Abstract
Noninvasive in vivo molecular imaging has developed over the past decade and involves nuclear (positron emission tomography [PET], gamma camera), magnetic resonance, and in vivo optical imaging systems. Most current in vivo molecular imaging strategies are “indirect” and involve the coupling of a “reporter gene” with a complementary “reporter probe.” Imaging the level of probe accumulation provides indirect information related to the level of reporter gene expression. Reporter gene constructs are driven by upstream promoter/enhancer elements; reporter gene expression can be constitutive, leading to continuous transcription and used to identify the site of transduction and to monitor the level and duration of gene (vector) activity. Alternatively, reporter gene expression can be inducible, leading to controlled gene expression, or reporter genes can function as a “sensor” to monitor the level of endogenous promoters and transcription factors. The development of versatile and sensitive assays that do not require tissue sampling will be of considerable value for monitoring molecular-genetic and cellular processes in animal models of human disease, as well as for studies in human subjects in the future. Noninvasive imaging of molecular-genetic and cellular processes will complement established ex vivo molecular-biologic assays that require tissue sampling, and will provide a spatial as well as a temporal dimension to our understanding of various diseases. Several examples of imaging endogenous biologic processes in animals using reporter constructs, radiolabeled probes, and PET imaging are reviewed (e.g., p53-dependent gene expression, T-cell receptor-dependent activation of T-lymphocytes, and preliminary studies of endogenous HIF-1α expression). Issues related to the translation of noninvasive molecular imaging technology into the clinic are also discussed.
Keywords
The extraordinary developments in both molecular/cellular biology and noninvasive imaging over the past 2 decades occurred largely in parallel with little direct interaction. This began to change less than a decade ago, however, when “reporter” gene technology was first applied to in situ imaging of tissue sections (Wick et al., 1989; Hooper et al., 1990; Welsh et al., 1997), and applied later to noninvasive, in vivo imaging. Three different noninvasive imaging technologies developed more or less in parallel: (1) magnetic resonance imaging (Weissleder et al., 1990, 1991, 1997; Kayyem et al., 1995; Louie et al., 2000); (2) nuclear imaging (quantitative autoradiography, gamma camera, and PET) (Tjuvajev et al., 1995, 1996, 1998; Gambhir et al., 1998, 1999); and (3) in vivo optical imaging of small animals (Mayerhofer et al., 1995; Bennett et al., 1997; Sweeney et al., 1999; Rehemtulla et al., 2000). These developments led to the term molecular imaging, which was coined in the mid 1990s. Molecular imaging provides visualization in space of normal as well as abnormal cellular processes at a molecular or genetic level rather than at the anatomic level.
Molecular imaging has its roots in both molecular biology and cell biology as well as in imaging technology. These disciplines have now converged to provide a well-established foundation for exciting new research opportunities and for translation into clinical applications. For example, established ex vivo molecular assays require invasive sampling procedures that preclude sequential studies in the same animal or in human subjects. Tissue sampling may not always adequately represent the biochemical or pathologic process under investigation due to tissue heterogeneity, which is especially characteristic of some tumors. Furthermore, temporal studies that employ molecular-biologic assays have required large numbers of animals that are killed at specific time points in order to achieve a statistically significant temporal profile. The development of sensitive imaging-based assays to monitor molecular-genetic and cellular processes in vivo would be of considerable value in the study of animal models of human disease (including transgenic animals), as well as for studies in human subjects. Noninvasive imaging of molecular-genetic and cellular processes will complement established ex vivo molecular-biologic assays, and imaging can provide a spatial as well as a temporal dimension to our understanding of various diseases.
Using noninvasive imaging instrumentation, molecular imaging has now been extended to the visualization of molecular processes in living animals and will soon be applied to patients with specific diseases. It is now possible to serially monitor molecular-genetic processes over time in the same animal, to assess such processes before and after a specific experimental intervention, to assess the effects and time-course of specific genetic alterations in transgenic animals, and to better assess treatment effects of new molecular-based therapies and drugs targeted to specific molecular or signal transduction steps. Recent progress in our understanding of the molecular-genetic mechanisms of many diseases and the application of new biologically based approaches in therapy are exciting new developments. New gene-based therapies can provide control over the level, timing, and duration of action of many biologically active transgene products by including specific promoter/activator regulatory elements in the genetic material transferred. Noninvasive imaging of molecular-genetic and cellular processes will accelerate these developments and lead to more effective therapeutic strategies. Methods are being developed for controlled gene delivery to various somatic tissues and tumors using novel gene constructs, for targeting vectors to specific tissues/organs, and for controlling gene expression using cell specific, replication-activated and drug-controlled expression systems (Harrington et al., 2000; Fussenegger, 2001; Nakagawa et al., 2001). A noninvasive, clinically applicable method for quantitatively imaging the expression of transduced genes in target tissue or specific organs would be of considerable value. It will facilitate the monitoring and evaluation of gene therapy in human subjects by defining the location(s), magnitude and persistence of gene expression over time.
