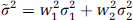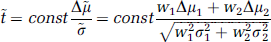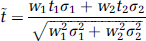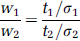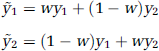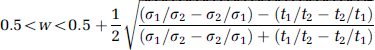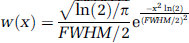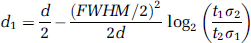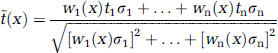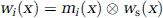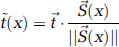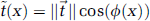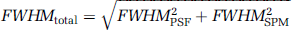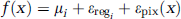Abstract
Voxelwise statistical analysis has become popular in explorative functional brain mapping with fMRI or PET. Usually, results are presented as voxelwise levels of significance (t-maps), and for clusters that survive correction for multiple testing the coordinates of the maximum t-value are reported. Before calculating a voxelwise statistical test, spatial smoothing is required to achieve a reasonable statistical power. Little attention is being given to the fact that smoothing has a nonlinear effect on the voxel variances and thus the local characteristics of a t-map, which becomes most evident after smoothing over different types of tissue. We investigated the related artifacts, for example, white matter peaks whose position depend on the relative variance (variance over contrast) of the surrounding regions, and suggest improving spatial precision with ‘masked contrast images’: color-codes are attributed to the voxelwise contrast, and significant clusters (e.g., detected with statistical parametric mapping, SPM) are enlarged by including contiguous pixels with a contrast above the mean contrast in the original cluster, provided they satisfy P < 0.05. The potential benefit is demonstrated with simulations and data from a [11C]Carfentanil PET study. We conclude that spatial smoothing may lead to critical, sometimes-counterintuitive artifacts in t-maps, especially in subcortical brain regions. If significant clusters are detected, for example, with SPM, the suggested method is one way to improve spatial precision and may give the investigator a more direct sense of the underlying data. Its simplicity and the fact that no further assumptions are needed make it a useful complement for standard methods of statistical mapping.
Introduction
Voxelwise statistical analysis has become popular in explorative functional brain mapping (Ashburner et al, 2003) and powerful tools for spatial normalization (Ashburner and Friston, 1999) and statistical analysis (Friston et al, 1995), including correction for multiple testing (Worsley et al, 1992; Friston et al, 1994) are publicly available. In the absence of an anatomically defined a priori hypothesis, statistical tests can be calculated for each voxel after spatial normalization and smoothing. Spatial smoothing is required to cope with interindividual functional anatomic variability that is not compensated by spatial normalization, and to improve the signal-to-noise ratio. Following the matched filter theorem (Rosenfeld and Kak, 1982) that states that the optimal smoothing kernel should match the signal to be detected, smoothing kernels of up to a FWHM (full-width at half-maximum) of 20 mm have been used. Smoothing with an FWHM of 10 to 15 mm is common in PET studies; often smaller kernels are applied in fMRI. The resulting t-maps (t = estimated parameter divided by its standard error) are masked at a certain threshold of voxel-level significance (often P < 0.001), and depicted as an overlay on corresponding anatomic sections, as maximum intensity projections or as surface projections. Those clusters that survive correction for multiple testing are frequently characterized by the coordinates of the maximum t-value and the associated ‘nearest gray matter’. Many of these techniques have originally been developed to detect cortical patterns of neural activation (e.g., with [15O]H2O PET or fMRI BOLD signal) and are now increasingly applied also in neuroreceptor/-transporter studies.
It is widely accepted that smoothing limits spatial selectivity and significant clusters are to be interpreted as regional results rather than anatomically precise information. However, little attention is being given to the fact that smoothing affects t-maps differently from ‘contrast images’ (contrast = linear combination of parameter estimates in the general linear model that is supposed to reflect the interesting physiologic parameter). While the latter change with FWHM in a way one might guess intuitively, t-statistics formed from the smoothed data are also affected by the nonlinear interaction of the filter kernel with the voxel variances, which becomes most evident after smoothing over different types of tissue (e.g., gray matter, white matter) with a different variability. Voxelwise maps of parameters such as binding potential have two main sources of variance–-measurement error (all sources of error associated with instrumentation, operator imprecision and the stochastic nature of isotope decay) and authentic between-subject physiologic variance. The physiologic variance can also be decomposed into two main sources–-variance from nuisance variables such as intersubject differences in nonspecific binding, and differences in receptor availability, the variable of interest. The variance attributable to between-subject differences in receptor availability can be considerable, leading to larger variance in gray matter tissues with high receptor density compared with other tissues, especially white matter. If, following smoothing, voxel by voxel statistical tests are performed, spatially inaccurate results may occur because the linear weighting scheme will redistribute voxel variances differently than it redistributes voxel intensities.
