Abstract
Background
Assays for cardiac troponin (cTn) have undergone improvements in sensitivity and precision in recent years. Increased rates of outliers, however, have been reported on various cTn platforms, typically giving irreproducible, falsely higher results. We aimed to evaluate the outlier rate occurring in patients with elevated cTnI using a contemporary and high-sensitivity assay.
Methods
All patients with elevated cTnI (up to 300 ng/L) performed over a 21-month period were assayed in duplicate. A contemporary assay (Abbott STAT Troponin-I) was used for the first part of the study and subsequently a high-sensitivity assay (Abbott STAT High-Sensitive Troponin-I) was used. Outliers exceeded a calculated critical difference (CD) (CD = z × √2 × SDAnalytical) where z = 3.5 (for probability of 0.0005) and critical outliers also were on a different side of the decision level.
Results
The respective outlier and critical outlier rates were 0.22% and 0.10% for the contemporary assay (n = 4009) and 0.18% and 0.13% for the high-sensitivity assay (n = 3878). There was no significant reduction in outlier rate between the two assays (χ2 = 0.034, P = 0.854). Fifty-six percent of outliers occurred in samples where cTn was an ‘add-on’ test (and was stored and refrigerated prior to assay).
Conclusion
Despite recent improvements in cTn methods, outliers (including critical outliers) still occur at a low rate in both a contemporary and high-sensitivity cTnI assay. Laboratory and clinical staff should be aware of this potential analytical error, particularly in samples with suboptimal sample handling such as add-on tests.
Introduction
Assays for cardiac troponin (cTn), the preferred biomarker for the diagnosis of acute myocardial infarction, have undergone improvements in sensitivity and precision in recent years particularly at concentrations near the decision level (99 th percentile of a normal population).1,2 New ‘high-sensitivity’ cTn assays have been introduced that detect cTn in a majority of normal subjects. 2 Increased rates of outliers, however, have been reported on various cTn platforms in both plasma3,4 and serum,3,5 typically giving an irreproducible, falsely higher cTn result in the first or second analysis.3–5 A previous study using a contemporary cTnI assay from the same manufacturer as our study, found the majority of outliers had values ≤260 ng/L. 4
We aimed to evaluate the outlier rate in patients with elevated cTnI levels, where outliers are more likely, using a contemporary and a high-sensitivity assay. Our approach was to assay elevated cTnI results (up to 300 ng/L) in duplicate over a 21-month period.
Methods
Study design
All cTnI requests performed at Canterbury Health Laboratories over a 21-month period were available for inclusion in the study. Samples were collected as lithium heparin plasma (BD Vacutainer® PST™ II Tube) (BD, Franklin Lakes, NJ, USA) and promptly centrifuged (2800 g for 10 min in a swinging-bucket centrifuge). A contemporary assay was used for the first part of the study (3 April 2012–23 April 2013) and subsequently a high-sensitivity assay was used (23 April 2013–28 December 2013). Values 30–300 ng/L (0.03–0.30 µg/L) for the contemporary assay and 16–300 ng/L for the high-sensitivity assay were assayed in duplicate. This was performed by automatic reflex testing with samples held at ambient temperature following the initial assay and then repeat testing (without re-centrifugation) occurring within 10 min. Where the repeat was substantially different from the initial result, the sample was re-assayed (if sample volume permitted) so the outlier value could be identified from the replicates.3–5
Reagents and materials
The contemporary assay used was the STAT Troponin-I (Abbott Diagnostics, Abbott Park, IL, USA) and the high-sensitivity assay was the Abbott STAT High Sensitive Troponin-I. Assays (reagent pack size 500) were both performed on either of two Abbott ARCHITECT i2000 SR analysers attached to an Abbott Accelerator Automated Processing System (APS).
The assay decision levels used in our laboratory are ≤30 ng/L (0.03 µg/L) for the contemporary assay and ≤16 ng/L in females and ≤34 ng/L in males for the high-sensitivity assay. The assay 10% coefficient of variation (CV) levels (and detection limits) are 32 ng/L (10 ng/L limit of blank) for the contemporary assay and 4.7 ng/L (1.9 ng/L limit of detection) for the high-sensitivity assay. Duplicate result data, extracted weekly, were available for 46/55 weeks of the study using the contemporary assay and 32/36 weeks using the high-sensitivity assay.
