Abstract
A novel organic anion transporter selectively expressed at the blood—brain barrier (BBB), originally designated BBB-specific anion transporter type 1 (BSAT1), and now classified as
Introduction
Drug penetration into the brain is determined by transport across the brain capillary endothelial wall, which forms the blood—brain barrier (BBB)
Materials and methods
Materials
The PCR-SelectTM cDNA subtraction kit was obtained from BD Biosciences Clontech (Palo Alto, CA, USA). [α32P]dCTP (3,000 Ci/mmol; 1Ci = 37 GBq), [3H]estradiol 17 β-
Isolation of Poly(A+) RNA
Poly(A+) RNA was isolated from rat brain capillaries, C6 and RG2 implanted rat brain tumor, and rat brain and peripheral tissues using a single step method described previously (Boado and Pardridge, 1991). RG2 and C6 brain tumors were produced in Fischer rats as described previously (Suzuki et al, 2004). The yield of poly(A+) RNA from rat brain capillaries isolated from a pool of 21 cerebral rat brain hemispheres was 3.2 μg, whereas the yield averaged 20 μg/g tissue of total brain and peripheral organs.
Suppression Subtractive Hybridization
Suppression subtractive hybridization was performed using the PCR-Select cDNA subtraction kit as reported previously (Li et al, 2001a). The SSH procedure was performed using rat brain capillary mRNA to produce tester cDNA and rat liver/kidney mRNA to produce driver cDNA. Double-stranded cDNA was synthesized using 1 μg of either tester or driver poly(A+) RNA. cDNA size was normalized by digestion with
Cloning of Full-Length cDNA
To isolate full-length cDNAs corresponding to clones identified in the SSH cloning, a rat brain capillary cDNA library in the pSPORT vector was produced as described previously (Li et al, 2001b). This library was screened with 32P-partial cDNAs isolated by SSH using colony hybridization (Boado et al, 2003a). Clones K-2 and LK-3 were isolated in the SSH cloning, and the DNA sequence of the partial clone indicated that the clone was novel and warranted further examination. The partial insert was released and labeled with [32P] and was used to screen 2.4 × 105 recombinants of the rat brain capillary pSPORT cDNA library. Forty positive clones were identified, and 25 of them were isolated for further characterization. The cDNA inserts were released by double digestion with
DNA Sequence Analysis
DNA sequencing of isolated clones was performed in both directions as described previously (Boado et al, 2003a). Initial DNA sequencing was performed with M13 forward and reverse primers, which are located in pSPORT at the 5′- and 3′-flanking regions of
Northern Blot Analysis
Two micrograms of poly(A+) RNA were resolved on 1.5% agarose/2.2 mol/L formaldehyde gels, and blotted onto GeneScreen Plus membranes. Northern blots were hybridized with 32P-labeled LAT1, 4F2hc, BSAT1, or β-actin cDNAs as described previously (Boado et al, 1999). cDNAs were labeled with [32P]dCTP by the random priming technique using the Megaprimer DNA labeling kit (Boado et al, 2003a). Autoradiograms were performed with Biomax MS film and exposed at −70°C for various periods. Films were scanned with an Umax PowerLook III scanner.
In Situ Hybridization
Real-Time Polymerase Chain Reaction Measurement of Rat Brain Capillary and Whole Brain mRNA
Poly(A+) RNA was isolated from rat brain capillaries or rat whole brain using a single step method as previously described (Boado and Pardridge, 1991). Reverse transcription (RT) was performed according to the Invitrogen SuperScript III First-Strand Synthesis System for PCR using 0.4 μg of poly(A+) RNA as the template and oliogodeoxythymidine as primer in a total volume of 21 μL per reaction. The RT product was diluted by 20-fold before real-time PCR assays. Real-time PCR was performed using the BioRad iQ SYBR Green Supermix according to the manufacturer's instruction. In each PCR reaction in 25 μL, 2 μL of the diluted RT product was used as the template. The primer sequences for rat β-actin (GenBank accession no 42475962) are 5′-GGGACGATATGGAGAAGATTTGG-3′ and 5′-TGAAGGTCTCAAACATGATCTGG-3′. The primer sequences for rat BSAT1 are 5′-AACCTCACTCCCAACATCTCTC-3′ and 5′- GTATCCACTTCACACTCATCGTTTA-3′.
Each primer pair was selected to amplify the sequence shared by contiguous exons of a target gene, to eliminate the possible interference of genomic DNA. Polymerase chain reaction was initiated by a 3-min incubation at 95°C, followed by 40 cycles consisting of 10 secs annealing at 58.3°C, 10 secs extension at 72°C, and 10 secs denaturing at 95°C. Melting curve of each PCR product was determined by measuring the ratio of single-strand DNA versus double-strand DNA at each 0.5°C of temperature increase from 55 to 94°C. Single peak in the melting curve with a signature
Western Blotting
A rabbit polyclonal antiserum against the C-terminus of rat BSAT1 was prepared using the synthetic peptide LQPKYWPGKETRL corresponding to amino acids 706 to 718 of the deduced amino acid sequence of BSAT1. The synthetic peptide was used to screen for antibody titer by enzyme-linked immunosorbent assay. Anti-BSAT1 antibodies were affinity purified after production of an affinity column by coupling the synthetic peptide to cyanogen bromide-activated Sepharose 4B. Rat brain capillary protein was resolved in a 12% sodium dodecyl sulfate-polyacrylamide gel electrophoresis under reducing conditions for Western blotting. Proteins on the gel were electroblotted onto a nitrocellulose membrane. Immunoblot analysis was performed with the affinity-purified anti-BSAT1 antibodies either without or with absorption of the antibody by the synthetic peptide. Biotinylated goat anti-rabbit IgG was used as secondary antibody. Results were visualized with the ABC Elite detection kit as described previously (Pardridge et al, 1990). Western blot analysis with the affinity-purified anti-rat BSAT1 antibody showed that the antibody reacted with a single 55-kDa immunoreactive protein in rat brain capillaries, and that this signal was completely removed by absorption of the anti-BSAT1 antibody with the BSAT1 C-terminal synthetic peptide. However, it is believed that this represents a sequence-specific artifactual cross-reaction of this C-terminal peptide antibody with a non-BSAT1 protein, because the same 55-kDa immunoreactive species was detected in multiple tissues, which do not express the BSAT1 mRNA.
