Abstract
Molecular imaging is a relatively new discipline, which developed over the past decade, initially driven by in situ reporter imaging technology. Noninvasive in vivo molecular–genetic imaging developed more recently and is based on nuclear (positron emission tomography [PET], gamma camera, autoradiography) imaging as well as magnetic resonance (MR) and in vivo optical imaging. Molecular–genetic imaging has its roots in both molecular biology and cell biology, as well as in new imaging technologies. The focus of this presentation will be nuclear-based molecular–genetic imaging, but it will comment on the value and utility of combining different imaging modalities. Nuclear-based molecular imaging can be viewed in terms of three different imaging strategies: (1) “indirect” reporter gene imaging; (2) “direct” imaging of endogenous molecules; or (3) “surrogate” or “bio-marker” imaging. Examples of each imaging strategy will be presented and discussed. The rapid growth of in vivo molecular imaging is due to the established base of in vivo imaging technologies, the established programs in molecular and cell biology, and the convergence of these disciplines. The development of versatile and sensitive assays that do not require tissue samples will be of considerable value for monitoring molecular–genetic and cellular processes in animal models of human disease, as well as for studies in human subjects in the future. Noninvasive imaging of molecular–genetic and cellular processes will complement established ex vivo molecular–biological assays that require tissue sampling, and will provide a spatial as well as a temporal dimension to our understanding of various diseases and disease processes.
Keywords
Introduction
“Molecular imaging” is a term that developed in the 1990s, with roots that go back to in situ visualization of target molecules and biological processes (in situ optical imaging) [1–4]. Molecular–genetic imaging has evolved and has become a more broadly defined term over the past decade; it includes studies that were previously described as “gene imaging” or “reporter gene imaging”, and now relates to other aspects of molecular and cell biology. Recent advances have included the development of noninvasive in vivo molecular imaging in animals, including transgenic animals.
The involvement of three different noninvasive, in vivo imaging technologies developed more or less in parallel: (1) magnetic resonance (MR) imaging [5–9]; (2) nuclear imaging (QAR, gamma camera, and positron emission tomography [PET]) [10–14]; and (3) optical imaging of small animals [15–17]. These developments led to the term “in vivo molecular imaging,” which was coined in the mid 1990s. Molecular–genetic imaging has its roots in both molecular biology and cell biology as well as in imaging technology. These disciplines have now converged to provide a well-established foundation for exciting new research opportunities and for translation into clinical applications. For instance, established ex vivo molecular assays require invasive sampling procedures that preclude sequential studies in the same animal or in human subjects. Furthermore, temporal studies that employ molecular–biological assays have required large numbers of animals that are sacrificed at specific time points in order to achieve a statistically significant temporal profile. The development of sensitive imaging-based assays to monitor molecular–genetic and cellular processes over time can provide a better understanding of disease development and progression in both animal models of human disease (including transgenic animals). Noninvasive imaging of molecular–genetic and cellular processes will complement established ex vivo molecular–biological assays, and imaging can provide a spatial as well as a temporal dimension to our understanding of various disease processes.
The principles of reporter gene imaging have been applied to all three noninvasive imaging technologies and these developments have occurred more or less in parallel. Each imaging technology has specific advantages and limitations. For example, in vivo optical imaging is relatively inexpensive and can be performed repetitively in the same animal. Current optical imaging technology is largely limited to imaging small animals (e.g., mice or rats) [15–17], although this is likely to change rapidly with the development of new optical technologies and the inclusion of tomographic acquisition of optical data. MR has the advantage of very high spatial resolution and excellent soft tissue imaging characteristics. MR imaging can be performed repetitively over time in the same animal or human subject; it also provides the opportunity to perform a magnetic spectral analysis (MRS) of a particular tissue region to characterize its chemical composition as well as to identify specific molecular species. Initial studies have clearly demonstrated the feasibility of performing molecular imaging studies using MR imaging technology [5–9], although MR is currently more limited with respect to the number and variety of molecular and reporter gene assessments that can be performed. Nuclear imaging has the benefit of high sensitivity and localization of radioactivity in 3-D space; it can be used to directly image an assortment of biological processes as well as the expression of different reporter genes with highly specific radiolabeled probes; quantitative information can be obtained from the images and the imaging paradigms are directly transferable to large animals and human subjects [10–14]. However, nuclear imaging technology is very expensive and requires a multidisciplinary team (e.g., it requires a cyclotron and radiochemistry resources for PET imaging).
Although it may appear somewhat presumptuous to imply that current noninvasive imaging technologies (MR, PET, gamma camera, etc.) can image molecular events that occur within cells, it has already been shown that it is possible to image transcriptional regulation of endogenous gene expression [18]. Needless to say, current PET or gamma camera instruments that are used to image animals and patients do not visualize individual cells, much less molecules. What is so exciting about this emerging new field relates to the novel imaging paradigms that are being developed. These paradigms can be successful within the inherent spatial resolution limits of existing imaging systems providing the volume element (voxel) is relatively homogenous.
Recent progress in our understanding of the molecular–genetic mechanisms of many diseases and the application of new biologically based approaches in therapy are exciting new developments. New genebased therapies can provide control over the level, timing, and duration of action of many biologically active transgene products by including specific promoter/activator regulatory elements in the genetic material transferred. Noninvasive imaging of molecular–genetic and cellular processes will accelerate these developments and lead to more effective therapeutic strategies. Methods are being developed for controlled gene delivery to various somatic tissues and tumors using novel gene constructs, for targeting vectors to specific tissues/organs, and for controlling gene expression using cell-specific, replication-conditional, and drug-controlled expression systems [19–21]. A noninvasive, clinically applicable method for quantitatively imaging the expression of transduced genes in target tissue or specific organs would be of considerable value. It will facilitate the monitoring and evaluation of gene therapy in human subjects by defining the location(s), magnitude, and persistence of gene expression over time.
It should also be noted that the field of in vivo molecular imaging has enjoyed exceptional growth over the past 5 years. This rapid growth is due to several factors. There was an established base of imaging technology (nuclear, MR, and optical), established programs in molecular and cell biology, and a convergence of these disciplines developed during this period. In addition, there have been several NCI-sponsored initiatives. “Cancer imaging” was identified as one of six “extraordinary scientific opportunities” by NCI in 1997–1998. Subsequent funding initiatives from NCI have provided a major stimulus to further the development of in vivo molecular imaging. Substantial resources have been made available to the research community through NCI's Small Animal Imaging Resources Program (SAIRP) and the In Vivo Cellular and Molecular Imaging Centers (ICMIC) program. Further interaction with other programs, such as the Mouse Models of Human Cancers Consortium (MMHCC), is being encouraged and NCI is attempting to coordinate the yearly meetings of the SAIRP, ICMIC, and MMHCC programs. Similar funding initiatives have been developed by other NIH Institutes and by the Department of Energy (DOE). Furthermore, a new NIH institute—the Institute for Biomedical Imaging and Engineering—has recently been formed to expand the breadth of the imaging community in the United States.
Imaging Strategies
Three molecular imaging strategies—“direct,” “indirect,” and “surrogate”—will be described, and examples of each will be discussed. “Direct” imaging of endogenous genes and molecules can be defined in terms of a probe–target interaction, whereby the resultant image of probe localization and magnitude (image intensity) is directly related to its interaction with the target molecule, epitope, or enzyme. “Indirect” molecular-gene imaging is more complex and involves multiple components. One example of indirect imaging that is now being widely used is “reporter gene imaging.” This indirect imaging paradigm usually includes a “marker/reporter gene,” a complementary “marker/reporter probe” and imaging technology that can visualize the spatial distribution of the probe (that identifies the site of reporter gene expression). In the case of PET, quantitative information about the level of reporter gene expression can also be obtained. “Surrogate” imaging that reflects downstream effects of one or more endogenous molecular–genetic processes is also being explored. This latter approach is particularly attractive for potential translation into clinical studies in the near-term. Examples of direct, indirect, and surrogate imaging using radiolabeled probes and PET to visualize endogenous molecular processes, such as the regulation of endogenous gene expression, will be discussed below.
Direct Imaging
Examples of the direct imaging paradigm are common in nuclear medicine. PET imaging of receptor density/occupancy using small radiolabeled molecular probes is widely used, particularly in neuroscience research. Imaging cell surface-specific antigens or epitopes with radiolabeled antibodies is another well-established example of direct molecular imaging. More recently, multidrug resistance (MDR) involving P-glycoprotein (Pgp), a transmembrane protein acting as an ATP-dependent drug efflux pump, has been extensively studied. Other MDR-associated proteins include the MDR-associated protein (MRP1), the lung resistance protein (LRP), and the breast cancer resistance protein (BCRP). It has been shown that the efflux process and level of MDR activity can be imaged in both animals and human subjects using small radiolabeled probes [22–25]. The above examples represent some of the first “molecular imaging” applications, and they continue to be used in both animal and clinical nuclear medicine research.
Imaging cell-specific surface antigens or epitopes with radiolabeled antibodies was one of the first examples of “molecular imaging.” Tumor (target) cell imaging with radiolabeled proteins dates back over 30 years. More recently, the visualization of various tumor cell surface receptors using radiolabeled regulatory peptides is increasingly being pursued in oncology research. These regulatory peptides are small, readily diffusible, and potent natural substances with a wide spectrum of receptor-mediated actions. High-affinity receptors for these peptides are frequently overexpressed in many cancers and represent molecular targets for cancer diagnosis and therapy. A recent review summarizes peptide-based radiopharmaceuticals, which are presently commercially available or are in advanced stages of clinical testing [26].
