Abstract
Cyclooxygenase-3 (COX-3), a new acetaminophen-sensitive isoform of the COX family, has recently been cloned from canine tissues. Canine COX-3 apparently is identical to the full-length form of COX-1, with the exception that the COX-3 mRNA retains intron 1. Additionally, COX-3 mRNA expression is high in the brain. We investigated the expression of the putative rat COX-3 mRNA in primary cultures of neurons, astrocytes, endothelial cells, pericytes, and choroidal epithelial cells from the rat brain. Specific RT-PCR primers were designed to detect putative rat COX-3 mRNA, and the RT-PCR products were sequenced and compared to the known sequence of the rat COX-1 gene. Our results demonstrate that the mRNA of the putative COX-3 is expressed in all of the cell types except neurons. Cerebral endothelial cells showed the highest COX-3 expression. Whereas COX-2 expression increased several-fold after lipopolysaccharide (LPS) challenge, COX-1 and COX-3 expression did not change significantly, suggesting that cells constitutively express COX-3. In summary, we report, for the first time to our knowledge, that the putative COX-3 mRNA is detectable in rats and is differentially expressed in various cell types from rat brain, as well as that its expression is not stimulated by LPS.
A new isoform of the cyclooxygenase (COX) family has been identified recently (Chandrasekharan et al., 2002). This new isoform, named COX-3, has been cloned and characterized from canine tissues and is putatively an acetaminophen-sensitive splice variant of COX-1. Canine and human COX-3 apparently are identical to the full-length form of COX-1, with the exception that the COX-3 mRNA retains intron 1 in the coding sequence (Chandrasekharan et al., 2002). Additionally, COX-3 mRNA expression is particularly high in the brain, although the cell types responsible for expression were not investigated (Chandrasekharan et al., 2002). Given the importance of COX isoforms in the physiologic and pathologic functioning of the cerebral circulation and brain and the dependence upon rodent models in many studies, we investigated the expression of COX-3 mRNA in primary cultures of neurons, endothelial cells, pericytes, astrocytes, and choroidal epithelial cells from the cerebral cortex of the rat. The present study had several aims. First, we determined whether COX-3 mRNA was present in these cell types from rat cortex. Second, if present, we compared the expression of COX-3 mRNA with levels of mRNA for COX-1. Third, we evaluated whether COX-3 mRNA expression was constitutive or inducible with lipopolysaccharide (LPS) stimulation. To address these issues, specific RT-PCR primers were designed to detect putative COX-3 mRNA, and the RT-PCR products were sequenced and compared with the known sequence of the rat COX-1 gene.
METHODS
Animals
Wistar rats were obtained from Harlan (Indianapolis, IN, U.S.A.). All animal experiments were approved by the Animal Care and Use Committee of Wake Forest University School of Medicine.
Rat cortical neuronal culture
Primary rat cortical neurons were cultured using the method of Hampson et al. (2000). Brain cortical pieces from E18 fetuses were incubated with dispase I (2 U/mL, Roche, Mannheim, Germany) for 35 minutes at 37°C. After digestion, the cells were dissociated by gentle trituration and plated at a density of 106 cells/cm2 onto poly-D-lysine coated dishes (Becton-Dickinson, San Jose, CA, U.S.A.) in plating medium consisted of 60% Dulbecco's modified Eagles medium (DMEM, Gibco BRL, Grand Island, NY, U.S.A.), 20% F-12 HAM (Gibco BRL), and 20% horse serum (Gibco BRL) supplemented with glutamine (0.5 mM; Sigma, St Luis, MO, U.S.A.). After cell attachment, the plating medium was replaced by Neurobasal medium (Gibco BRL) supplemented with B27 (Gibco BRL, 2%), L-glutamine (Sigma, 0.5 mM), β-mercaptoethanol (Gibco BRL, 55μM), and potassium chloride (Sigma, 25 mM). Medium was changed every third day. Cultures consisted of more than 98% of neurons verified by positive immunostaining for microtubule-associated protein-2 (Becton-Dickinson) and negative immunostaining for glial fibrillary acidic protein (GFAP, Chemicon, Temecula, CA, U.S.A.).
