Abstract
To understand the molecular processes of continuous vasospasm of cerebral arteries after subarachnoid hemorrhage, mRNA differential display and screening of cDNA expression array were performed to identify genes that are differentially expressed in vasospastic arteries of canine two-hemorrhage models. The expression levels of 18 genes were found to be upregulated, and those of two genes to be down-regulated. Of these, 12 represent known genes or homologues of genes characterized previously, and the other eight genes are not related to any sequences in the databases. The known genes include five upregulated inflammation-related genes encoding monocyte chemotactic protein-1, cystatin B, inter-α-trypsin inhibitor family heavy chain-related protein, serum amyloid A protein, and glycoprotein 130, suggesting that inflammatory reaction may be involved in the development of cerebral vasospasm. The upregulation of three known genes encoding stress-related proteins of vascular endothelial growth factor, BiP protein, and growth-arrest and DNA-damage–inducible protein may be involved in possible cell survival in the damaged arteries. A full-length cDNA for the unknown clone DVS 27, whose expression was most highly upregulated, was isolated from the cerebral artery cDNA library by hybridization. Characterization of these genes should help to clarify the molecular mechanism of continuous cerebral vasospasm after subarachnoid hemorrhage.
Cerebral vasospasm after subarachnoid hemorrhage, which is the most serious complication causing cerebral ischemia in patients, is a complex pathologic process characterized by continuous contraction of arterial smooth muscle for several weeks and morphologic changes in the arterial wall such as cell proliferation and necrosis. Several molecular biological techniques have been applied in the studies of cerebral vasospasm. After exposure to periarterial blood, arterial walls were found to increase the synthesis of procollagen types I and III, possibly in response to the increased levels of transforming growth factor-β (Kasuya et al., 1993). Expression of soluble guanylate cyclase was shown to be diminished in basilar arteries in the canine two-hemorrhage model (Kasuya et al., 1995). The levels of endothelin B receptor mRNA expression were shown to increase in vasospastic arteries in monkeys after subarachnoid hemorrhage (Hino et al., 1996). However, most of the previous studies focused on some known genes encoding related factors such as vasoconstriction agonists or antagonists.
A single cell is estimated to express approximately 15,000 genes, several hundred of which are thought to play some tissue-specific roles in the cell. However, because most of them are unknown and have not been characterized, it is important to identify the unknown genes which could be involved in the pathogenesis of cerebral vasospasm. In this study, as a first step to better understand the whole molecular process of cerebral vasospasm, the two molecular biological techniques of cDNA expression array and mRNA differential display (Liang and Pardee, 1992) were applied to identify differentially expressed genes during cerebral vasospasm.
MATERIALS AND METHODS
Cerebral vasospasm models
The experimental protocol used for animals was evaluated and approved by the Institutional Animal Care and Use Committee of Tokyo Women's Medical University. Care of the animals and surgical procedures were performed according to the standards of Tokyo Women's Medical University Protocol on Laboratory Animals.
Twenty-one mongrel dogs weighing between 12 and 23 kg were anesthetized with intravenous sodium pentobarbital (25 mg/kg). In 13 of the dogs used, the cisterna magna was punctured percutaneously, and 0.3 mL/kg of CSF was removed by spontaneous egress. Subsequently, 0.5 mL/kg of fresh autologous arterial nonheparinized blood was injected into the cisterna magna at a rate of 2 mL/minute. The dogs were tilted with tail up for 15 minutes. The catheter then was removed, and the animals were allowed to recover.
For the cDNA expression array study, 6 of the 21 dogs were randomly selected and assigned to the two groups, D-0 and D-7, of three each. For mRNA differential display, the other 15 dogs were divided into three groups with five animals in each group. Group D-0 comprised control animals, killed on day 0 without cisternal blood injection. Group D-2 animals were killed on day 2 after the first injection on day O. Group D-7 animals were killed on day 7 after injections on days 0 and 2. Anigiography of cerebral arteries of each animal was performed on days 0, 2, and 7, and diameters of the basilar arteries were measured on angiographic film for evaluation of vasospasm. Statistical comparisons of arterial diameters between groups were evaluated by analysis of variance and Scheffe's for pairwise comparisons.
RNA preparation
The animals were killed by injection of 100 mg/kg pentobarbital, exsanguinated, and perfused with 1,500 to 2,000 mL normal saline. Total RNAs were extracted from individual basilar arteries from each group using TRIzol (GIBCO BRL, Gaithersburg, MD, U.S.A.) according to the manufacturer's instruction. Possible traces of genomic DNA contaminating RNA preparations were removed by DNase I (Promega, Madison, WI, U.S.A.) digestion.
Total RNA also was extracted from middle cerebral artery, brain, muscle, heart, lung, liver, kidney, small intestine, and testis tissue for the analysis of tissue distribution of mRNA.
cDNA expression array
Fifty μg of total RNA prepared from D-0 and D-7 groups was purified to poly (A) RNA. The cDNA probes were synthesized from the poly (A) RNA in the presence of [α−32P] dATP (3,000 Ci/mmol; Amersham, Arlington, IL, U.S.A.) using random primers and M-MLV reverse transcriptase (Clontech, Palo Alto, CA, U.S.A.). The two Atlas Human cDNA Expression Array membranes (Clontech) were prehybridized at 42°C for 30 minutes in solution containing 50% formamide, 5 × standard sodium phosphate ethylenediamine tetraacetic acid (SSPE), 10× Denhardt's solution, 1.4% sodium dodecyl sulfate (SDS), and 0.1 mg/mL salmon testes DNA. The two membranes then were separately hybridized at 42°C for 16 hours in the same solution with the 32P-labeled cDNA probes prepared from D-0 and D-7 samples. The membranes were washed at room temperature with 0.1 × standard saline citrate and 0.1% SDS and then at 60×C for 30 minutes, and exposed to x-ray film for 3 days with an intensifying screen. After confirming equal intensities of hybridization signals for several housekeeping genes between autoradiograms for D-0 and D-7, differentially expressed genes were evaluated by comparing intensities of specific hybridizing signals.
mRNA differential display
The cDNA was synthesized from 1 μg total RNA using one of five oligo (dT) primers T12AG, T12CG, T12GG, T12GC, and T12AC. The cDNA from at least four individual basilar arteries from each group was subjected to the same experiment to confirm reproducibility. The oligo (dT) primer was end-labeled with [γ−32P] ATP (6,000 Ci/mmol). One twentieth of the cDNA generated was amplified by polymerase chain reaction (PCR) in the presence of labeled oligo (dT) primer and 1 of 21 upstream arbitrary primers. One hundred five primer pairs were used to screen mRNA from groups D-0, D-2, and D-7. The PCR conditions used were 25 cycles of denaturation at 94°C for 30 seconds, annealing at 40°C for 1 minute, and extension at 72°C for 30 seconds. The radiolabeled PCR products were separated on 5% or 6% denaturing polyacrylamide gels, and the dried gels were exposed to x-ray film for 12 to 16 hours.