IMAGING STRATEGIES
Two imaging strategies—“direct” and “indirect”—are currently the most widely used. Direct molecular imaging can be defined in terms of a probe-target interaction, whereby the resultant image of probe localization and magnitude (image intensity) is directly related to its interaction with the target epitope or enzyme. These strategies are based on imaging the target directly, usually with a target-specific probe. Indirect molecular imaging is a little more complex in that it may involve multiple components. One example of indirect imaging that is now being widely used is “reporter imaging,” which usually includes a “reporter gene” and a “reporter probe.” The reporter gene product can be an enzyme that converts a reporter probe to a metabolite that is selectively trapped within transduced cells. Alternatively, the reporter gene product can be a receptor or transporter that “irreversibly traps” the probe in transduced cells. Indirect reporter imaging paradigms are currently more widely used in molecular imaging and will be discussed in greater detail below.
Direct imaging strategies are common in nuclear medicine and include monoclonal antibody targeting of a particular cell membrane epitope, or imaging the activity of a particular enzyme (e.g., hexokinase) with an enzyme-specific probe (e.g., deoxyglucose), or imaging the activity of a particular transporter with a transporter-specific probe. Imaging cell surface specific antigens or epitopes with radiolabeled antibodies is an example of direct molecular imaging that has developed over the past 30 years. PET imaging of receptor density/occupancy using small radiolabeled molecular probes has also been widely used, particularly in neuroscience research. These examples represent some of the first molecular imaging applications used in clinical nuclear medicine research.
A more recent direct imaging strategy involves the development of antisense and aptomer oligonucleotide probes that specifically hybridize to target mRNA or proteins in vivo. Radiolabeled antisense probes are RASONs that have been developed to directly image endogenous gene expression at the transcriptional level. RASONs are small oligonucleotide sequences that are complementary to a small segment of target mRNA or DNA and could potentially target any specific mRNA or DNA sequence. In this context, imaging specific mRNAs with RASONs produces “direct” images of specific molecular-genetic events. Some efficacy for gamma camera and PET imaging endogenous gene expression using RASONs has been reported (Phillips et al., 1997; Tavitian et al., 1998; Cammilleri et al., 1996; Dewanjee et al., 1994). Nevertheless, RASON imaging has several serious limitations, including: (1) low number of target mRNA/DNA molecules per cell; (2) limited tracer delivery (poor cell membrane and vascular permeability, cannot penetrate blood-brain barrier); (3) poor stability (degradation by H-RNAse); (4) slow clearance (slow washout of nonbound oligonucleotides); and (5) comparatively high background activity and low specificity of localization (low target/background ratios). Imaging specific RASON targets in the body is complicated, and interpretation of the images must be approached with caution.
Indirect molecular imaging is currently the most widely used strategy for radionuclide-based imaging (Tjuvajev et al., 1999; Yu et al., 2000) as well as for optical (Mayerhofer et al., 1995; Bennett et al., 1997; Sweeney et al., 1999; Rehemtulla et al., 2000) and magnetic resonance (Weissleder et al., 1997; Louie et al., 2000) imaging. Most indirect molecular imaging paradigms involve the use of reporter-transgene technology and specific probes to produce an image that reflects reporter gene expression. Reporter gene imaging initially used optical technology that frequently required post-mortem tissue sampling and processing (e.g., betagalactosidase assay). More recent studies have emphasized noninvasive imaging techniques involving live animals (and soon human subjects). Noninvasive reporter gene imaging involves a reporter transgene (e.g., HSV1-tk gene) placed under the control of upstream promoter/enhancer elements. These promoter/enhancer elements can be “always turned on” with constitutive promoters (e.g., LTR, RSV, CMV), or they can be “sensitive” to activation by specific endogenous transcription factors (factors that bind to and activate specific promoter-enhancer elements). Several noninvasive imaging paradigms have been described, and it has recently been shown that transcriptional regulation of endogenous (host tissue) gene expression can be imaged using both nuclear (PET) and optical (fluorescence) imaging (Doubrovin et al., 2001; Ponomarev et al., 2001), and combined PET and bioluminescence dual-reporter imaging will be reported in the near future.