Rat cerebral astrocyte culture
Rat cerebral astrocyte cultures were prepared as previously described (Kis et al., 2002). Cortical pieces from P1 neonatal Wistar rats were mechanically dissociated in astrocyte culture medium (DMEM supplemented with 10% fetal bovine serum and antibiotics). Dissociated cells were seeded into cell culture flasks. To obtain type I astroglia, confluent cultures were shaken at 37°C overnight. The purity of astrocytes was checked by immunostaining for GFAP, and the cells were used at passage 2.
Rat cerebral endothelial cell culture
Primary rat cerebral endothelial cells (CECs) were isolated as previously described (Kis et al., 1999) and were seeded onto collagen type IV and fibronectin coated dishes. The endothelial culture medium consisted of DMEM supplemented with 20% fetal bovine plasma derived serum (Animal Technologies Inc., Tyler, TX, U.S.A.), 2 mM glutamine, 1 ng/mL basic fibroblast growth factor, 100 μg/mL heparin, 5 μg/mL vitamin C, and antibiotics. Confluent cultures (days 4 to 5 in vitro) consisted of more than 95% of RCECs verified by positive immunohistochemistry for von Willebrand factor and negative immunochemistry for GFAP and α-smooth muscle actin.
Rat cerebral microvascular pericyte culture
Pure cultures of rat cerebral pericytes were obtained by prolonged culture of primary RCEC preparations. Pericyte survival and proliferation was favored by selective culture conditions using uncoated dishes, and DMEM was supplemented with 10% fetal bovine serum and antibiotics. Pericytes were characterized by their large size, branched morphology, positive immunostaining for α-smooth muscle actin, and absence of von Willebrand factor and GFAP staining.
Rat choroidal epithelial cell culture
Rat choroidal epithelial cell cultures were prepared from 2-week-old (20 to 30 g) Wistar rats. A pool of choroid tissues was incubated with collagenase type 2 (Worthington, Lakewood, NJ, U.S.A.; 350 U/mL) and DNase (Sigma; 10 μg/mL) for 1 hour at 37°C. After digestion, cells were dissociated by gentle trituration and were plated in culture medium [DMEM supplemented with 10% fetal bovine serum, 2 mM L-glutamine, 10 ng/mL epidermal growth factor (Gibco BRL), 50 μg/mL endothelial cell growth supplement (Sigma), and antibiotics] onto uncoated culture dishes and cultured for 40 minutes to allow attachment of contaminating cells, mostly fibroblasts. The unattached epithelial cells were collected and cultured in Primaria dishes (Becton-Dickinson). The cell culture continued without disturbance for 3 days after seeding to ensure good attachment. The medium was changed every second day thereafter for the duration of the cell culture. The cultures reached confluence in 6 to 7 days.
LPS treatment
When the cultures reached confluence, the cells were incubated with 100 ng/mL LPS (Sigma; Escherichia coli O26:B6) in the regular culture media at 37°C for different periods of time. Neuronal cultures were not incubated with LPS. After the incubations, cells were washed twice with phosphate buffered saline, and total RNA was isolated by SV Total RNA Isolation System (Promega, Madison, WI, U.S.A.). For each series of experiments, cells were from at least three independent cultures.
RT-PCR
RT-PCR experiments were performed in an Eppendorf Mastercycler thermocycler (Brinkmann Instruments Inc., Westbury, NY, U.S.A.). From each sample, 300 pg of total RNA was reverse-transcribed and amplified using AccessQuick single-tube coupled RT-PCR system (Promega) with gene specific primers targeting COX isoforms. α-actin primers (Promega) were also included in the RT-PCR reaction to normalize our RT-PCR results with an expected product length of 285 base pairs. In our preliminary experiments, we used different cycle numbers and different amounts of total RNA to titrate the optimal condition for RT-PCR amplification to be in the linear range. We used 24 cycles for COX-1 and COX-2 and 40 cycles for COX-3 amplification. In control experiments, when reverse transcription was omitted, no amplification was observed.