After autoradiography, the intensity of each band was compared among groups D-0, D-2, and D-7. Differentially expressed bands were excised from the gels, reamplified by PCR using the same set of primers, and subcloned into pGEM-3Z (Promega). The cDNA inserts then were sequenced using an Applied Biosystem DNA Sequencer model 377 with a Taq DyeDeoxy Termination Cycle Sequencing Kit (Perkin Elmer, Norwalk, CT, U.S.A.).
Reverse transcription-polymerase chain reaction
Total RNA (1 μg) from arteries of groups D-0, D-2, and D-7 or from several tissues were reverse-transcribed using oligo (dT) primer. One-twentieth of the cDNA synthesized was amplified by PCR in the presence of 5 μCi of [α−32P] dCTP (6,000 Ci/mmol) and a pair of specific primers. The PCR conditions used were as follows: 18 to 31 cycles of denaturation at 94°C for 45 seconds; annealing at 58°, 60°, or 62°C for 45 seconds; and extension at 72°C for 45 seconds. The PCR products were separated on a 6% nondenaturing polyacrylamide gel, and the dried gel was exposed to x-ray films for 3 to 6 hours. The relative ratio of expression patterns of mRNA was evaluated by direct measurement of radioactivity in each band using a scintillation counter.
The dog sequences corresponding to the human genes demonstrated in cDNA expression array were determined. The cDNA prepared from dog middle cerebral arteries were amplified by PCR using the degenerate oligo primers, which were designed based on the human and mouse nucleotide sequences retrieved from databases and directly sequenced as described earlier. The dog sequences, which represented over 90% identity to the human sequences, subsequently were used to design specific PCR primers.
To evaluate the expression levels of the DVS 27 mRNA, quantitative reverse transcription (RT)-PCR was performed with real time TaqMan technology (Holland et al., 1991) using a Sequence Detection System model 7700 (Perkin Elmer) according to the manufacturer's instructions. The PCR primers used were a sense primer of 5′-ATGAATCAGGTGACGGT-GTTG-3′ and an antisense primer of 5′ -CTCCAGGAT-CAGTCTTGCATT-3′. The detection probe labeled with fluorescence was 5′-ATGCCAACAACAAGGAACACTCTGTG-3′. The human nucleotide sequence used to design the oligo primers and probe was obtained from the expressed sequence tag (accession no. AB024518), which encodes a possible human counterpart of canine DVS 27.
Additional 5′ -upstream cDNA sequences
The dog cerebral artery cDNA library in λZAP II (Stratagene, LaJolla, CA, U.S.A.) was constructed using mRNA from middle cerebral arteries according to the manufacturer's instructions. To obtain more 5'-upstream sequence, the cDNA library was directly amplified by PCR using the vector primer and antisense primer specific to each cDNA sequence. The PCR products were subcloned and sequenced as described earlier. To isolate the full-length cDNA for DVS 27, the cDNA library was screened by hybridization with the 32P-labeled cDNA fragments according to the standard method. Hybridization conditions were 50% formamide, 5 × SSPE, 2 × Denhardt's solution, 0.1% SDS, 0.1 mg/mL salmon testes DNA, and 1 × 106 cpm/mL of 32P-labeled probe at 42°C for 16 to 20 hours; washing conditions were −0.1 × standard saline citrate and 0.1% SDS at room temperature for 1 hour and then at 60°C for 30 minutes. The cDNA segments in the λ clones obtained were subcloned and sequenced.
Cell culture and immunohistochemistry
The kidney-derived COS-7 cells were cultured in Dulbecco's modified Eagle's medium (GIBCO BRL) containing 10% fetal bovine serum at 37°C under a humidified condition of 95% and 5% CO2. The cells were transiently transfected with pCMV-DVS-27 by electropolation and grown on the chamber slide containing appropriate media. After a 48-hour culture, cells were fixed with 3% formamide, washed with phosphate-buffered saline, and incubated in the mouse anti-HA primary antibody (Boehringer Mannheim, Mannheim, Germany) at 37°C for 30 minutes. After washing with phosphate-buffered saline, the cells were incubated in the secondary antibody of fluorescein isothiocyanate-conjugated anti-mouse IgG (Jackson ImmunoResearch Laboratories, West Grove, PA, U.S.A.) at 37°C for 30 minutes. Localization of proteins within the cells was analyzed using fluorescence microscopic methods.
Human umbilical artery smooth muscle cells (HUASMC) were cultured in Dulbecco's modified Eagle's medium containing 10% fetal bovine serum and human umbilical vein endothelial cells (HUVEC) in Humedia EB-2 (Kurabo, Osaka, Japan) supplemented with 20% human serum. The cells, passage 3 or 4, were incubated in serum-free medium, which was followed by treatments with inflammation-related stimuli, including 10 μg/mL lipopolysaccharide, 10 ng/mL tumor necrosis factor-α, 10 ng/mL interleukin (IL)-1α, 10 ng/mL IL-1β, 10 ng/mL interferon-γ for 8 hours, and 100 ng/mL IL-6 for 16 hours. Total RNA was extracted from the cells for quantitative RT-PCR.
RESULTS
Cerebral vasospasm model
Figure 1 shows the change in vessel caliber of basilar arteries after cisternal injection of blood. A significant reduction in vessel caliber was recognized on angiograms in the D-2 and D-7 groups compared with D-0 baseline (P = 0.01 and P < 0.0001, respectively). Arterial spasm in D-7 group was significantly more intense than that in D-2 (P = 0.001).