REPORTER GENE IMAGING
A common feature of all reporter vectors is the cDNA expression cassette containing the reporter transgene(s) of interest (e.g., HSV1-tk). The advantage and versatility of reporter vectors is that the design and arrangement of the expression cassette can be varied (Naylor et al., 1999). For example, the reporter transgene(s) can be driven by any promoter/enhancer sequence of choice. The promoter can be constitutive (leading to continuous transcription), or it can be inducible (leading to controlled expression). The promoter can also be cell specific, allowing expression of the transgene to be restricted to certain cells and organs. The paradigm for quantitative imaging of transgene expression involves several steps, including the initiation of transcription (that can be controlled by specific promoter/enhancer elements), the process of DNA transcription and stabilization of mRNA, and subsequent translation of mRNA into the gene product (a protein). In this manner, the reporter expression cassette can be designed to provide information about endogenous gene regulation, mRNA stabilization, and specific protein-protein interactions.
A general paradigm for gene imaging using radiolabeled probes was initially described in 1995 (Tjuvajev et al., 1995) and is diagrammatically shown in Fig. 1. This paradigm requires the selection of an appropriate combination of reporter/marker transgene and reporter/marker probe. It is important to note that imaging transgene expression is independent of the vector used to transfect/transduce target tissue, namely, any of several currently available vectors can be used (e.g., retrovirus, adenovirus, adeno-associated virus, lentivirus, liposomes, etc.). The reporter transgene can encode for an enzyme (e.g., HSV1-tk) that selectively metabolizes the radiolabeled probe and results in its entrapment and accumulation in the transduced cell. It may be helpful to consider this reporter-imaging paradigm as an example of an in vivo enzymatic radiotracer assay that reflects reporter gene expression. Enzymatic amplification of the signal (e.g., level of radioactivity) facilitates imaging the location and magnitude of reporter gene expression. Viewed from this perspective, reporter gene imaging using HSV1-tk is similar to imaging hexokinase activity with FDG.
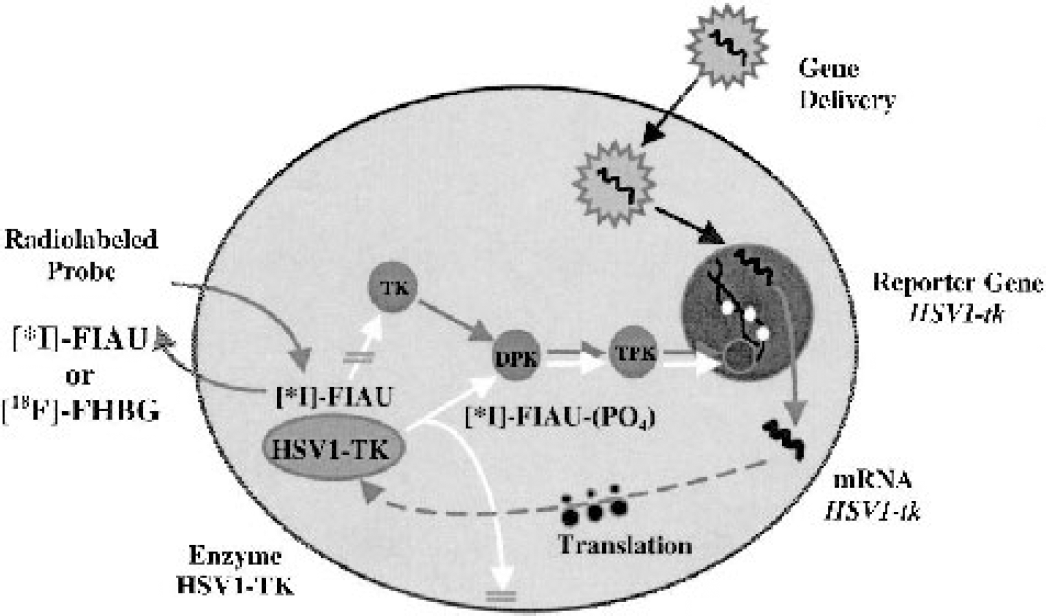
Schematic for imaging HSV1-tk reporter gene expression with reporter probes FIAU and FHBG. The herpes simplex virus 1 thymidine kinase (HSV1-tk) gene complex is transfected into target cells by a vector. Inside the transfected cell, the HSV1-tk gene is transcribed to HSV1-tk mRNA and then translated on the ribosomes to a protein (enzyme), HSV1-TK. After administration of a radiolabeled probe (e.g., 5-iodo-2'-fluoro-2'deoxy-1-β-
Wild-type HSV1-tk (Tjuvajev et al., 1998) or a mutant HSV1-tk gene, HSV1-sr39tk (Gambhir et al., 2000), are the reporter genes most commonly used in current molecular imaging studies using radiolabeled probes and PET imaging. The HSV1-tk and HSV1-sr39tk gene products are proteins (enzymes) that have less substrate specificity than mammalian thymidine kinase 1 (TK1). The viral kinases phosphorylate a wider range of compounds, including acycloguanosines (e.g., ACV, GCV, FHBG) and 2'-fluoro-nucleoside analogues of thymidine (e.g., FIAU). This difference between mammalian and viral TK enzymes permits the development and use of radiolabeled probes that are phosphorylated to a significantly greater extent by HSV1-tk or HSV1-sr39TK in comparison to mammalian TK1.