The design of specific primers for rat COX-3 detection was based upon the hypothesis that the rat COX-3 mRNA is identical to the full length form of COX-1, with the exception that it retains intron 1, like in the dog. The sense primer (5′-TGGAACGCTAGGCTCAACT; 1310570 to 1310588 bases of GeneBank NW043716) was designed to bind to intron 1, and the antisense (5′-AGAGGGCAGAATGCGAGTAT; 501 to 520 bases of GeneBank S67721) was designed to bind to exon 5 of the rat COX-1 gene. The expected length of the RT-PCR product was 497 base pairs.
The sequence for the COX-1 sense primer was 5′-CATCCATCTACTCCCAGAGTCATGAG (32 to 57 bases of GeneBank S67721). The antisense sequence was 5′-GAGGGCTGGGGATAAGGTTGGACCGC (415 to 440 bases of GeneBank S67721), with the expected PCR product length of 409 base pairs. The sequence for the COX-2 sense primer was 5′-GCTGTGCTGCTCTGCGCTTGCCCTGGC (137 to 163 bases of GeneBank S67722), and the antisense sequence was 5′-GATCTGGACGTCAACACGTATCTCATG (418 to 444 bases of GeneBank S67722), with the expected product length of 308 base pairs. All of the PCR products were identified by size and sequence. Automated DNA sequencing of the PCR products was performed on a ABI Prism 377 DNA sequencer (Applied Biosystems, Foster City, CA, U.S.A.).
RESULTS
COX-3 expressed in rat brain cells except in neurons
Our RT-PCR experiments resulted in a PCR product in all rat brain cell types we examined (Fig. 1A). In the case of astrocytes, CECs, cerebral microvascular pericytes, and choroidal epithelial cells, the PCR product size matched with the expected COX-3 PCR product size, whereas in neurons, the PCR product was approximately 50 base pairs shorter. We found the highest COX-3 expression in CECs. The sequence analysis of the PCR products of astrocytes, CECs, choroidal epithelial cells, and cerebral microvascular pericytes showed 99% homology with the putative rat COX-3 mRNA. The PCR product from neurons did not match with the putative COX-3 sequence. The sequence analysis of the PCR product from neurons showed 92% homology to a yet unidentified mRNA transcribed from chromosome 4.