Stable Transfection of Human Embryonic Kidney 293 Cells
The full-length (fl) BSAT1 cDNA was released from the pSPORT vector by double digestion with
A second human embryonic kidney 293 stably transfected cell line was established with a rat BSAT1 expression construct lacking both the 5′- and 3′-UTR of the rBSAT1 transcript. The 2.0-kb BSAT1 orf was obtained from pSPORT-BSAT1 by PCR cloning with Stratagene PfuUltra Hotstart DNA polymerase, forward primer (5′-phosphate-AATGGACACTTCATCCAAAGAAAATGC) and a reverse primer (5′-phosphate-CTTAAAGTCGGGTCTCCTTGCCTG). The PCR-generated BSAT1 orf cDNA was subcloned in the pcDNA3.1(+) vector at the EcoR V site to form the pcDNA-BSAT1(orf) plasmid named clone 994. Positive clones with the correct orientation of the BSAT1(orf) were identified by restriction endonuclease mapping (i.e.,
Influx Assays in Stably Transfected 293 Cells
293-BSAT1(orf) or 293-BSAT1(fl) cells were washed with physiological buffer (PB: 118 mmol/L NaCl, 25 mmol/L HEPES (4-(2-hydroxyethyl)-1-piperazineethanesulfonic acid), pH 7.4, 4.9 mmol/L KCl, 1.4 mmol/L K2HPO4, 1.2 mmol/L MgCl2, 1 mmol/L CaCl2, and 11 mmol/L
[125I]Thyroxine Efflux in Stably Transfected 293 Cells with Prealbumin Trap Technique
The isolation of the efflux kinetics in cells preloaded with [125I]T4 is complicated by the rapid re-entry of the isotope into the cell. The re-entry of T4 was eliminated by the addition of human prealbumin to the medium. Human plasma prealbumin is a high-affinity binder of T4, and this rapid binding prevents any further influx of T4 into the cell. 293-BSAT1(orf) cells were plated in Biocoat collagen I-coated 24-well cluster dishes and grown to confluency. The cells were washed with warm physiological buffer (118 mmol/L NaCl, 25 mmol/L HEPES, 4.9 mmol/L KCl, 1.4 mmol/L K2HPO4, 1.2 mmol/L MgCl2, 1 mmol/L CaCl2, 11 mmol/L glucose, pH 7.4) and incubated with 200 μL of physiological buffer containing 3.26 μCi/mL 125I-T4 (34 nmol/L) at 37°C for 15 mins. Efflux was initiated by adding 10 μL of 2 μg/μL of prealbumin (2 μmol/L final concentration) to the medium, and was terminated by the rapid addition of 2.5 mL of ice-cold physiological buffer. After three washes with ice-cold physiological buffer, the cells in each well were solubilized with 200 μL 1N NaOH by overnight incubation at room temperature. Uptake result was determined by γ-counting and protein assay of the lysates.
In Vitro Transcription and Cross-Linking of BSAT1 RNA to Polysome Proteins
The full-length rat BSAT1 cDNA was obtained by
Results
Rat brain capillaries were purified free of adjoining brain tissue (Figure 1A). The RNA isolated from the brain capillaries was used to produce the subtracted rat brain capillary cDNA library using SSH (Materials and methods). Southern blotting with subtracted tester cDNA (Figure 1B, top panel) or unsubstracted cDNA (Figure 1B, bottom panel) allowed for the isolation of individual clones that were highly enriched at the BBB, such as the clone in well E3 (Figure 1B). This clone comprises a 0.8 kb cDNA insert (Figure 1C). The insert was 32P-labeled, and was used to screen a rat brain capillary cDNA library composed of full-length transcripts. DNA sequence analysis of this clone, designated BSAT1 (Materials and methods), showed the transcript was composed of 2,736 nucleotides (nt), with a 374 nt 5′-UTR, a 2,148 nt coding region, a stop codon, a 204 nt 3′-UTR, and a 17-mer poly A region (GenBank accession no. AF306546). The protein is predicted to be composed of 716 amino acids including the methionine initiation codon. The protein is predicted to lack a signal peptide, to be composed of 12 transmembrane regions, which include five predicted extracellular-projected N-linked glycosylation sites, and intracellular N- and C-termini. The human homologue of rat BSAT1, SLCO1C1, maps to chromosome p12.2. There is 84% identity in the amino acid sequence of human and rat BSAT1.