Quality Control (QC) samples used for the contemporary assay were Thermo Scientific MAS Cardioimmune XL controls level 1, 2, 3 (Thermo Fisher Scientific, Melbourne, Australia) with overall CV for both analysers of 29% (36 ng/L), 10% (200 ng/L) and 8% (704 ng/L) and for the high-sensitivity assay Liquichek Cardiac Markers Plus Control LT level low, 1, 2, 3 (Bio-Rad Laboratories, Hercules, CA, USA) with overall CV of 12% (16.6 ng/L), 9% (52.6 ng/L), 7% (1237.3 ng/L) and 7% (6732.9 ng/L). QC data were extracted from Unity Real Time® (Bio-Rad Laboratories).
Statistical analysis
Outliers exceeded a critical difference (CD) (CD = z × √2 × SDAnalytical) where z = 3.5 (for CD with probability of 0.0005) and critical outliers also were on a different side of the decision level.3,4 SD was interpolated from QC data (i.e. linear regression of overall SD versus concentration)3–5 and given for the lowest concentration of the pair. 3 The results below the lowest QC were assumed to have the same SD as that QC3,5 and the highest QC sample for the high-sensitivity assay (6732.9 ng/L) was not included. Singlet measurement outliers were those outliers identified to occur in the initial result.3,5
Chi-square test statistic was used to compare outliers between the two assays and to the expected outlier number (at P = 0.0005) and statistical analysis was performed using software from SigmaPlot (Systat Software, San Jose, CA, USA).
Results
The respective outlier and critical outlier rates were 0.22% (9/4009) and 0.10% (4/4009) for the contemporary assay and 0.18% (7/3878) and 0.13% (5/3878) for the high-sensitivity assay (Figure 1). There was no significant reduction in the proportion of outliers between