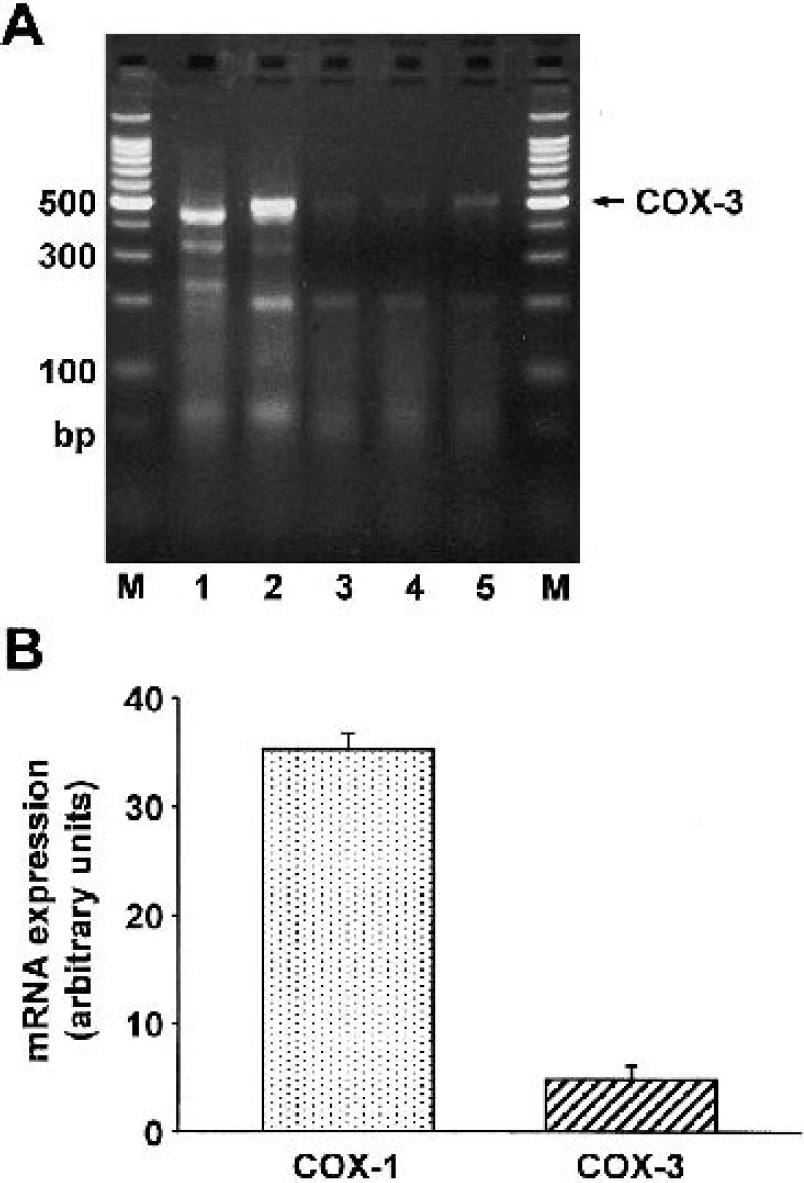
Putative COX-3 mRNA expression in cultured rat brain cells.
In CECs, we also examined the expression level of COX-3 compared with COX-1. During this experiment, we used the same cycle number during the RT-PCR for both COX-1 and COX-3 detection. In CECs, COX-3 mRNA is present at approximately 13% of the level of COX-1 mRNA (Fig. 1B). Note that we used a semiquantitative RT-PCR method; thus the resulting comparison is only approximate.
COX-3 shows constitutive expression in rat brain cells
We evaluated the expression of all three COX isoforms after LPS stimulation in CECs, astrocytes, choroidal epithelial cells, and pericytes. In CECs, LPS stimulation increased the expression of COX-2 by approximately sixfold, whereas the expression levels of COX-1 and COX-3 did not change (Fig. 2). We observed the similar expression pattern of COX isofroms in astrocytes: COX-2 expression increased (control, 100% ± 8; LPS 1 hour, 373 ± 22%∗; LPS 2 hours, 371 ± 84%∗; LPS 3 hours, 276 ± 28%∗; LPS 6 hours, 101 ± 16%; LPS 12 hours, 120 ± 19%; LPS 24 hours, 159 ± 17%; mean ± SD, n = 3, ∗ P < 0.05), whereas COX-1 (control, 100% ± 11; LPS 1 hour, 90 ± 10%; LPS 2 hours, 84 ± 10%; LPS 3 hours, 94 ± 15%; LPS 6 hours, 88 ± 11%; LPS 12 hours, 92 ± 9%; LPS 24 hours, 108 ± 14%; mean ± SD, n = 3) and COX-3 (control, 100 ± 6%; LPS 1hour, 101 ± 20%; LPS 2 hours, 96 ± 14%; LPS 3 hours, 109 ± 15%; LPS 6 hours, 95 ± 13%; LPS 12 hours, 102 ± 12%; LPS 24 hours, 111 ± 8%; mean ± SD, n = 3) expression were not influenced by LPS challenge. LPS treatment stimulated the COX-2 expression in choroidal epithelial cells (control, 100 ± 21%; LPS 1 hour, 292 ± 28%∗; LPS 3 hours, 472 ± 48%∗; LPS 6 hours, 388 ± 44%∗; LPS 12 hours, 324 ± 36%∗; LPS 24 hours, 280 ± 32%∗; mean ± SD, n = 3, ∗ P < 0.05), but it had no significant effect on COX-1 (control, 100 ± 9%; LPS 1 hour, 108 ± 11%; LPS 