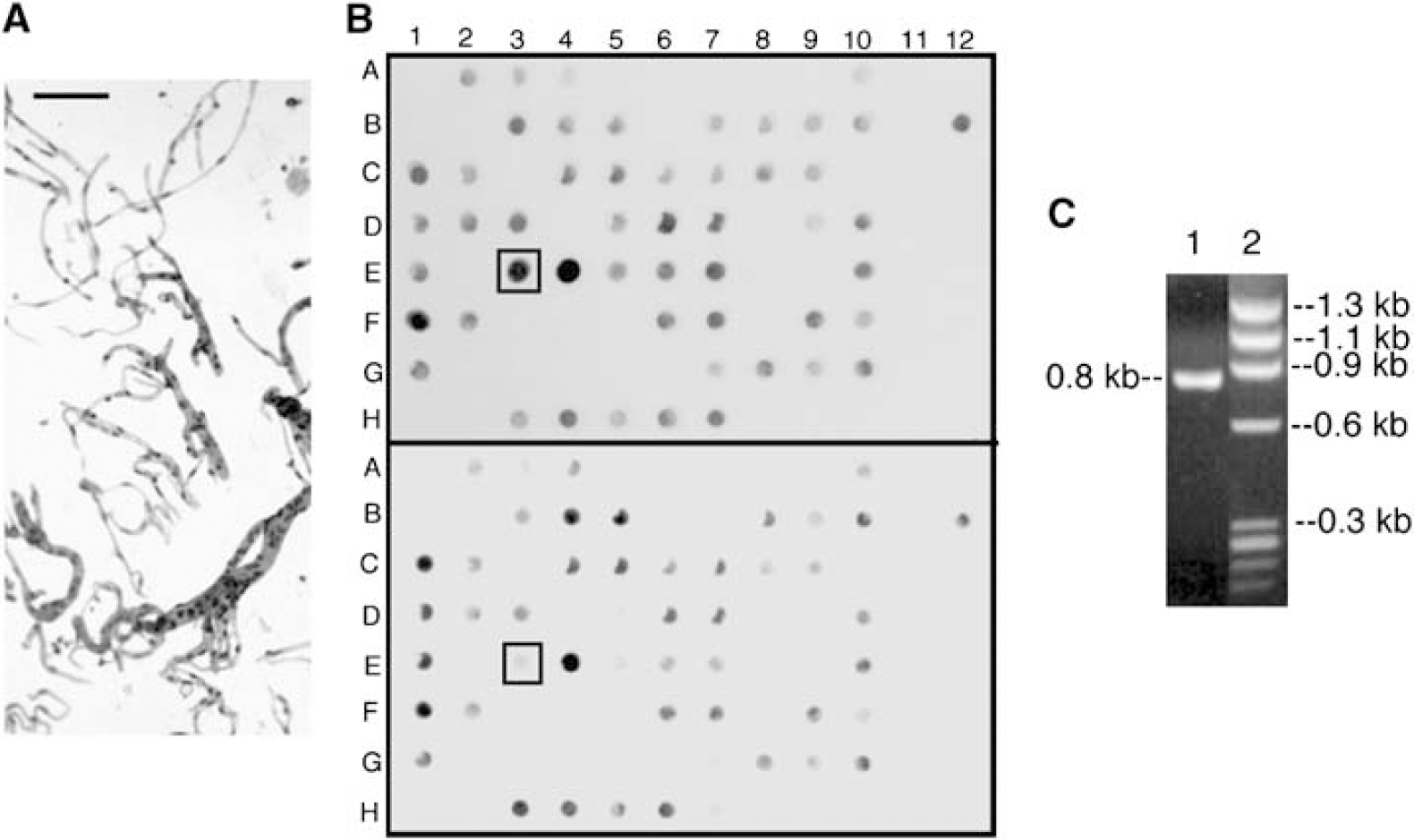
(
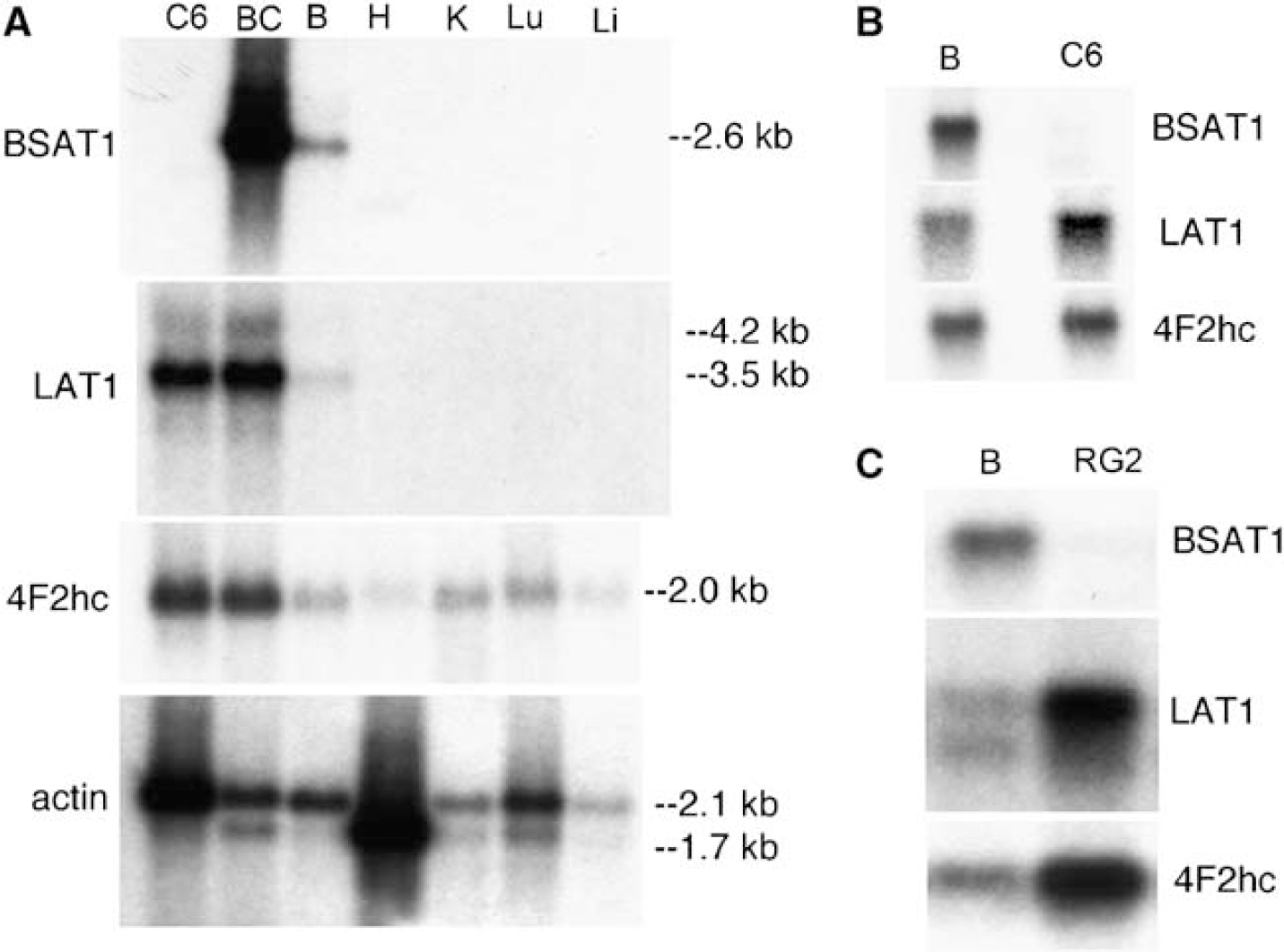
Northern blot analysis with poly(A+) RNA isolated from rat organs shows the BSAT1 mRNA is not detectable in rat heart, kidney, lung, or liver, whereas actin and 4F2hc mRNA is readily detected in these organs (Figure 2A). The BSAT1 mRNA is highly abundant at the rat BBB, and the signal is overexposed with a 16 h exposure of the film (Figure 2A). The low BSAT1 signal in whole rat brain suggests the transcript is preferentially expressed at the BBB, and this was confirmed by