Alternatively, a reporter gene can encode for an extracellular or intracellular receptor that “irreversibly” binds or transports a radiolabeled or paramagnetic probe. The hD2R is an example of such a reporter gene (MacLaren et al., 1999). This was a very clever strategy because hD2R expression is largely limited to the striatal-nigral system of the brain, and because an established radiolabeled probe, FESP, has been extensively used to image striatal-nigral D2 receptors in human subjects (Barrio et al., 1989). Similarly, the hSSTR2 gene has been suggested as a potential reporter gene for human studies (Rogers et al., 1999, 2000), because hSSTR2 expression is largely limited to carcinoid tumors. There is also a complementary radiolabeled somatostatin analogue ([111In]DTPA-octreotide) that can be used for imaging hSSTR2 expression (Ur et al., 1993). Radiolabeled octreotide has also been approved for administration to patients with carcinoid tumors. Both of these reporter systems have distinct benefits with respect to initiating molecular/reporter imaging in human subjects. Receptor expression on the surface of cells, however, is a complex process and involves intracellular trafficking and cell membrane expression that is likely to be altered under different conditions and different disease states. It remains to be shown whether imaging receptor-based reporter systems (e.g., the hD2R and hSSTR2 reporter gene systems) will provide a consistent and reliable measure of reporter gene expression. In either case, the level of probe accumulation (level of radioactivity) must be shown to be proportional to the level of gene expression.
IMAGING ENDOGENOUS BIOLOGIC PROCESSES
Reporter-gene imaging is being used to visualize transcriptional and posttranscriptional regulation of target gene expression, as well as specific intracellular protein-protein interactions. Several examples from studies performed in our laboratory will be described, including imaging p53-dependent gene expression, imaging TCR-dependent activation of T lymphocytes as well as some preliminary data on imaging endogenous HIF-1α expression.
p53
Imaging transcriptional regulation of endogenous genes in living animals (and potentially in human subjects) using noninvasive imaging techniques can provide a better understanding of normal and cancer-related biologic processes. A recent article from our group was the first to show that p53-dependent gene expression can be imaged in vivo with PET and by in situ fluorescence (Doubrovin et al., 2001). A retroviral vector (Cis-p53/TKeGFP) was generated by placing the herpes simplex virus type 1 thymidine kinase (tk) and enhanced green fluorescent protein (egfp) fusion gene (TKeGFP, a dual-reporter gene) under control of a p53-specific response element. DNA damage-induced upregulation of p53 transcriptional activity was demonstrated and correlated with the expression of p53-dependent downstream genes (including p21). These findings were observed in U87 (p53 +/+) cells and xenografts, but not in SaOS (p53 -/-) cells. This was the first demonstration that a Cis-reporter system (Cis-p53/TKGFP) was sufficiently sensitive to image endogenous gene expression using noninvasive nuclear (PET) imaging (Fig. 2a). The PET images corresponded with upregulation of genes in the p53 signal transduction pathway (p53-dependent genes) in response to DNA damage induced by BCNU chemotherapy (Fig. 2b). PET imaging of p53 transcriptional activity in tumors using the Cis-p53TKGFP reporter system could be used to assess the effects of new drugs or other novel therapeutic paradigms that are mediated through p53-dependent pathways. For example, specific p53 gene therapy strategies that are based on p53 over-expression (Merritt et al., 2001) could be monitored by noninvasive imaging.