3 hours, 116 ± 19%; LPS 6 hours, 108 ± 14%; LPS 12 hours, 95 ± 12%; LPS 24 hours, 90 ± 25%; mean ± SD, n = 3) or COX-3 expression (control, 100 ± 14%; LPS 1 hour, 119 ± 19%; LPS 3 hours, 121 ± 16%; LPS 6 hours, 103 ± 14%; LPS 12 hours, 109 ± 12%; LPS 24 hours, 96 ± 9%; mean ± SD, n = 3). In pericytes, LPS stimulation resulted in an interesting time response curve with two peaks at 3 and 24 hrs in COX-2 expression (control, 100 ± 19%; LPS 1 hour, 283 ± 27%∗; LPS 2 hours, 362 ± 35%∗; LPS 3 hours, 384 ± 17%∗; LPS 6 hours, 169 ± 26; LPS 12 hours, 354 ± 54%∗; LPS 24 hours, 383 ± 79%∗; mean ± SD, n = 3, ∗ P < 0.05). The COX-1 and COX-3 expressions in pericytes were not influenced by LPS (COX-1: control, 100 ± 28%; LPS 1 hour, 103 ± 5%; LPS 2 hours, 95 ± 4%; LPS 3 hours, 90 ± 5%; LPS 6 hours, 80 ± 3%; LPS 12 hours, 89 ± 4%; LPS 24 hours, 96 ± 9%; mean ± SD, n = 3; COX-3: control, 100 ± 14%; LPS 1 hour, 103 ± 6%; LPS 2 hours, 114 ± 19%; LPS 3 hours, 89 ± 25%; LPS 6 hours, 94 ± 32%; LPS 12 hours, 93 ± 17%; LPS 24 hours, 115 ± 15%; mean ± SD, n = 3).
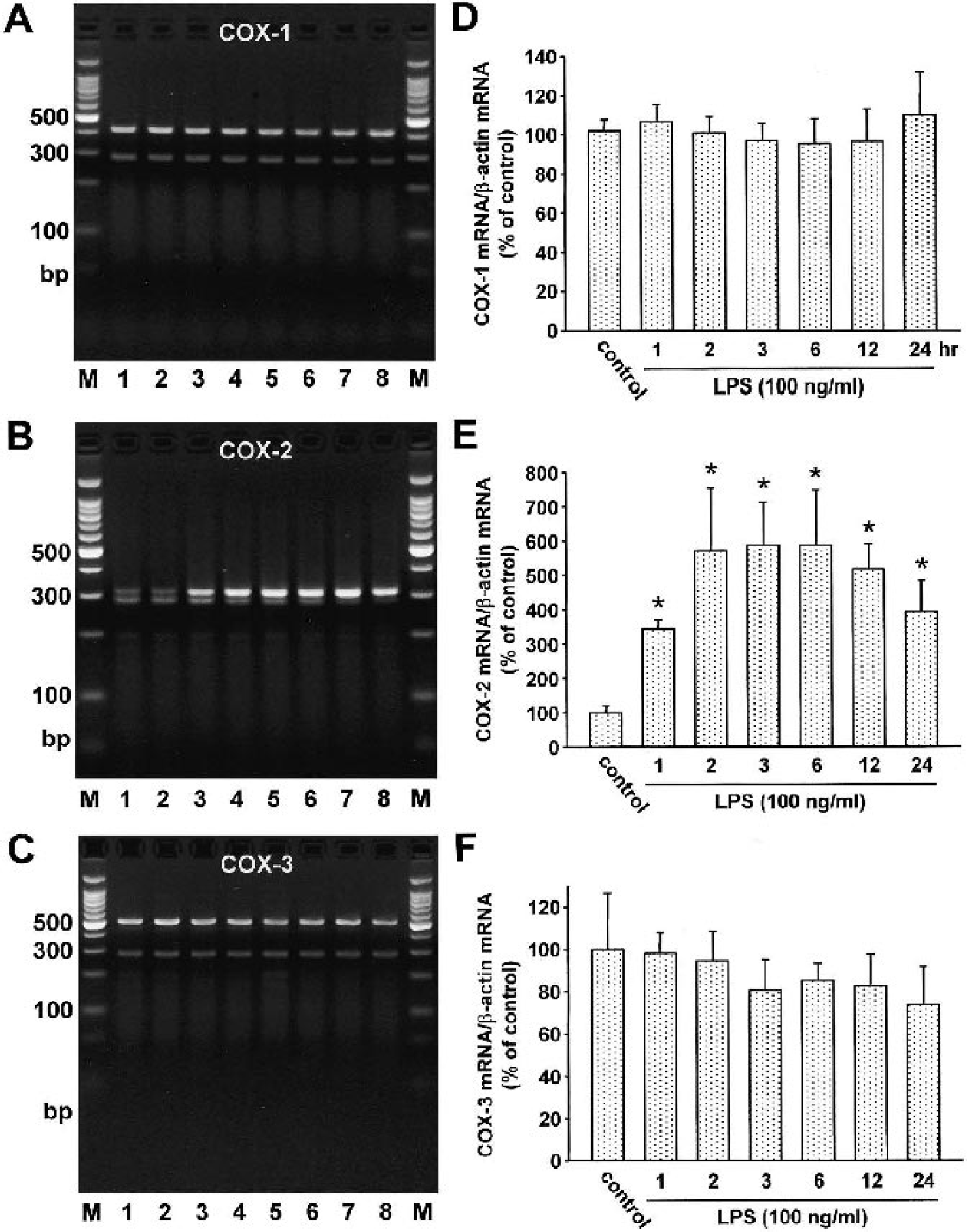
Effect of LPS stimulation on the expression of COX isoforms in cultured rat cerebral endothelial cells. Primary cultures of rat CECs were incubated with LPS (100 ng/mL) for 1, 2, 3, 6, 12, and 24 hours; total RNA was then isolated and subjected to RT-PCR analysis using specific primers detecting COX-1, COX-2, and COX-3 mRNA. β-actin primers were also included in each RT-PCR reaction. The expected molecular weights of the PCR products were 409 bp for COX-1, 308 bp for COX-2, 497 bp for COX-3, and 285 bp for β-actin. Representative gel electrophoresis of RT-PCR products for COX-1
DISCUSSION
The existence of COX-3 as a brain-specific COX isoform, which is mainly responsible for pain and fever, has already been speculated for years (Botting, 2000). Very recently, Chandrasekharan et al. (2002) successfully isolated and cloned COX-3 from the dog. The canine COX-3 is identical to the full-length form of