. (b) Segregation of individuals according to the onset of infection. PCA plots show distinct segregation between acute dengue patients from healthy and recovered individuals. These 106 miRNAs reflect the responses upon the onset of infection. (c) Dot plots of miRNAs differentially expressed in recovered dengue individuals. Expression of miR-1270 and miR-1307-3p was seen to be distinctly suppressed in dengue recovered patients only.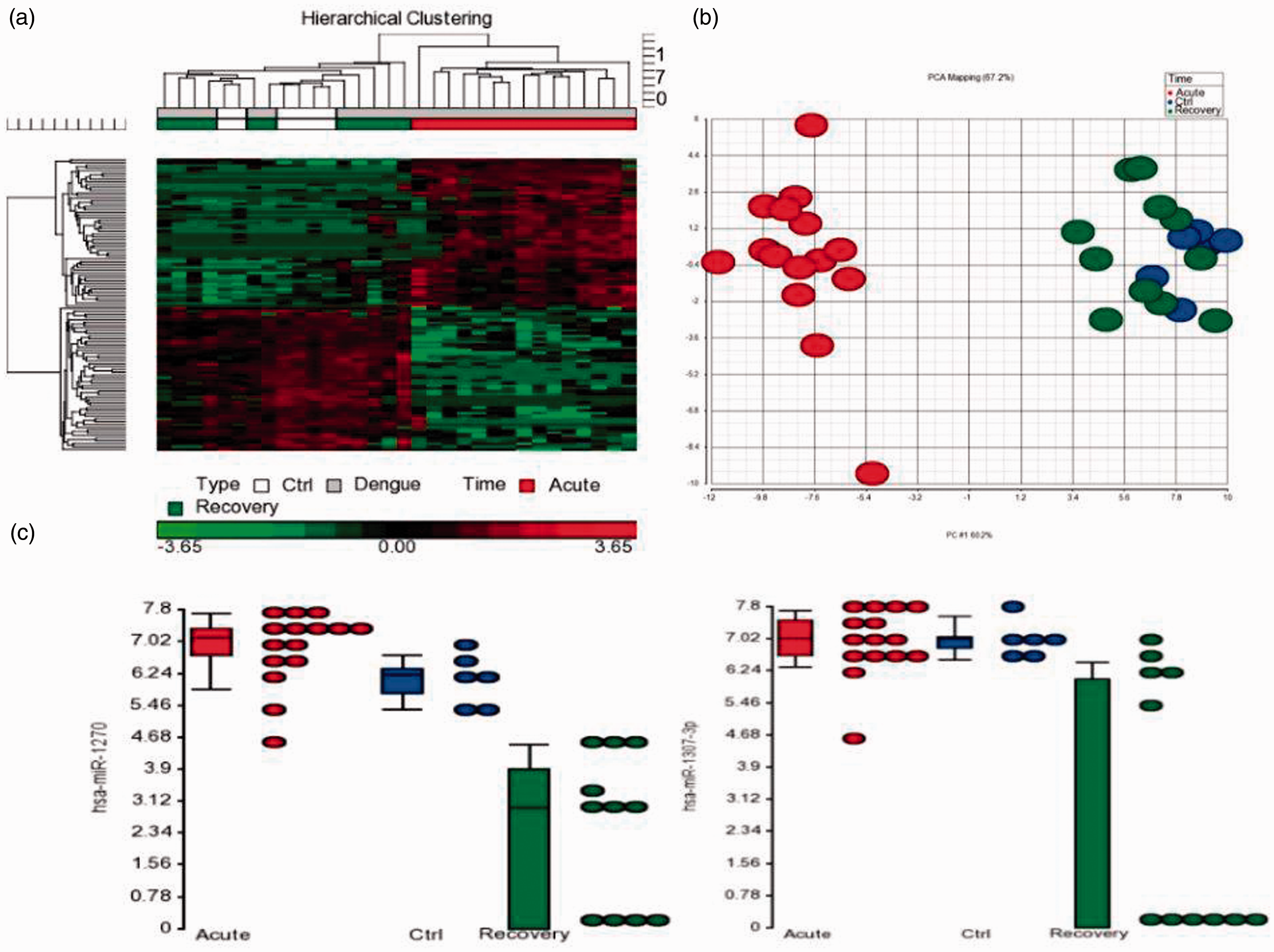
Distinction of miRNAs differentially expressed in response to acute DENV infection
Receiver operating characteristic analysis for selected miRNAs.