real-time PCR. The BSAT1 mRNA in the rat brain capillary compartment was enriched 19-fold relative to whole brain, whereas the β-actin mRNA content in both compartments was comparable (Table 1). The expression of the LAT1 is also confined to the BBB; however, LAT1 is expressed in C6 rat glioma cells, whereas BSAT1 is not expressed in cultured C6 cells (Figure 2A).
BSAT1 and β-actin mRNA in rat brain capillary and whole brain
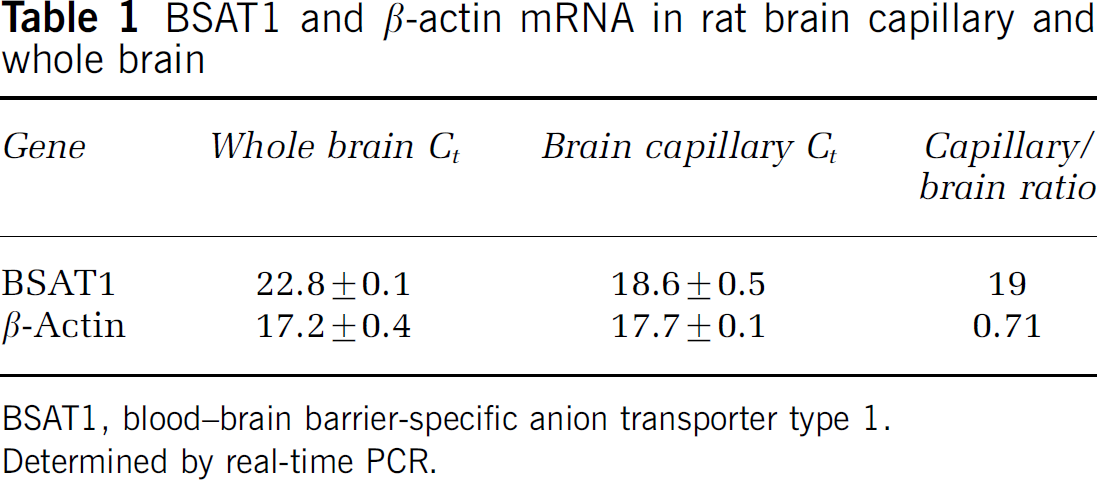
BSAT1, blood-brain barrier-specific anion transporter type 1.
Determined by real-time PCR.