COX-1, with the exception that it retains intron 1. This new isoform is most abundant in the cerebral cortex and shows higher sensitivity towards acetaminophen than COX-1 and COX-2. It is speculated that the retention of intron 1 might change the way the enzyme molecule is folded or the conformation of its active site, which can explain its different enzymatic properties. The discovery of COX-3 provides a possible explanation for the efficacy of acetaminophen (paracetamol, Tylenol) in treating pain and fever. Acetaminophen is generally categorized as a nonsteroidal antiinflammatory drug (NSAID) because of its potent analgesic and antipyretic actions; however, it lacks the other typical actions of NSAIDs, such as antiinflammatory and antithrombotic activity and gastrotoxicity. Acetaminophen administered into the cerebral ventricles of conscious cats reduced fever (Feldberg et al., 1973), and the analgesia induced by acetaminophen can be attributed to an action in the central nervous system (Alloui et al., 2002). The work of Flower and Vane (1972), who showed that brain COX is more sensitive to inhibition by acetaminophen than spleen COX, also supports the general consideration that acetaminophen acts centrally.
We hypothesized that COX-3 is also expressed in the rat, and, according to the known sequence of the rat COX-1 gene, we designed a primer set to detect the rat COX-3. The sense primer was targeted against the sequence of intron 1, which is part of the COX-3 mRNA, and the antisense primer binds to the exon 5 of the COX-1 gene. This primer set selectively amplifies only the COX-3 but not the COX-1 mRNA. We carried out our RT-PCR experiments in primary cultures of rat brain cells because Chandrasekharan et al. (2002) found the highest expression in the cerebral cortex. However, they did not examine specific cell types expressing COX-3 mRNA. Thus our primary cultures gave us an opportunity to selectively study the expression of COX-3 in different cell types of the brain.