The aim of this paper was to demonstrate this effect and to suggest a way of combining t-maps, t-thresholds of significance and contrast images to ‘masked contrast images’ that can be used for presentation and that may allow for a more precise localization of significant effects than t-maps alone. The suggested algorithm was applied to data from a previously published PET study with [11C]Carfentanil that showed subcortical brain regions with increased μ-opiate receptor availability in abstinent alcoholics.
Materials and methods
Theory
Smoothing is an averaging process in which the intensity at a given voxel is replaced by a weighted average (i.e., a linear combination) of the values of voxels in some spatial neighborhood of that voxel. Common smoothing methods such as Gaussian filtering attribute the most weight to the (pretransformation value of) transformed voxel itself and are symmetric in the sense that the weights then decrease as a continuous function of spatial distance from the transformed voxel location, without any preferred direction (isotropic). A very simple example of such a scheme would consist of a bivariate statistic sampled from two subject groups (i.e., two variables per subject), but analyzed following a transformation such that for each subject, the transformed data associated with each variable is a linear combination of the original two variables. The weighted sum associated with one of the samples, say ỹ would be ỹ = w1y1 + w2y2 where yi and wi are the original samples and weights (i = 1, 2) with standard deviations σ1and σ2. If y1 and y2 are uncorrelated, the variance of ỹ is given by
A contrast, for example, a between-group difference Δμ can be tested by a t-statistic; the t-statistics on the transformed variable ỹ, can be expressed in terms of contrasts Δμ1 and Δμ2 on each of the original variables y1 and y2:
where const depends only on sample size. On replacing Δμi with tiσi/const, one obtains
Assuming t1, t2 > 0, there is a ratio of weights w1 and w2 for which
When the applied weights are proportional to 1/(relative variance), the maximal t-value will be associated with the variable having the smaller relative variance, and thus the ordering of the t-values may be reversed compared with their presmoothed values if the sample with the higher t-value is also the sample with the higher relative variance. The contrast itself is not similarly affected–-if Δμ1 > Δμ2 and w1 > w2 then the same ordering is preserved in the transformed contrast.
We now seek the range of weights that will lead to a reversal of t-statistic ordering. Given the symmetry of the smoothing kernel and appropriate normalization of the weights (i.e., w1 + w2 = 1), ỹ1 and ỹ2 can be expressed as
and, if y1 and y2 are uncorrelated, the corresponding t-values
Next, we extend the discussion to a setting in which the data is defined on a continuous spatial domain and smoothing has been performed. As commonly applied, Gaussian smoothing is linear, isotropic and stationary (independent on position), and so can be represented by convolution ỹ(x) = f(x) ⊗ w(x) with w(x) being the kernel function in the shape of a normal probability density function:
where FWHM denotes the ‘full-width at half-maximum’ of the smoothing kernel.
As a very idealized model for Gaussian smoothed data with two gray matter regions that are small compared with the FWHM of the smoothing kernel and surrounded by white matter with negligible absolute variability, let y1 and y2 be two point sources, separated by a distance d. With d1 being the distance from source one to a given voxel on the line segment between source one and source two, the distance of this voxel to source two is d2 = d–d1 and the weights w1, w2 attributed to source one and source two are given by equation (4). Together with equation (2), one obtains the position of the maximum t-value:
For a small FWHM, this is the middle of both point sources. With increasing FWHM, the maximum shifts towards the point source with the lower relative variance.
For imaging data, which can be thought of as a discrete lattice representation of a variable defined on an underlying spatially continuous domain, we expand equation (1) to account for n samples y1, y2 … yn that contribute to a smoothed voxel ỹ(x) (x = three-dimensional voxel coordinate) with the weights w1(x), w2(x), …, wn(x):
The assumption for this equation is that all original samples yi are uncorrelated, which, of course, is not the case for measured voxels. We therefore use yi to represent independent components of an image rather than voxels. For example, a homogeneous region i with the spatial extent being represented as a voxel mask mi(x) enters equation (6) as one single sample yi, and the weight wi (x) with which it contributes to a smoothed voxel ỹ(x) can be obtained from a convolution of the voxel mask with the smoothing kernel ws(x):
Note that the mi may overlap, corresponding to different regions sharing some component of signal; that is, a voxel x may be a part of one homogenous set of voxels with respect to some component of the signal, and a different set with respect to another component.