(
Human 293 cells were permanently transfected with the pcDNA3.1 vector expressing either the rat BSAT1(orf), pcDNA/BSAT1(orf), or the full-length rat BSAT1 mRNA, pcDNA/BSAT1(fl), and [3H]E2G was used as a model organic anion substrate for both cell lines. There was minimal uptake of the [3H]E2G by 293 cells exposed to only lipofectamine, and the
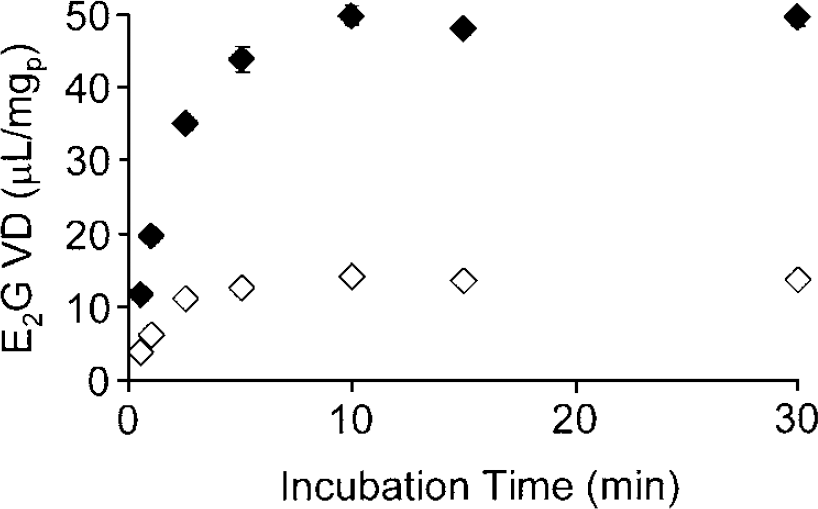
Time course of 3H-E2G uptake at 37°C in 293 cells permanently transfected with the pcDNA/BSAT1(orf) (closed diamonds) or the pcDNA/BSAT1(fl) (open diamonds). Data are mean ± s.e. (
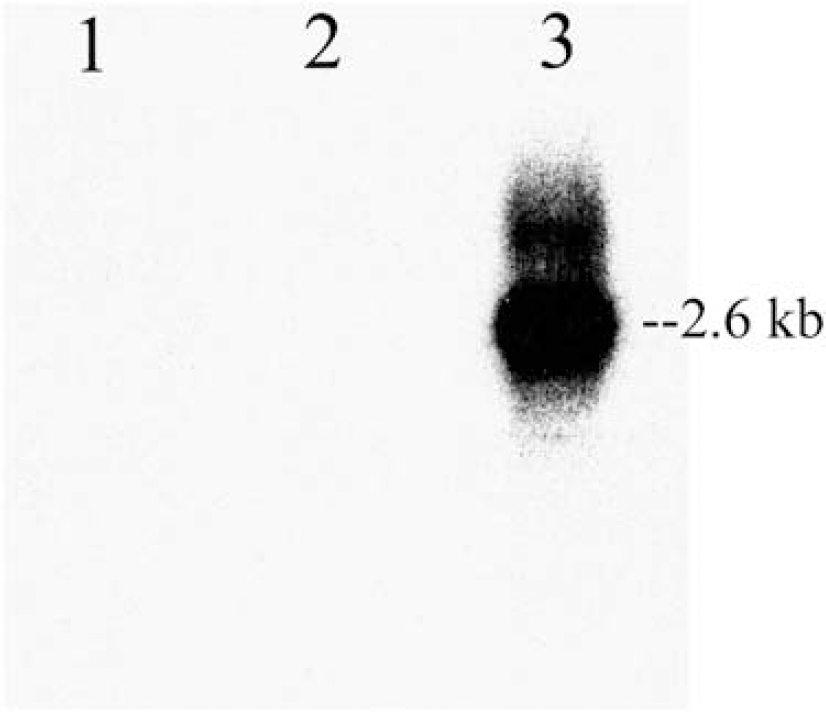
Northern blot of polyA(+) RNA probed with the 32P-rat BSAT1 cDNA. RNA was isolated from control 293 cells (lane 1), 293 cells transfected with the empty pcDNA3.1 vector (lane 2), and 293 cells transfected with the pcDNA/BSAT1(fl) (lane 3). The film was exposed to the blot for 16 h at −70°C.
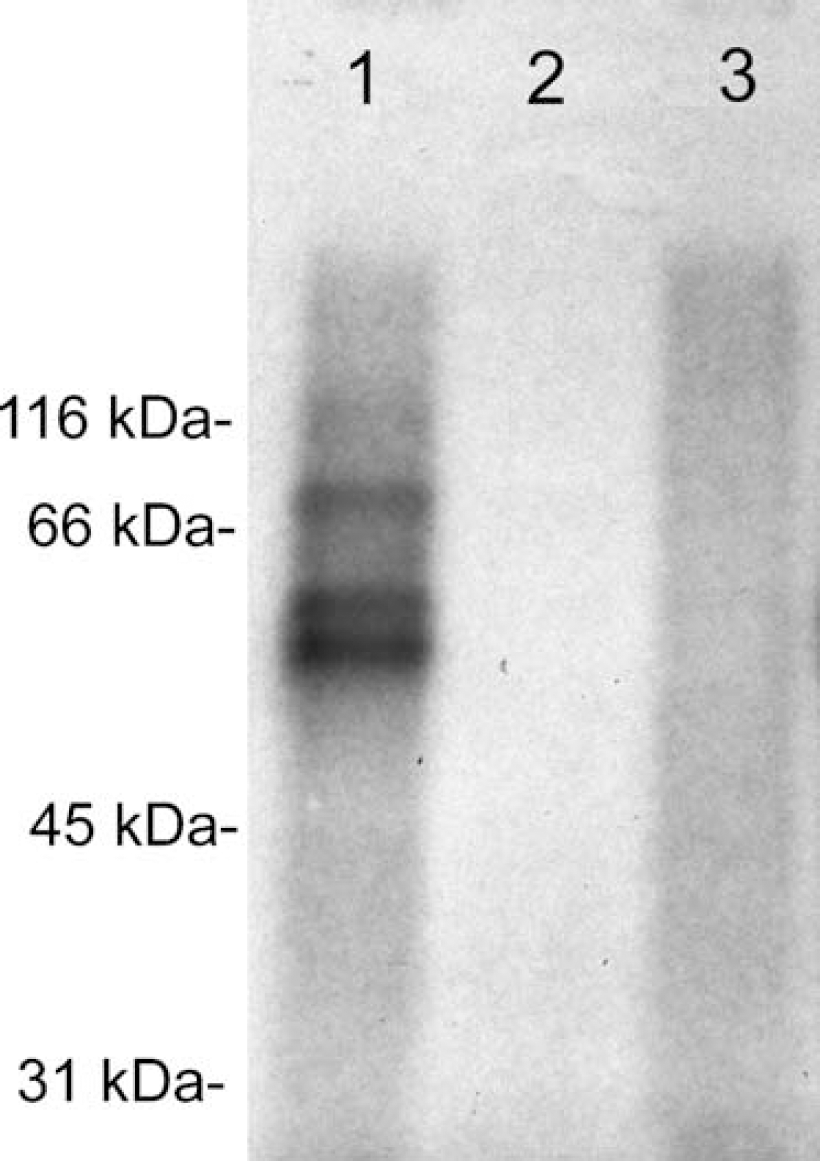
Ultraviolet light cross-linking of brain endothelial proteins to 32P-labeled full-length BSAT1 cRNA. RNA (2.0 × 105 c.p.m.) was incubated with 2 μg protein from either the P100 polysome fraction (lane 1) or the S100 cytosolic fraction (lane 2) of bovine brain capillary endothelial cells, or buffer (lane 3), for 10 mins at 30°C. After RNase T1 digestion, the cRNA/protein mixture was resolved by electrophoresis in a 10% sodium dodecyl sulfate-polyacrylamide gel electrophoresis gel. Film autoradiography of the gel shows RNAse protection by three proteins selectively localized to the polysome fraction of the cell.