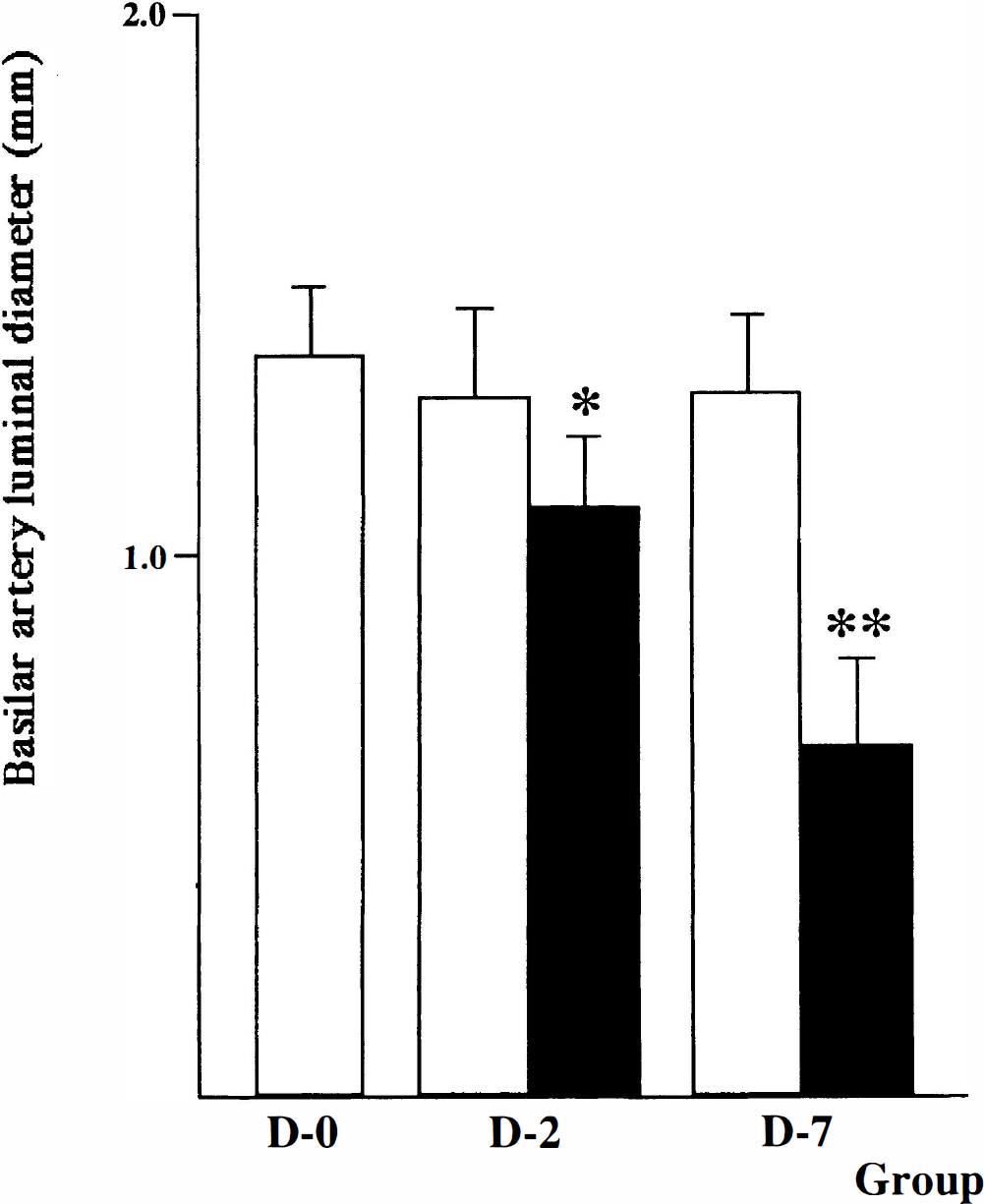
Basilar artery luminal diameter on angiograms in group D-0, D-2, and D-7 dogs. A significant reduction in vessel caliber was observed in angiograms between the D-2 and D-7 groups and D-0 baseline (*P = 0.01 and **P < 0.0001, respectively). In the D-7 group, arterial spasm was significantly more intense than in the D-2 group (P = 0.001). All values were given as mean ± SD (eight dogs per groups D-0 and D-7 and five dogs per group D-2). Open bar, baseline; closed bar, vasospasm.
Differentially expressed mRNA identified
The cDNA Atlas Array examined includes 588 human known cDNA. High stringent hybridization indicated that 49 (8.3%) of them were found to be expressed in dog basilar arteries; a representative hybridization pattern is shown in Fig. 2a. The dots of six housekeeping genes for ubiquitin, β-actin, glyceraldehyde-3-phosphate dehydrogenase, α-tubulin, 23-kDa highly basic protein, and ribosomal protein S9 generated equal intensities of hybridizing signals in both membranes for D-0 and D-7 groups, which could normalize mRNA abundance between samples. Of 49 genes, 4 were upregulated in the basilar arteries with vasospasm: the genes for gadd45 (growth arrest and DNA-damage-inducible protein 45), monocyte chemotactic protein-1, neuromodulin, and vascular endothelial growth factor (VEGF). Their differential expression patterns were confirmed by RT-PCR (Fig. 3A); the dog-specific primers and PCR conditions used are shown in Table 1 (Accession No. AU052021-AU052024).
Nucleotide sequences of dog-specific primers for RT-PCR and PCR conditions

bp, base pairs; MCP, monocyte chemotactic protein; VEGF, vascular endothelial growth factor.