The t-value of a smoothed voxel can also be expressed by using the vector notation for equation (6):
where
where φ(x) is the angle between
For reasons of simplicity, we do not distinguish between the point spread function wPSF(x) of the scanner and an additional smoothing kernel applied during image postprocessing wSPM(x), since, in a typical voxelwise analysis, the former is much smaller and its contribution is negligible. However, one can easily account for both steps of smoothing by defining ws(x) as the combined kernel ws(x) = wPSF(x)⊗wSPM(x). Assuming wPSF(x) to have the shape of a bell-curve (as wSPM(x)), the full-width half-maximum of ws(x) is given by
One-dimensional Simulations
All simulations in this paper were calculated with matlab (Mathworks, Natick, MA, USA). To allow for Monte Carlo simulations with small sample sizes, we did not use equation (6) in the one-dimensional simulation, since equation (6) refers to the true population contrast and standard deviation Δμ and σ and not their estimates from small samples (here, the latter are denoted
Assuming uncorrelated Gaussian noise εpix(x) in each pixel (measurement error) and interindividual variability εreg i in each region, f(x) was calculated for two groups of ‘subjects’ (N = 2*1000) from
The corresponding group difference Δμ(x) and the standard deviation
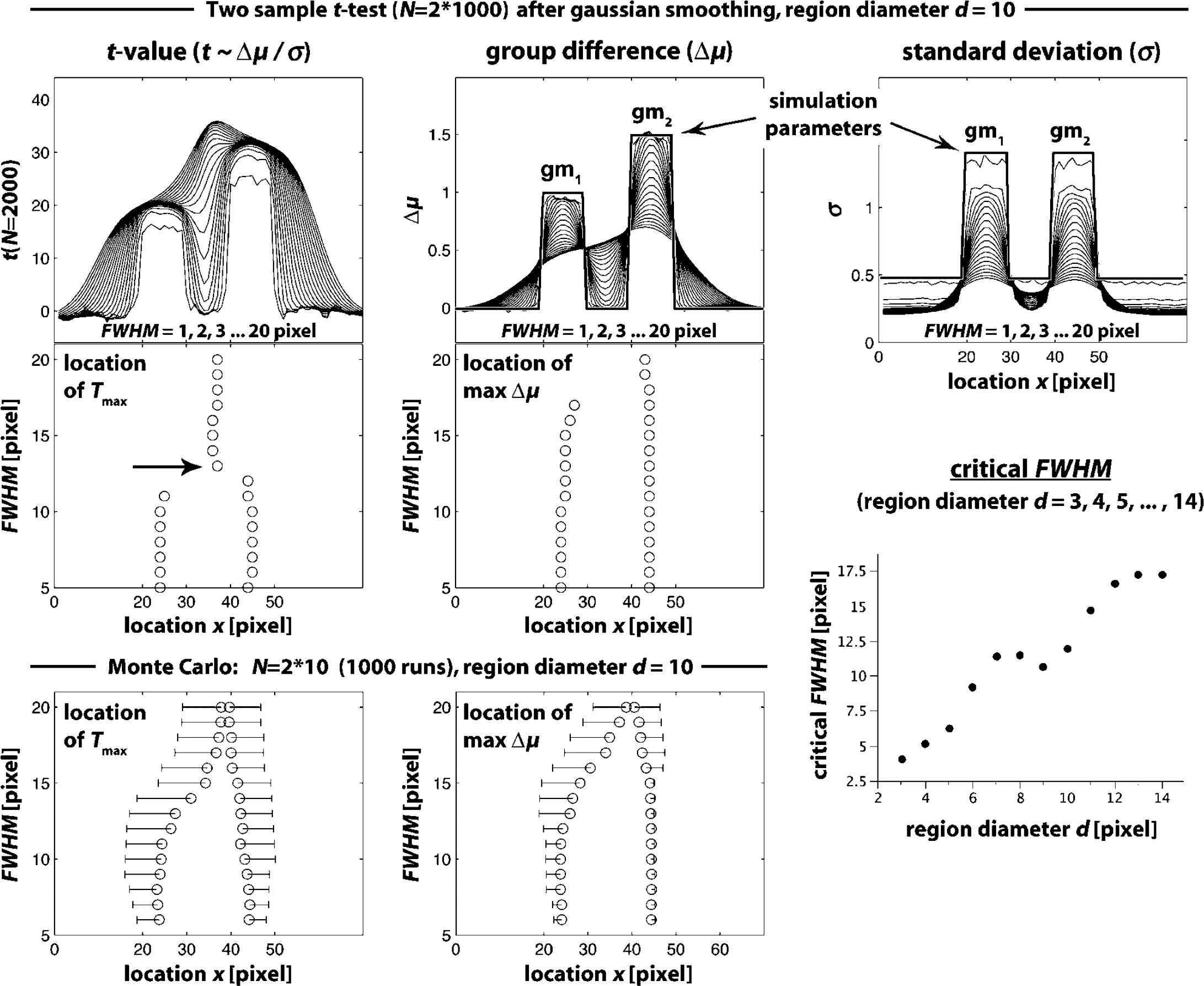
Effect of one-dimensional smoothing on t (x),
To further illustrate the artifacts in Figure 1, we chose some modified profiles, including the (idealized) assumption of σ = 0 in the white matter between gm1 and gm2. The corresponding Δμ(x) and σtotal(x) are shown in Figure 2. Unless mentioned explicitly, the parameters σreg and σpixwere the same as in Figure 1.