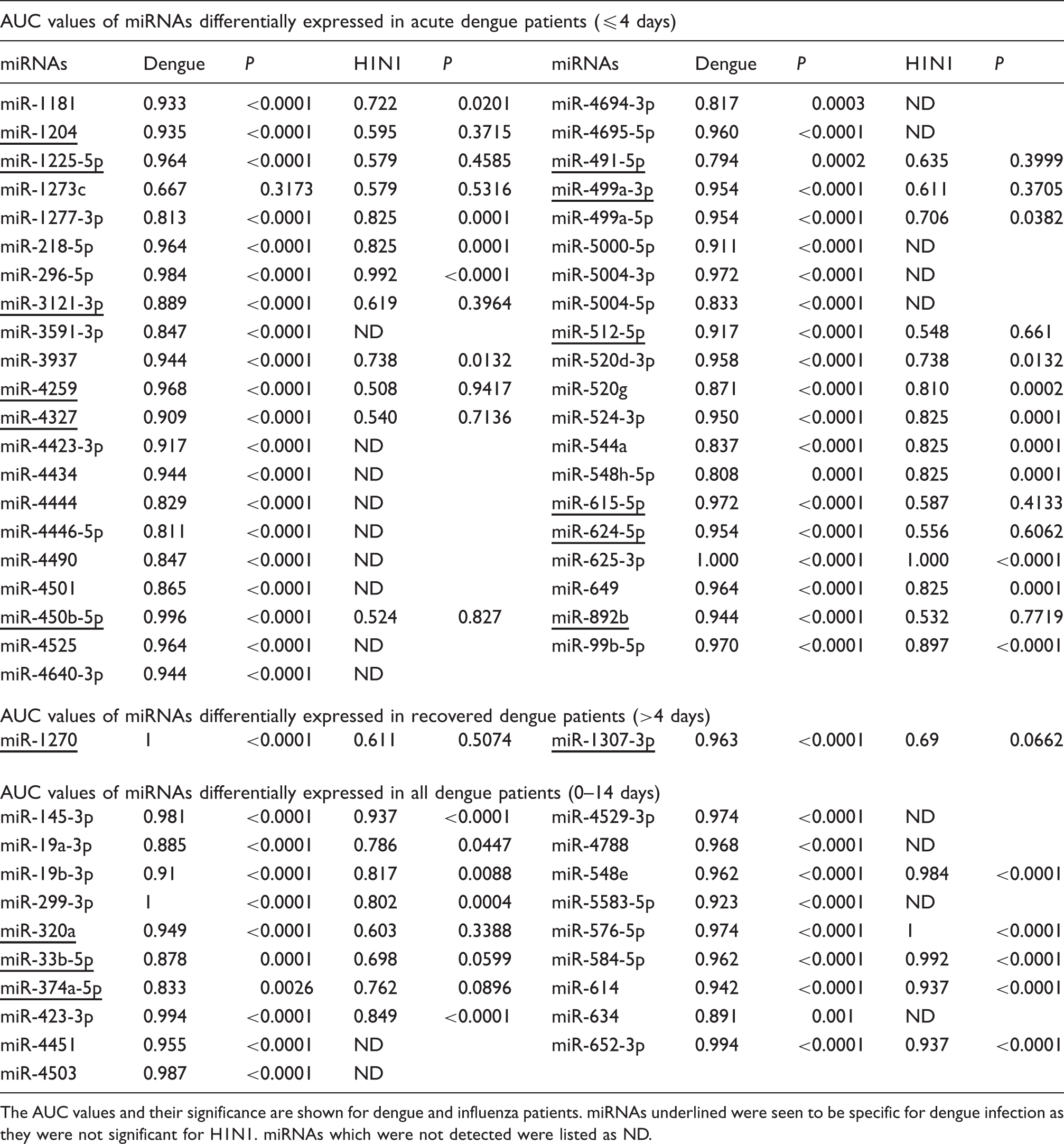
The AUC values and their significance are shown for dengue and influenza patients. miRNAs underlined were seen to be specific for dengue infection as they were not significant for H1N1. miRNAs which were not detected were listed as ND.