ClustalW alignment of nucleotide sequence of 5′-UTR of human and rat BSAT1 mRNA.
The saturation of E2G influx was measured at 2.5 mins, which is within the initial rate part of the time course of cell uptake (Figure 4). The E2G influx into the 293/BSAT1(orf) cells was saturable, with
Summary of kinetic parameters of BSAT1-mediated transport into transfected 293 cells
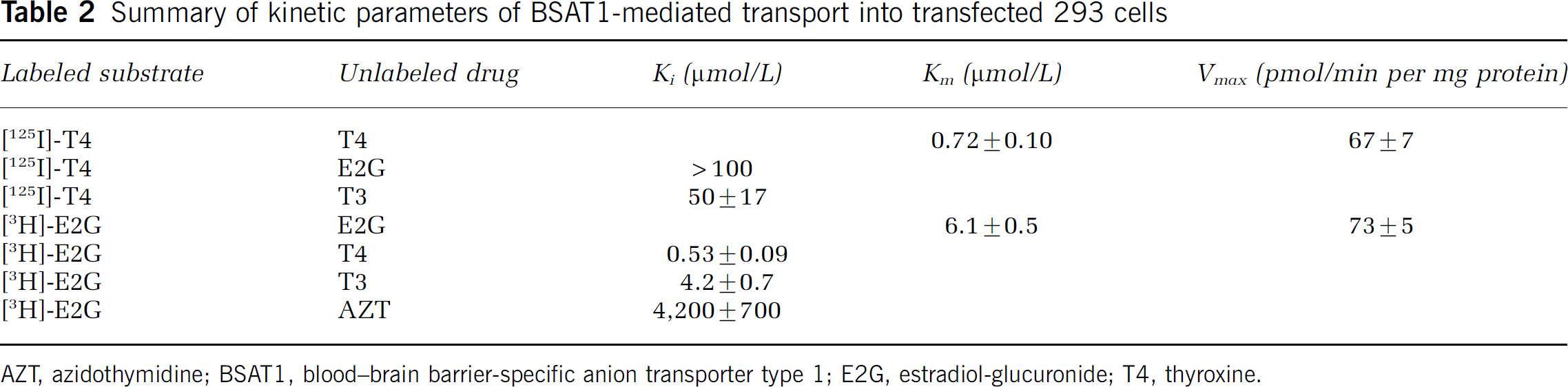
AZT, azidothymidine; BSAT1, blood-brain barrier-specific anion transporter type 1; E2G, estradiol-glucuronide; T4, thyroxine.
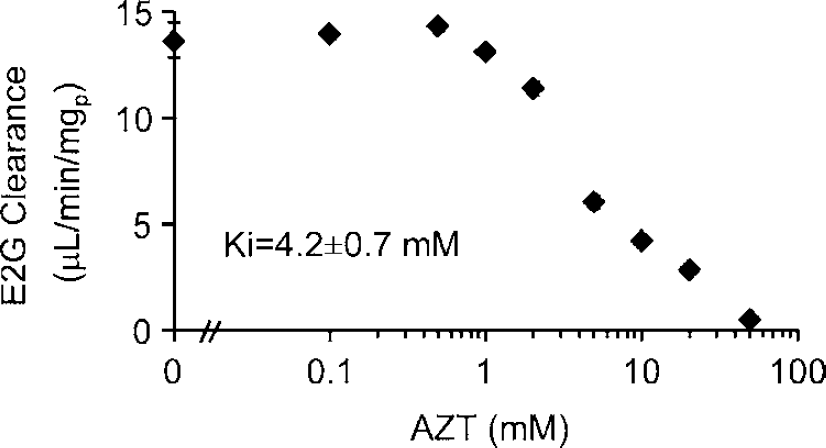
Competition of uptake of 3H-E2G transport by unlabeled AZT in 293 cells permanently transfected with the pcDNA/BSAT1(orf). Data are mean ± s.e. (
Using the prealbumin trap technique to monitor the efflux of [125I]T4 from 293-BSAT1(orf) cells, the efflux was rapid and characterized by a rate constantof 0.082 ± 0.007 min−1, which is equivalent to a half-time of 8.5 ± 0.7 min (Figure 9). Thyroxine efflux was accelerated in the presence of 50 μmol/L E2G in the medium, which showed an efflux rate constant of 0.286 ± 0.032 min−1, which is equivalent to a half-time of 2.4 ± 0.3 min (Figure 9). The identical efflux experiment was performed with 293 cells transfected with the empty expression plasmid, and medium E2G had no effect on T4 efflux from these control cells. The rate constant of T4 efflux was 0.109 ± 0.007 and 0.101 ± 0.009 min−1 in the cells exposed to either 0 or 50 μmol/L E2G, respectively.