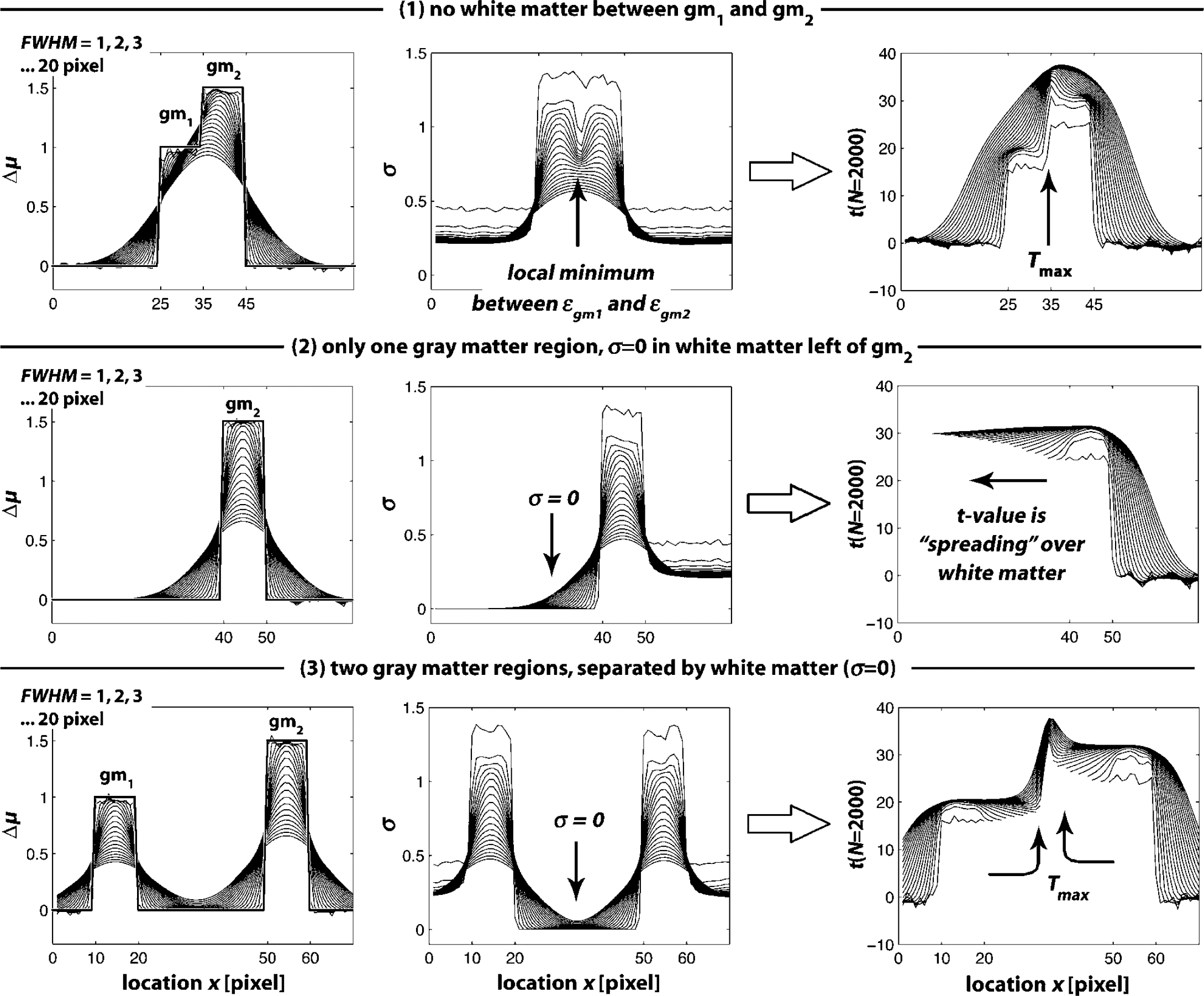
Effects from white matter. The simulation from Figure 1 was modified to illustrate the effects from white matter. Top row: without white matter, smoothing leads to a local minimum of
Two-dimensional Simulation
We used a two-dimensional model of three square gray matter regions (5 × 5 pixel each, Figure 3) and surrounding white matter with the same parameters σreg (between-subject variability) and σpix (measurement error) as in Figure 1. The group difference Δμ was zero in the gray matter region in the middle and was Δμ = 1 in the other two regions. With FWHM = 14 pixel, weights wi(x) were calculated from equation (7) for all ‘anatomic’ regions (three gray matter regions, one white matter region, standard deviation σreg) and additionally for each pixel (standard deviation σpix). The weighted standard deviations
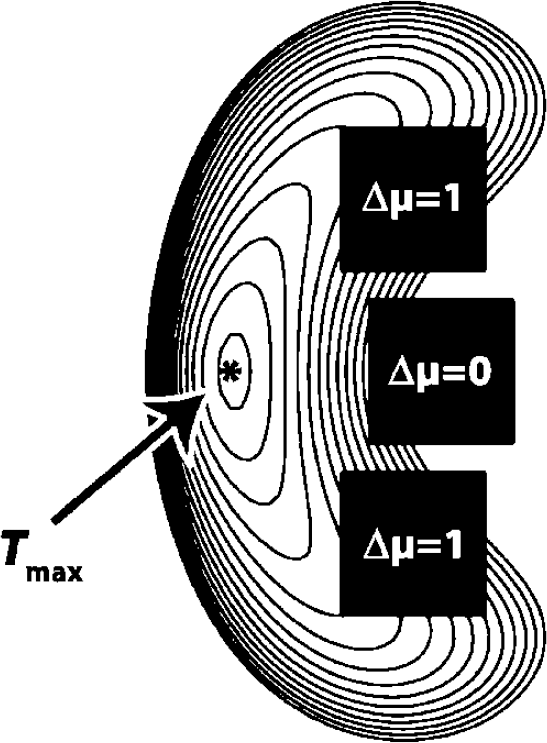
Two-dimensional simulation (worst case): t-isocontours from three gray matter regions (5 × 5 mm = 5 × 5 pixel) and surrounding white matter, smoothed with FWHM = 14 mm. All gray matter regions had the same variance, yet only the upper and the lower region had a positive contrast (Δμ = 1). Note that the nearest gray matter of Tmax is the region with Δμ = 0.