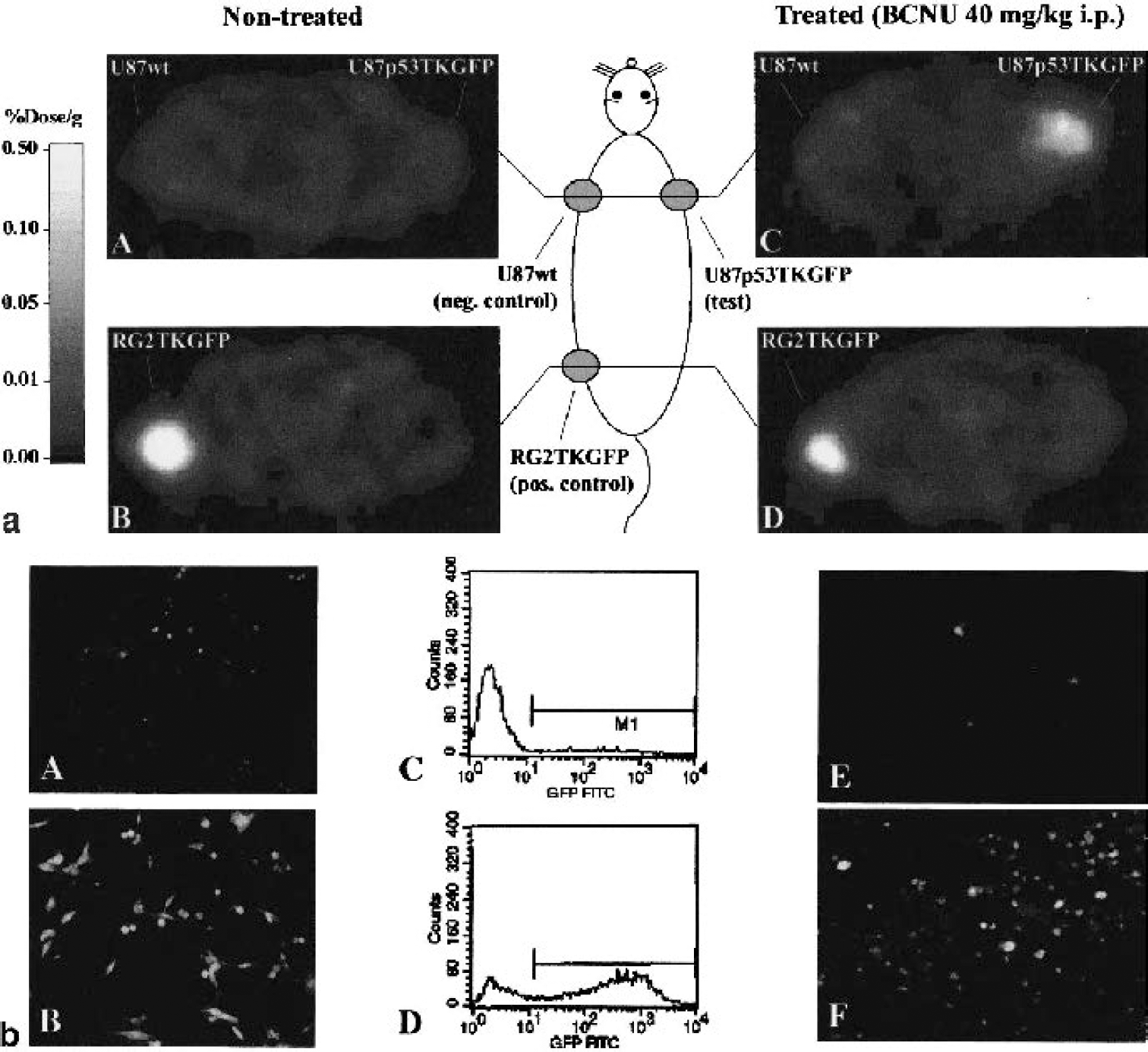
(a) PET imaging of endogenous p53 activation (adapted from Doubrovin et al., 2001). Transaxial PET images (GE Advance tomograph) through the shoulder (A, C) and pelvis (B, D) of two rats are shown (upper panel); the images are gray-scaled to the same radioactivity scale (percent dose per gram). An untreated animal is shown on the left, and an animal treated with an alkylating anticancer drug, carmustine (BCNU), is shown on the right. Both animals have three subcutaneous tumor xenografts: U87p53TKGFP (test) in the right shoulder, U87 wild-type (negative control) in the left shoulder, and RG2TKGFP (positive control) in the left thigh. The nontreated animal on the left shows localization of radioactivity only in the positive control tumor (RG2TKGFP); the test (U87p53TKGFP) and negative control (U87wt) tumors are at background levels. The BCNU-treated animal on the right shows significant radioactivity localization in the test tumor (right shoulder) and in the positive control (left thigh), but no radioactivity above background in the negative control (left shoulder). (b) Validation of Cis-p53/TKGFP reporter system in cell cultures (left and middle panels; adapted from Doubrovin et al., 2001). Fluorescence microscopy and FACS analysis of a transduced U87p53/TKGFP cell population in the noninduced (control) state (A, C), and 24 hours after a 2-hour treatment with BCNU 40 μg/mL (B, D). Assessment of Cis-p53/TKGFP reporter system in U87p53/TKGFP subcutaneous tumor tissue (right panel; adapted from Doubrovin et al., 2001). Fluorescence microscopy images of U87p53/TKGFP subcutaneous tumor samples obtained from nontreated rats (E) and rats treated with 40 mg/kg BCNU intraperitoneally (F).
It should also be pointed out that the dual reporter construct (TKeGFP–fusion gene) provides the opportunity for multimodality (both nuclear and optical imaging) imaging of endogenous gene expression in vivo. The TKeGFP reporter gene could be introduced into other reporter assay systems to assess other molecular-biologic pathways. It should also be possible to use the TKeGFP reporter gene in transgenic animals; this will facilitate the monitoring and assessment