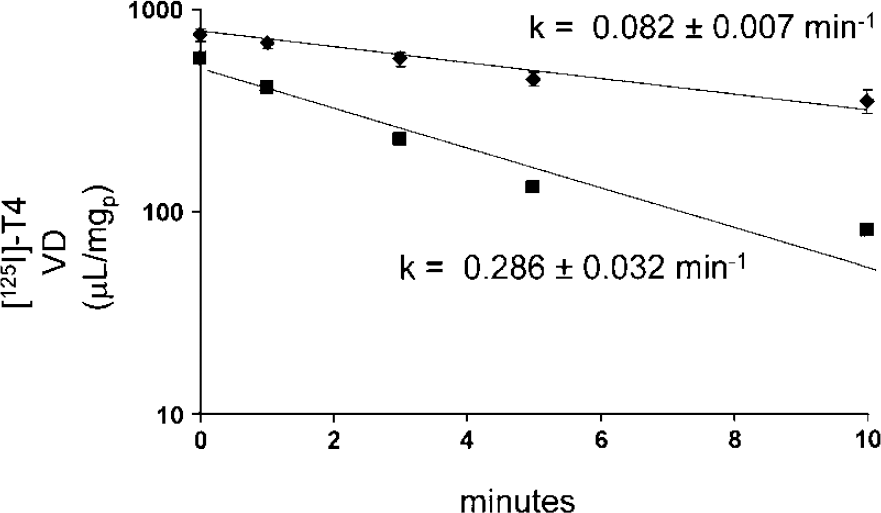
Thyroxine efflux from preloaded 293-BSAT1(orf) cells is monitored with the prealbumin trap technique (Materials and methods). The cells are exposed to fresh medium containing [125I]T4 for 15 mins, and either 0 (closed diamonds) or 50 μmol/L E2G (closed squares). The efflux is monitored starting with the addition of prealbumin to the medium at zero time, and the T4 efflux is linear for 10 mins. The addition of E2G to the medium accelerates the efflux of T4, as indicated by the 3.5-fold increase in T4 efflux rate constant.
Discussion
The results of these studies are consistent with the following conclusions. First, the SSH procedure allows for the rapid cloning of partial cDNAs corresponding to BBB-selective genes (Figure 1), and the availability of the brain capillary cDNA library allows for cloning of the full-length cDNAs corresponding to the BBB-selective gene. Second, one such gene, originally designated BSAT1 (Li et al, 2001a), is highly expressed at the BBB, and is not expressed in major rat peripheral tissues (Figure 2A). Third, unlike another BBB-selective gene, LAT1, which is expressed in rat glial tumors, the BSAT1 gene is not expressed in rat glial cells either in cell culture or as rat brain tumors (Figure 2B). Fourth, ISH shows the continuous microvascular immune staining (Figure 3), which indicates BSAT1 is expressed in the capillary endothelial cell. Fifth, the production of stably transfected 293 cell lines with the BSAT1(orf) results in the transport of organic anions, including E2G and T4 (Table 2). Sixth, the transporter mediates the efflux of T4 from cells, and this efflux is accelerated by extracellular E2G (Figure 9). Seventh, the expression of the transporter is suppressed in cells expressing the full-length BSAT1 mRNA as compared with the BSAT1(orf) (Figures 4 and 5), and this may be due to the interaction of the BSAT1 mRNA with specific polysome proteins (Figure 6), and conserved sequence within the 5′-UTR of the BSAT1 mRNA (Figure 7).
BSAT1 was originally isolated in a BBB genomics program using the SSH methodology (Li et al, 2001a). DNA sequence analysis indicated BSAT1 was an orphan transporter member of the oatp gene family. Sugiyama et al (2003) showed that E2G and T4 were ligands for BSAT1. This finding is confirmed by this study, which also shows a marked reduction in E2G transport via 293/BSAT1(fl) cells as compared with 293/BSAT1(orf) cells (Figure 4), despite high expression of the full-length BSAT1 mRNA (Figure 5). These observations suggest there is post-transcriptional regulation of BSAT1 gene expression, which produces low transporter in cells despite high mRNA levels. Such translational repression would be mediated by the binding of cellular proteins to specific sequences within the 5′- or 3′-UTR of the BSAT1 mRNA, similar to that previously showed for other BBB transporters, including the GLUT1 glucose transporter mRNA (Tsukamoto et al, 1996), or the LAT1 amino acid transporter mRNA (Boado et al, 2003b). The selective binding of brain endothelial polysome proteins to the full-length BSAT1 cloned RNA is showed in Figure 6. A triplet of polysome proteins bind to the BSAT1 RNA, whereas there is no binding of cytosolic proteins (Figure 6). The observation that there is minimal expression of the full-length rat BSAT1 mRNA in human 293 cells suggests there may be nucleotide conservation between the 5′-UTR of the human and rat BSAT1 mRNA. Blast2 analysis of the 374 nucleotide sequence of the rat BSAT1 mRNA 5′-UTR with the human genome indicates there is 77% sequence conservation between rat and humans approximately 350 nucleotides corresponding to nucleotides 8 to 350 of the rat BSAT1 mRNA 5′-UTR, where the methionine initiation codon starts at nucleotide 375 (Figure 7). Placement of cis-regulatory sequence within the 5′-UTR is supported by the observation that there is no phylogenetic conservation of BSAT1 mRNA sequence in the 3′-UTR of the transcript (Results), although the role of the 3′-UTR has not been investigated experimentally.
