Abstract
The electrophoretic pattern of laddered DNA fragments which has been observed after cerebral ischemia is considered to indicate that neurons are dying by apoptosis. Herein the authors directly demonstrate using ligation-mediated polymerase chain reaction methods that 99% of the DNA fragments produced after either global or focal ischemia in adult rats, or produced after hypoxia-ischemia in neonatal rats, have staggered ends with a 3' recess of approximately 8 to 10 nucleotides. This is in contrast to archetypal apoptosis in which the DNA fragments are blunt ended as seen during developmental programmed cell death in dying cortical neurons, neuroblastoma, or thymic lymphocytes. It is not simply ischemia that results in staggered ends in DNA fragments because ischemic myocardium is similar to archetypal apoptosis with a vast majority of blunt-ended fragments. It is concluded that the endonucleases that produce this staggered fragmentation of the DNA backbone in ischemic brain must be different than those of classic or type I apoptosis.
Despite its initial description as long ago as 1980 (Wyllie et al., 1980), an electrophoretic laddered pattern of DNA fragments remains the premier biochemical hallmark of an apoptotic mode of cell death (Bortner et al., 1995; Choi, 1996; Walker and Sikorska, 1997). However, some exceptions have been noted, for example in neonatal brain after excitotoxicity (van Lookeren Campagne et al., 1995), that prevent the sole use of this criterion as conclusive evidence of apoptosis. This electrophoretic pattern arises from the cleavage of the linker DNA between nucleosomes in chromatin, resulting in oligomeric fragments of unit size of approximately 180 base pairs (bp). The occurrence of such internucleosomal cleavage has been documented in a wide variety of cells and tissues induced to die by a myriad of insults. The endonuclease responsible for the ordered scission of the DNA backbone remains unknown, but several candidates such as DNase I or II and Nuc 18 have been proposed over the years (Bortner et al., 1995; Walker and Sikorska, 1997).
Neurons have the capacity to die by apoptosis and do so in huge numbers during fetal and early postnatal development as supernumerary precursor neuroblasts are culled during synapse formation. The programmed cell death of such immature neurons is considered to be apoptotic because they exhibit both the morphologic characteristics of marginated chromatin and karyorrhexis (Lo et al., 1995) and the biochemical characteristic of laddered DNA fragmentation (Ferrer et al., 1994; Blaschke et al., 1996; Portera-Cailliau et al., 1997a).
To date there have been numerous reports showing electrophoretic patterns of laddered DNA fragments from either ischemic brain or other neurodegenerative situations (reviewed by Choi, 1996; Chopp and Li, 1996; Linnik, 1996; MacManus and Linnik, 1997), which has led to the conclusion that cerebral ischemia induces apoptosis of brain cells. This is despite the fact that, in general, ischemic neurons do not display the morphologic characteristics of apoptosis (Deshpande et al., 1992; van Lookeren-Campagne and Gill, 1996; Petito et al., 1997), although there are exceptions (Endres et al., 1998; States et al., 1996), particularly in the boundary zone on the edge of the infarcted brain after a focal ischemic insult (Charriault-Marlangue et al., 1996; Li et al., 1995). The laddered DNA fragmentation seen after cerebral ischemia has been considered apoptotic for several reasons. First, the fragments appear as oligomers of approximate unit size of 180 to 200 bp, which is similar to classic apoptosis (reviewed by Choi, 1996; Chopp and Li, 1996; Linnik, 1996; MacManus and Linnik, 1997). Second, the fragmentation is different from the random cleavage of necrosis (Tominaga et al., 1993; MacManus et al., 1995). Third, there is a clear involvement of the apoptotic proteolytic cascade of caspases enzymes after cerebral ischemia (Endres et al., 1998; Namura et al., 1998). In a similar vein, oligonucleosomal DNA fragmentation indicative of apoptosis has been described in tissue from ischemic rodent and human heart (Fliss and Gattinger, 1996; Veinot et al., 1997).
Other than noting the common oligonucleosomal size of apoptotic fragments in all dying cells studied, the nature of the ends of the DNA fragments has received scant attention. Alnemri and Litwack (1990) demonstrated that apoptotic cultured human lymphocytes displayed internucleosomal cleavage with blunt-ended double-strand breaks. This has been confirmed in rat thymus by use of ligation-mediated polymerase chain reaction (LM-PCR) methods (Staley et al., 1997). Contradictory views have been presented using DNA polymerase (Peitsch et al., 1993) or synthetic oligonucleotides in an in situ hybridization histochemical approach with paraffin-embedded sections of rat thymus (Didenko and Hornsby, 1996). Although the latter methods are not quantitative, it was suggested that the majority of fragment ends in thymus has single-base 3' overhangs, although some blunt-ended fragments were also present (Didenko and Hornsby, 1996). Alternatively, because DNA from glucocorticoid-treated thymic lymphocytes was labeled by the Klenow fragment of DNA polymerase (which lacks nick-translation activity), it was argued that some ends produced during apoptosis must have 3'-recessed ends (Peitsch et al., 1993).
The situation with respect to the ends of DNA fragments in ischemic neurons has not been studied, but neurons dying during fetal development clearly contain oligonucleosomal DNA fragments in which a majority are blunt ended (Blaschke et al., 1996). However the ability of in situ end-labeling by the Klenow fragment of DNA polymerase I to detect DNA scission in neurons in brain sections from ischemic adult animals suggests the presence of 3'-recessed ends (Tobita et al., 1995; Tagaya et al., 1997). We herein directly examine the nature of the ends of DNA fragments appearing after either global or focal cerebral ischemia in adult animals, or hypoxia-ischemia in neonatal animals, and find them all different from those present in either ischemic myocardium, apoptotic neurons in developing brain, or apoptotic cultured cortical neurons and human neuroblastoma. It is concluded that the endonuclease responsible for degrading DNA into oligonucleosomal fragments in ischemic brain is not the same as that operating during classic apoptosis.
MATERIALS AND METHODS
Cerebral ischemia
Adult rat transient global ischemia by two-vessel occlusion. All procedures involving animals received institutional approval and met the guidelines of the Canadian Council on Animal Care. Transient cerebral global ischemia by two-vessel occlusion was produced in pentobarbital-anesthetized male Sprague Dawley rats (350 to 400 g) by bilateral carotid artery occlusion combined with hypovolemic arterial hypotension for 12 minutes with careful maintenance of normal brain temperature (37.5° to 38°C) as described previously (MacManus et al., 1995). After 48 hours of reperfusion when DNA fragmentation is maximal (MacManus et al., 1995), the brain was sampled after sacrifice by decapitation under anesthesia with 3% fluothane, immediately chilled, and dissected on a cold plate (0°C), and the striatum was frozen and stored at −80°C until assay.