Similarly, radiolabeled oligonucleotide antisense and aptomer probes have been developed to specifically hybridize with target mRNA or proteins for direct in vivo imaging. Antisense strategies involve the use of radiolabeled oligonucleotides (RASONs). RASONs are small oligonucleotide sequences that are complementary to a small segment of target mRNA or DNA, and could potentially target any specific mRNA or DNA sequence. RASON probes are being used to directly image endogenous gene expression at the transcriptional level. Aptomers are also small oligonucleotide sequences that can be used to target specific proteins. In this context, imaging specific mRNAs and proteins with RASONs provides a direct image of molecular–genetic events. Some efficacy for gamma camera and PET imaging endogenous gene expression using RASONs has been reported [27–30]. Nevertheless, RASON imaging has several serious limitations, including: (a) low number of target mRNA/DNA molecules per cell; (b) limited tracer delivery (poor cell membrane and vascular permeability, cannot penetrate the blood–brain barrier [BBB]); (b) poor stability (degradation by H-RNAse); (c) slow clearance (slow washout of nonbound oligonucleotides); (d) comparatively high background activity and low specificity of localization (low target/background ratios). Imaging specific RASON targets in the body is complicated and interpretation of the images must be approached with caution.
Another example of a commonly used direct imaging paradigm is the visualization of glucose utilization using fluorodeoxyglucose (FDG) and PET [31,32]. Clinical FDG PET images largely reflect an enzymatic reaction; namely, the activity of hexokinase—the enzyme that converts glucose and FDG to glucose-6-phosphate and FDG-6-phosphate, respectively. Since phosphorylated FDG does not significantly cross the cell membrane and is relatively stable in the cell, the radiolabeled probe accumulates in the cell over time. FDG transport does influence the early PET images, but is likely to have only a small influence on images obtained after 60 min. This is because glucose transport across most cell membranes is more rapid than the rate of phosphorylation by hexokinase and glucose transport in most cells is equilibrative (bidirectional and nonconcentrative). PET imaging and the measurement of FDG accumulation in tissue (e.g., tumor) can be used to provide information about hexokinase activity and can be used to derive a measure of glucose utilization [33–35]. FDG PET imaging is a widely used example of “direct” molecular imaging; it involves a small radiolabeled probe (FDG) that is selectively metabolized (phosphorylated) and trapped in the cell by a single endogenous enzymatic reaction (hexokinase). Since mitochondria hexokinase is the dominant rate-determining step for glucose utilization in most cells and the FDG–hexokinase enzymatic reaction determines the rate and magnitude of FDG accumulation (an enzymatic radiotracer assay), PET FDG images can be used to measure glucose utilization in the tissue.
On the other hand, FDG imaging has been routinely used as a “surrogate” imaging method for assessment of tumor localization and degree of malignancy, because many tumor have high levels of glucose utilization (Warburg effect). For example, FDG could be used as a “surrogate” marker for imaging the activities of IGFR-PI3 kinase-Akt-hexokinase(+ Glut1) and HIF1a-hexokinase(+ Glut1) pathways.
Indirect Imaging
Indirect imaging of endogenous gene expression is currently a widely used strategy. The paradigm was initially demonstrated using in situ optical imaging technology and reporter transgenes [1–3,36]. In this paradigm, the reporter transgene (e.g., Escherichia coli β-galactosidase) product is an enzyme (β-galactosidase) that cleaves a nonchromogenic galactoside (X-gal or β-gal) to a chromogenic precipitate that is localized to the site of reporter gene expression and appears blue on light microscopy [3,36]. This technology, however, requires tissue sampling and processing for the β-galactosidase assay. Noninvasive, indirect imaging paradigms in animals (that can also be translated to human subjects) will be described in subsequent sections of this review.
PET-based, Indirect reporter gene imaging. Many PET-based molecular imaging paradigms are “indirect”; many use reporter-transgene technology and complementary radiolabeled probes. Current PET-based reporter gene imaging paradigms fall into two categories: enzyme-based and receptor-based. The most widely used PET “reporter genes” are transcribed and translated to an enzyme; the enzyme selectively converts a complementary “reporter probe” to a metabolite that is trapped within transduced cells. The receptor-based reporter gene imaging paradigm involves a reporter gene product that is a receptor that “irreversibly” traps the reporter probe in transduced cells.
Modern molecular techniques provide the opportunity to design specific reporter gene constructs, where the reporter gene is placed under the control of different promoter/enhancer elements. These promoter/enhancer elements regulate reporter gene expression. For instance, they can be “always turned on” by constitutive promoters (such as LTR, RSV, CMV, PGK, EF1, etc.). Alternatively, the promoter/enhancer elements can be constructed to be “inducible” and “sensitive” to activation and regulation by specific endogenous transcription factors and promoters (factors that bind to and activate specific enhancer elements in the promoter region of the reporter vector construct leading to the initiation of reporter gene transcription). Promoter activation can be tissue specific, where enhancer elements are activated by promoter/transcription factors that are selectively expressed in specific tissue (e.g., PSMA promoter that is selectively active in prostate cancer cells [37], the albumin promoter expressed selectively in liver [38] or the CEA promoter selectively expressed in colorectal cancer [39]). Inducible reporter genes can also be constructed to be sensitive to specific endogenous molecular processes, including the regulation of endogenous gene expression [40], the activity of specific signal transduction pathways [18], as well as specific protein–protein interactions [41], and posttranscriptional regulation of protein expression [42].
A general paradigm for reporter gene imaging using PET imaging technology has been described [12,14]; the paradigm is illustrated in Figure 1. The herpes simplex virus type 1 thymidine kinase gene (HSV1-tk) with specific upstream promoter/enhancer elements is transfected into target cells by a vector. It is important to note that imaging transgene expression is independent of the vector used to transfect/transduce target tissue; namely, any of several currently available vectors can be used (e.g., retrovirus, adenovirus, adeno-associated virus, lentivirus, liposomes, etc.). Inside transfected cells, the HSV1-tk gene is transcribed to HSV1-tk mRNA and is then translated on the ribosomes to a protein (enzyme), HSV1-TK. After administration of a complementary radiolabeled reporter probe (5-iodo-2′-fluoro-2′deoxy-1-β-

Schematic for imaging HSV1-tk reporter gene expression with reporter probes FIAU and FHBG. The HSV1-tk gene complex is transfected into target cells by a vector. Inside the transfected cell, the HSV1-tk gene is transcribed to HSV1-tk mRNA and then translated on the ribosomes to a protein (enzyme), HSV1-TK. After administration of a radiolabeled probe and its transport into the cell, the probe is phosphorylated by HSV1-TK (gene product). The phosphorylated radiolabeled probe does not readily cross the cell membrane and is “trapped” within the cell. Thus, the magnitude of probe accumulation in the cell (level of radioactivity) reflects the level of HSV1-TK enzyme activity and level of HSV1-tk gene expression.
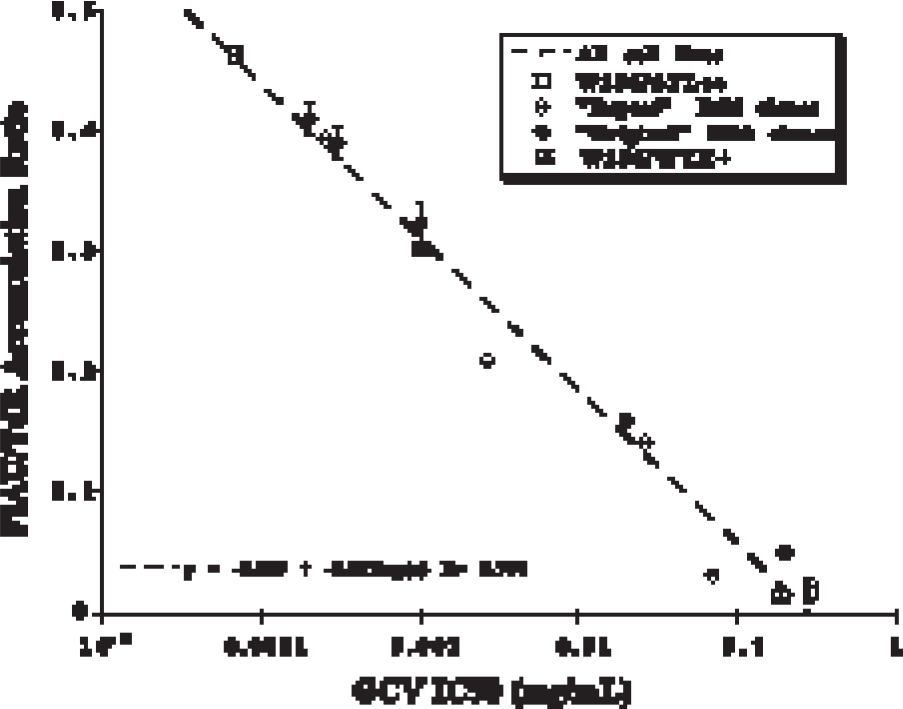
Comparative assays of HSV1-tk expression. The radiotracer accumulation ratio (FIAU/TdR) which normalizes FIAU uptake for cell proliferation (as measured by radiolabeled thymidine, TdR) is compared to a functional assay of HSV1-tk expression; namely, sensitivity (IC50) to the antiviral drug, GCV. Four different sets of stably transduced cell lines (clones) expressing HSV1-tk were studied. A highly reproducible relationship between these two assays is demonstrated which is independent of cell line and retroviral transduction vector. Figure was adapted from Tjuvajev et al. [10].
Most reporter imaging studies using radiolabeled probes and PET imaging involve wild-type HSV1-tk or mutant HSV1-sr39tk reporter genes [43]. The HSV1-tk and HSV1-sr39tk gene products are proteins (enzymes) that have less substrate specificity than mammalian thymidine kinase 1 (TK1). They phosphorylate a wider range of compounds, including acycloguanosines (e.g., acyclovir, ACV; ganciclovir, GCV; FHBG) and 2′-fluoro-nucleoside analogues of thymidine (e.g., FIAU). This difference in enzyme specificity permits the development and use of radiolabeled probes that are phosphorylated to a significantly greater extent by HSV1-TK or HSV1-sr39TK in comparison to mammalian TK1. An example of imaging HSV1-tk expression in an animal model with [124I]FIAU and PET is shown in Figure 3.