of newly cloned genes or novel signal transduction pathways. Another advantage of the dual reporter system is the ability to compare the images of reporter gene expression obtained with PET, gamma camera, or autoradiography with corresponding in situ GFP fluorescence images. The comparison between GFP fluorescence and autoradiographic images, coupled with histology of corresponding tissue sections, provides for spatial and quantitative assessments of reporter gene expression at the microscopic as well as macroscopic level.
NUCLEAR FACTOR OF ACTIVATED T-CELLS
T-cell activation is an essential component of the immune response in many normal and disease states. The objective of a recent study in our laboratory (Ponomarev et al., 2001) was to monitor and assess TCR-dependent activation of T cells in vivo using noninvasive PET imaging. A retroviral vector (Cis-NFAT/TKeGFP) was generated by placing the fusion gene (TKeGFP) under control of the nuclear factor of activated t-cells (NFAT) response element. A human T-cell leukemia cell line (Jurkat) that expresses a functional TCR was transduced with the Cis-NFAT/TKGFP reporter vector and used in these studies. Known activators of T cells (anti-CD3 and anti-CD28 antibody) produced significantly higher levels of TKeGFP reporter gene expression (increased GFP fluorescence, increased levels of HSV1-tk mRNA, and increased [14C]FIAU accumulation in vitro) in Cis-NFAT/TKeGFP+ Jurkat cells, in comparison to nontreated or nontransduced cells. In mice with focal Cis-NFAT/TKeGFP+ Jurkat cell infiltrates, similar results were observed in the microPET images (Fig. 3a), and the in vivo fluorescence images (Fig. 3b). A strong correlation between TKeGFP expression and upregulation of T-cell activation markers (CD69 and IL-2 production) was demonstrated both in vitro and in vivo. These results demonstrated that (1) activation of the NFAT signal transduction pathway occurs after TCR stimulation, and (2) PET imaging of T-lymphocyte activation in tumors following TCR engagement is feasible using the described TKeGFP-based Cis-reporter system. This imaging paradigm could be used to assess the efficacy of novel antitumor vaccines and adoptive immunotherapy.