[11C]Carfentanil-PET
Measured PET data presented in this paper originate from a previously published PET study in abstinent alcoholics (Heinz et al, 2005). In 20 alcoholics, abstinent for 2 to 3 weeks, and in nine healthy control subjects, radioactivity distribution in the brain was measured with a GE Advance PET-scanner 0 to 66 min after intravenous injection of 700 MBq [11C]Carfentanil, a highly selective μ-opiate receptor ligand. Stereotactically normalized parametric images of receptor availability (Vʺ3) were calculated with Logan's linearization and the occipital cortex as a reference region with negligible specific binding. For details, see Heinz et al (2005).
SPM Analysis
With SPM2, Vʺ3 images were smoothed with a 12-mm Gaussian kernel, and a voxelwise two-sample t-test was calculated. Unlike previously published (Heinz et al, 2005), we did not mask out white matter. We applied a voxel-level threshold of P = 0.001 (uncorrected) and confirmed that both striatal suprathreshold clusters survived correction for multiple comparisons with SPM's small volume correction and a mask for the striatal volume of interest (12.1 cm3). The analysis was repeated with a voxel-level threshold of P =0.05 (uncorrected) to obtain the corresponding t-threshold for later use.
Masking Algorithm for Contrast Images
To use contrast images for presentation, just as t-maps with significant clusters being displayed over corresponding anatomic sections, a masking algorithm is needed. Instead of using the original t-isocontours, which are affected by the investigated artifacts, we used a corresponding contrast-threshold that we had calculated separately for each cluster.
The following data were available from the SPM analysis:
t-maps (file: spmT_0002.img). Contrast images (file: con_0002.img), containing the smoothed ΔVʺ3. A list of clusters (coordinates of Tmax) that survived SPM's correction for multiple testing. t-thresholds for P = 0.001 and P = 0.05.
The mask that was applied to SPM's contrast images was calculated as illustrated in Figure 4. For each significant region, we first obtained the original SPM-cluster from t-maps with a region growing algorithm that starts at Tmax and includes all voxels with a t-value above the original threshold (here P = 0.001). The mean contrast in this cluster was subsequently used for thresholding and all contiguous voxels that met both the ΔVʺ3 threshold and satisfied P < 0.05 (to ensure some statistical evidence) were included in the resulting region.