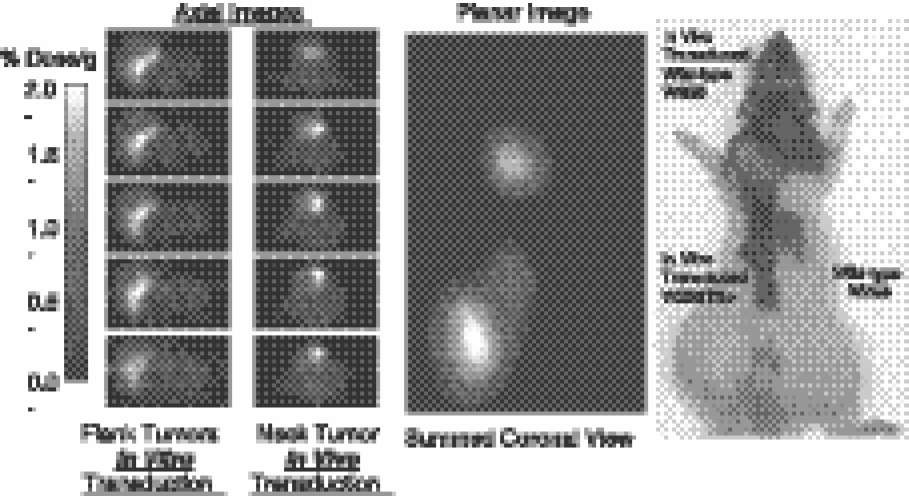
[124I]FIAU PET imaging of HSV1-tk gene expression. Three tumors were produced in rnu/rnu rats. A W256TK+ tumor (positive control) was produced from stably transduced W256TK+ cells and is located in the left flank, and wild-type W256 tumors was produced in the dorsum of the neck and in the right flank (negative control). The neck tumor (wild type) was inoculated with 106 gp-STK-A2 vector-producer cells (retroviral titer: 106–107 cfu/mL) to induce HSV1-tk transduction of the tumor in vivo. No carrier added [124I]FIAU (25 μCi) was injected intravenously 14 days after gp-STK-A2 cell inoculation and PET imaging was performed 30 hr later. Localization of radioactivity is clearly seen in the left flank tumor (positive control) and in the in vivo transduced neck tumor (test), but only low background levels of radioactivity were observed in the right flank tumor (negative control). Figure was adapted from Tjuvajev et al. [12].
A reporter gene can also encode for an extracellular or intracellular receptor or transporter (e.g., hD2R, hSSTR2, or hNIS) that binds or transports a radiolabeled or paramagnetic probe. The human dopamine 2 receptor (hD2R) [44], the human somatostatin receptor subtype-2 (hSSTR2) [45,46], and the human sodium iodide symporter (hNIS) [47–49] genes have been suggested as potential reporter genes for human studies. All three human genes have limited expression in the body; hD2R expression is limited to the striatal–nigral system of the brain, high hSSTR2 expression is largely limited to carcinoid tumors, and hNIS expression is limited largely to thyroid, stomach, and salivary glands. This approach is a very clever strategy because there are established complementary radiolabeled probes for each of these reporter genes: 3-(2′-[18F]fluoroethyl)spiperone (FESP) for hD2R imaging [50] (Figure 4), [111In]DTPA-octreotide (a complementary radiolabeled somatostatin analogue) for hSSTR2 imaging [51], and radiolabeled iodide or pertechnetate for hNIS imaging [49,52]. Furthermore, these probes are approved for human administration, and the three reporter systems have distinct benefits with respect to initiating molecular/reporter imaging in human subjects. However, receptor and transporter expression on the surface of cells is a complex process and involves intracellular trafficking and cell membrane incorporation that is likely to be altered under different conditions and different disease states. It remains to be shown whether imaging receptor and transporter-based reporters (e.g., the hD2R, hSSTR2, and hNIS systems) will provide a consistent and reliable measure of reporter gene expression under variable stress or altered conditions. In any case, the level of probe accumulation (level of radioactivity) must be shown to be proportional to the level of gene expression.

PET and autoradiographic images of fluoroethylspiperone following Ad-D2 R and Ad-Bgal virus administration. Nude mice were injected via the tail vein with (A) 9 × 109 pfu of Ad-Bgal virus (negative control) or (B) 9 × 109 pfu Ad2R virus (test). Two days after virus administration, both mice were injected via the tail vein with FESP (200 uCi, 200 ul). Three hours after the FESP injection, the animals were imaged with micro-PET A whole-body coronal projection image of the [18F] activity distribution is displayed on the left. The liver outline is shown in white. The second images from the left are coronal sections, approximately 2 mm thick, from the micro-PET. After their PET scans, the mice were killed, frozen, sectioned, and photographed (third image from left); the images on the right are autoradiographs of the tissue sections. The color scale represents the percentage of injected dose per gram of tissue (%ID/g). Images are displayed on the same quantitative color scale to allow signal intensity comparisons among the panels. Figure was adapted from MacLaren et al. [44] and was provided by S. Gambhir
Surrogate Imaging
“Surrogate marker” probes that reflect downstream effects of one or more endogenous molecular–genetic processes are also being explored. This approach is particularly attractive for potential translation into clinical studies in the near-term. This is because existing radiopharmaceuticals and imaging paradigms may be useful for monitoring downstream effects of changes in specific molecular–genetic pathways in diseases such as cancer. The “surrogate marker” approach is very likely to be less specific and more limited with respect to the number of molecular–genetic processes that can be imaged. Nevertheless, it benefits from the use of radiolabeled probes that have already been developed and studied in human subjects. Thus, the translation of “surrogate marker” imaging paradigms into patients will be far easier than either the reporter transgene or direct imaging paradigms outlined above. However, it remains to be shown whether there is a sufficiently high correlation between “surrogate marker” imaging and direct molecular assays that reflect the activity of a particular molecular–genetic pathway of interest. Very few studies have attempted a rigorous correlation between “surrogate marker” imaging and transcriptional activity of a particular gene, or posttranscriptional processing of the gene product, or the activity of a specific signal transduction pathway that is targeted by a particular drug. The application of “surrogate marker” imaging for monitoring treatment response is gaining increasing attention, particularly as it relates to the development and testing of new pathway-specific drugs. For example, the assessment of noncyto-toxic, cytostatic drugs such as the antiangiogenic class of drugs poses particular problems for imaging assessments based on tumor volume. Thus, surrogate imaging markers of tumor vascularity are being proposed for assessing antiangiogenesis treatment response. Whether imaging “surrogate markers” will be of value for assessing treatment directed at other molecular–genetic abnormalities in tumors (e.g., signaling by EGFR, p53, c-Met, HIF-1, etc.) remains to be demonstrated.
Applications of PET-based Reporter Gene Imaging
Monitoring Gene Therapy
A noninvasive, clinically applicable method for imaging the expression of successful gene transduction in target tissue or specific organs of the body would be of considerable value for monitoring and evaluating gene therapy in human subjects [53].
PET imaging could define the location, magnitude, and persistence of gene expression over time. Targeting gene therapy to particular tissue (e.g., tumor) or specific organs of the body is an increasingly active area of research. Several issues that are important for clinical optimization of gene therapy remain unresolved in many current clinical protocols: (1) Has gene transduction or transfection been successful? (2) Is the distribution of the transduced or transfected gene localized to the target organ or to the target tissue, and is the distribution in the target optimal? (3) Is the level of transgene expression in the target organ or tissue sufficient to result in a therapeutic effect? (4) Does the transduced or transfected gene localize to any organ or tissue at sufficient levels to induce unwanted toxicity? (5) In the case of combined prodrug–gene therapy protocols, when is transgene expression m maximum (optimal) and when is the optimal time to initiate treatment with the prodrug? (6) How long does transgene expression persist in the target and other tissues?
We and others have proposed that noninvasive imaging techniques using selected reporter gene and reporter probe combinations will provide a practical and clinically useful way to identify successful gene transduction and expression in patients undergoing gene therapy. One could argue that biopsies of target tissue or blood sampling and analysis of transgene expression, or assay for secretable marker (e.g., CEA, AFP, hCGT) could be performed and that imaging is not critical. However, imaging provides some clear advantages, including: (a) the ability to repeatedly assess gene expression over time, especially when multiple sequential biopsies are not feasible, (b) the absence of any perturbation of the underlying tissue which occurs with biopsy procedures, and (c) the ability to obtain 3-D spatial information in the entire body as well as target organs and tumors, which could be of considerable value when addressing toxicity issues.
Therapeutic Gene Imaging
HSV1-tk has the advantage of being both a “therapeutic gene” (when combined with GCV treatment) and a “reporter gene” (using an appropriate radiolabeled probe, such as FIAU or FHBG). This combination allows for direct imaging of the therapeutic gene product (HSV1-tk), and can be used to define the location, magnitude, and duration of HSV1-tk gene expression. Experimental validation of this approach has been demonstrated in animal models of colorectal metastases to the liver treated with adenoviral-mediated HSV1-tk gene transfer and GCV (“suicide” gene therapy) [39,54,55], or treatment with conditionally replicating, oncolytic herpes viruses that constitutively express the HSV1-tk gene [56,57]. However, most therapeutic genes do not lend themselves to direct imaging of their transgene product. Furthermore, the development and validation of a new probe and a new imaging paradigm specific to each therapeutic transgene of interest would be a very costly and time-consuming endeavor. Many therapeutic gene products do not readily lend themselves to radionuclide assessments and it may not even be possible in some cases. It is therefore more reasonable to consider alternative strategies for “indirect” imaging of therapeutic gene expression that use established reporter gene–reporter probe combinations.
Reporter Gene Imaging
Several indirect reporter gene imaging strategies have been described, and they can be used to image different therapeutic genes. These strategies take advantage of using established reporter gene–reporter probe combinations, and can achieve the objective of monitoring therapeutic gene expression without directly imaging/measuring the therapeutic gene product. One strategy uses a “fusion gene” containing cDNA from both the reporter and therapeutic genes [54]; a second strategy uses an “internal ribosome entry site (IRES)-linked” construct for proportional therapeutic and reporter gene expression [46,58,59]; a third strategy uses “multiple vectors,” where one or more separate vectors carry a therapeutic gene(s) and another vector carries the reporter gene [60]; and a fourth strategy uses multiple promoters for expression of the two genes (e.g., HSV1-tk and NeoR) [10], although this latter strategy may be less reliable, because of the higher probability of promoter interference.