Our results demonstrate that the mRNA of the putative COX-3 is expressed in rat brain cells, except in neurons. Among the cells we studied, the CECs showed the highest COX-3 expression. Whereas COX-2 expression increased several-fold after LPS challenge, COX-3 expression did not change significantly, suggesting that cells constitutively express COX-3 similarly to COX-1. However, it is possible that COX-3 mRNA levels are able to increase under other various conditions.
In basal cell culture conditions, we detected COX-2 expression in all cell types. These results are in accordance with others that showed constitutive COX-2 expressions in neurons (Schiltz and Sawchenko, 2002), astrocytes (Hirst et al., 1999), choroidal epithelial cells (Maslinska et al., 1999), and CECs (Parfenova et al., 1997). On the other hand, the COX-2 levels that we detected in astrocytes, choroidal epithelial cells, CECs, and pericytes may be artificially higher than those detected in vivo because of the stimulating effect of the culture medium (Luo et al., 1998). We do not know any study that shows constitutive COX-2 expression in pericytes. Pericytes may also have constitutive COX-2 expression like other cells of the brain. However, it is possible that the basal COX-2 expression in pericytes in our experimental condition is caused by the stimulating effect of the culture medium.
At this time, we do not know whether the rat COX-3 mRNA codes a functional enzyme, such as has been reported in the dog. Retention of intron 1, which is 98 base pairs in rat, would be expected to shift the remaining COX-3 sequence out of frame with respect to the open reading frame of the COX-1 and result in a completely different protein. However, it is possible but untested that factors such as a different initiation site or alternative downstream splicing may restore the reading frame of the proposed COX-3 protein and result in a fully functioning, acetaminophen-sensitive COX-1 variant. It is also possible that the COX-3 mRNA, although not coding for a functioning COX variant, may be involved in modulation of the translation of mRNA for COX-1, such as has been reported to occur with microRNAs (Lagos-Quintana et al., 2001).
The physiologic role of a putative acetaminophen-sensitive COX-3 is unclear. On the surface, a COX-3 variant that is enriched in CECs may account for the well-described antifever effects of acetaminophen. CECs are a central player in mediating the fever response from the periphery to the brain via increased prostaglandin synthesis (Matsumura et al., 1998), and we found the highest COX-3 expression in these cells. However, the deletion of the COX-2 but not COX-1 gene, which also encodes COX-3, in mice blunts pyresis (Li et al., 1999), which argues against this view. In addition, LPS treatment did not increase COX-3 expression in CECs or astrocytes. Schwab et al. (2003) assume that the COX-3 discovery might be a clue to the analgesic effect of acetaminophen. However, our finding showing the lack of COX-3 expression in neurons makes this hypothesis implausible. Nonetheless, the COX-3 isoform has only recently been described, and insufficient time has passed to fully explore its physiologic role.
We believe that there are several aspects of our findings that suggest that COX-3 may be functionally important. First, COX-3 mRNA is prevalent in CECs and represents levels greater than 10% of those for COX-1 mRNA. We are unaware of other abundant mRNA variants that fail to play a role in cell functioning or yield functional proteins. Second, there is a cell-specific regulation of COX-3 mRNA, such that there is differential expression in various cell types from brain. Thus the CECs show a high level of expression for COX-3 mRNA, whereas we were unable to detect expression in neurons despite the normal expression of COX-1 mRNA. Other cell types from brain, such as astrocytes, pericytes, and choroidal epithelial cells, showed detectable but lower levels of COX-3 mRNA than CECs. Third, COX-3 joins an increasing group of COX isoforms. The different selectivity of these isoforms to various pharmacologic agents allows independent modulatory function.
In summary, we report here for the first time to our knowledge that the putative COX-3 mRNA is detectable in rats and is differentially expressed in various cell types from rat brain, as well as that its expression is not stimulated by LPS.