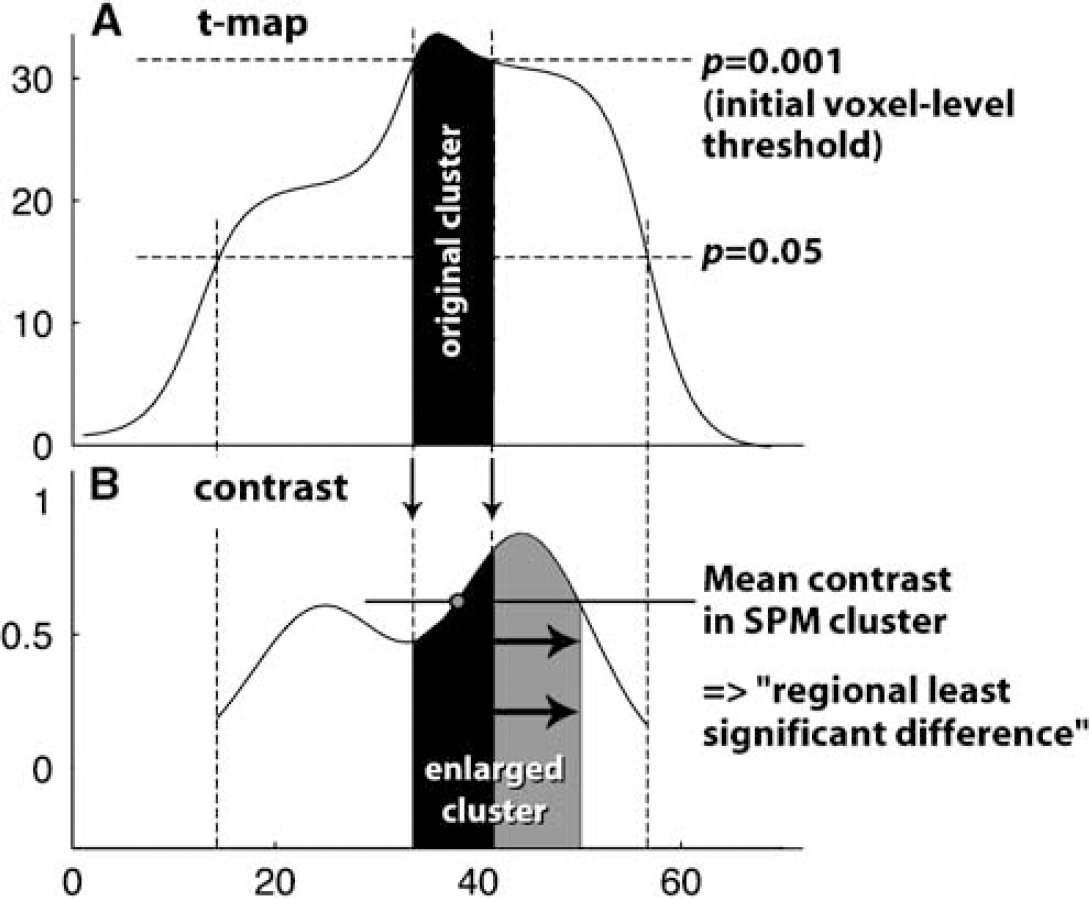
Suggested algorithm to create masked contrast images. (
Results
Simulations
The effect of Gaussian smoothing on t(x),
Results from three modified profiles are depicted in Figure 2:
Without white matter between gm1 and gm2 (Figure 2, top row), smoothing resulted in a local minimum in In white matter with a hypothetical variance σtotal = 0 (Figure 2, middle row) and a Δμ = 0, the standard deviation and When white matter with σtotal = 0 was surrounded by two gray matter regions (Figure 2, bottom row), smoothing resulted in a peak t(x) in the middle of gm1 and gm2, which first occurred when both ‘spreading’ t-values met exactly in the middle. For higher FWHM, Tmax shifted towards gm2 (the region with the lower relative variance). Interestingly–-unlike in the simulation without white matter (Figure 2, top row)–-interindividual variability εreg (in addition to the measurement error εpix) was not needed for this effect.
In the two-dimensional simulation (Figure 3), the maximum t-value was shifted to the left by the gray matter region in the middle (with Δμ = 0). Note that in the chosen setting, Tmax was still closer to the gray matter region in the middle than to the regions with a positive contrast.
[11C]Carfentanil PET
SPM analysis of [11C]Carfentanil PET-data confirmed significantly increased μ-opiate receptor availability in the bilateral ventral striatum. Without masking out white matter, the maximum t-value in the left hemisphere was found between ventral striatum and frontal cortex (Figure 5, top left), close to a local minimum of the unsmoothed ΔVʺ3. Masked contrast images (Figure 5, bottom) were more symmetrical, including more voxels in the middle of the left ventral striatum where the maximum ΔVʺ3 was found.