Adult rat transient focal ischemia. Mature male spontaneously hypertensive (SHR, 225 to 250 g) Wistar rats (Harlan Sprague-Dawley Inc., Indianapolis, IN, U.S.A.) were fasted for 18 hours before transient focal ischemia was produced by temporarily clipping the middle cerebral artery (for 2 or 3 hours) as described by Buchan et al. (1992). This procedure reduced regional cerebral blood flow to 15% of basal level during the ischemic period and returned it to 95% of basal level on reperfusion. Brain temperature was controlled to 37.5° ± 0.5°C both during anesthesia and after recovery. After 24 hours of reperfusion when DNA fragmentation is easily detectable, rats were sacrificed, the brains were rapidly removed, and the right cortex was dissected on a cold plate and stored at −80°C.
Neonatal rat hypoxia-ischemia. A period of cerebral hypoxia-ischemia was produced in 7-day-old Wistar rats using a well-established neonatal model of hypoxia-ischemia similar to that described previously (MacManus et al., 1997b). Briefly, animals were anesthetized with isoflurane (4% for induction, 1.5% to 2.5% for maintenance), and the right carotid artery was isolated and ligated. A 2- to 3-hour recovery period with the dam was followed by 2 hours in a humidified hypoxic chamber (8% oxygen + 0.3% isoflurane for sedation; remainder nitrogen) that was maintained at 35.5°C, resulting in an axillary temperature of 36.5° to 37°C. The animals were then returned to the dam and 18 hours after the end of hypoxia, when DNA fragmentation was obvious (MacManus et al., 1997b), were reanesthetized with pentobarbital (40 mg/kg) before sacrifice. The brain was quickly removed and the cortex, striatum, and thalamus were dissected and frozen in isopentane (–60°C) and subsequently stored at −75°C until assay.
Cardiac ischemia
Cardiac ischemia was produced by coronary artery occlusion for 45 minutes in male Sprague-Dawley rats under halothane anesthesia as described previously (Fliss and Gattinger, 1996). After removal of the suture, reperfusion was continued for 4 hours, at which time the ischemic cardiac tissue was dissected after delineation by negative staining with Evans blue dye, rapidly frozen, and stored at −80°C.
Cultured cells
Fetal rat (E18) cortical neurons were cultured on poly-l-lysine-coated 35-mm tissue culture dishes (Gibco-BRL Life Technologies, Burlington, Ontario, Canada), and apoptosis was induced by staurosporine by 7 hours as described previously (MacManus et al., 1997a). Human SH-SY5Y neuroblastoma cells were cultured in RPMI 1640 supplemented with nonessential amino acids (Eagle's), 2 mmol/L 1-glutamine, 20 µg/mL gentamycin sulfate, and 15% (v/v) fetal bovine serum, and apoptosis was induced by staurosporine by 24 hours of treatment. Apoptotic rat thymic lymphocytes were produced by treatment with 1 µmol/L dexamethasone for 6 hours.
DNA isolation and electrophoresis
The total DNA was extracted and purified by precipitation from brain samples or from cultured cells by standard methods, and then 3'-end labeled using terminal transferase before electrophoresis as described (MacManus et al., 1995). The final DNA was resuspended in TE buffer (10 mmol/L Tris, 1 mmol/L EDTA, pH 8.0) to which was added 4 µL of a mixture to a final concentration of 10% glycerol, 0.2% sodium dodecyl sulfate, 0.02% bromophenol blue, and 20 mmol/L EDTA, and heated at 65°C for 10 minutes. The oligonucleosomal DNA fragments were electrophoretically separated on a 1.8% Synergel (Diversified Biotech Inc., Boston, MA, U.S.A.) in TAE electrophoresis buffer (40 mmol/L Tris, 1 mmol/L EDTA, pH 8.0 with acetic acid). The gels with DNA size markers were run for 16 to 18 hours at 30 V at room temperature and autoradiographed (MacManus et al., 1995).
Purification of discrete mononucleosomal DNA. When the discrete mononucleosomal DNA fragment was required, the total DNA (without 3'-end labeling) from ischemic adult or neonatal brain and ischemic heart was initially electrophoresed as outlined above, the gel was stained with ethidium bromide, and the monosome-sized fragment was excised from the gel. This discrete DNA fragment was isolated from the gel medium using BioRad Prep-A-Gene DNA purification matrix (BioRad Inc., Mississauga, Ontario, Canada) following the manufacturer's instructions.
Ligation-mediated PCR for blunt-ended DNA
The feasibility of using LM-PCR to detect the presence of blunt-ended DNA fragments (Staley et al., 1997) was demonstrated in our hands using λ-DNA (purchased from Gibco-BRL) and pBR322 plasmid DNA (isolated from Escherichia coli HB 101 grown in LB medium), which was purified using a QIAamp Plasmid Kit (QIAGEN Inc., Chatsworth, CA, U.S.A.) according to the manufacturer's instructions. Both blunt-ended and 3'-recessed fragments were produced from these purified DNA samples. A blunt-ended fragment (604 bp) was obtained from λ-DNA by restriction with Stu I (Pharmacia, Ste-Anne-de-Bellevue, Quebec, Canada), and a fragment (564 bp) with a 4-nucleotide (nt) 3'-recessed end by restriction with Hind III (Gibco BRL, Burlington, Ontario, Canada). pBR322 was used to obtain a 1-nt 3'-recessed fragment by restriction with Mva I (MBI Fermentas, Hamilton, Ontario, Canada), and a 315-bp, 2-nt 3'recessed fragment by restriction with Taq I (Gibco-BRL). In some experiments any existing 3'-recessed ends were blunt ended by fill in using Klenow polymerase I. Briefly, DNA was purified using a silica gel matrix (BioRad), and 2-µg aliquots were treated with 5 U of Klenow polymerase (Gibco BRL) with the four deoxyribonucleotides at pH 8.0 for 15 minutes at 30°C, followed by inactivation at 75°C for 10 minutes. The excess deoxyribonucleotides were removed by QIAGEN spin-columns (QIAGEN Inc.), and the DNA was recovered for electrophoresis.