The use of “fusion gene” technology provides a single transcription cassette for coexpression of two genes that are “linked” in a fixed, definable manner. One segment of the fusion gene cDNA corresponds to the “therapeutic” gene and another segment corresponds to the “reporter” gene. Transcription of the fusion gene occurs under the control of upstream promoter/enhancer regulatory elements of choice (e.g., constitutive expression or expression only in response to a specific activator that is present only in targeted organs or tumors). Translation of mRNA proceeds to yield a single “fusion protein” (the gene product) containing both reporter and therapeutic amino acid sequences. The fusion protein is a single hybrid of the two individual proteins. Thus, information obtained by imaging the “reporter” component will provide corresponding information about the “therapeutic” component [54] (Figure 5). However, there are a number of potential disadvantages of the fusion gene approach. The fusion construct may not result in a functional gene product. This could be due to a change in the conformational structure of the native protein or result in an alteration in the subcellular localization of the fusion protein, or to a loss in activity of its “therapeutic component,” or to a loss in activity of its “reporter component.” Modulation of the fusion mRNA or a change in the clearance (breakdown) of the fusion protein may also be sufficiently different compared to the two native proteins (gene products of the two native genes). Such differences could have a significant impact on the level of the fusion gene product, and thereby, on the level of its biological activity. Fusion proteins are larger than each of the corresponding native proteins and are more likely to generate an immunological response. Thus, fusion gene technology cannot be generalized and may not be widely applicable in clinical imaging of therapeutic gene expression. However, when a fusion gene product is functional and nonimmunogenic, it may be a very useful construct for monitoring therapeutic gene expression.

CD/TK fusion gene assays in vitro and in vivo. Functional coexpression of the CD and HSV1-tk subunits of the CDglyTK fusion gene product (fusion protein) in different single cell-derived clones of W256CDTK+ cells (in vitro experiments, left panel). Functional coexpression of the CD and HSV1-tk subunits of CDglyTK fusion gene product (fusion protein) in different subcutaneous W256CDTK+ tumors (in vivo experiments, right panel). CD enzyme activity is plotted against FIAU accumulation (radiotracer assay for HSV1-TK enzyme activity). Figure was adapted from Hackman et al. [54].
The use of “cis-linked” genes with an IRES element within a single bicistronic transcription cassette provides an alternative approach for achieving proportional coexpression of two genes. The IRES element enables translation initiation within the bicistronic mRNA, thus permitting gene coexpression by cap-dependent translation of the first cistron and cap-independent, IRES-mediated translation of the second cistron [61,62]. Successful imaging studies in animals using IRES constructs have been performed and indicate that proportional coexpression can be achieved [46,58,59] (Figure 6). However, cap-dependent translation of the first cistron has been shown to be several-fold greater than cap-independent, IRES-mediated translation of the second cistern. Although gene product assays have shown proportional expression over a wide range in transduced cells and xenografts, it remains to be demonstrated whether IRES-based vector expression is a reliable indicator of transgene coexpression in different tissues. Namely, that the half-life of each encoded protein remains proportional in different tissues and organs of the body. This is necessary when noninvasive imaging is used to assess organ (tissue) specificity, as well as the level and duration of transgene expression.

Imaging Cis-linked HSV1-tk and LacZ gene coexpression. Gamma camera images of [131I]FIAU accumulation in HSV1-tk transduced W256STLEO and RG2STLEO subcutaneous tumors (derived from HSV1-tk transduced clonal cell lines) 24 hr after tracer administration; the images in Panel A reflect HSV1-tk expression. The white dashed line outlines the contours of the animal and tumors with relatively low levels of activity, and wild-type control tumors (on the right thigh) with the lowest level of activity. Some residual bladder activity is still present in RG2STLEO tumor-bearing rats. Note the difference in [131I]FIAU accumulation between different tumors. The [131I]FIAU images of HSV1-tk expression were also converted into parametric images of lacZ gene expression. The parametric scaling of lacZ gene expression is based on the relationship defined in Panel D and is indicated by the color-coded intensity bar showing units of β-galactosidase activity (U/g) in Panel A. The levels of HSV1-tk gene expression measured in different tumors (%dose/g [131I]FIAU) were plotted against measures of lacZ gene expression (U/g) obtained in corresponding tumor samples for different W256SLEO (Panel B) and RG2STLEO (Panel C) clonal tumors. The relationship between the two measures was defined by regression analysis. The same relationship was observed when the RG2STLEO and W256STLEO data were combined (Panel D). Figure was adapted from Tjuvajev et al. [58].
The use of “multiple vectors” is another approach for achieving coexpression of one or more therapeutic genes and a reporter gene. In this paradigm, the genetic backbone of each vector (e.g., adenovirus) is identical, except for inclusion of a therapeutic gene or a reporter gene. Each vector would have identical envelope characteristics and identical promoter/enhancer elements driving the expression of the therapeutic and reporter transgenes. A multiple vector cocktail containing definable (different) combinations of the individual vectors can be administered. Key requirements for this approach are that: (1) the multiple vectors transfect and transduce target organs and tissue proportionate to their ratio in the administration cocktail; (2) the therapeutic and reporter genes are coexpressed proportionally in all target organs and tissue; and (3) the proportionality of coexpression is constant over a wide range of expression levels. A recent report has provided encouraging results in support of using multiple vectors (adenovirus) to deliver reporter and therapeutic genes to target organs [60] (Figure 7).
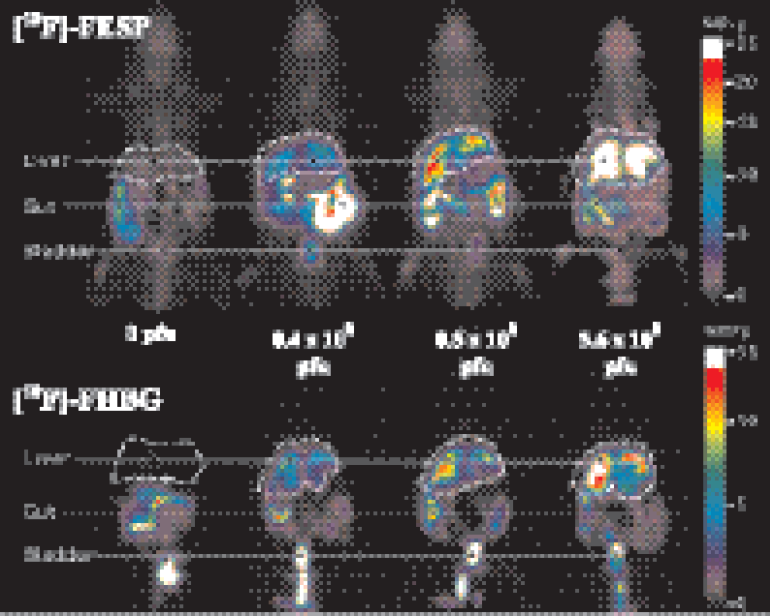
Micro-PET whole body [18F]FESP and [18F]FHBG images of Ad-CMV-D2R and Ad-CMV-HSV1-sr39tk localization. The objective of this study was to compare the imaging characteristics and intensity of two different reporter systems (involving two different radiolabeled probes) that are transfected by two different adenoviruses which have the same backbone. Equal viral titers of Ad-CMV-D2R and Ad-CMV-HSV1-sr39tk were injected into the tail vein of nude mice. After 3 days, the mice were injected with [18F]FHBG and imaged by micro-PET. The next day, the same mice were injected with [18F]FESP and were reimaged with micro-PET. The images illustrate the biodistribution and degree of [18F]FHBG and [18F]FESP accumulation in individual nude mice coinjected with different, but equivalent titers of Ad-CMV-HSV1-sr39tk and Ad-CMV-D2R. From left to right, the mice received 0, 0.4 × 108, 0.5 × 108, and 3.6 × 108 pfu. The upper panel illustrates biodistribution of [18F]FESP and the lower panel that of [18F]FHBG. Images are the average of several whole-body slices containing the liver. The images are scaled based on the injected dose. Figure was adapted from Yaghoubi et al. [60] and was provided by S. Gambhir
PET-based Reporter Gene Imaging of Biological Processes
Reporter gene imaging is being used to visualize endogenous biological processes, including transcriptional and posttranscriptional regulation of target gene expression, as well as specific intracellular protein–protein interactions. Imaging transcriptional regulation of endogenous genes in living animals using noninvasive imaging techniques is providing a clearer understanding of normal and cancer-related biological processes. These new imaging paradigms can be directly translated into human studies using similar reporter constructs, radiolabeled reporter probes, and PET imaging. Several examples are described below.
p53. Imaging transcriptional regulation of endogenous genes in living animals (and potentially in human subjects) using noninvasive imaging techniques is feasible. A recent article from our group [18] was the first to show that p53-dependent gene expression can be imaged in vivo with PET and by in situfluorescence. A retroviral vector (Cis-p53/TKeGFP) was generated by placing the HSV1-tk and enhanced green fluorescent protein (egfp) fusion gene (TKeGFP, a dual-reporter gene) under control of a p53-specific response element. DNA damage-induced up-regulation of p53 transcriptional activity was demonstrated and correlated with the expression of p53-dependent downstream genes (including p21). These findings were observed in U87 (p53 +/+) cells and xenografts, but not in SaOS (p53 -/-) cells. This was the first demonstration that a Cis-reporter system (Cis-p53/TKGFP) was sufficiently sensitive to image endogenous gene expression using noninvasive nuclear (PET) imaging (Figure 8). The PET images corresponded with up-regulation of genes in the p53 signal transduction pathway (p53-dependent genes) in response to DNA damage induced by BCNU chemotherapy. PET imaging of p53 transcriptional activity in tumors using the Cis-p53TKGFP reporter system could be used to assess the effects of new drugs or other novel therapeutic paradigms that are mediated through p53-dependent pathways. For example, specific p53 gene therapy strategies that are based on p53 overexpression [63] could be monitored by noninvasive imaging.