the two assays (χ2 = 0.034, P = 0.854). The observed outlier numbers were > 4 times (χ2 = 3.277, P = 0.070) and >3 times (P = 0.1764 using Fisher’s exact test) higher than predicted (at probability of 0.0005) for the contemporary and high-sensitivity assays, respectively, although this did not reach statistical significance.
Comparison of cTnI results from duplicate analysis of patient samples using (a) a contemporary generation assay (concentration 30–300 ng/L, n = 4009) and (b) a high-sensitivity assay (concentration 16–300 ng/L, n = 3878). Outliers are shown as open circles.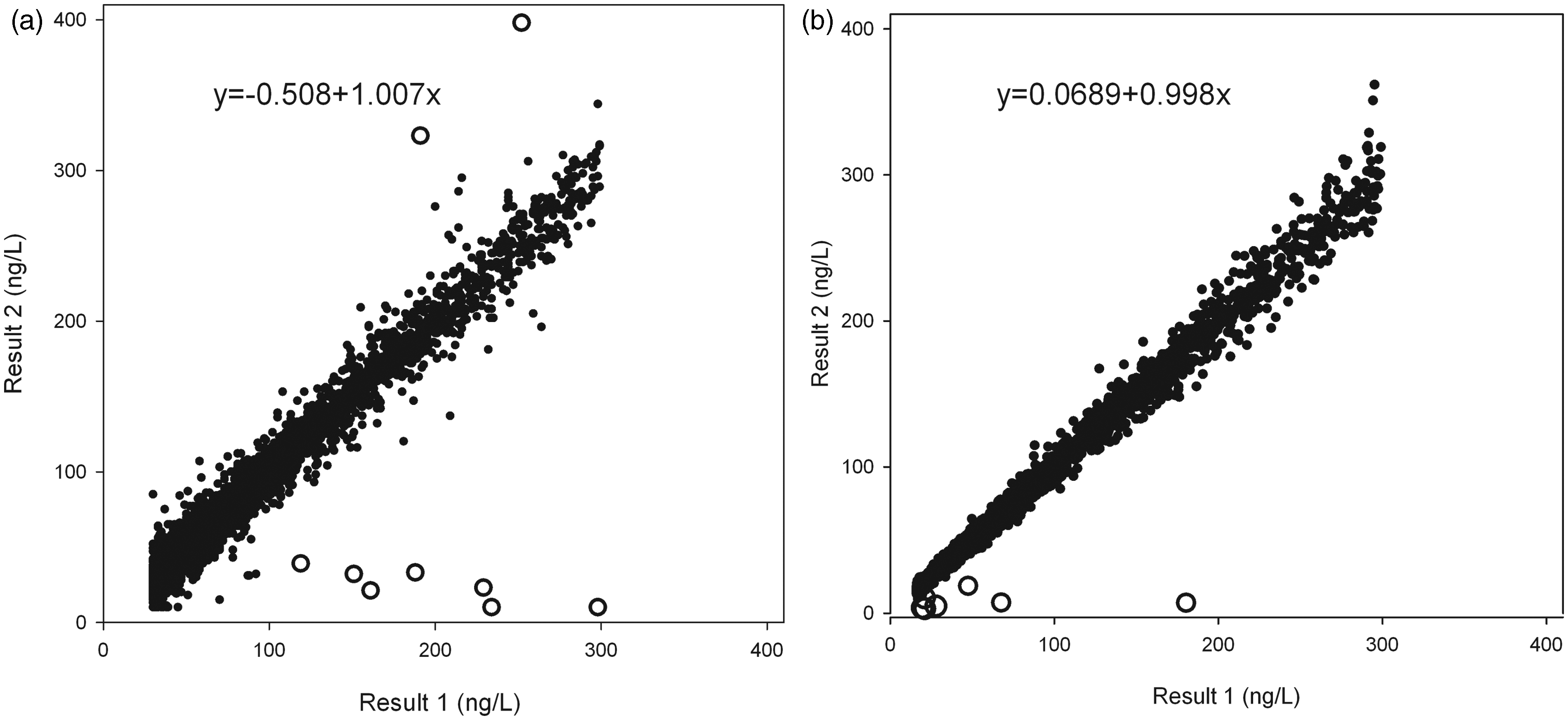
Replicate results for 16 outlier cTnI results identified by repeat analysis of patient samples using a contemporary generation (n = 4009) and a high-sensitivity (n = 3878) assay.
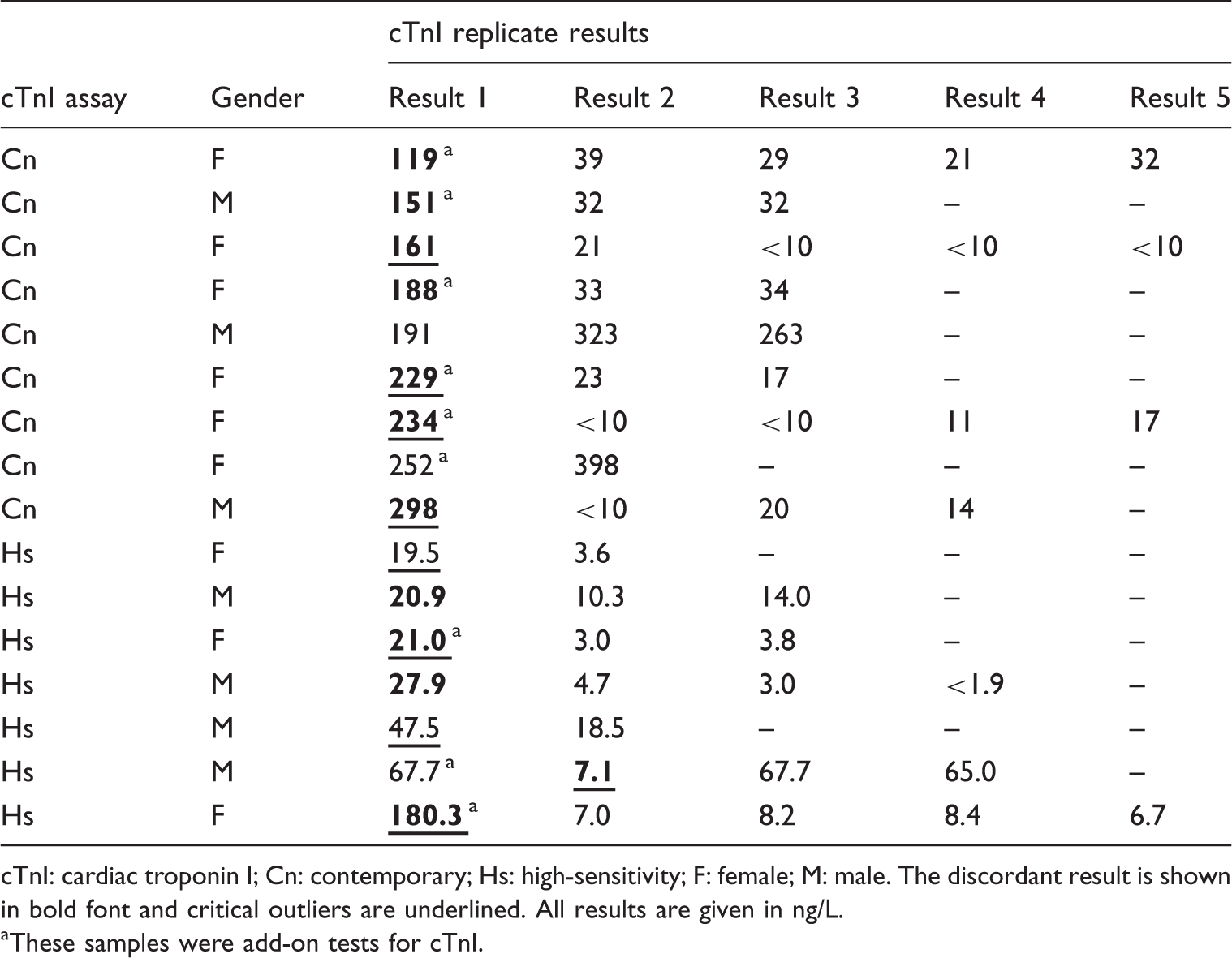
cTnI: cardiac troponin I; Cn: contemporary; Hs: high-sensitivity; F: female; M: male. The discordant result is shown in bold font and critical outliers are underlined. All results are given in ng/L.
These samples were add-on tests for cTnI.
Fifty-six percent (9/16) of all outliers (six with contemporary assay, three with high-sensitivity assay) were add-on tests (defined as a delay in the request for cTn resulting in samples being stored and refrigerated for any length of time prior to assay). The median delay between collection and request for cTnI was 5.58 h (range 1.48–8.73 h). Consequently, from 30 May 2013, our laboratory (and study) sample handling protocol was modified to include re-centrifugation of an aliquot of plasma sample in an Eppendorf Minispin microcentrifuge (Eppendorf, Hamburg, Germany) at 13,000 rpm for 2 min, for all cTnI add-on requests prior to assay. Following this modification only one of the four subsequent outliers was an add-on test; however, in this sample the re-centrifugation step was not followed.