The LM-PCR procedure was performed on DNA from both the test samples with blunt and recessed ends described above, and the various tissue samples as outlined (Staley et al., 1997). Briefly, two oligonucleotides (24 bp: 5'-AGCACTCTCGAGCCTCTCACGCA-3'; 12 bp: 5'-TGCGGTGAGAGG-3') were synthesized (Expedite Nucleic Acid Synthesis System, PerSeptive Biosystems Inc. Framingham, MA, U.S.A.) and gel purified. The oligonucleotides were annealed (5 nmoles each) and ligated to 200 ng of target DNA using 1 µL of 20 mmol/L ATP, 1 µL of 20 mmol/L dithiothreitol, and 4 U T4 DNA ligase (New England BioLabs, Mississauga, ON, Canada) and incubated at 16°C overnight, and the enzyme was inactivated by heating at 75°C for 10 minutes. The PCR reaction was carried out with fivefold serial dilutions of target DNA (1000-8 pg) with 65 pmol of the 24-bp oligonucleotide and 3 U of Taq polymerase (Gibco-BRL), 2 mmol/L MgCl2 with 0.324 mmol/L deoxyribonucleotides in 67 mmol/L Tris, 16 mmol/L (NH4)2SO4 10 mmol/L β-mercaptoethanol, and 0.1 mg/mL bovine serum albumin, pH 8,8, in a 50-µL volume. The amplification cycles used were 1 cycle of 72°C for 3 minutes, 94°C for 5 minutes, and 77°C for 3 minutes; 25 cycles of 94°C for 1 minute and 77°C for 3 minutes; and 1 cycle of 77°C for 2 minutes. An aliquot of the samples were run on standard agarose gels (1.8%) as described above.
The LM-PCR method was also used to estimate the length of the 3' recess by determining the size difference between the largest fragment amplified from a monosome blunt ended by filling in the recess with Klenow polymerase and the smallest fragment amplified from a monosome with the recess digested back to a blunt end with the single-strand-specific S1 nuclease, For the largest fragment, the recessed end was filled in using 0.5 U of Klenow DNA polymerase (Gibco BRL) in 20 µL with 33 µmol/L deoxyribonucleotides in 50 mmol/L Tris, 10 mmol/L MgCl2, and 50 mmol/L NaCl, pH 8.0, for 15 minutes at 30°C. For the smallest fragment, the 5' overhang was removed using 1 µL of S1 nuclease from Aspergillus oryzae (Pharmacia) in a pH 5.5 buffer (0.2 mol/L NaCl, 50 mmol/L NaOAc, and 1 mmol/L ZnSO4 in 0.5% glycerol) for 10 minutes at 37°C. Both samples were precipitated after the reaction with NaOAc/ethanol. The LM-PCR amplification reaction was performed on the two blunt-ended monosomal fragments as described above. The amplified fragments were electrophoresed on a nondenaturing 8% acrylamide gel (40: 1 acrylamide monomer to bis-acrylamide) at 120 V overnight using 20-bp and 50-bp DNA size markers until the bromophenol blue dye had migrated three quarters of the gel length. The gel was stained with ethidium bromide and photographed with type 55 film (Polaroid).
RESULTS
As demonstrated in the Chun laboratory (Staley et al., 1997), a method of LM-PCR can be designed that will only amplify blunt-ended DNA fragments, In Fig. 1 we confirmed this specificity using a blunt-ended fragment of λ-DNA and showing that a 3'-recessed end of only 1 nt was sufficient to prevent PCR amplification. Blunt ending the fragments, which contained either 1-, 2-, or 4-nt recessed ends by use of Klenow DNA polymerase to fill in the missing nucleotides, now permitted amplification (Fig. 1).
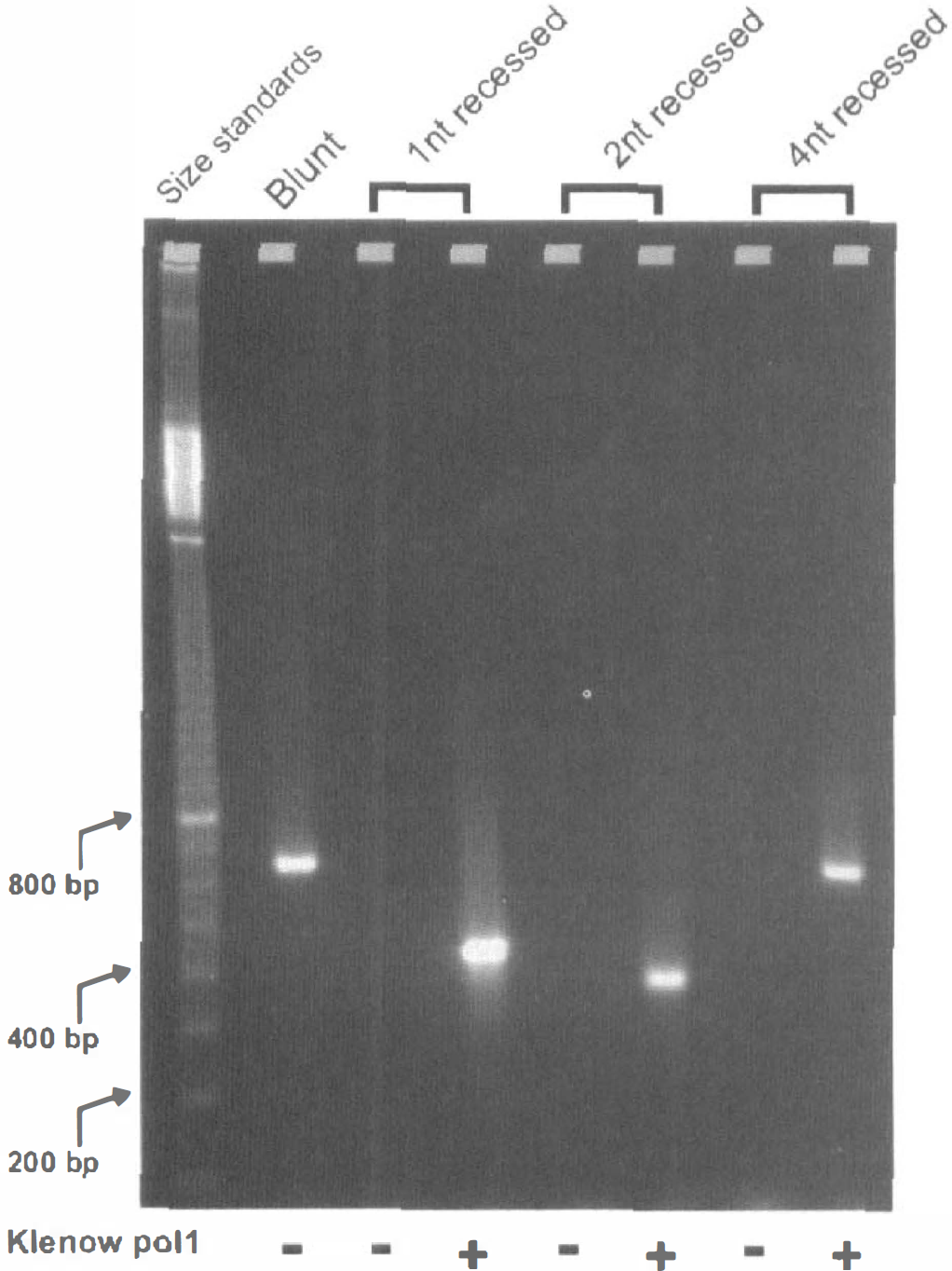
Demonstration of the absolute requirement for blunt-ended DNA fragments in amplification by LM-PCR. A blunt-ended fragment of λ-DNA amplified as expected, but a fragment with only a 1-nt 3' -recessed staggered end was as efficient as 2- or 4-nt recessed in blocking PCR amplification. Blunt ending these staggered ends by pretreatment with Klenow polymerase 1 to fill in any existing 3' recess subsequently permitted the LM-PCR reaction to proceed.