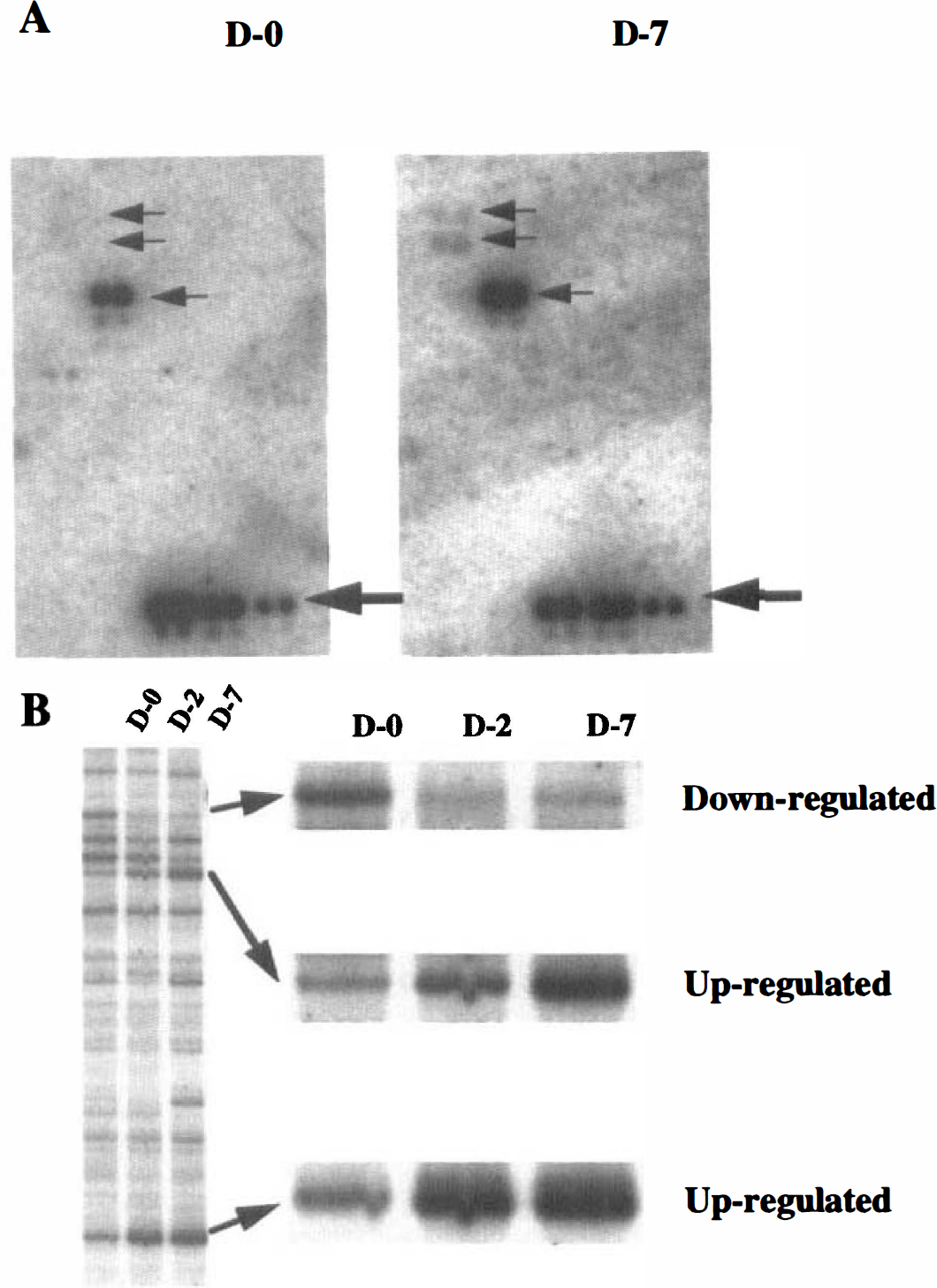
Identification of differentially expressed mRNA.
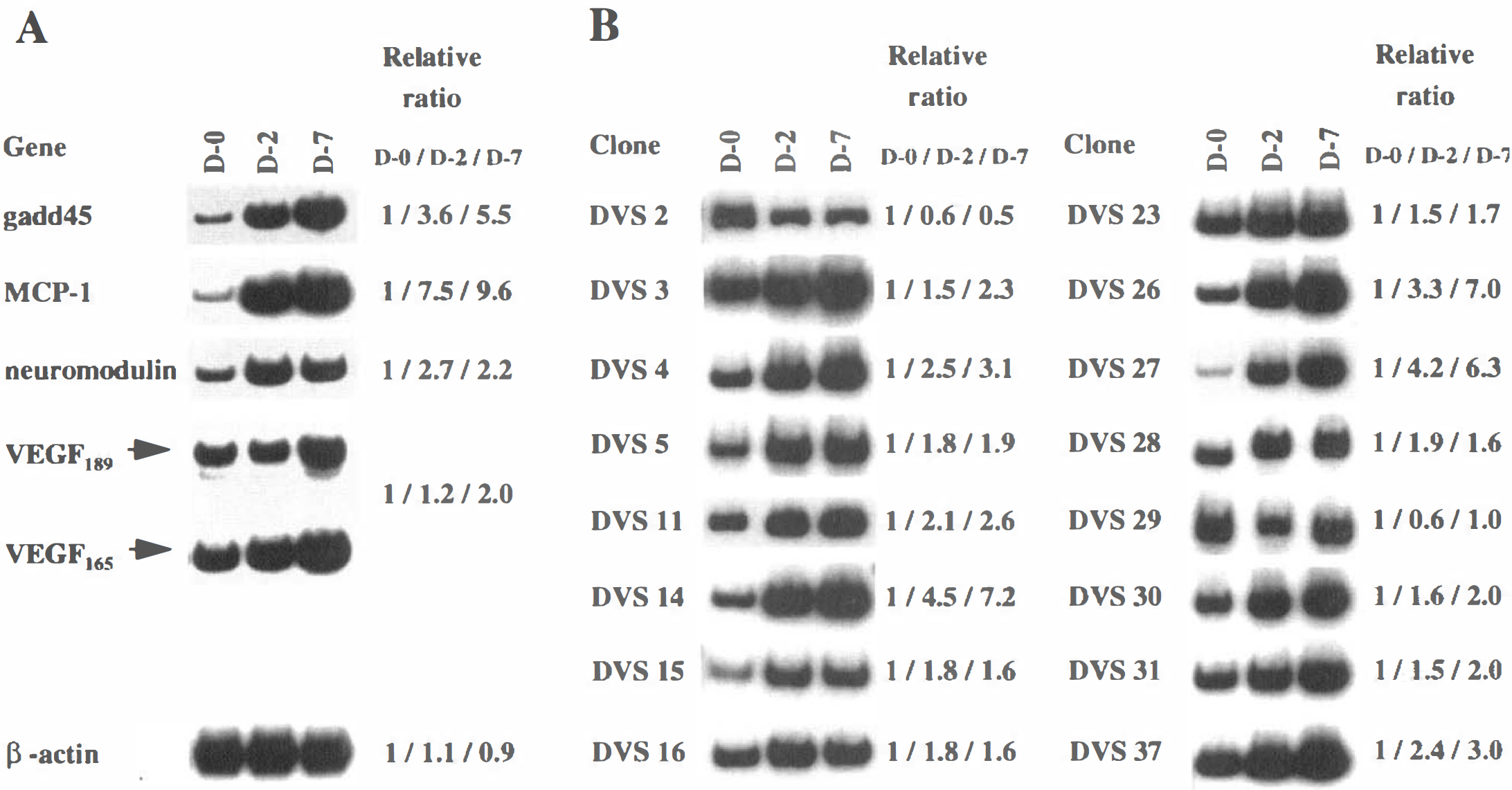
Forty differentially expressed bands in groups D-2 and D-7 were identified by mRNA differential display; a representative pattern of display is shown in Fig. 2B. The bands, designated DVS 1 through 40, were sequenced, generating 26 nonredundant sequences. Also, RT-PCR confirmed the differentially expressed patterns of 16 of the 26 clones obtained (Fig. 3B); the PCR primer pairs used are shown in Table 1 (accession no. AU052006-AU052020). Three clones (DVS 14, 26, and 27) were more highly upregulated in group D-7 than in group D-2, and in group D-2 than in group D-0; six clones (DVS 3, 4, 11, 30, 31, and 37) were more mildly upregulated. Five clones (DVS 5, 15, 16, 23, and 28) were upregulated in groups D-2 and D-7 equally; one clone (DVS 2) was downregulated in groups D-2 and D-7, and one clone (DVS 2 9) was downregulated in only group D-2.