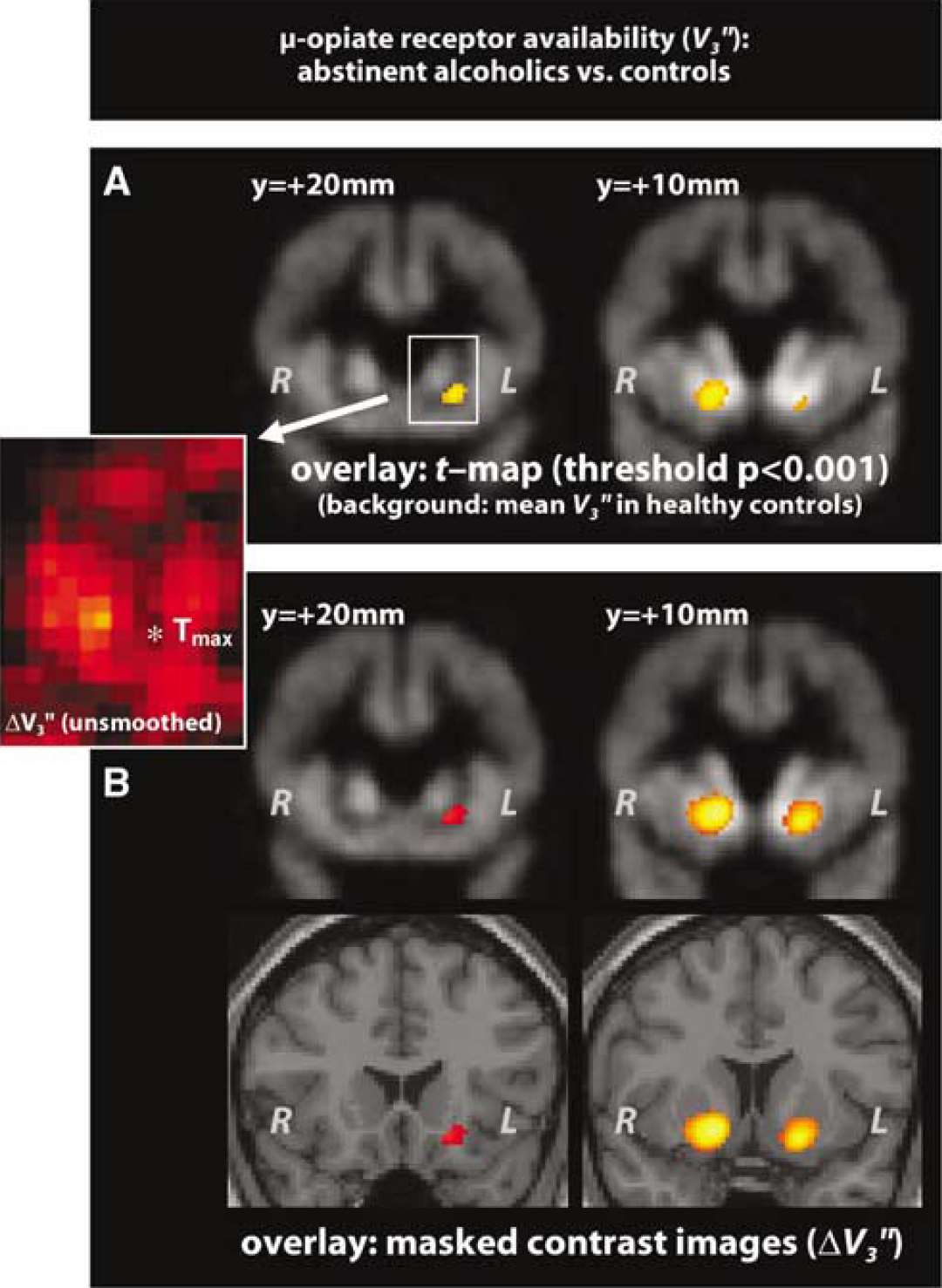
Results from a PET-study with [11C]Carfentanil. μ-opiate-receptor availability Vʺ3 in abstinent alcoholics was compared with that in healthy controls. (
In the right striatum, group differences were stronger and the standard deviation showed less local variation. Accordingly, masked contrast images were very similar to the original t-map.
Discussion
In addition to the loss of resolution that is inherent in spatial smoothing, the nonlinear interactions with the voxel variances (which affect t-maps, but not contrast images) can result in sometimes-counterintuitive artifacts, as demonstrated in this paper. These effects include a displacement of Tmax away from a region with a higher relative variance, and a displacement of Tmax towards the middle of two or more regions with a positive contrast and independent error terms. In the latter case, a displaced peak differs from a smooth, combined peak in contrast images in that it manifests at much lower levels of smoothing, and is spatially narrowed in a manner such that it may be mistakenly identified as an additional finding. The position of this peak depends on the relative variances of the surrounding regions so that the ‘nearest gray matter’ algorithm may not always determine the correct anatomic region.
This can be understood by looking at two effects:
smoothing causes the independent error terms of two or more neighboring gray matter regions to be averaged, resulting in a minimum variance between these regions white matter with little statistical noise and low absolute interindividual variability ‘inherits’ the t-value of adjacent gray matter, and peaks in t-maps may become broader than the corresponding peaks in contrast images. This not only leads to a higher uncertainty of the position of Tmax (as demonstrated in the Monte Carlo simulation), white matter also may serve as a ‘bridge’ between two or more interacting gray matter regions.
In our one-dimensional simulation, Gaussian smoothing with an FWHM in the same order of magnitude as the distance of the involved regions was already critical. Depending on the actual structure, this effect may even be stronger after three-dimensional smoothing. While such a t-shift is less likely to affect functional mapping of the cortex, it may seriously impair localization of subcortical findings, as for example, in the presented PET study with [11C]Carfentanil, where a maximum t-value was observed in the white matter between ventral striatum (where higher μ-receptor availability was expected) and the frontal cortex (where alcoholics also displayed a higher μ-receptor availability, though not significant). Smaller misplacements may just be an aesthetical concern or may raise the question of how to present the results in a convincing way. Larger misplacements may get masked out (white matter mask) at the expense of sensitivity. Findings may also go undetected when the Tmax shifts into a neighboring gray matter region with lower relative variance. Finally, misinterpretations may occur in brain areas with more than two gray matter regions, as in our two-dimensional (worst case) simulation, where the ‘nearest gray matter’ algorithm would not have detected the two regions that contributed to Tmax. Likewise, one has to be careful when assigning peaks to functional subunits, for example, of the thalamus or the ventral striatum.