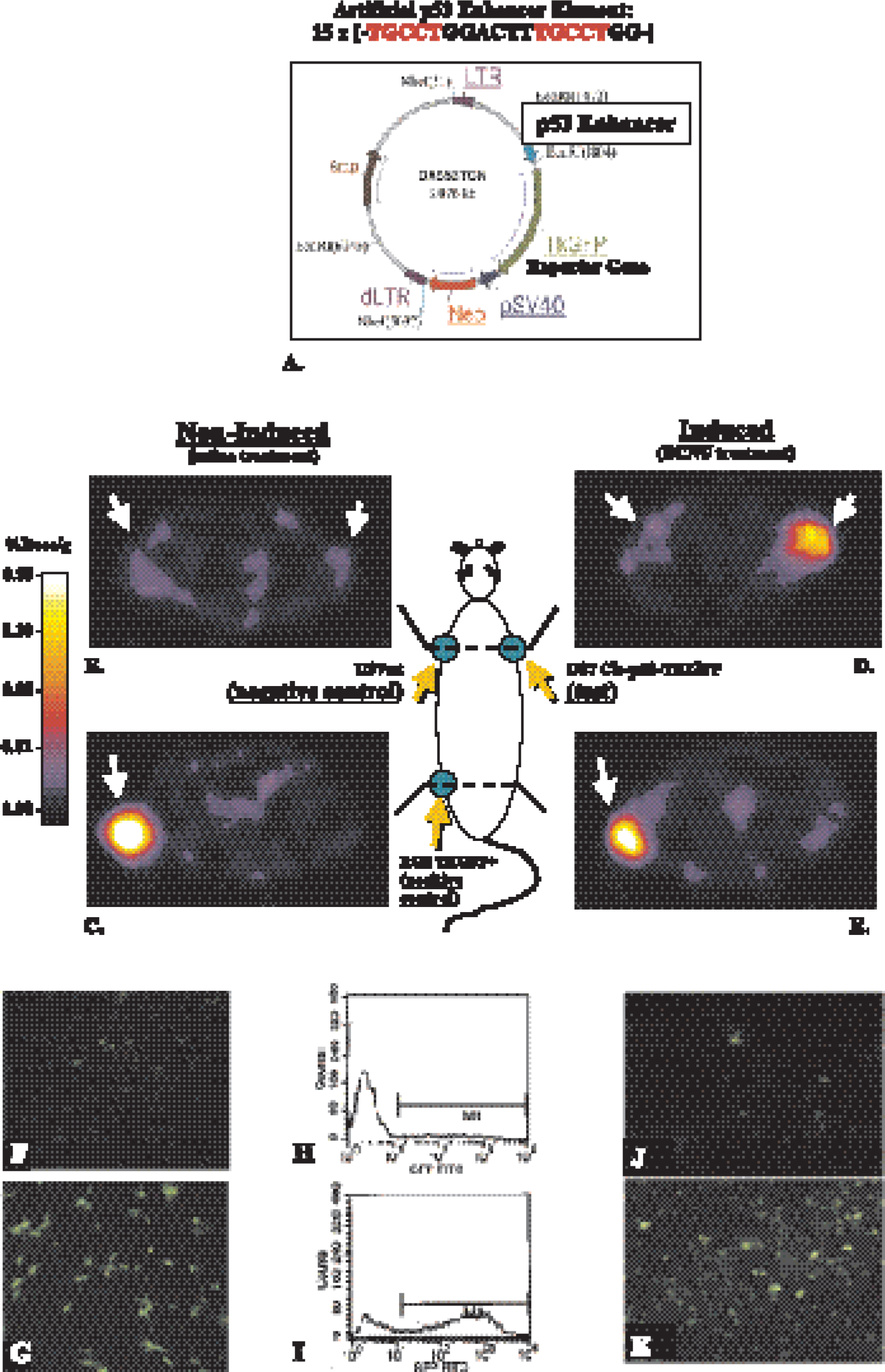
PET imaging of endogenous p53 activation and validation of Cis-p53/TKGFP reporter system in cell cultures and sampled tumor tissue. The p53-sensitive reporter vector (A) contains an artificial p53-specific enhancer that activates expression of the TKGFP reporter gene. Transaxial PET images (GE advance tomograph) through the shoulder (B, D) and pelvis (C, E) of two rats are shown (upper panel); the images are color-coded to the same radioactivity scale (%dose/g). An untreated animal is shown on the left, and a BCNU-treated animal is shown on the right. Both animals have three subcutaneous tumor xenografts: U87p53TKGFP (test) in the right shoulder, U87 wild-type (negative control) in the left shoulder, and RG2TKGFP (positive control) in the left thigh. The nontreated animal on the left shows localization of radioactivity only in the positive control tumor (RG2TKGFP); the test (U87p53TKGFP) and negative control (U87wt) tumors are at background levels. The BCNU-treated animal on the right shows significant radioactivity localization in the test tumor (right shoulder) and in the positive control (left thigh), but no radioactivity above background in the negative control (left shoulder). Fluorescence microscopy and FACS analysis of a transduced U87p53/TKGFP cell population in the noninduced (control) state (F, H), and 24 hr after a 2-hr treatment with BCNU 40 μg/ml (G, I). Fluorescence microscopy images of U87p53/TKGFP subcutaneous tumor samples obtained from nontreated rats (J) and rats treated with 40 mg/kg BCNU ip (K). Figure was adapted from Doubrovin et al. [18].
It should also be pointed out that the dual-reporter construct (TKeGFP—fusion gene) provides the opportunity for multimodality (both nuclear and optical imaging) imaging of endogenous gene expression in vivo. The TKeGFP reporter gene could be introduced into other reporter assay systems to assess other molecular-biological pathways. It should also be possible to use the TKeGFP reporter gene in transgenic animals; this will facilitate the monitoring and assessment of newly cloned genes or novel signal transduction pathways. Another advantage of the dual-reporter system is the ability to compare the images of reporter gene expression obtained with PET, gamma camera, or autoradiography with corresponding in situ GFP fluorescence images. The comparison between GFP fluorescence and autoradiographic images, coupled with histology of corresponding tissue sections, provides for spatial and quantitative assessments of reporter gene expression at the microscopic as well as macroscopic level.
Nuclear Factor of Activated T-cells (NFAT). T-cell activation is an essential component of the immune response in many normal and disease states. The objective of a recent study in our laboratory [40] was to monitor and assess T-cell receptor (TCR)-dependent activation of T-cells in vivo using noninvasive PET imaging. A retroviral vector (Cis-NFAT/TKeGFP) was generated by placing the fusion gene (TKeGFP) under control of the nuclear factor of activated T-cells (NFAT) response element. A human T-cell leukemia cell line (Jurkat) that expresses a functional TCR was transduced with the Cis-NFAT/TKGFP reporter vector and was used in these studies. Known activators of T-cells (anti-CD3 and anti-CD28 antibody) produced significantly higher levels of TKGFP reporter gene expression (increased GFP fluorescence, increased levels of HSV1-tk mRNA, and increased [14C]FIAU accumulation in vitro) in Cis-NFAT/TKGFP+ Jurkat cells, in comparison to nontreated or nontransduced cells. In mice with focal Cis-NFAT/TKGFP+ Jurkat cell infiltrates, similar results were observed in the micro-PET images (Figure 9 top), and the in vivo fluorescence images (Figure 9 bottom). A strong correlation between TKGFP expression and up-regulation of T-cell activation markers (CD69 and IL-2 production) was demonstrated both in vitro and in vivo. These results demonstrated that: (1) activation of the NFAT signal transduction pathway occurs after TCR stimulation, and (2) PET imaging of T-lymphocyte activation in tumors following TCR engagement is feasible using the described TKGFP-based Cis-reporter system. This imaging paradigm could be used to assess the efficacy of novel antitumor vaccines and adoptive immunotherapy.
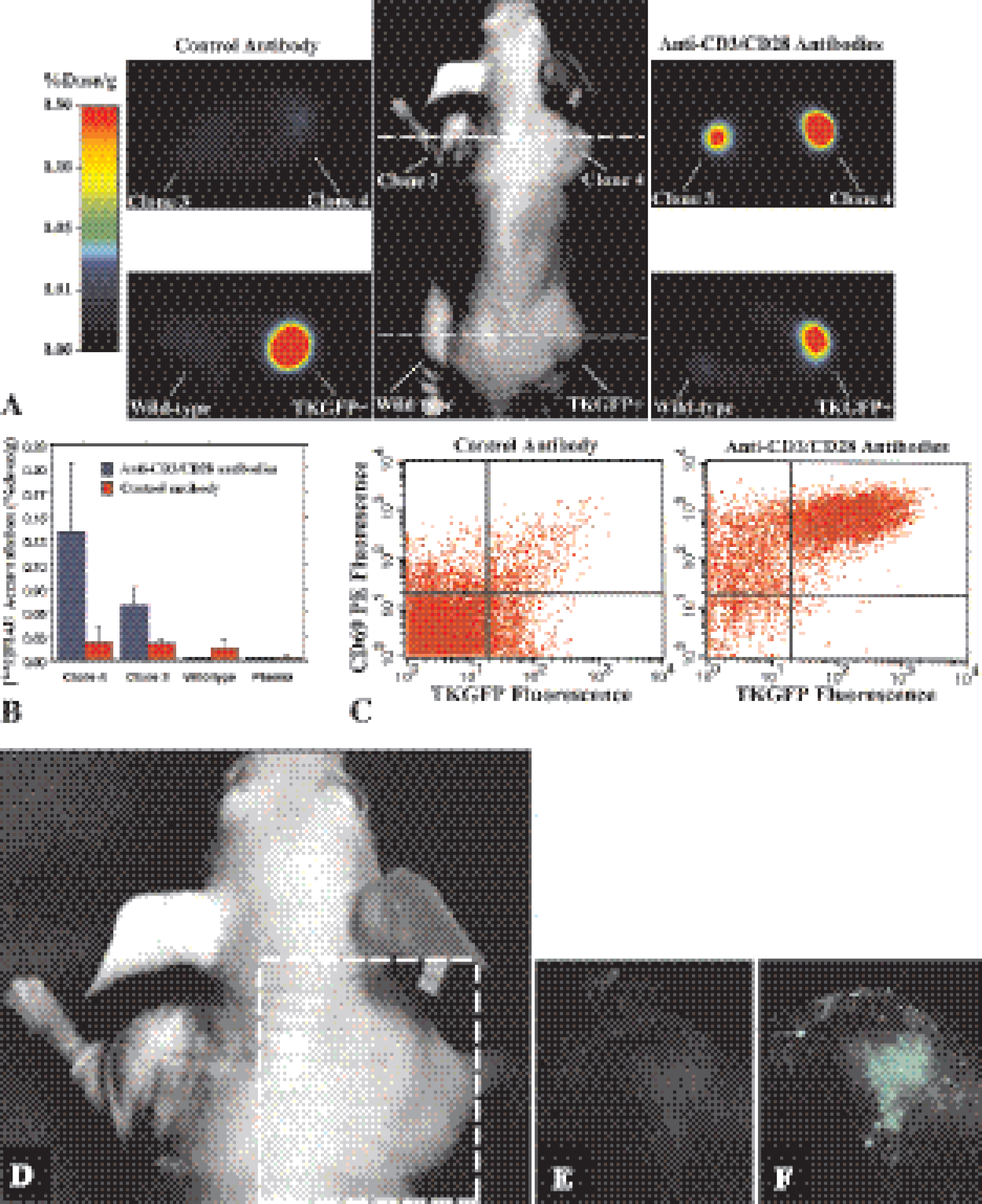
Imaging NFAT-TKGFP reporter system activity with PET and optical fluorescence, plus assessments in tissue samples and optical fluorescence imaging of the NFAT-TKGFP reporter system activity in vivo. Photographic image of a typical mouse bearing different subcutaneous infiltrates (middle panel); transaxial PET images (GE advance tomograph) of TKGFP expression in a mouse treated with control antibody (left panel) and anti-CD3/CD28 antibodies (right panel) were obtained at the levels indicated by the dashed lines (A). [124I]FIAU accumulation (%dose/g) in tissue samples of the Jurkat/dcmNFATtgn clone 3 and 4 infiltrates, wild-type Jurkat infiltrates, and blood plasma, obtained after PET imaging (B). FACS profiles of TKGFP and CD69 expression in a tissue sample from the same Jurkat/dcmNFATtgn clone 4 infiltrate that was imaged with PET (C). Images of TKGFP fluorescence in a subcutaneous Jurkat/dcmNFATtgn (clone 4) infiltrate were obtained in the same animal (D); first before (E) and then after treatment (F) with anti-CD3/CD28 antibodies. Figure was adapted from Ponomarev et al. [40].