Database analysis
The sequences of 16 clones obtained by mRNA differential display were compared with those in the nucleotide databases using the BLAST network service
(Altschul et al., 1990). The sequences of six clones were highly homologous to those of human known genes, whereas the other 10 clones were not related to any known sequences in the databases (Table 2). The DVS 3, 4, 11, 14, 26 and 30 clones had 90%, 82%, 84%, 64%, 85%, and 95% identity to human genes for glycoprotein 130 (gp130), inter-α-trypsin inhibitor family heavy chain-related protein (IHRP), protein disulfide isomerase-related protein P5 (PDI-RP P5), acid sphingomyelinase-like phosphodiesterase, serum amyloid A protein, and BiP protein, respectively.
Summary of differentially expressed clones identified by mRNA differential display
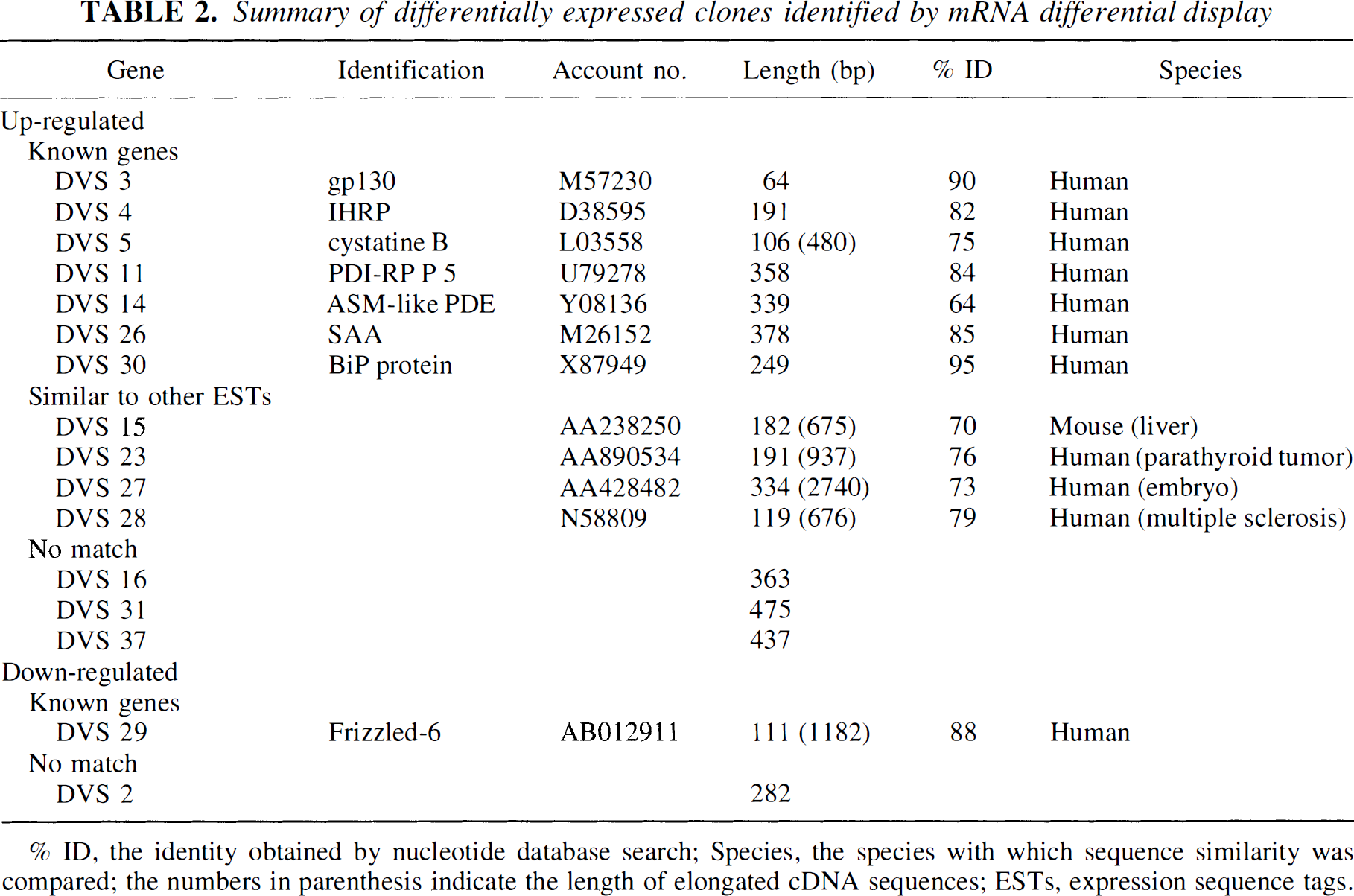
% ID, the identity obtained by nucleotide database search; Species, the species with which sequence similarity was compared; the numbers in parenthesis indicate the length of elongated cDNA sequences; ESTs, expression sequence tags.
Because the partial sequences obtained ranged from 100 to 500 base pairs (bp), it was necessary to obtain the upstream coding sequence to predict the role of proteins encoded. The six additional sequences for unknown genes (DVS 5, 15, 23, 27, 28, and 29) were obtained by PCR amplification of a dog cerebral artery cDNA library and compared with those in the nucleotide and peptide databases using BLASTN and BLASTX programs, respectively. Of these 5′ -sequences, those for DVS 5 and 29 showed 75% and 88% identity to human mRNA for cystatin B and Frizzled-6, respectively. The other four sequences were not related to any known genes in the databases (Table 2).
Tissue distribution of differentially expressed mRNA
Tissue distribution of eight unknown mRNA obtained was examined by RT-PCR. Although they were widely expressed, the clones DVS 2, 23, 27, and 37 showed relatively high expression in cerebral arteries (Fig. 4).