miRNA expression reflects complications observed in DENV infection
miRNAs in dengue complications.
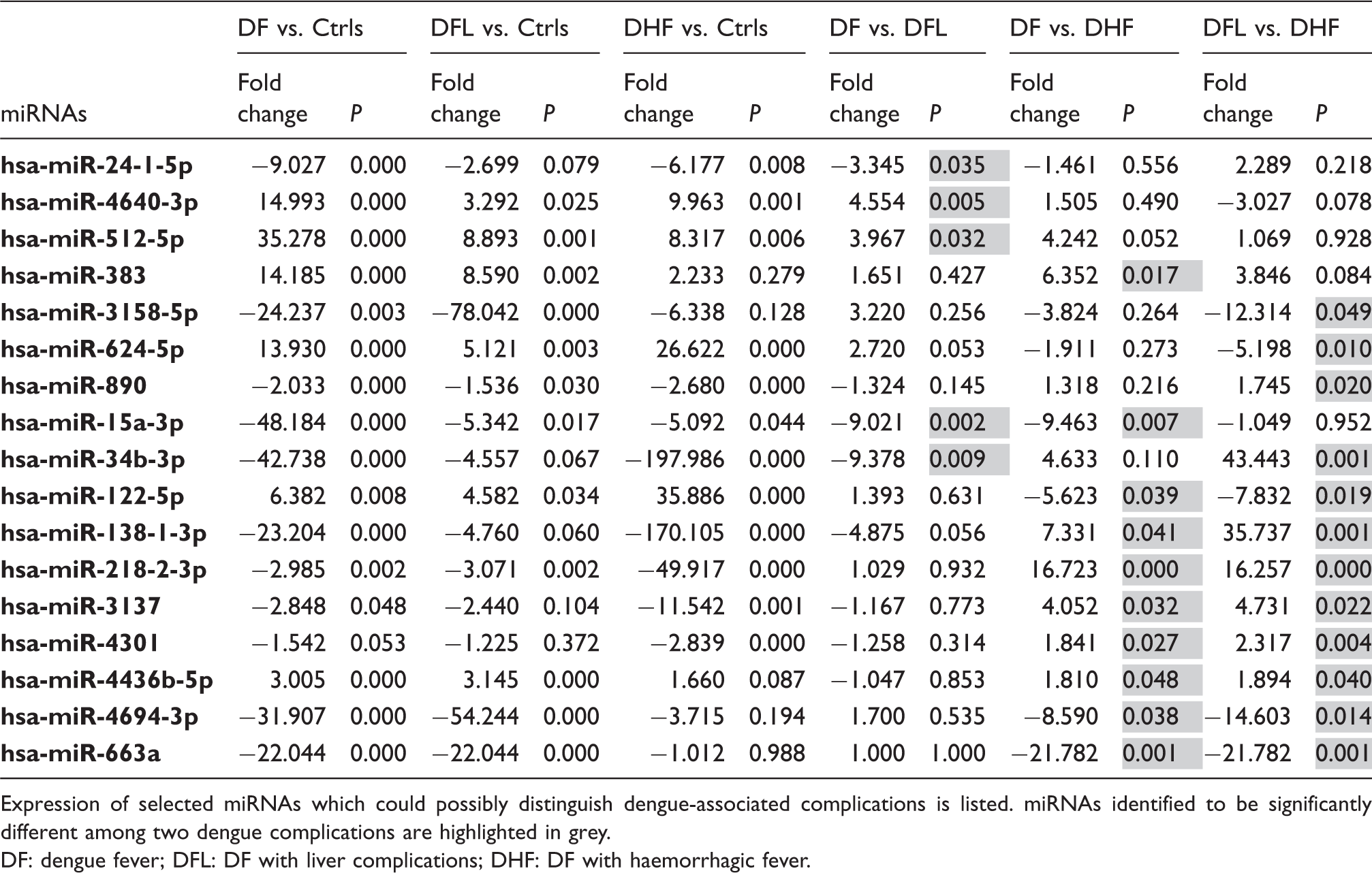
Expression of selected miRNAs which could possibly distinguish dengue-associated complications is listed. miRNAs identified to be significantly different among two dengue complications are highlighted in grey.