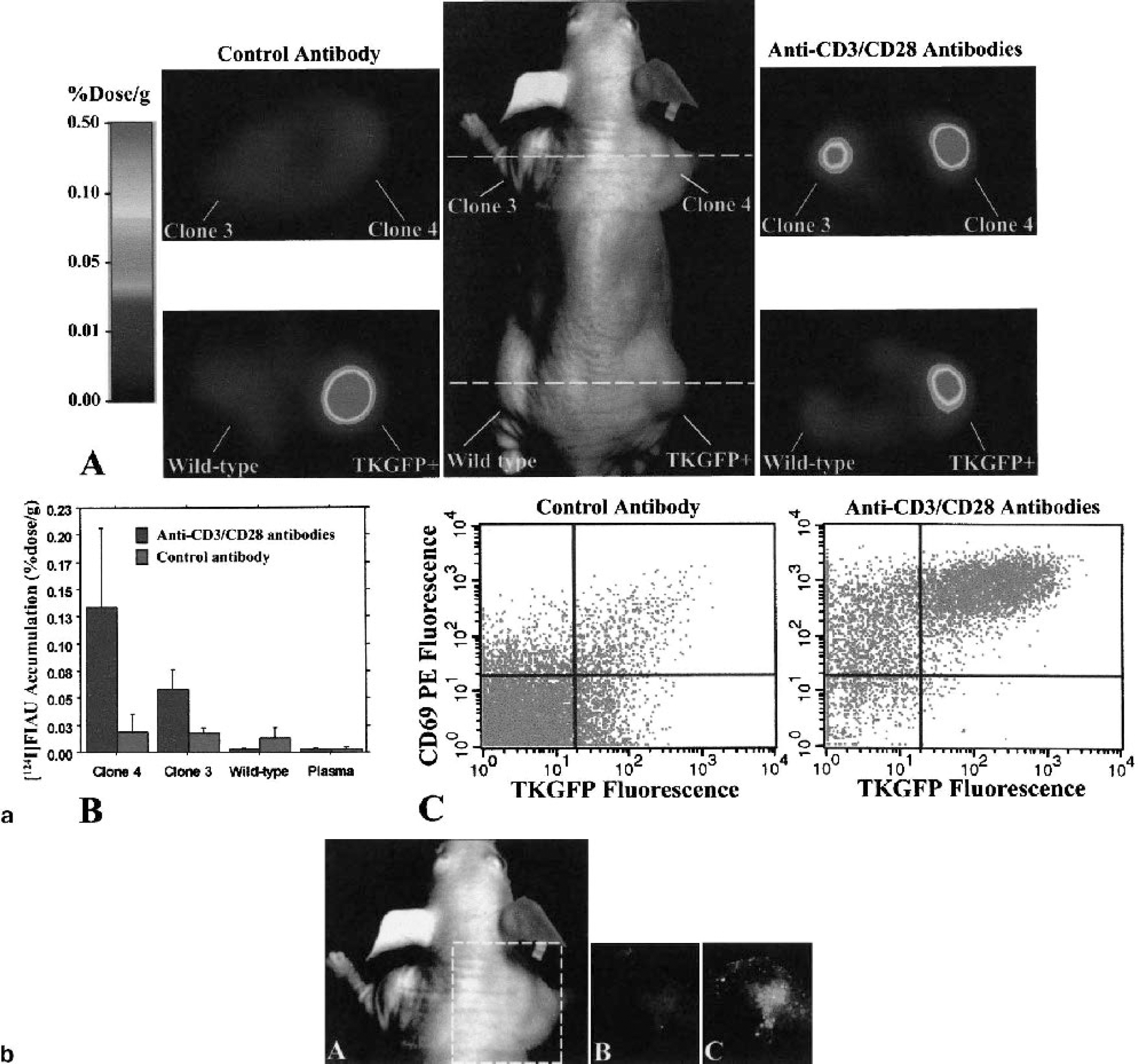
(a) Imaging NFAT-TKGFP reporter system activity with [124I]FIAU and PET plus assessments in tissue samples. Photographic image of a typical mouse bearing different subcutaneous infiltrates (middle panel); transaxial PET images (GE Advance tomograph) of TKGFP expression in a mouse treated with control antibody (left panel) and anti-CD3/CD28 antibodies (right panel) were obtained at the levels indicated by the dashed lines (A). [124I]FIAU accumulation (percent dose per gram) in tissue samples of the Jurkat/dcmNFATtgn clone 3 and 4 infiltrates, wild-type Jurkat infiltrates, and blood plasma, obtained after PET imaging (B). FACS profiles of TKGFP and CD69 expression in a tissue sample from the same Jurkat/dcmNFATtgn clone 4 infiltrate that was imaged with PET (C).