When this LM-PCR method was applied to extracts of adult thymic lymphocytes (Fig. 2A), we were able to corroborate its ability to easily detect laddered DNA fragments from apoptotic thymic lymphocytes. It should be noted that the size of the individual rungs of such laddered DNA fragments is increased by 48 bp because of addition of the PCR primer oligonucleotide to each end of the amplified fragment (Staley et al., 1997). Pretreatment of the DNA fragments with DNA polymerase to blunt end any existing recessed ends gave no increase in the intensity of band staining, suggesting that the vast majority of the fragments from the thymic lymphocytes were indeed blunt ended. To our surprise, there were few detectable laddered DNA fragments from ischemic adult brain, whether striatum (Fig. 2B) or hippocampus (not shown) after transient global ischemia, or cortex (Fig. 2C) or dorsolateral striatum (not shown) after transient focal ischemia. However, pretreatment of these brain DNA fragments with DNA polymerase permitted their amplification and ready detectability in the agarose gels (Fig. 2B and C), indicating that the majority of these fragments from ischemic adult brain were not blunt ended, but rather had ends with at least a l-nt 3' recess. The similarity in the intensity of the band staining produced from 8 pg of target DNA which had been polymerase pretreated to the intensity produced from 1,000 pg of target DNA without treatment (Fig. 2B and C) led to the conclusion that more than 99% of the fragments induced by cerebral ischemia in the adult were not blunt ended. On the other hand, the fragments from ischemic myocardium did appear to be blunt ended (Fig. 2D) and were as similarly insensitive to polymerase pretreatment as those from apoptotic thymic lymphocytes, indicating that the majority of the DNA fragments in injured myocardium had different ends compared with those in the ischemic adult brain.