Hypoxia-inducible Factor 1 (HIF-1). Tissue hypoxia has been recognized as a significant component of tumors, and is associated with malignant progression and resistance to radiation therapy. Cellular responses to hypoxia are mediated largely through a transcriptional activator, HIF-1α. A heterodimer (HIF-1) is formed in response to hypoxia, which binds to the hypoxia-responsive element (HRE) in the enhancer region of many target genes (erythropoietin, VEGF, glycolytic enzymes, and other stress response genes) and initiates gene transcription. Work in our laboratory is being directed to the development of a noninvasive imaging paradigm to monitor and assess the level of HIF-1 activity during tumor development and growth. This work in progress has been presented in abstract form only [64]. We developed a dual-reporter vector containing a “sensor” reporter element (Cis-HRE/TKGFP) and a “beacon” reporter element (CMV-RFP/XPRT) that is constitutively expressed. This vector was used to transduce C6 and RG2 glioma cells, and provides the opportunity to perform both optical and radioactivity-based imaging. Optical, radioactivity, and molecular assays were performed to assess the activity of the reporter systems; optical imaging of transduced (CMV-RFP/XPRT–Cis-HRE/TKGFP) C6 cells growing as spheroids in culture is shown in Figure 10.

Optical imaging of hypoxia. The HIF-1a reporter vector is shown in the left panel. There are two elements in this reporter construct: one is a “sensor” element that consists of an artificial HIF-1α enhancer/promoter controlling the expression of TKeGFP gene (a fusion between the HSV1-tk and the enhanced green fluorescent protein, eGFP, genes) and the other is a “beacon” element that consists of the SV40 promoter that constitutively expresses the Red1/XPRT gene (a fusion between the Red1 fluorescent protein and E. coli xanthine phosphoribosyl transferase genes). Spheroids produced from transduced C6 cells are shown; red fluorescence is seen in a confocal image of each spheroid demonstrating constitutive expression of the Red1/XPRT fusion gene (beacon element). Green fluorescence is seen in the center of the larger spheroid indicating hypoxia-inducible expression of the TKeGFP gene (sensor element). Figure was adapted from Serganova et al. [64].
Transcriptional activation of the HIF-1α signaling pathway was also observed after 16 hr of exposure to 2% O2 and to CoCl2 (20–100 μM). The hypoxic/treated cells exhibited high levels of TKGFP reporter protein as measured by fluorescence (FACS) and [14C]FIAU accumulation in comparison to control nontreated cells. The CoCl2-induced increase in HIF-1α resulted in a corresponding up-regulation of VEGF and TKGFP gene expression (mRNA and protein levels). Noninvasive PET imaging studies of the HIF-1α signaling pathway using [124I]FIAU have been performed and the results are very encouraging. This novel reporter system, Cis-HRE-TKGFP, for monitoring hypoxia by fluorescence and nuclear imaging, should provide a valuable resource for monitoring the time course of hypoxia during tumor development, growth, and treatment (e.g., radiation, and antiangiogenesis chemotherapy).
Transforming Growth Factor Beta (TGF-β). The cyto-kine TGF-β plays a dual role in tumorigenesis. It stimulates the proliferation of mesenchymal cells while inhibiting the growth of most normal epithelial cells. Work in our laboratory is being directed to the development of a noninvasive imaging paradigm to monitor and assess the activity of TGF-β signal transduction pathways during tumorigenesis. This work in progress has been presented in abstract form only [65]. A Cis-TGF-β/TKGFP reporter vector was developed containing both AML1 and Smad consensus binding sites (Igα-Smad-AML-TKGFP reporter gene). MDA-MB-231 cells were stably transduced with the Igα-Smad-AML-TKGFP reporter system and were exposed to various concentrations of the human TGF-β1 for 48 hr; the level of TKGFP expression was assayed by fluorescence microscopy, quantified by FACS and a radiotracer assay ([14C]FIAU and [3H]dThd).
Transcriptional activation of the TGF-β signaling pathway was observed after 16 hr of exposure to minimal concentrations of TGF-β1 (2 ng/ml) and nutrient depletion. TGF-β1 treated cells exhibited high levels TKGFP reporter protein as measured by FACS, and significantly higher levels of the FIAU accumulation were observed compared to baseline, nonstimulated cells. The level of FIAU accumulation measured in stimulated Igα-Smad-AML-TKGFP transduced cells indicates that successful imaging of TGF-β induced transcriptional activity with 124FIAU and PET is feasible. The ability to image the activity of the TGF-β signaling pathway will be used to assess novel antitumor drugs that act through inhibition of this pathway.
Imaging Weak Promoters. Several attempts have been made to image reporter gene expression driven by weak promoters (e.g., CEA, PSE, etc.), but the results were usually hampered by the poor transcriptional activity of such promoters. In the original study, Qiao et al. [39] validated methods to enhance the transcriptional activity of the carcinoembryonic antigen (CEA) promoter using a trans- or so-called two-step transcriptional amplification (TSTA) system [66]. To increase promoter strength while maintaining tissue specificity, a recombinant adenovirus was constructed which contained a TSTA system with a tumor-specific CEA promoter driving a transcription transactivator, which then activates a minimal promoter to drive expression of the HSV1-tk suicide/reporter gene. This ADV/CEA-binary-tk system resulted in equal or greater cell killing of transduced cells by GCV in a CEA-specific manner, compared with GCV killing of cells transduced with a CEA-independent vector containing a constitutive viral promoter driving HSV-tk expression (ADV/RSV-tk). It was demonstrated in vitro that the expression of the HSV1-tk gene mediated by the CEA-specific TSTA system was about 250-fold higher than that from the cis-CEA promoter alone. To test the imaging capacity of the ADV/CEA-binary-tk system for monitoring CEA-specific transgene expression in tumor-bearing animals, [131I]FIAU and gamma camera imaging was used to assess adenovirus-mediated HSV-tk gene expression in vivo. After injection of the ADV/binary-tk virus into CEA-expressing liver tumors, [131I]FIAU accumulation was observed only in the area of CEA-positive tumors. Importantly, there was significantly less HSV1-tk expression in adjacent liver tissue following ADV/binary-tk injection than after the constitutively expressing ADV/RSV-tk virus. A dose escalation therapeutic trial in these animals demonstrated significantly reduced hepatoxicity following GCV administration with the ADV/binary-tk virus compared to the ADV/RSV-tk virus, with equal antitumor efficacy.
In another study, Iyer et al. [67] validated methods to enhance the transcriptional activity of the androgen-responsive prostate-specific antigen promoter (PSE) using a similar TSTA approach to amplify expression of firefly luciferase, and mutant herpes simplex virus type 1 thymidine kinase (HSV1-sr39tk) in a prostate cancer cell line (LNCaP) [67]. A ~50-fold (luciferase) and a ~12-fold (HSV1-sr39tk) enhancement of transgene expression were demonstrated by using this TSTA system while retaininig tissue selectivity. A cooled charge-coupled device (CCD) optical imaging system was used to visualize the signal generated from the luciferin–luciferase reporter system in living mice implanted with LNCaP cells that were transfected ex vivo. Further improvements of the androgen-responsive TSTA system for reporter gene expression was made using a “chimeric” TSTA system that uses duplicated variants of the prostate-specific antigen (PSA) gene enhancer to express GAL4 derivatives fused to one, two, or four VP16 activation domains [67]. The resulting activators were transduced into prostate carcinoma cells with reporter templates bearing one, two, or five GAL4 binding sites upstream of firefly luciferase. The activity of the luciferase reporter gene in transfected cell extracts and in live nude mice was monitored using an in situ luciferase assay and a cooled CCD imaging, respectively. It was found that luciferase expression in prostate cancer cells in vitro can be varied over an 800-fold range, and that a single plasmid bearing the optimized chimeric GAL4–VP16 enhancer expressed the firefly luciferase reporter at 20-fold higher levels than the cytomegalovirus enhancer. A very encouraging result was the demonstration that the TSTA system was androgen concentration sensitive, suggesting a continuous rather than binary reporter response. However, as observed with the CEA-TSTA reporter system above, the in vivo imaging comparison of the TSTA and Cis-reporter systems showed substantially less dramatic differences than that obtained by the in vitro analyses. Only a ~5-fold difference in response to androgen stimulation was observed between PSE TSTA and Cis-reporter systems in the in vivo firefly luciferase optical imaging experiments.