HYPOXIA-INDUCIBLE TRANSCRIPTION FACTOR 1
Tissue hypoxia has been recognized as a significant component of tumors and is associated with malignant progression and resistance to radiation therapy. Cellular responses to hypoxia are mediated largely through a transcriptional activator, hypoxia-inducible transcription factor 1-α (HIF-1α). A heterodimer (HIF-1) is formed in response to hypoxia, which binds to the hypoxia response element (HRE) in the enhancer region of many target genes (erythropoietin, VEGF, glycolytic enzymes, and other stress response genes) and enhances gene transcription. Work in our laboratory is being directed to the development of a noninvasive imaging paradigm to monitor and assess the level of HIF-1 activity during tumor development and growth. This work-in-progress has been presented in abstract form only (Serganova et al., 2002). We developed a dual reporter vector containing a “sensor” reporter element (Cis-HRE/TKGFP) and a “beacon” reporter element (CMV-RFP/XPRT) that is constitutively expressed; this vector was used to transduce C6 and RG2 glioma cells. Optical, radioactivity, and molecular assays were performed to assess the activity of the reporter systems.
Transcriptional activation of the HIF-1α signaling pathway was observed after 16 hours of exposure to 2% O2 and to CoCl2 (20–100 μmol/L). The hypoxic/treated cells exhibited high levels of TKGFP reporter protein as measured by fluorescence (FACS) and [14C]FIAU accumulation in comparison to control nontreated cells. The CoCl2-induced increase in HIF-1α resulted in a corresponding upregulation of VEGF and TKGFP gene expression (mRNA and protein levels). Noninvasive PET imaging studies of the HIF-1α signaling pathway using [124I]FIAU have been performed, and the results are very encouraging. This novel reporter system, Cis-HRE-TKGFP, for monitoring hypoxia by fluorescence and nuclear imaging should provide a valuable resource for monitoring the time course of hypoxia during tumor development, growth, and treatment (e.g., radiation, and antiangiogenesis chemotherapy).
CONCLUSIONS
The opportunities for molecular imaging (and biomedical imaging as a whole) indeed look bright. Noninvasive reporter gene imaging is a very exciting strategy that can be fully exploited in experimental animals. Reporter gene imaging applications, however, will be more limited in patients owing to the necessity of transducing target tissue with specific reporter constructs. Ideal vectors for targeting specific organs or tissue (tumors) in patients do not exist at this time, although vector development is a very active area of human gene therapy research. Each new vector requires extensive and time-consuming safety testing prior to government approval for human administration. Similarly, government approval is required for human administration of new radiopharmaceuticals and radiolabeled probes. The translation of molecular imaging research into patient studies and clinical application must proceed stepwise and must be carefully monitored.
The benefits of noninvasive monitoring (imaging) of transgene expression in gene therapy protocols are substantial and provide a stimulus to proceed with vigor. The ability to visualize transcriptional and posttranscriptional regulation of endogenous target gene expression, as well as specific intracellular protein-protein interactions in patients will provide the opportunity for new experimental venues for research in patients. For example, it may be possible to image a drug's effect on a specific regulatory or signal transduction pathway in an individual patient's tumor. The development of versatile and sensitive assays that do not require tissue samples will be of considerable value for monitoring molecular-genetic and cellular processes in animal models of human disease, as well as for studies in human subjects in the future. Noninvasive imaging of molecular-genetic and cellular processes will complement established ex vivo molecular-biologic assays that require tissue sampling, and imaging will provide a spatial as well as a temporal dimension to our understanding of various diseases.
At the moment, the requirement of reporter gene transduction vectors that target specific organs or tissue (tumors) is the Achilles' heel of this technology. Nevertheless, we remain optimistic; the tools and resources largely exist, and we should be able to perform limited gene imaging studies in carefully selected patients in the near future.