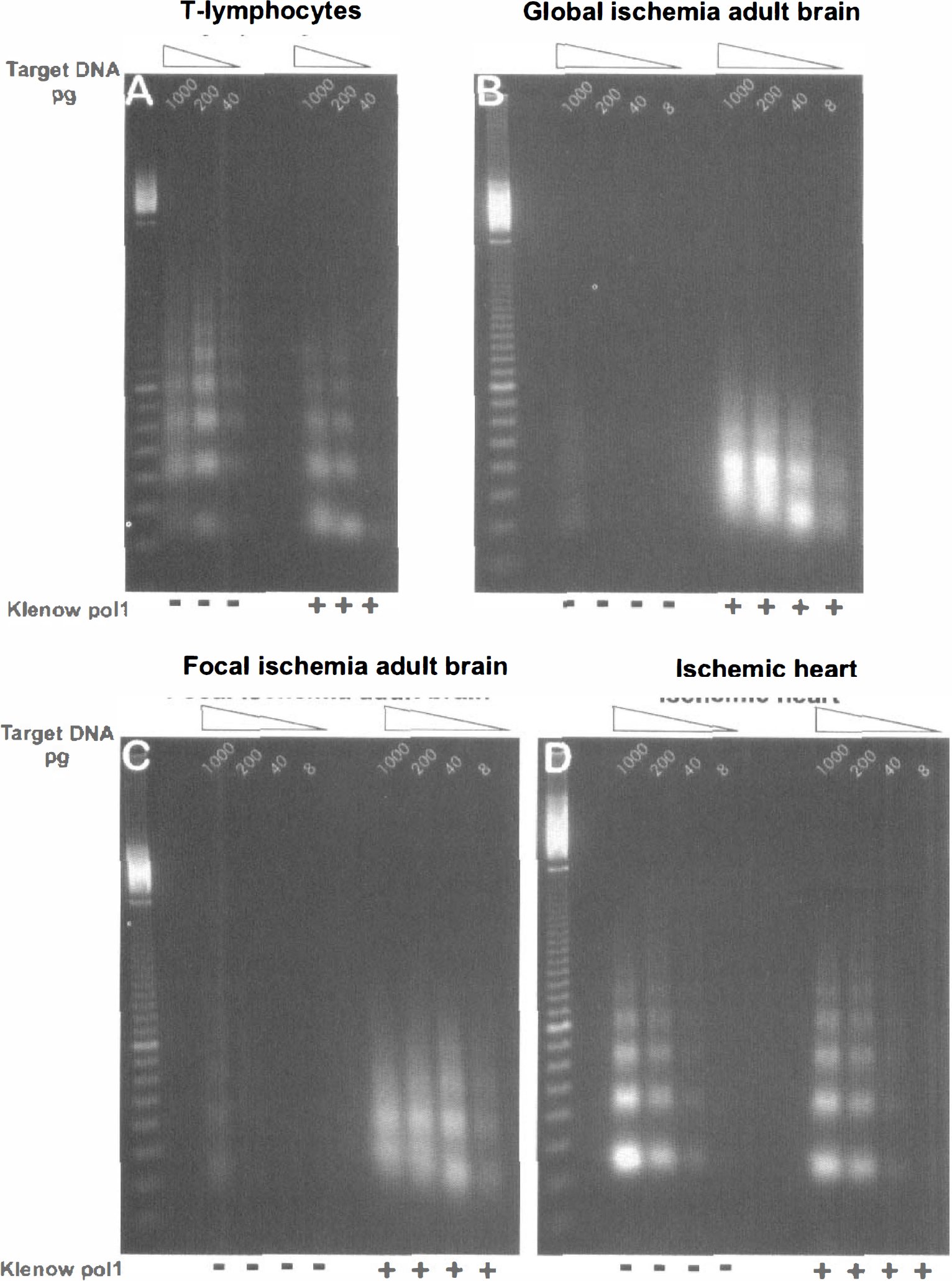
Ligation-mediated PCR demonstrates that the majority of laddered DNA fragments from ischemic adult brain are not blunt ended, Fivefold serial dilutions of target DNA (1000-8 pg DNA) purified from dexamethasone-treated thymic lymphocytes, striatum 48 hours after two-vessel occlusion, cortex 24 hours after middle cerebral artery occlusion, and ischemic heart with 4 hours of reperfusion were subjected to PCR with and without prior Klenow polymerase treatment. (
The finding of fragments with recessed ends was extended to ischemic brain from day 7 neonatal animals subjected to hypoxia-ischemia (Fig. 3). In DNA isolated from either hypoxic-ischemic cortex, striatum, or thalamus, the similarity in staining intensity of the amplified fragments from 8 to 40 pg of blunt-ended target DNA to that from 1,000 pg of original DNA again led to the conclusion that the majority of the fragments were not blunt ended. In contrast, when immature neurons from the neocortex of E18 rats were cultured and induced to undergo apoptosis by staurosporine treatment, the ensuing oligonucleosome-sized DNA fragments were easily detectable and were unaffected by blunt ending, suggesting that few, if any, recessed ends were present (Fig. 4A). Similar blunt-ended DNA fragments were found to predominate in apoptotic neuroblastoma cells (Fig. 4B).
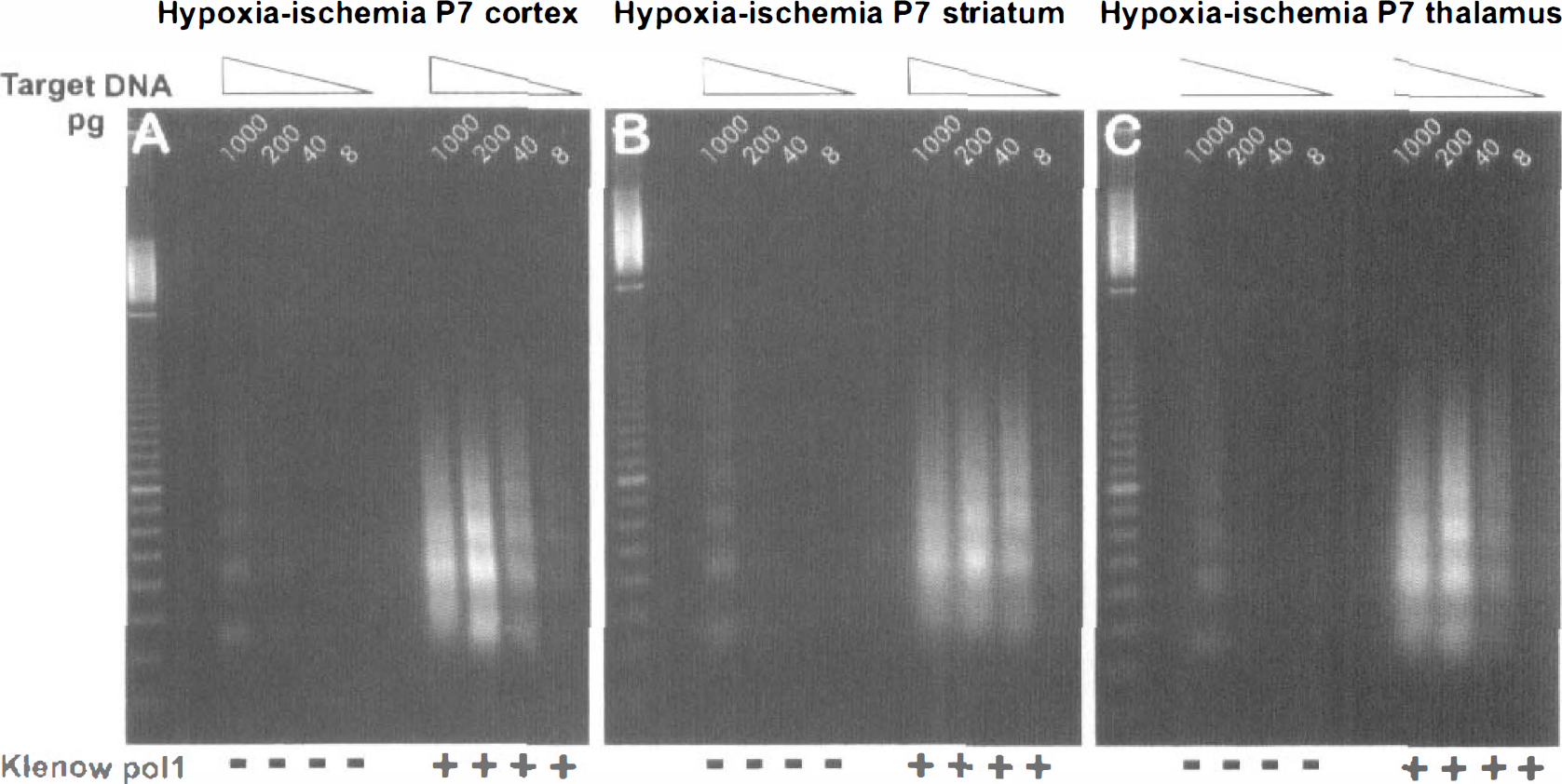
The majority of laddered DNA fragments from ischemic neonatal brain are not blunt ended. Fivefold serial dilutions of target DNA (1000-8 pg DNA) purified from cortex (