DF: dengue fever; DFL: DF with liver complications; DHF: DF with haemorrhagic fever.
Pathway analysis of acute phase miRNAs
Among the 41 miRNAs specifically (P ≤ 0.0005) altered in the acute phase of DENV infection, 27 were detected in influenza patients as well (Table 1). Of these except miR-1273c, expression of 12 miRNAs (miR-450b-5p, -491-5p, -499a-3p, -512-5p, -615-5p, -624-5p, -892b, -1204, -1225-5p, -3121-3p, -4259 and -4327) were seen to be specific to dengue whereas the remaining 14 miRNAs (miR-99b-5p, -218-5p, -296-5p, -499a-5p, -520d-3p, -520g, -524-3p, -544a, -548h-5p, -625-3p, -649, -1181, -1277-3p and -3937) were seen to be a consequence of general response to viral infection. Thus to determine the differences in the targets of these clusters of miRNAs, a pathway analysis was performed using Diana prediction software. 16 The cluster of 12 miRNAs specific for DENV infection showed that a total of 73 pathways were significantly altered with PI3K-AKT (P < 0.0005) being the most affected pathway. This was followed by Jak-Stat and cytokine–cytokine receptor signalling. Analysis using the 14 miRNAs cluster commonly altered in both dengue and influenza, showed that the adherens junction pathway was the most significantly (P < 0.0005) altered followed by Ubiquitin-mediated proteolysis (Supplementary Figure 3).
Discussion
Blood miRNA profiles of dengue patients reflect reports on the cellular studies
Limited vector surveillance and control measures coupled with the lack of an effective vaccine have resulted in the resurgence of dengue, thus making DENV infection a worldwide threat. Although dengue pathophysiology has been extensively studied, very little information is available on the changes occurring in the global miRNA expression patterns upon infection. Hence, using global miRNA expression (microarray) analysis of blood of dengue patients, we performed a detailed analysis of the changes occurring in the host miRNA transcriptome upon acute dengue infection till the recovery phase. Despite the limited number of samples, we were able to show that extensive changes in the expression of blood miRNAs profiles in infected patients. The changes in miRNA expression were vastly different between patients in their acute infection stage (≤ 4 days) compared to those in their recovery state (> 4 days). Infection of peripheral blood mononuclear cells with DENVs showed increased expression for several miRNAs including miR-625-3p, -767-5p and-1200. 17 Qi et al. 17 reported that the expression of miR-625-3p, -767-5p and -1200 was increased by 8.9, 3.2 and 6.9 fold respectively. Reflecting this, we also observed that expression of miR-625-3p, -767-5p and -1200 were increased by 16.8, 3.2 and 10.8 fold in total blood of dengue-infected patients. Wu et al. 18 reported that miR-146a facilitated DENV replication by reducing interferon expression. Although expression of miR-146a was not detected in the blood of our dengue patients, miR-146b, an isomiR of miR-146a, was seen to be highly expressed. Furthermore, increased expression of miR-299-3p was associated with replicative senescence of endothelial cells, 19 a phenomenon also seen with DENV infection. 20 This was also evident in our blood profiling data suggesting that the patients’ blood profiles were highly relevant and reflective of the pathology.