One outlier sample had a borderline elevated icterus index but no other outliers had haemolysis, icterus or lipaemia indices above the manufacturer’s quoted thresholds for interference (data available for 14/16 samples). The outliers occurred on both of our analysers (6/12 identifiably false results occurred on each). The most common source of referrals of the outlier samples was the Emergency Department, accounting for 7/16 outlier samples.
In the study data collection period, there were 35,071 cTnI requests (20,487 for the contemporary assay and 14,584 for the high-sensitivity assay) with 46.5% being female and mean patient age 67.2 years (SD 16.3). The proportions of abnormal results using the above decision levels (and results >300 ng/L) were 29.6% (6059/20,487) (11.8% (2421/20,487)) for the contemporary assay and 32.8% (4783/14,584) (11.7% (1713/14,584)) for the high-sensitivity assay. Precision profiles for both assays from our study duplicates data are available in the Data Supplement that accompanies the online version of this article at http://acb.sagepub.com/.
Discussion
Our study found outliers occurred at similar low rates in both assays. This was supported by our data showing outlier (and critical outlier) rates of 0.22% (0.10%) and 0.18% (0.13%) in samples with elevated cTnI results (up to 300 ng/L) using a contemporary and high-sensitivity assay, respectively. The singlet measurement outlier rates in our study were 0.17% for the contemporary assay and 0.10% for the high-sensitivity assay.