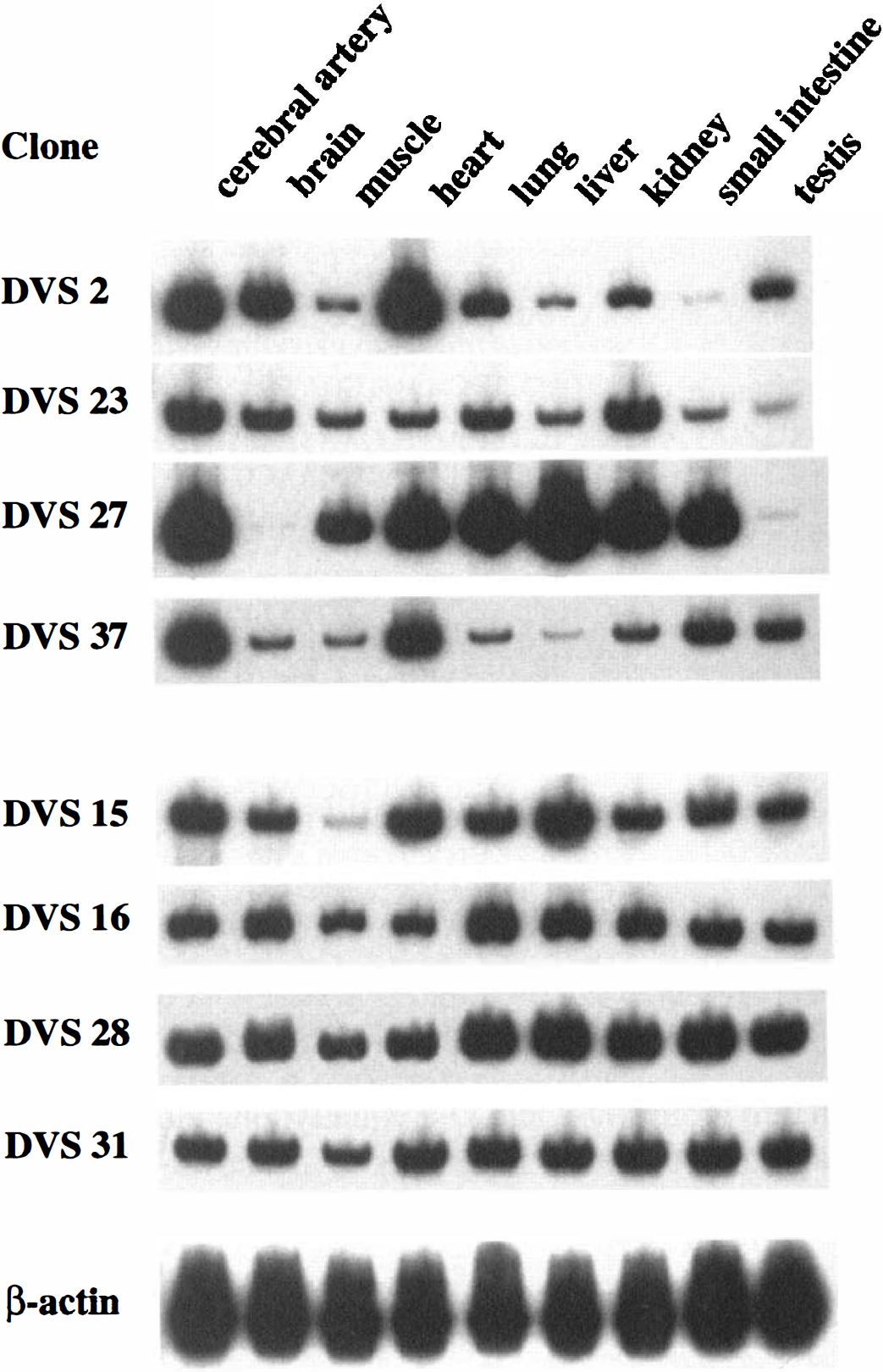
Tissue distribution of eight unknown mRNA. Reverse transcription-PCR was performed using RNA prepared from cerebral artery, brain, skeletal muscle, heart, lung, liver, kidney, small intestine, and testis. Four clones, DVS 2, 23, 27, and 37, were relatively highly expressed in cerebral artery, and the other four clones, DVS 15, 16, 28, and 31, were equally expressed in tissues examined. The RNA loading was normalized by the expression levels of β-actin mRNA.
Isolation and characterization of a full-length cDNA for DVS 27
The entire cDNA sequence for the DVS 27, whose expression was most highly increased among the unknown clones, was obtained by screening of the cDNA library. One hundred three positive clones were isolated, of which four clones, ADVS 27-17, λ DVS 27-18, λ DVS 27–37, and λ DVS 27–39, were sequenced. The composite nucleotide sequence of 2,740 bp encoded a protein of 263 amino acids (molecular weight = 30,182) (accession no. AB024517) (Fig. 5). Northern blot analysis using liver RNA showed a 2.7-kb band, indicating that the cDNA obtained is full length. The search with the open reading frame sequence using programs of BLASTX at NCBI and MOTIF at http://www.motif.genome.ad.jp showed no significant match in peptide databases. To predict the role of the unknown protein encoded, immunohistochemical analysis was performed. The nucleotide sequence encoding the HA epitope of YPYDVPDYA was introduced in frame at the 3′ -end in the expression vector pCMV6b to generate pCMV-DVS-27. The COS-7 cells were transiently transfected with pCMV-DVS-27, and localization of the protein within the cells was immunohistochemically analyzed with anti-HA antibody using fluorescence microscopic study. Strong signals for the protein were observed predominantly around the perinuclear membrane on the inside face of the nucleus, suggesting that the DVS 27 gene encodes a nuclear protein (Fig. 6).

Composite nucleotide sequence of DVS 27 cDNA and predicted amino acid sequence of the protein. The amino acid sequence is shown in single-fetter code below the nucleotide sequence. Nucleotide and amino acid numbers are shown at the right end.