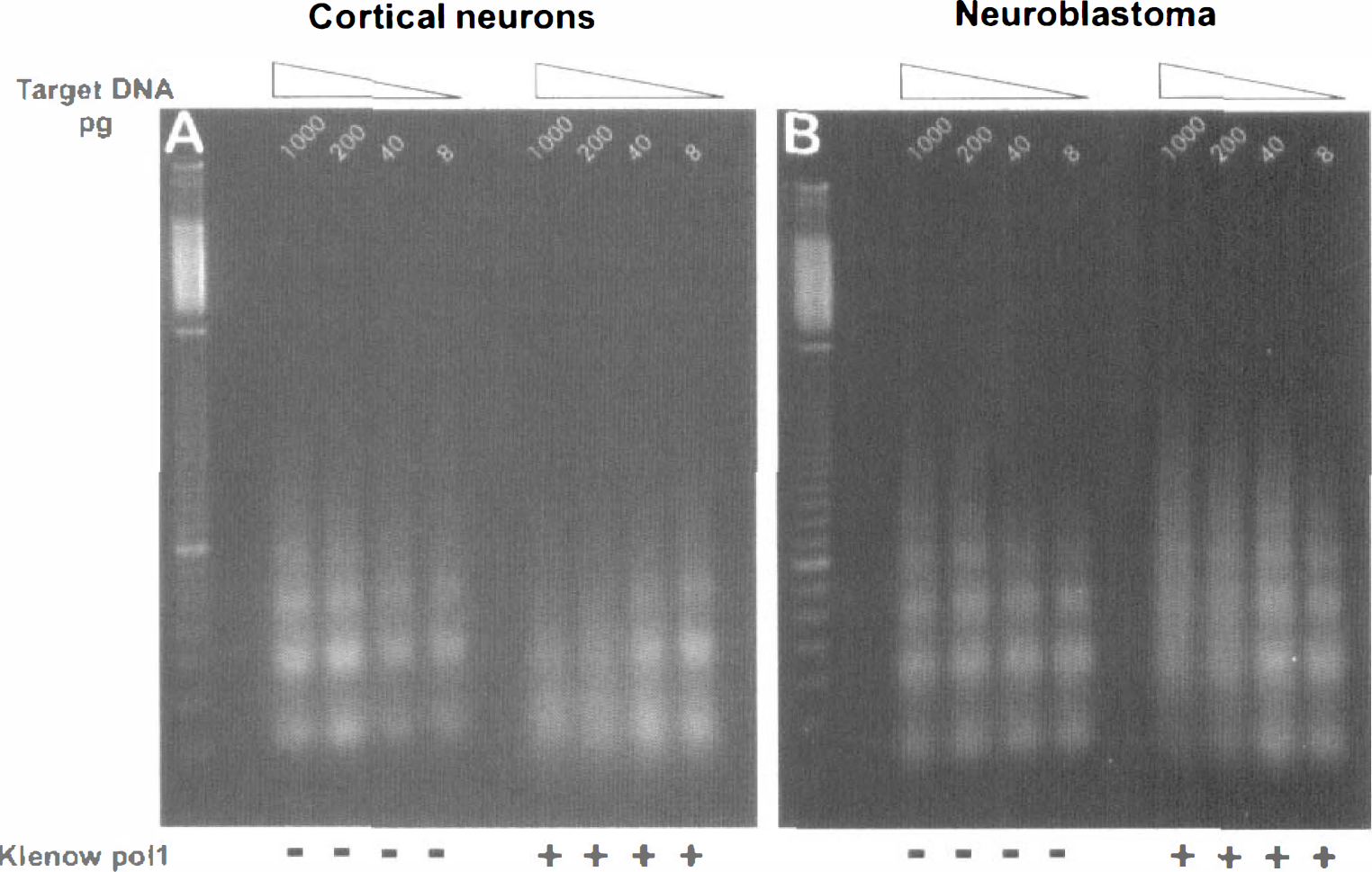
The majority of laddered DNA fragments from apoptotic neurons or neuroblastoma cells are blunt ended. Serial dilutions of target DNA (1000-8 pg DNA) purified from staurosporine-treated E18 cortical cultures (
Finally, the LM-PCR method was used to determine the length of the 3' recess in the DNA fragments from ischemic brain. To simplify the subsequent analysis, we first isolated the monosome-sized DNA fragment from ischemic adult and neonatal brain and from ischemic myocardium, and then determined the size difference between the largest double-stranded monosomal fragment obtained by blunt ending (filling in the recess) with Klenow polymerase and the smallest double-stranded fragment obtained with blunt ends obtained by removal of any unpaired 5' overhang with the single-strand-specific S1 nuclease. The monomeric DNA does not electrophorese in tight bands because it is heterogeneous, but Fig. 5 still shows that a difference of approximately 20 bp exists in the mean size of the monosomal DNA fragments from either ischemic adult or neonatal brain treated with polymerase compared with those treated with nuclease. It was concluded that a 3'-recess at either end of the double-strand DNA fragment of approximately 8 to 10 nt would explain such a finding. It could be argued that nuclease digestion would always lead to a smaller DNA fragment because in addition to removal of any overhanging unpaired DNA, it was possible that SI nuclease digestion could reduce fragment size by cleaving into the double-stranded region. This was not observed under the conditions used because the fragments from ischemic heart, which were considered blunt ended, did not change in size after treatment with S1 nuclease or polymerase (Fig. 5).