Imaging Protein-Protein Interactions. Recent work demonstrating a proof of principle for monitoring protein–protein interactions in vivo was reported by Ray et al. [41] and Luker et al. [68]. The interaction of two proteins, ID and MyoD, was the focus of one study [41]. To modulate the expression of these two proteins, the NF-κB promoter was used to drive expression of the id-gal4 and/or myoD-vp16 fusion genes, and tumor necrosis factor a (TNF-α) was used to induce the NF-κB promoter. The reporter construct contained five GAL4 DNA-binding sites driving the expression of the firefly luciferase (fl) reporter gene. (Note that fl refers to the gene and FL, to the enzyme.) Transcription of fl leads to FL, which is quantitatively imaged after intraperitoneal administration of
In the other study, protein-protein interactions could be imaged in vivo with PET and by fluorescence imaging using a variant of HSV1-tk/GFP dual-reporter fusion gene [68]. It is well known that TAg, which is the transforming protein of SV40 virus, binds constitutively to p53 and blocks transactivation of p53 target genes, leading to transformation of mammalian cells. In this study, the authors used an approach similar to that described previously by Ray et al. [41], but they used a somewhat different reporter system. To enable inducible expression of the hybrid interacting proteins in vitro and in vivo, the authors stably transfected HeLa-Gal4 reporter cells with a reverse tetracycline-responsive transactivator expressed from a cytomegalovirus promoter and a plasmid expressing either Gal4-BD-p53 and VP16-TAg or Gal4-BD-p53 and VP16-CP from a bidirectional, tetracycline-regulated promoter. A hybrid protein of VP16 and polyoma virus coat protein (VP16-CP), which does not bind to p53, was used as a negative control. The expression of a chimeric dual-reporter gene in which the GFP was fused with the HSV1-sr39tk gene was driven by an enhancer element containing multiple GAL4 DNA binding sites.
A similar dual-reporter fusion gene, combining wild-type HSV1-tk and enhanced green fluorescent protein (HSV1TKeGFP), was previously described [57] and has been used in several reporter gene imaging studies [18,40,64,65]. Luker et al. [68] introduced a mutation in the nuclear localizer signal (NLS) of the HSV1-sr39tk gene (mutant nlsHSV1-sr39tk/GFP). The NLS mutation was introduced based on previous reports by Ponomarev et al. [69], which demonstrated that mutations in one of three putative NLS sites in HSV1-TK (or the addition of a nuclear export signal) resulted in pancellular (or only cytoplasmic redistribution) of the GFP-fused enzyme [69,70]. The NLS mutation also resulted in a greater uptake of 124I-FIAU due to better metabolic availability of these fluorescently tagged mutant reporter enzymes.
To determine the feasibility of imaging protein–protein interactions in living animals, Luker et al. [68] produced xenografts from their transduced cell lines (TAg and CP cells) in nu/nu mice. After the xenografts grew to an appropriate size, the mice were treated with doxycycline for 48 h to induce expression of the hybrid proteins. To visualize expression of the Gal4-mNLS-sr39TK-EGFP reporter at 48 hr, [18F]FHBG was injected intravenously and micro-PET imaging was performed 1 hr later. Accumulation of radiotracer was imaged in the TAg xenograft, but not in the CP xenograft, showing that specific interaction of p53 and TAg could be detected by PET imaging. Based on region-of-interest values from the micro-PET images, the uptake of 18F-FHBG was 5.5-fold greater in TAg xenografts compared to that in CP xenografts.
These initial results are very encouraging, but at the same time, they are also limited because they demonstrate only constitutive interactions of the known pairs of interacting proteins. Further studies will be required to validate the approach described in both studies with respect to the sensitivity and dynamic range of these reporter systems for monitoring the induction and inhibition of “endogenous” protein–protein interactions. Namely, protein–protein interactions that occur as a result of a change or modulation in activity of a corresponding signal transduction pathway (as opposed to induction of the “bait” and “target” protein expression by either the NFkB- or the tetracycline-dependent systems). For example, such studies are being conducted to assess the interactions of different Smad-signaling proteins in regulation of TGF-β signaling pathways in various tumors during therapy with novel anticancer drugs.
Imaging Posttranscriptional Regulation of Gene Expression. Human cells exposed to antifolates show a rapid increase in the levels of the enzyme dihydrofolate reductase (DHFR). Several studies indicate that DHFR enzyme binds to its own mRNA in the coding region, and that inhibition of DHFR by MTX releases the DHFR enzyme from the mRNA. Consequently, this release results in an increase in translation to DHFR protein [71–73]. In addition to the described translational regulation of DHFR in cancer cells exposed to MTX, increased levels of DHFR also result through DHFR gene amplification, a common mechanism of acquired resistance to this drug. In contrast to rapid translational modulation of DHFR, gene amplification occurs in response to chronic exposure to antifolates, and elevated cellular levels of DHFR result from transcription of multiple DHFR gene copies.
In a recent report by Mayer-Kuckuk et al. [42] from MSKCC, this adaptive cellular response mechanism was used as a model to evaluate the efficacy of PET imaging of posttranscriptional regulation of gene expression. In this study, mouse cells were transfected with a reporter system that expresses a green fluorescent protein–DHFR fusion protein following exposure to an antifolate drug, trimetrexate (TMTX). The results demonstrated a specific and time-dependent increase in cellular levels of the fusion protein. Next, human HCT-8 and HCT-116 colon cancer cells were transduced to express a DHFR–HSV1TK fusion protein, and the cells were then treated with the DHFR inhibitor TMTX. Increased levels of the DHFR–HSV1TK fusion protein and an increase in GCV sensitivity by 250-fold were demonstrated. The level of fusion protein in antifolate-treated human tumor cells was increased in response to a 24-hr exposure of methotrexate, trimetrexate, as well as dihydrofolate. This effect depended on the concentration of the antifolate and was independent of fusion protein mRNA levels. These results indicated that the increase in DHFR–HSV1TK fusion protein was occurring at a translational level, rather than at the transcriptional level. Furthermore, this effect could be visualized by [124I]FIAU and PET imaging studies that were performed on nude rats bearing DHFR–HSV1TK-transduced HCT-8 xenografts. Animals treated with TMTX showed a higher accumulation of [124I]FIAU (reflecting a higher expression of the DHFR-HSV1TK fusion protein) than that in comparable nontreated animals. The imaging results were substantiated by a 2- to 4-fold increase of fusion protein levels in the transduced xenografts 24 hr after TMTX treatment (as measured by Western blotting) compared to that measured in the transduced xenografts of nontreated control animals.
The latter reports are very encouraging. However, the current approaches to imaging the regulation of endogenous gene expression or protein–protein interactions have several limitations. Namely, these approaches cannot fully identify the temporal kinetics of gene expression or protein–protein interaction. If as the result of a change in signal transduction activity the target gene expression or a protein–protein interaction will decrease or stop, it still may be the case that the expressed reporter protein will persist for a much longer time (which is determined by the half-life of the reporter protein). In protein–protein interaction reporter systems, a significantly longer time is required for the reporter protein expression because sufficient levels of both chimeric interactive proteins must be achieved for adequate transactivation of the reporter gene construct. Future studies to look at the half-lives of the fusion proteins, reporter mRNA, and different reporter proteins (e.g., destabilized short-lived reporters) will help to optimize the kinetics of different reporter systems.
Another important observation is that in several reporter systems, including cis- (e.g., p53 [18] or NFAT [40]), trans- or TSTA (e.g., CEA [39] or PSA [67]), and protein–protein interaction systems [41,68], the degree of in vivo activation is more than a log lower than that observed in vitro. These observations clearly demonstrate the differences between the in vitro and in vivo conditions that influence the regulation of gene expression and protein–protein interactions in various signaling pathways and emphasize the value of noninvasive in vivo imaging.
PET-based Imaging of Adoptive Therapy. Reporter gene imaging is also being used to visualize adoptively transferred cells (e.g., T-lymphocytes and stem/progenator cells). Genetically labeling cells to be used for adoptive therapy has been used previously and dates back to the late 1980s [74–79] and has particular advantages. Although reporter gene expression could be detected by PCR or FACS analysis of peripheral blood, the in vivo distribution of the infused cells in the body over time could not be addressed with this approach. The results of preliminary studies that monitor the trafficking, distribution, and activation of T-lymphocytes are described below.
Trafficking of T-lymphocytes. The long-term trafficking and localization of T-lymphocytes is an important component of the immune response and in the elimination of abnormal cells and infectious agents from the body. Passive (ex vivo) labeling of T-cells with radioactive isotopes can be unstable and does not account for proliferation of activated T-cells in the body. An objective of ongoing studies in our laboratory is to image the in vivo targeting and the accumulation of Epstein–Barr virus (EBV) lymphoma-specific cytotoxic T-cells (CTLs) in autologous and allogeneic HLA-matched EBV(+) B-lymphomas using PET and optical imaging modalities. This work has been presented in abstract form only [80]. CTLs specific for homogenous EBV-transformed B-lymphocytes were obtained and were stably transduced with a constitutively expressing dual-reporter gene (HSV1tk/eGFP fusion gene). Specific accumulation and localization of radioactivity was observed only in the autologous and allogeneic HLA-matched EBV(+) B-lymphoma and in the spleen; no localization was seen in the allogeneic HLA-matched, EBV(-) negative lymphoma and the HLA-mismatched EBV(+) lymphoma (Figure 11). Sequential imaging over 72 hr in another set of animals showed trafficking as well as targeting of the transduced and radiolabeled CTLs. These preliminary studies indicate that it may be feasible to isolate and transduce CTLs (and other immune-specific cells) with reporter constructs and then monitor their targeting and proliferation in the donor or HLA-matched recipient using noninvasive reporter gene PET imaging. Importantly, these imaging studies can be repeated in the same animal sequentially over time because the adoptively transferred CTLs have been genetically labeled with a reporter gene.