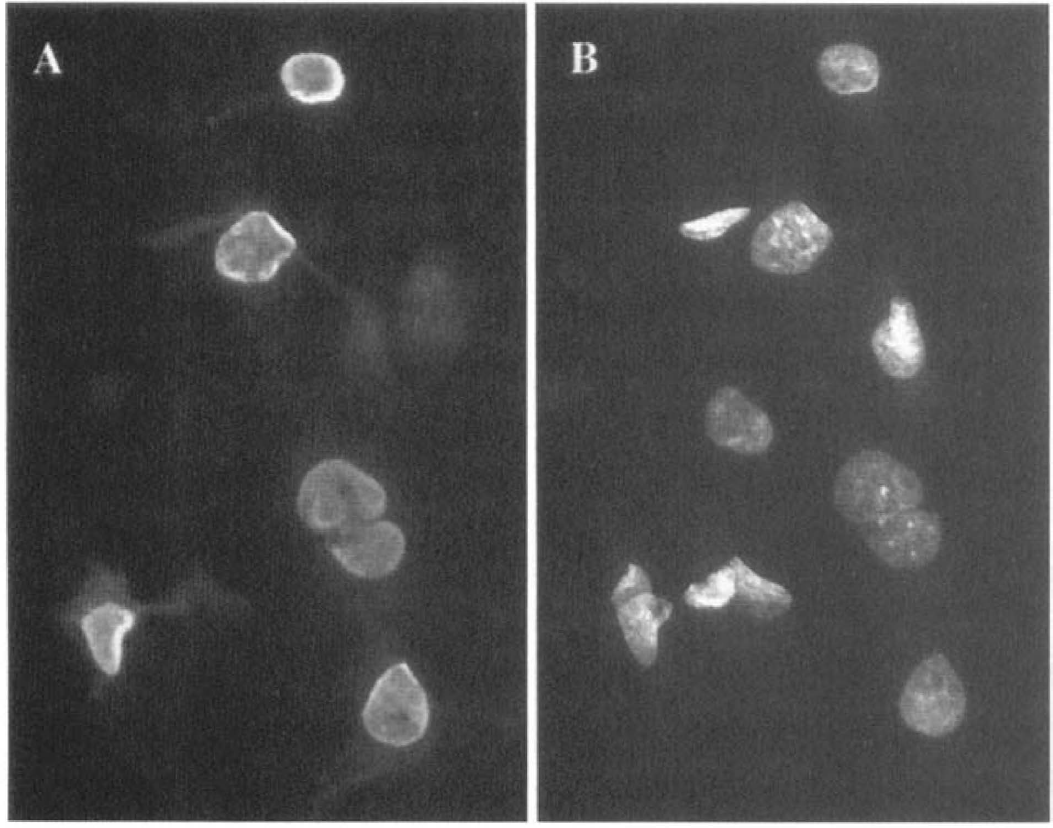
Intracellular localization of the protein encoded by DVS 27 mANA. The COS-7 cells were transfected with pCMV-DVS-27. Localization of proteins within the cells was analyzed using fluorescence microscopic study. Nuclear localization of the protein
To elucidate whether DVS 27 expression is associated with inflammatory reactions, we examined the level of DVS 27 mRNA in the presence of various inflammation-related stimuli in HUASMC and HUVEC cells. Quantitative RT-PCR using real time TaqMan technology showed that the expression levels of the DVS 27 mRNA in HUASMC cells were highly increased in response to IL-1α and IL-1β, and mildly increased in response to interferon (IFN)-γ (Fig. 7). Lipopolysaccharide, tumor necrosis factor (TNF)-α, and IL-6 did not significantly change the expression levels of the DVS 27 mRNA in the cells. In contrast, none of these stimuli has affected the expression levels of DVS 27 mRNA in HUVEC cells (data not shown), suggesting that the functional properties of the protein encoded by DVS 27 in smooth muscle cells may be different from those in endothelial cells in the vessels.
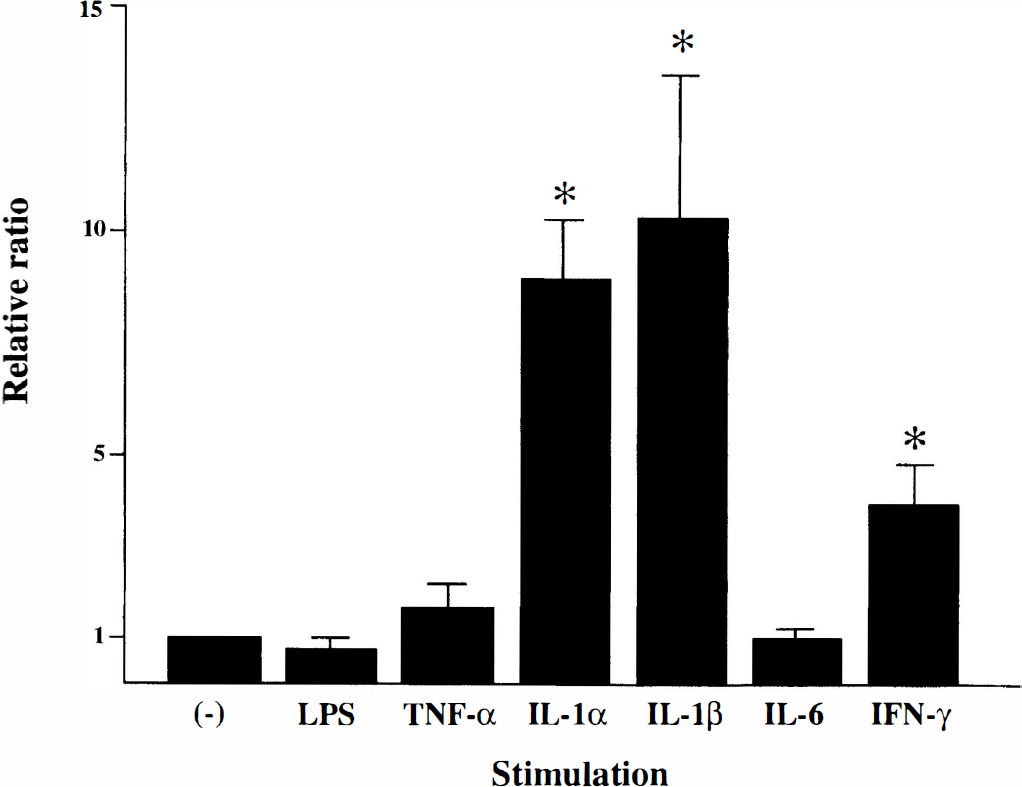
Expression levels of DVS 27 mANA in human umbilical artery smooth muscle cells (HUASMC) treated with inflammation-related stimuli. The expression levels of DVS 27 mANA were strongly upregulated by addition of interleukin (IL)-1α and IL-β, and mildly by interferon-γ (IFN-γ). Lipopolysaccharide (LPS), tumor necrosis factor-α (TNF-α), and IL-6 did not increase the expression of DVS 27 mANA in the cells. The expression levels were estimated by quantitative AT-PCA using real time TaqMan technology. The standard curve for relative quantity was constructed using the levels of glyceraldehyde-3-phosphate dehydrogenase (GAPDH) mANA as an internal control. Serial five twofold dilutions of each cDNA sample were analyzed, and all values were given as mean ± SD.* P < 0.0001, unpaired t test.
DISCUSSION
In this study, 12 genes differentially expressed during cerebral vasospasm were known. They can be classified into three groups according to their function: the genes for molecules associated with inflammation, genes whose expression can be induced by stress conditions such as hypoxia and hypoglycemia, and genes apparently not related to cerebral vasospasm.