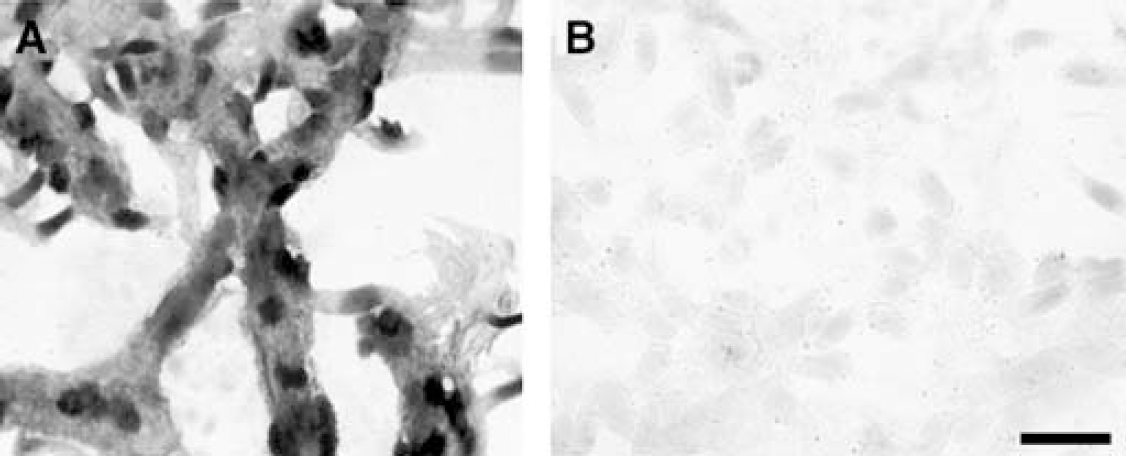
The hypothesis of translation repression of the BSAT1 mRNA may explain the unexpected finding of the very high level of BSAT1 mRNA in brain capillary endothelial cells (Figure 2). The high expression of BSAT1 mRNA at the BBB underlies the frequent detection of this transcript in BBB genomics programs (Li et al, 2001a; Enerson and Drewes, 2006). The level of the BSAT1 mRNA at the BBB is as high as any known transcript, and much higher than the level of the GLUT1 or LAT1 mRNA. It is hypothesized that the combined effects of a very high brain capillary endothelial level of the BSAT1 mRNA, in combination with translation repression, would allow for a very rapid activation of BSAT1 transporter activity via a post-transcriptional mechanism. The expression of the BSAT1 gene is highly focused to the BBB, as showed by the Northern blotting studies of rat organs in Figure 2A. The selective expression of the BSAT1 gene is further showed by the lack of BSAT1 gene expression in brain tumors (Figures 2B and 2C). The C6 and RG2 tumors are rat models of astrocytoma and glioblastoma multiform, respectively (Barth, 1998), and BBB-specific genes such as LAT1 are expressed in rat brain tumors (Boado et al, 1999). However, there is no BSAT1 gene expression in C6 or RG2 tumors (Figure 2). BSAT1 is expressed in the endothelium of the brain microvessel, based on the finding of continuous immunostaining in the ISH study (Figure 3).
The principal endogenous substrate of BSAT1 at the BBB is not known, although thyroxine has high affinity for this transporter (Tohyama et al, 2004), and these findings are extended in this study. E2G also is transported via BSAT1, and thyroxine inhibits E2G transport with a
BSAT1 is also an efflux system. The use of the prealbumin trap technique (Materials and methods) allowed for immediate restriction of influx of thyroxine, which is avidly bound by prealbumin. The prealbumin trap technique allows for accurate measurements of the kinetics of T4 efflux via BSAT1. The T4 efflux via BSAT1 is rapid with a half-time of 5.1 ± 0.2 min (Figure 9). Preloading the medium with unlabeled E2G accelerated the efflux of T4 from transfected cells by nearly fourfold (Figure 9). The design of the T4 efflux study takes advantage of the asymmetric kinetic properties of BSAT1 regarding T4 and E2G. The E2G in the medium has no effect on the T4 influx during the initial loading period, but the presence of E2G in the extracellular space accelerated the efflux of intracellular T4. These findings indicate that BSAT1 is a sodium-independent exchanger. The influx of E2G via BSAT1 accelerates the efflux of T4 via the transporter, since both ligands share binding sites on the same transporter protein.
There is no ultrastructural data that localizes BSAT1 to either the luminal or abluminal membrane of the brain capillary endothelial cell. However, it can be inferred that BSAT1 is an active efflux transporter at the BBB. This is because E2G is a high-affinity ligand for BSAT1 (Table 2), but sex steroid conjugates in blood do not enter brain (Pardridge et al, 1988). However, E2G is actively effluxed across the BBB from brain to blood (Sugiyama et al, 2001). Blood—brain barrier BSAT1 may function to mediate the efflux of iodothyronines, steroids, and other ligands that are conjugated via brain metabolism. In addition, BSAT1 may serve as a drug efflux system at the BBB. This study shows that AZT is transported via BSAT1 (Figure 8) with a