Recent studies have reported overall cTn outlier rates of 0.48–1.95% using Abbott cTnI assays in plasma samples 4 and singlet measurement outlier rates of 0.06–0.44% on various cTn platforms in serum samples. 5 One of these studies using the same assay manufacturer, platform and APS as our study found overall outlier rates with the contemporary assay of 1.95% (n = 7011) and after an enhanced instrument maintenance protocol 0.48% (n = 7089), and using the high-sensitivity assay 0.59%. 4 The critical outlier rates in their study were 0.51% and 0.37% using the contemporary assay but 0% using the high-sensitivity assay, although the sample size for their high-sensitivity assay study was smaller (n = 1522). 4 Another study using a contemporary cTnI assay from the same manufacturer in serum samples had an overall outlier rate of 0.21% (5/2391) (and singlet measurement outlier rate of 0.10%) but the duplicates were performed on a single instrument as a daily batch analysis. 5
Our study only evaluated for outliers in samples with elevated cTnI values (up to 300 ng/L) as previous studies have shown that the vast majority of cTn outliers give a falsely higher result3–5 (thus normal results were excluded) and using a contemporary Abbott cTnI assay the majority (91.1–96.4%) of outliers occur at ≤260 ng/L. 4 The outlier rates in our study, therefore, cannot be directly compared with the overall outlier rates reported in other studies. An estimate of our overall outlier rate, however, was made by applying our study outlier rates to all elevated results obtained during the study period (i.e. 6361 results were ≥30 ng/L for the contemporary assay, 5613 results were ≥16 ng/L for the high-sensitivity assay) and dividing this by the total number of cTnI requests. The resulting estimated overall (and overall singlet measurement) outlier rates are 0.07% (0.05%) and 0.07% (0.04%) for the contemporary and high-sensitivity cTnI assay, respectively. These rates are lower than those reported with similar assays4,5; however, in our study duplicates were performed on either of two Abbott Architect instruments (as is our current lab protocol); therefore, larger overall imprecision data (from both instruments) were used for CD calculation, potentially reducing outlier detection. In addition, 4/16 outliers in our study had insufficient replicate data to identify the outlier result, therefore, potentially underestimating the singlet measurement outlier rate.
Outliers are irreproducible, typically falsely higher results and currently their cause is unknown. Fifty-six percent of the outliers in our study were add-on tests that were stored and refrigerated prior to assay, suggesting suboptimal sample handling may contribute to the occurrence of outliers. This is consistent with a recent study indicating sample-related factors were the cause of outliers, as no outliers occurred in concurrently tested QC material samples (n = 4767). 4 Although one suggested explanation for the outliers is fibrin formation in the sample (i.e. ‘micro-clot’), 6 this is speculative and evidence for this being causative is sparse. In our laboratory, current practice is to re-centrifuge (in a microcentrifuge) all add-on cTnI requests prior to assay.
As assay imprecision is used to define outliers in this study, assays with greater precision will identify errors that are relatively smaller in size (as illustrated in Figure 1) and this should be taken into consideration when comparing outlier rates between methods. One recent study suggested a cTnI absolute change of ≥28 ng/L (at 3 h) could be used to ‘rule in’ acute coronary syndrome in patients presenting to the Emergency Department with recent onset chest pain. 7 We applied this absolute cut-off to our duplicate data and found the outlier rate using the high-sensitivity assay of 0.77% (30/3878) was significantly lower (χ2 = 27.5, P = <0.001) than the contemporary assay rate of 2.24% (90/4009) in our samples with elevated cTnI.
Outliers are a potential source of analytical error that can have significant impact on clinical decision making. The occurrence of outliers is not detected in assay QC imprecision data as QC material may be unaffected 4 (and outliers are usually excluded from %CV calculation 3 ); therefore, outliers may only be detected with repeat testing (although this is associated with increased reagent costs and lengthened turn-around times).
In conclusion, despite recent improvements in cTn assay precision and sensitivity, irreproducible outliers (including critical outliers) still occur at a low rate in both a contemporary and high-sensitivity cTnI assay. Laboratory and clinical staff should be aware of this potential analytical error in cTn assays and have a protocol in place to identify it –particularly in samples with suboptimal sample handling (such as add-on tests).
Footnotes
Declaration of conflicting interests
None.
Funding
This research received no specific grant from any funding agency in the public, commercial, or not-for-profit sectors.
Ethical approval
Not required.
Guarantor
PMG.
Contributorship
All authors were involved in designing the study. JBR researched the literature. JBR and SS analysed the data. JBR wrote the first draft of the manuscript. All authors reviewed and edited the manuscript and approved the final version of the manuscript.