miRNAs specific for acute DENV infection
We also observed extreme upregulation of miR-296-5p in acute dengue patients with no significant changes in recovered dengue patients (Supplementary Table 3). Increased expression of miR-296-5p was also seen in Enterovirus 71 infection and it was shown to inhibit viral replication. 21 Furthermore, we had previously seen that miR-625-3p was upregulated in influenza patients whereas miR-423-3p was downregulated. 15 As a viral infection will trigger a cascade of defence and immune responses, we attempted to establish that the changes seen in the miRNA expression were specific to dengue. Thus, using existing data of influenza patients (another RNA virus), we compared the specificity of the miRNA expression to distinguish dengue patients from those infected with influenza virus as a type of a non-specific viral infection. We found that 27 out of the selected 41 acute (dengue) phase miRNAs were also detected in influenza patients. Using receiver operating characteristic curves, expression patterns of 12 miRNAs (miR-450b-5p, -491-5p, -499a-3p, -512-5p, -615-5p, -624-5p, -892b, -1204, -1225-5p, -3121-3p, -4259 and -4327) were seen to be significant and specific for acute dengue cases.
While not all miRNAs in this cluster have had their functional roles established, we observed strong correlation between the expression patterns of several miRNAs with possible implications for DENV infection.
miR-1204, (together with miR-1205, miR-1206, miR-1207 and miR-1208) is encoded from PVT1 a non-protein coding locus of more than 300 kb and transcription of PVT1 is regulated by p53 which binds at about 1200 bp downstream from the PVT1 transcriptional start site (Figure 4).
22
Thus, an increase in p53 expression will induce over-expression of these miRNAs and this was seen in our profiling data. All four miRNAs except miR-1205 were significantly upregulated in dengue patients. Presence of p53 was considered to be a prerequisite for efficient induction of apoptosis in the infected cells.
23
Induction of apoptosis via p53 upregulation is considered as a survival mechanism to prevent the spread of the virus. Nasirudeen et al.
23
showed that active dengue viral replication-induced apoptosis in various cell lines via the upregulation of p53 expression. One of the possible mechanisms by which p53-induced apoptosis occurred was via miR-1204 as over-expression of miR-1204 resulted in cell death which was reversed with the use of miR-1204 inhibitors.
22
Implication of miRNA expression in cell survival. A schematic representation showing the effect of (dengue specific) miR-1204 and miR-491-5p expression on the survival of dengue-infected cells.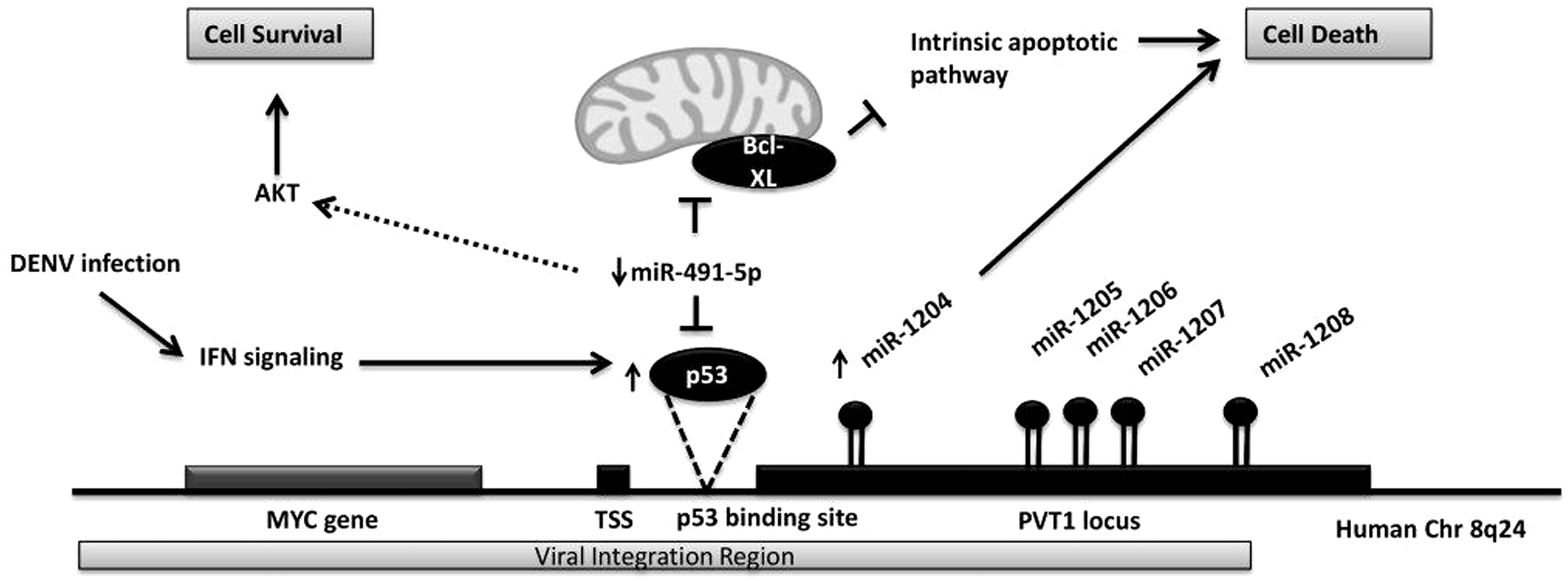