Micro-PET imaging of T-cell migration and targeting. Xenograft layout (left panel) of a mouse bearing xenografts generated from an autologous Epstein -Barr virus transformed B-lymphoma cell line (EBV(+) BLCL) (T-1), an allogenic HLA-A0201 matched EBV(+) BLCL (T-2), an HLA-mismatched EBV(+) BLCL (T-3), and an HLA-A0201 matched EBV(-) ALL (T-4) tumors in the left and right shoulders and the left and right thighs, respectively. Axial micro-PET images through the shoulders (upper-middle panel) and thighs (lower-middle panel) from a single representative animal. In situ fluorescence image (right panel) of an autologous EBV(+) BLCL xenografts showing a nest of infiltrating green fluorescent cells. Images were obtained 8 days after intravenous injection of CTLs previously transduced with the HSV1tk/eGFP dual-reporter gene, and 1 day after intravenous administration of [14I]FIAU. Figure was adapted from Doubrovin et al. [80].
When combined with the imaging of NFAT-mediated activation of T-cells (described earlier) [40], noninvasive imaging using PET would allow for monitoring the trafficking, proliferation, and antigen-specific activation of T-cells in antitumor vaccination trials. A dual-reporter system (containing a “constitutive or beacon” reporter to identify the site, number, and duration of T-cell location, plus an “inducible or sensor” reporter to identify the site, intensity, and duration of T-cell activation—e.g., NFAT expression) could be used for genetic labeling of T-cells prior to adoptive therapy. This dual-reporter system could address several important issues related to adoptive T-cell therapies. For instance, is there substantial proliferation of adoptively transferred T-cells at the target site, or does activation and proliferation occur at other sites (e.g., specific lymphoid organ sites), followed by migration and localization to the target site?
This question could potentially be addressed in a quantitative manner by repetitive PET imaging of the dual-reporter system described above, repetitive imaging in the same animal or subject over time.
Issues for the Future
Molecular imaging has its roots in both molecular biology and cell biology as well as in imaging technology. This rapid growth is due to the established base of three in vivo imaging technologies (nuclear, MR, and optical), and to the established programs in molecular and cell biology. However, it is the convergence of these disciplines that is at the heart of this success story and is the wellspring for further advances in this new field. The development of versatile and sensitive assays that do not require tissue samples will be of considerable value for monitoring molecular–genetic and cellular processes in animal models of human disease, as well as for studies in human subjects. Noninvasive imaging of molecular – genetic and cellular processes will complement established ex vivo molecular–biological assays that require tissue sampling, and would provide a spatial as well as a temporal dimension to our understanding of various diseases.
Continued success in the future depends on bringing the imaging disciplines closer together (as we have seen in the complementary benefits of dual-modality reporters—optical and nuclear—imaging paradigms), as well as further involvement with our molecular biology and cell biology colleagues. Complementary instrumentation for imaging both animals and patients provides a unique opportunity for substantive translational research. Animals bearing transduced xenografts containing specific reporter constructs and animals transduced with viral vectors bearing similar reporter constructs currently provide the main resource for in vivo molecular imaging research. Transgenic animals expressing reporter constructs systemwide or reporter constructs that are expressed only in certain organs or cell types have been developed and are a very active area of current research. Noninvasive imaging can assist in these studies by non-invasive assessments of the activation or suppression of specific endogenous genes, the activity of specific signal transduction pathways and specific protein–protein interactions. This information could be useful in defining and monitoring cell transformation to a more malignant state, or whether a specific drug effectively targets a specific signal transduction pathway. Studies designed to test these hypotheses are now being developed.
Reporter gene imaging studies will be more limited in patients compared to that in animals due to the necessity of transducing target tissue with specific reporter constructs and the use of transgenic animals bearing the reporter constructs. Ideal vectors for targeting specific organs or tissue (tumors) do not yet exist, although this is a very active area of human gene therapy research. Each new vector requires extensive and time-consuming safety testing prior to regulatory approval for human administration. Similarly, approval of new radiolabeled probes for imaging the expression of new reporter systems with PET or gamma camera will be required. The translation of molecular imaging research from animal “proof of principle” experiments into patient studies and clinical application will be measured and will develop more slowly under careful monitoring.
Two alternative molecular imaging approaches are being explored: (1) the development of better “direct” imaging probes, and (2) the use and validation of “surrogate” imaging markers that reflect the activity of specific molecular processes or the activity of a specific signal transduction pathway. The “direct” molecular imaging motif builds on established radiochemistry–nuclear medicine relationships. This needs to be expanded through the development of new relationships and focused interactions among molecular/cellular biologists, chemists, and pharmacologists, as well as among chemists, radiochemists, imagers, and clinicians. The next generation of direct molecular imaging probes will come from better interactions between pharmaceutical companies, academia, and hospital centers. Such interactions are now being pursued and the objective is to develop and evaluate new compounds for imaging; compounds that target specific molecules (e.g., DNA, mRNA, proteins) or activated enzyme systems in specific signal transduction pathways. The “indirect surrogate” imaging motif benefits from the use of established imaging probes (e.g., FDG), but will require extensive validation (involving established ex vivo molecular–biological assays) that correlate the level of endogenous gene expression or the activity of a specific signal transduction pathway to specific physiological or biochemical processes that can be imaged using established (or new) radiolabeled probes (e.g., FDG). In either case, considerable effort will have to be expended to validate and insure that the surrogate and direct imaging motifs provide valid and reliable information.
The primary applications of noninvasive in vivo marker/reporter gene imaging are likely to be: (i) quantitative monitoring of transduction efficacy in gene therapy and animal research protocols by imaging the location and extent of transgene expression; (ii) non-invasive assessments of endogenous molecular events in transduced tissue using different reporter gene imaging technologies; (iii) monitoring cell trafficking and replication of immune-competent cells following ex vivo transduction; (iv) repetitive, quantitative evaluation of reporter gene expression over time in both patients and transgenic animals during longitudinal experiments. Applications involving therapeutic genes will involve the use of bicistronic and fusion gene vectors that express therapeutic and reporter gene products from a common transcript, or the use of multiple transfer vectors where each vector expresses a different therapeutic or reporter gene.
Noninvasive reporter gene imaging will have both preclinical and direct clinical applications in the study of various diseases where different molecular events need to be monitored by whole body imaging over time. These include: (1) assessment of trafficking for different cell populations (i.e., bone marrow transplantation, T-cell and NK-cell vaccines, stem cell therapy, etc.) and their activation, proliferation, or differentiation; (2) assessment of the efficacy and specificity of various gene therapy vectors (i.e., nonreplicating and replicating adenoviral and herpesviral vectors, AAV, lentiviral, plasmid/liposomal vectors, etc.); and (3) indirect monitoring of various endogenous genes and activities of different signal transduction pathways (i.e., oncogene activity, growth factor receptor-mediated, proapoptotic, or differentiation-inducing signaling, etc.).
Many applications of in vivo reporter gene imaging will develop over the next several years. Three applications related to cancer therapy are likely to be introduced into the clinic in the near future. One application involves visualization of therapeutic gene expression following transduction of primary tumors and/or metastases with “suicide” genes (i.e., HSV1-tk, CD), which sensitize the transduced tumor to cytotoxic prodrugs (i.e., GCV, 5FC). In this therapeutic/imaging protocol, PET will be performed to assess the efficacy and tumor specificity of vector delivery and therapeutic gene expression; optimization of viral vector dose, mode, and timing of administration could be performed and correlated with treatment response.
Another application involves intratumoral administration of different types of conditionally replicating oncolytic viruses that constitutively overexpress the HSV1-tk gene. In this paradigm, PET imaging of HSV1-tk expression would be performed to assess the efficacy and tumor specificity of oncolytic virus delivery, replication, and lateral spread through the tumor. PET imaging of HSV1-tk reporter gene expression could potentially visualize the wave of the oncolytic virus spreading through the tumor and identify the presence of virus in adjacent or remote normal tissues.
A third application could involve the administration of tumor-specific CTLs following ex vivo transduction with a reporter gene under control of an NFAT-promoter. PET imaging of reporter gene expression could be used to visualize the sites of T-cell activation upon recognizing tumor-specific antigens. This could provide for improvements in antitumor sensitization procedures and optimization of T-cell dose and schedule of administration. A similar approach could be used for imaging the distribution and engraftment of stem cells in the treatment of various diseases, including cancer.
Future research directions should also be aimed at the development of novel nonimmunogenic reporter genes for human applications; development of novel reporter gene–reporter substrate combinations that would be suitable for CNS imaging (with probes that cross the BBB easily); the development of reporter systems for imaging key oncogenic signal transduction pathways that are involved in tumorigenesis, angiogenesis, tumor progression, invasion, organ/tissue-specific metastasis, and tumor maintenance. Development of transgenic reporter mice that allows for imaging the endogenous expression/activity of various genes and signal transduction pathways would significantly further our understanding of many incurable diseases including cancer. The ability to noninvasively image transgenic animals will also foster the development of novel and more effective drugs to treat these diseases.
We remain optimistic; the tools and resources largely exist and we should be able to perform limited gene imaging studies in patients in the near future. The advantages and benefits of noninvasive imaging to monitor transgene expression in gene therapy protocols are clear. The ability to visualize transcriptional and posttranscriptional regulation of endogenous target gene expression, as well as specific intracellular protein–protein interactions in patients will provide the opportunity for new experimental venues in patients. Optimistically, they could include the potential to image the malignant phenotype of an individual patient's tumor at a molecular level and to monitor changes in the phenotype over time. The potential to image a drug's effect on a specific signal transduction pathway in an individual patient's tumor provides the opportunity for monitoring treatment response at the molecular level. At the moment, this requires the use of reporter gene transduction vectors that target specific organs or tissue (tumors), and this will initially limit the translation and application of reporter gene technology to patients. However, direct and surrogate molecular imaging may begin to fill this gap over the next decade.