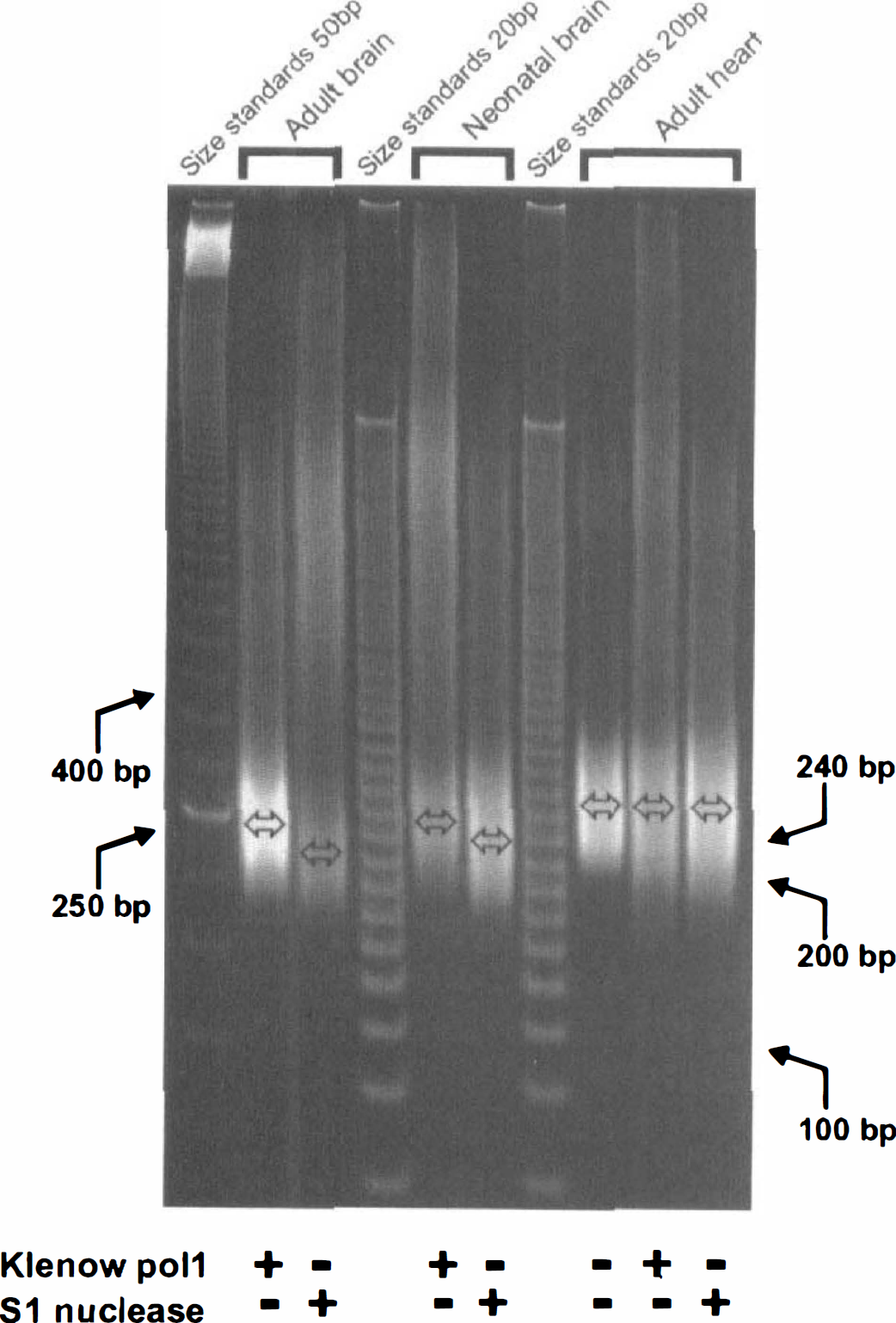
The size of the 3' recess in DNA fragments from ischemic adult and neonatal brain is 8 to 10 nt. The monosomally sized DNA fragment isolated from total DNA of ischemic adult or neonatal brain and ischemic heart was blunt ended by treatment with either Klenow polymerase or single-strand-specific S1 nuclease before amplification by LM-PCR. The center of the DNA bands is indicated for each treatment, and is approximately 20 bp lower after nuclease treatment from both ischemic adult and neonatal brain. The size of the monosomal DNA from ischemic heart was unaffected by either treatment, confirming its blunt-ended nature, a fact that also demonstrates the specificity of the S1 nuclease reaction.
DISCUSSION
This study addressed the question of whether all electrophoretically separated laddered DNA fragments are similar. The laddered fragments from adult ischemic brain have been termed “apoptotic” by ourselves (e.g. MacManus et al., 1995, 1997b) and many others (reviewed by Choi 1996; Chopp and Li, 1996; Linnik, 1996; MacManus and Linnik, 1997) based solely on similar oligomeric patterns of 180 to 200 bp unit size. Such oligonucleosomal electrophoretic patterns are regarded as one of the hallmarks of an apoptotic mode of cell death (Wyllie et al., 1980). However, on closer inspection in the current study, these laddered DNA fragments from ischemic neonatal or adult brain were shown to be different from those found during programmed cell death in immature brain, or classic apoptosis exemplified by either glucocorticoid-treated thymocytes or staurosporine-treated cultured primary neurons or neuroblastoma. Despite the huge amount of work done in the last several years demonstrating laddered DNA fragmentation patterns in myriad cell types, relatively little information has emerged as to the nature of the ends of these fragments. Dying thymocytes produced by glucocorticoid treatment remain one of the primary models for studies on apoptosis, and evidence has been presented that the DNA fragments appearing after such treatment are blunt ended (Alnemri and Litwack, 1990), as are fragments in novobiocin-treated chicken erythrocytes (Villeponteau et al., 1986). Similar conclusions were reached using a semiquantitative LM-PCR method that depends on the presence of blunt-ended fragments, although a small proportion of the thymocyte DNA fragments may have 3'-recessed ends (Staley et al., 1997). The presence of some such 3'-recessed ends in glucocorticoid-treated thymic lymphocytes had previously been deduced by labeling with the Klenow fragment of DNA polymerase (Peitsch et al., 1993). However, a completely opposite conclusion was reached in another recent study, in which it was suggested that the majority of fragment ends in thymus have single-base 3' overhangs, although some blunt-ended fragments were also present (Didenko and Hornsby, 1996). The methods used to generate these latter findings are not quantitative, so the LM-PCR method appears to produce the most reliable information owing to its semiquantitative nature. However, this leads to another dilemma in that if it is accepted that blunt ends predominate in this classic model of apoptosis, then the endonuclease that produces them has few counterparts. Most known endonucleases, especially the ones touted to be involved in apoptosis such as DNase I and II, or Nuc18 (Bortner et al., 1995; Walker and Sikorska, 1997), do not generate blunt-ended fragments (Lutter, 1977; Soliner-Webb et al., 1978; Staley et al., 1997); for example, DNase I and II produce double strand breaks with 3- or 4-nt 3'-overhang ends, not 3'-recessed ends (Soliner-Webb et al., 1978).