Molecules associated with inflammation
Five genes encoded monocyte chemotactic protein (MCP)-1, cystatin B, IHRP, serum amyloid A protein, and gp130, which were closely related to inflammatory events.
Monocyte chemotactic protein-1 is a chemotactic factor specific for monocytes and is produced in monocytes, vascular endothelial cells, smooth muscle cells, and fibroblasts by various stimuli such as lipopolysaccharide, IL-1, TNF-α, and IFN-γ (Leonard and Yoshimura, 1990; Colotta et al, 1992; Rollins et al., 1990; Shyy et al., 1990, Sica et al., 1990). In the current study, the expression of MCP-1 was strongly upregulated, suggesting that chemotaxis of monocytes is activated and that infiltrated macrophages play an important role in the development of cerebral vasospasm. This is supported by a previous observation that macrophages are abundant in the subarachnoid space, especially around the main cerebral arteries in rats that survived for 2 to 5 days after subarachnoid hemorrhage (Kubota et al., 1993). Cystatin B is one of the cysteine protease inhibitors and functions as a protector against damage by lysosomal proteinase released into the cytoplasm. The cystatin B levels have been shown to be markedly increased in sera from patients with inflammatory diseases (Hopsu-Havu et al., 1982). When activated by silicon dioxide, the cultured macrophages were shown to excrete cystatin B into the media (Hapsu-Havu et al., 1984). Thus, higher expression
levels of cystatin B in vasospastic arteries than in normal arteries suggest that inflammatory reactions related to the activated macrophages are involved in the pathogenesis of cerebral vasospasm. Also known as plasma kallikrein-120, IHRP is a member of the family of serine protease inhibitors (Saguchi et al., 1996). High levels of IHRP expression suggest that the acute-phase proteins also might be highly synthesized and excreted by cells such as monocytes in response to inflammatory stimuli. The acute-phase response cascade usually is activated by a broad spectrum of inflammatory mediators, such as IL-1, IL-6, and tumor necrosis factor-α, released by macrophages (Baumann and Gauldie, 1994). These cytokines have been shown to subsequently induce expression of mRNA for serum amyloid A protein, one of the major acute phase proteins (Ramadori et al., 1988; Sellar et al., 1991). An interleukin-6 receptor complex (IL-6R) is constituted by gp130; IL-6R and gpl30 are ligand-binding and nonligand-binding but signal-transducing chains, respectively. A ligand-bound form of IL-6R can interact with gp130 extracellularly to transfer IL-6 signals into the cytoplasm (Taga et al., 1989). Because Saito et ai. (1992) reported that gp 130 and IL-6R mRNA in murine liver were upregulated in vivo by IL-6 stimulation, the regulatory system may be mediated by IL-6 and IL-6R, and gp130 also may be activated in a similar manner in cerebral vasospasm.
Cerebral vasospasm can be inhibited by some antiinflammatory reagents in canine cerebral vasospasm models (Chyatte et al., 1983). The expression of intercellular adhesion molecule-1, a ligand for the leukocyte adhesion receptor, increased in the endothelial and medial layer of the cerebral arteries after subarachnoid hemorrhage in rats, and monoclonal antibodies against intercellular adhesion molecule-1 attenuated cerebral vasospasm after experimental subarachnoid hemorrhage in rabbits (Bavbek et al., 1998; Handa et al., 1995). Ono et al. (1998) report the successful inhibition of cerebral vasospasm in the rabbit subarachnoid hemorrhage model using decoy synthetic DNA fragments with a high affinity to nuclear factor-κB, a transcription factor activated in inflammatory responses. The current results are consistent with these observations that inflammatory reactions are closely related to the development of cerebral vasospasm, suggesting that the functional significance of other known chemokines and their receptors also should be assessed.
Stress-induced molecules
The expression of three upregulated genes for VEGF, BiP protein, and gadd45 is known to be induced under various stress conditions. It is known that VEGF is a potent and highly specific mitogen for vascular endothelial cells. A hypoxic condition strongly induces the expression of VEGF mRNA in a variety of cell types, including myocyte, glioma cells, fibroblasts, and skeletal muscle myoblasts (Ladoux and Frelin, 1993; Shweiki et al., 1992). Also, VEGF is upregulated in response to hypoglycemic stress in C6 cells, a clonal glial cell line derived from a rat glial tumor (Shweiki et al., 1995), and in a human monocytic cell line (Satake et al., 1998). The BiP protein, also known as glucose-regulated protein 78, was first discovered as a mammalian protein that could be specifically induced in cultured cells by starvation of glucose (Shiu et al., 1977) and by hypoxic conditions (Sciandra et al., 1984). The expression level of gadd45 mRNA also is induced by the hypoxic conditions (Price and Calderwood, 1992). Thus, the upregulated levels of VEGF, BiP protein, and gadd45 mRNA in the current study suggest that the vasospastic vessels are exposed to hypoxic and hypoglycemic stress conditions.
The BiP protein localized in the endoplasmic reticulum facilitates the translation, folding, and assembly of proteins under normal conditions as a molecular chaperon, preventing the formation of malfolded protein aggregates for cell survival under stress conditions. Also, PDI is one of the resident proteins in the endoplasmic reticulum and catalyzes the formation of native disulfide bonds in the biosynthesis of secretory proteins (Freedman, 1989). Because the highly conserved thioredoxin-domains between PDI-RP P5 and PDI suggest that they are functionally related (Hayano and Kikuchi, 1995), the upregulated expression level of BiP protein and PDI-RP P5 mRNA in cerebral vasospasm may contribute to the cell survival in arterial walls under stress conditions.
Although the other three genes (neuromodulin, acid sphingomyelinase-like phosphodiesterase, and Frizzled-6) also were differentially expressed in cerebral vasospasm, the functional significance of these molecules currently is unknown.
One of the unknown genes, DVS 27, whose expression was most highly upregulated in vasospastic arteries, has been cloned and partially characterized by analyses of intracellular localization of the protein and changes of the expression levels in response to inflammatory stimuli. Although its functional role still is unknown, the DVS 27 gene was found to encode a nuclear protein that could be involved in inflammatory events. Thus, such an approach will be useful, at least in part, to predict the role of proteins encoded by the unknown genes.
Further characterization of the known and unknown genes obtained in this study should improve understanding of the molecular processes and mechanism of continuous cerebral vasospasm after subarachnoid hemorrhage.
Footnotes
Acknowledgments:
The authors thank Dr. A. Kawashima, Dr. A. Sasahara, and Mr. H. Miyazaki for excellent assistance in preparation of the animals.