miR-491-5p was also shown to be a regulator of p53. 24 Intriguingly, it targets both p53 a tumour suppressor and Bcl-XL an anti-apoptotic gene. Nonetheless, the repressive effect of miR-491-5p was stronger on Bcl-XL than on p53. In doing so, miR-491-5p would activate the mitochondrial apoptotic pathway and eventually indirectly inhibit the P13K/AKT survival pathway. We observed downregulation of miR-491-5p in dengue patients suggesting upregulation of Bcl-XL expression which will in turn inhibit the mitochondrial apoptotic pathway and activate the P13K/AKT survival pathway. Incidentally, pathway analyses for the entire cluster of 12 miRNAs also showed that the P13K/AKT pathway was the most significantly regulated. However, it is highly possible that downregulation of miR-491-5p would also relive its suppression on p53 which could in turn induce miR-1204 expression by triggering PVT1 transcription. Since miR-1204 is known to induce apoptosis, the balance between the two processes could ultimately determine the final outcome of the cell fate.
miR-512-5p was also highly expressed in acute dengue patients, and it targets the nucleotide-binding oligomerization domain containing 2 (NOD2) gene. 25 It is an important immune response gene involved in various roles such as induction of autophagy, antiviral responses as well as T-cell activation. 26 NOD2 suppression leads to repression of NF-κB, JNK and MAPK pathways as well as interleukins and cytokines. 25 MAPK signalling and cytokine–cytokine receptor interactions were also some of the significantly altered pathways in our analysis. Furthermore, suppression of NF-κB activation has been shown in dengue infection, which will result in reduced cytokine expression. 14 Concurrently, activation of P13K is essential for NF-κB activation. 27
Increased expression for monocyte chemotactic protein 2 (MCP-2), interferon gamma-induced protein 10 (IP-10) and tumour necrosis factor-related apoptosis-inducing ligand (TRAIL) has been observed in acute dengue patients. Increased expression of these three markers was noted in the cellular level as well as serum samples. 28 Predictive bioinformatics analyses revealed that MCP-2 was targeted by dengue-specific miRNAs, selected in this study. MCP-2 was predicted to be targeted by miR-450-5p and miR-499-3p. Interestingly, miR-499-3p was predicted to have three binding sites in the MCP-2 3′UTR and its expression was downregulated in our patient profiles, suggesting a greater possibility of interaction between these two transcripts.