Our direct demonstration that the majority of laddered DNA fragments produced after either global or focal cerebral ischemia in adult brain has 3'-recessed ends is novel. However, it must be said that the use of Klenow DNA polymerase 1 (i.e., without the confounding activity of nick translation) to successfully label brain sections from ischemic animals had already suggested that such 3'-recessed ends were present (Tobita et al., 1995; Tagaya et al., 1997). The massive programmed cell death of neurons during late fetal and early neonatal development is considered to be apoptotic on the basis of both morphologic and DNA fragmentation criteria (Ferrer et al., 1994; Lo et al., 1995; Blaschke et al., 1996; Portera-Cailliau et al., 1997a). Both cultured cortical neurons (MacManus et al., 1997a) or human neuroblastoma SH-SY5Y (Posmantur et al., 1997) induced to die by staurosporine treatment also exhibit morphologic and biochemical hallmarks of classic apoptosis. The laddered DNA fragments from both these apoptotic models were likewise found to be blunt ended (Fig. 4), in addition to those from the archetypal apoptotic adult thymocytes induced to die by glucocorticoid (Fig. 2A). In contrast, the majority of oligonucleosomal DNA fragments from ischemic adult or neonatal brain are not blunt ended but contain 3'-recessed ends (Fig. 2B and C, and 3). In the global model of ischemia at the 48-hour reperfusion time chosen, we conclude that the majority of DNA fragments originate in striatal neurons (MacManus et al., 1995), but in both the adult focal and the neonatal hypoxia-ischemia models, with the relatively long periods of ischemia, cell death is nonselective, and it is concluded that the DNA fragments arise from both neurons and glia in the infarcted tissue. It remains to be seen whether mild ischemia, which produces visible apoptotic neurons after much longer reperfusion times than used in the current study (Du et al., 1996; Endres et al., 1998), gives rise to DNA fragments with blunt or staggered ends. On the other hand, the majority of laddered DNA fragments from ischemic myocardium are blunt ended (Fig. 2D), indicating that not all ischemic tissue generates DNA fragment ends different from archetypal apoptosis. This similarity of the DNA fragmentation of ischemic heart to classic apoptosis supports the conclusions in animal and human ischemic myocardium in which both morphologic and biochemical evidence of apoptosis has been observed (Fliss and Gattinger, 1996; Veinot et al., 1997).
There has been intense debate about whether the neuronal death after an episode of cerebral ischemia is necrotic or apoptotic. Morphologic evidence has been lacking at the ultrastructural level in both global and focal models of cerebral ischemia (Deshpande et al., 1992; van Lookeren-Campagne and Gill, 1996; Petito et al., 1997), although after focal ischemia apoptotic morphology has been reported primarily in the inner border zone of the ischemic infarct (Li et al., 1995) and in the hippocampus after global ischemia (Nitatori et al., 1995). At the simple light microscopic level some evidence of chromatin margination and karyorrhexis has been observed both within and outside the infarct zone (Charriault-Marlangue et al., 1996; Endres et al., 1998; Li et al., 1995; States et al., 1996) that is reminiscent of the appearance of dying neurons during programmed cell death in fetal and postnatal development (Lo et al., 1995). In addition to the morphologic criteria and the biochemical hallmark of oligonucleosomal DNA fragmentation discussed above, other biochemical hallmarks of apoptosis have been reported after cerebral ischemia, particularly the expression of proapoptotic genes such as p53, bax, and myc (reviewed by Koistinaho and Hokfelt 1997; MacManus and Linnik, 1997) and the activation of proteolytic caspases (Endres et al., 1998; Namura et al., 1998).
The current finding that the low-molecular weight DNA fragments appearing in neonatal or adult brain after ischemia are different from archetypal apoptosis adds support to the suggestion, found with the high-molecular weight DNA fragments (MacManus et al., 1997b), that ischemic cell death of neurons is not by apoptosis as originally defined, i.e., type I (Wyllie et al., 1980; Clark, 1990). However, it is not by necrosis either, because such degeneration in brain leads to random fragmentation of DNA that appears as electrophoretic smears (Tominaga et al., 1993; MacManus et al., 1995). Indeed, the distinction between apoptosis and necrosis may be somewhat artificial and needs to be replaced by an expanded view, with more or less of the core cell-death machinery being engaged, and perhaps controlled by the cellular ATP level (Leist and Nicotera, 1997). In a most thorough study on excitotoxicity in immature and adult brain, the provocative suggestion was made that a continuum exists in the damaged brain between apoptosis and necrosis (Portera-Cailliau et al., 1997a, 1997b). It has also been suggested that the occurrence of apoptosis or necrosis is dependent on the severity of the insult determined by the duration or level of the ischemia, or whether cell death is in the core or periphery of the damaged territory (Du et al., 1996; Endres et al., 1998; Li et al., 1995; Matsushita et al., 1998). In addition, it may be that characteristics of both can occur in the same cell, as in a study of brain trauma in which evidence was presented for apoptosis with nuclear characteristics but necrosis with cytoplasmic characteristics (Colicos and Dash, 1996). Thus, attempting to distinguish between the black and white definitions of cell death by apoptosis or necrosis is an oversimplification and not sufficient to describe the processes involved in cerebral ischemic cell death. Instead, responses to different types of injury and their divergence from archetypal type I apoptosis and developmental programmed cell death will need to be assessed independently.
Our studies will evolve to identify the endonuclease that leaves the footprint of both 3'- and 5'-OH end groups in high-molecular weight DNA fragments (MacManus et al., 1997b) and the footprint of low-molecular weight laddered DNA fragments with 8- to 10-nt 3'-recessed ends (Figs. 2, 3 and 5). The combination of endonucleases that cause such complex scission of the DNA backbone needs to be identified before any mechanism of cell death, whether apoptotic, necrotic, or particularly ischemic, can be formulated and targeted for therapeutic intervention.
Footnotes
Abbreviations used
Acknowledgments
The authors thank Debbie Gattinger, Jackie Webster, and Sarosindevi Bascaramurty for their excellent technical assistance; Julie Leblanc for generously providing the apoptotic lymphocytes; Christine Murray and Paul Morley for cultured cortical neurons; and Teena Walker for cultured SH-SY5Y neuroblastoma. The authors also thank J. Chun (University of California San Diego) for assistance with a protocol for LM-PCR.