In the literature, different smoothing kernels are usually compared with regard to the sensitivity of detecting significant regions (Hopfinger et al, 2000). Using a spatially stationary noise model, Worsley et al (1996) also describe the effect on the local characteristic of the t-map, notably a broadened peak between the two original maxima. However, the most striking effects, including a narrowly focused but shifted peak as in our simulations, only occur when estimating a local variance (Friston et al, 1991) instead of a pooled variance across voxels (Worsley et al, 1992), and when smoothing across different types of tissue. To our knowledge, these effects have not yet been investigated in the literature.
Several methods have been suggested to improve spatial precision of statistical maps. Clearly, reanalyzing the data with a smaller smoothing kernel (Worsley et al, 1996) would have reduced the described artifacts, but at the expense of statistical significance. Another approach is to incorporate spatial a priori assumptions. Davatzikos et al (2001) have described an atlas-based adaptive smoothing to avoid smoothing across anatomic boundaries, which theoretically prevents the described artifacts. However, including a priori information either increases the complexity of the analysis or limits the advantages of a voxel-based (no a priori assumptions) over traditional ROI analysis. Accordingly, ‘classical statistical maps’ (Penny and Friston, 2004) are still the most popular approach for explorative imaging.
While all established methods have in common that the reported values are statistical measures related to a level of confidence by which a ‘true effect’ was observed, the method suggested in this paper follows a different rationale. We do rely on established methods of statistical mapping (in particular those implemented in SPM2, however, other methods as a starting point are also possible) to detect significant regions, but once a cluster is considered significant, we suggest attributing voxelwise color-codes to the estimated parameter itself instead of the level of confidence by which it is greater than zero.
The limitations of the presented method are those of any voxel-based analysis. They are powerful tools for explorative imaging, but they are not necessarily a replacement for traditional region-of-interest (ROI) analysis when a regional a priori hypothesis exists and when an ROI definition is anatomically and physiologically justified. Comparing voxel-based with ROI-based analyses is not subject of this paper, but it should be mentioned that the proposed method was primarily developed for assessment of subcortical regions for which an ROI analysis may indeed be a more straightforward approach. In voxel-based analyses, the type I error must be reduced by using a rather conservative voxel-level threshold (e.g., P < 0.001), which may result in a loss of statistical power. Applying such a single t-threshold as a first step has shown to be a simple and powerful method to detect significant clusters (e.g., by considering the cluster size). However, such a t-threshold is not necessarily the best choice to determine the outline of a neurobiologic effect, which leaves room for other methods of masking such as the one presented in this paper. Using the contrast for color-coding, it may seem natural to use it also to define the outline of a displayed region. Indeed, we have shown that this may provide further information about the locus of, for example, a group difference, albeit in an exploratory form outside the framework of hypothesis testing. It should be noted that our masked contrast images differ from the original thresholded t-maps only if there is a local variation in the variance of the smoothed voxels. The suggested method can therefore be understood as an attempt to stick closely with the current standards while removing the described smoothing artifacts.
We believe that adding a further step to the analysis by going from thresholded t-maps to masked contrast images does not substantially increase the overall complexity of the analysis. On the contrary, it may give the investigator a more direct sense of the underlying data and may improve spatial precision when a significant region occurs. The simplicity of the suggested method and the fact that we do not make further assumptions make it a useful complement for established methods for statistical mapping.
Footnotes
Appendix A
We seek the values of w, 0.5 < w < 1, such that
Note that DIF(0.5) = 0, and DIF(1) = t1 – t2 > 0, and DIF is a differentiable function of w. The derivative, evaluated at w = 0.5 is
and this will be negative (i.e., DIF(w) will be negative immediately to the right of w = 0.5) when σ1/t1 > σ2/t2.
Therefore, if this condition pertains and DIF(w) has exactly one root in the open interval (0.5, 1), it must be the case that DIF is negative between 0.5 and the root, and positive between the root and 1, and the ordering of the t-map will be reversed compared with the original values on the interval 0.5 < w < root.
We therefore seek to show that DIF(w) has exactly one root between 0.5 and 1. DIF = 0 is equivalent to
with
This leads to a cubic equation in w (the 4th power term cancels). Noting that 0.5 is a solution of equation (A1), the remaining two solutions of equation (A1) are the roots of the quadratic
These roots, in terms of the original variables, are
Finally, given the conditions on t1 t2,