Significant changes in the expression of miR-99b-5p, -218-5p, -296-5p, -499a-5p, -520d-3p, -520g, -524-3p, -544a, -548h-5p, -625-3p, -649, -1181, -1277-3p and -3937 were seen for both dengue and influenza, yet the processes regulated by these 14 miRNAs were vastly different from those regulated by the cluster of 12 miRNAs. Pathway analyses of the cluster of 14 miRNAs yielded adherens junction and ubiquitin-mediated proteolysis as the top hits. In fact, a recent study on chikungunya infection also showed that adherens junction and ubiquitin-mediated proteolysis were significantly altered pathways. 29 The similarities in the pathway prediction highly implicates that these 14 miRNAs are involved in functional roles which are common upon viral infection.
miRNAs involved in dengue complications
DF ranges from mild to severe and involves several complications. However, no accurate markers are available to predict the outcome of the disease at the early stages of infection. Thus, we attempted to identify miRNAs that were differentially expressed within dengue patients with various complications. A panel of 17 miRNAs was identified with significantly different expression among patients with dengue and warning signs (Table 2). The expression patterns of the miRNAs implicated in dengue complications showed varying degrees of dysregulation. For instance, miR-218-2-3p was downregulated in DF, DFL and DHF patients by 2.9, 3.1 and 49.9 fold, respectively. Similarly, miR-122-5p was upregulated by 6.3, 4.6 and 35.8 fold in these patient groups respectively. These observations are in line with findings of Ubol et al. 30 The authors reported on the genome-wide transcriptomes of peripheral blood mononuclear cells collected from children with acute phase DF or acute phase DHF. Analysis of the data revealed that the differentially altered genes between DF and DHF did not show opposite expression patterns but rather various degree of upregulation or downregulation with respect to each other. 30 This was also seen in several of the dengue marker genes reported by Silva et al. 31 The authors reported that CFD, MAGED1, PRDX4, PSMB9 and FCGR3B to be differentially expressed in DHF patients compared to DF patients. In an independent report Nascimento et al. 32 also showed that MAGED1, PSMB9 and FCGR3B were differentially altered between DF and DHF patients. However, expression patterns of these genes in the two studies were inversely correlated.
While the events leading to DHF in patients infected with DENV have not been clearly elucidated, it is often thought to be mediated by the infected individual’s immune response rather than the effect of the virus. Severe dengue patients generally exhibited higher neutrophil-associated proteins when compared to milder dengue patients. 33 If this is so, then regulating expression of immune response genes during the acute phase of the infection is critical. This can be achieved by regulating the corresponding miRNAs which control the expression these genes. This study highlights the acutely changing miRNA profiles following DENV infection, and thus unlocks the possibility of further exploring and regulating its crucial targets in relation to dengue pathophysiology.
Conclusions
In summary, our study reports on the changes occurring in the blood miRNA transcriptome of dengue patients that have not been previously described. A cluster of 12 miRNAs were identified to be specific for acute dengue and several of these miRNAs been reported as regulators of crucial genes involved in DENV infection and pathogenesis, thus emphasizing the importance of these miRNAs in dengue infection. Concurrently, we have highlighted 14 miRNAs with possible implications in viral infection. Further exploration of the functional roles of these miRNAs will enable greater understanding of the dengue pathology and subsequently its related complications.
Footnotes
Acknowledgements
Our sincere thanks to the patients and clinical support staff from the National University of Singapore for their contributions.
Declaration of conflicting interests
PAT has received research support from Baxter, ADAMAS, Fabentech, GSK and Sanofi-Pasteur. He has also received honoraria from Astra-Zeneca, Novartis and Johnson and Johnson. All the other authors report no potential conflicts.
Funding
This work was supported and funded by the Singapore National Medical Research Council [R-172-000-211-213].
Ethical approval
The study protocol was approved by the National Healthcare Group’s Domain Specific Institutional Review Board (NUS-IRB Ref Code: 09/641).
Guarantor
PAT.
Contributorship
KJ, PAT and AA conceived and designed the research. PAT contributed in patient recruitment/management and clinical diagnosis. JMA contributed in sample collection and clinical data compilation. CSC extracted all the blood RNA and carried out the qPCR validation analysis/studies. AA performed the miRNA microarray and analysis. SS, AA and KJ were involved in analysis and interpretation of data, drafting the manuscript or revising it critically for important intellectual content. All authors contributed in writing the paper. All authors read and approved the final manuscript.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
