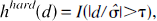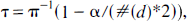Abstract
A new method is introduced for the analysis of multiple studies measured with emission tomography. Traditional models of statistical analysis (ANOVA, ANCOVA and other linear models) are applied not directly on images but on their correspondent wavelet transforms. Maps of model effects estimated from these models are filtered using a thresholding procedure based on a simple Bonferroni correction and then reconstructed. This procedure inherently represents a complete modeling approach and therefore obtains estimates of the effects of interest (condition effect, difference between conditions, covariate of interest, and so on) under the specified statistical risk. By performing the statistical modeling step in wavelet space, the procedure allows the direct estimation of the error for each wavelet coefficient; hence, the local noise characteristics are accounted for in the subsequent filtering. The method was validated by use of a null dataset and then applied to typical examples of neuroimaging studies to highlight conceptual and practical differences from existing statistical parametric mapping approaches.
Keywords
This article concludes a trilogy of papers devoted to the use of the wavelet transform (WT) for the spatial modeling of images obtained with emission tomography (ET). The first of the series (Turkheimer et al., 1999) described a unified framework for the analysis of single emission images and derived statistical maps in stationary noise conditions (homogeneous variance). The second article (Turkheimer et al., 2000) considered the spatio-temporal modeling of dynamic ET studies and developed a combined application of kinetic modeling and WT that allowed the optimal estimation, in the least square sense, of parametric maps in nonstationary noise conditions (heterogeneous variance).
This article builds upon this bulk of work and develops a framework for the statistical analysis of multiple ET images in nonstationary noise conditions. As it will be clear in the next sections, the theoretical basis of such a method already resides in the previous two manuscripts. Therefore, the purpose of the following sections is to clarify the use of the WT for this particular application and highlight the conceptual differences between current approaches in image space and WT methods. This task is pursued by theoretical means and applications to real datasets.
Let Y (s, i) be a set of ET scans indexed by i, where s:s = (sx, sy, sz)εN3 spans the pixels of the digitized images. In general, i:i = (k, h, j) εN3 as Y (s, i) may be measured in different conditions (index j) in different subjects (index h) in repeated occasions (index k). Y (s, i) are acquired from emission tomographs and reconstructed from sinograms or planar projections at their highest resolution using back-projection with a ramp filter. It is useful to consider Y (s, i) as projected in the same anatomic space so that each spatial coordinate s will correspond to the same anatomic location for all images in the set.
The measurement of the set Y (s, i) was aimed to obtain inferences on the spatial distribution P (s) of a certain parameter P that is a function of the conditions under which the data-sequence was acquired. For example, in cerebral blood flow (CBF) studies of brain function, the pattern of interest P(s) is the difference in CBF between baseline and an activation condition. This difference can be estimated by use of a general linear model (GLM) in which the effect of each condition is inscribed as an additive term and other effects of no interest (subject effects, temporal effects, scans global counts, and so on) are also included as additive terms or covariates (Friston et al., 1995).
The mathematic relation between the data and the effect of interest will be condensed here into a function labeled κ(). κ() in ET is usually derived from linear models (ANOVA or ANCOVA) where the set Y (s, i) is modeled in terms of conditions and covariates related to I = (k, h, j) (Friston et al., 1995).
Alternatively, the model for the sequence Y (s, i) can be extracted from the data themselves in the form of, say, a set of eigenvectors extracted from a Principal Component Analysis (PCA;Barber, 1980; Moeller et al., 1987; Friston et al., 1993); in this case, the pattern P (s) of interest is the regional distribution of the contribution of each eigenvector to the pixel variance.
The application of κ() models the dependency of the data from the experimental setting and accommodates the dimension of Y (s, i) indexed by i. However, a complete modeling approach aims to the estimation of the parameter of interest, in this instance the pattern P (s); to do this, all dimensions of the data must be considered. Therefore it is also necessary to model the spatial dimension indexed by s.
Fig. 1 sketches a common approach to solving such a problem in neuroimaging. In this scheme, spatial modeling is approached by two separate steps, filtering and statistical thresholding. Filtering commonly consists of the application of monoresolution filters (for example, gaussian filters) to produce the filtered set YF(s, i). However, such a simple modeling approach does not allow any kind of control over the statistical properties, bias and variance, of the derived map P̂ (s) = κ(YF(s)). Therefore, such a map is usually accommodated in the form of a statistical parametric map (SPM) in which the parametric pattern P̂ (s) is normalized by correspondent estimates of its standard errorσ̂(s) to obtain a map of statistical scores (for example, t scores, z scores, and so on) (Friston et al., 1991; Worsley et al., 1992). This is preliminary to the subsequent thresholding procedure that consists of feeding the statistical scores into the appropriate cumulative distribution function be ϕ() that allows the control of the statistical risk of false positives in the entire SPM. The correspondent P value map is then derived as P̂p(s) = 1 – ϕ(P̂ (s), σ̂(s)). Those coordinates s where P̂p(s) > τ (τ is an operator selected threshold, usually 0.1, 0.05, and so on) are then identified as those locations where parameter P has a statistically significant value. To capture varying patterns of P (s), a number of ϕ() have been proposed that are sensitive to spatial distributions of the signal that range from focal and sparse to nonfocal and distributed (Friston et al., 1994; Worsley et al., 1995). Alternatively, such sensitivity may be achieved by iterating the filtering and thresholding steps at different resolutions where ϕ() is derived to obtain control of statistical risk over the searched scales (Poline and Mazoyer, 1994; Worsley et al., 1996).
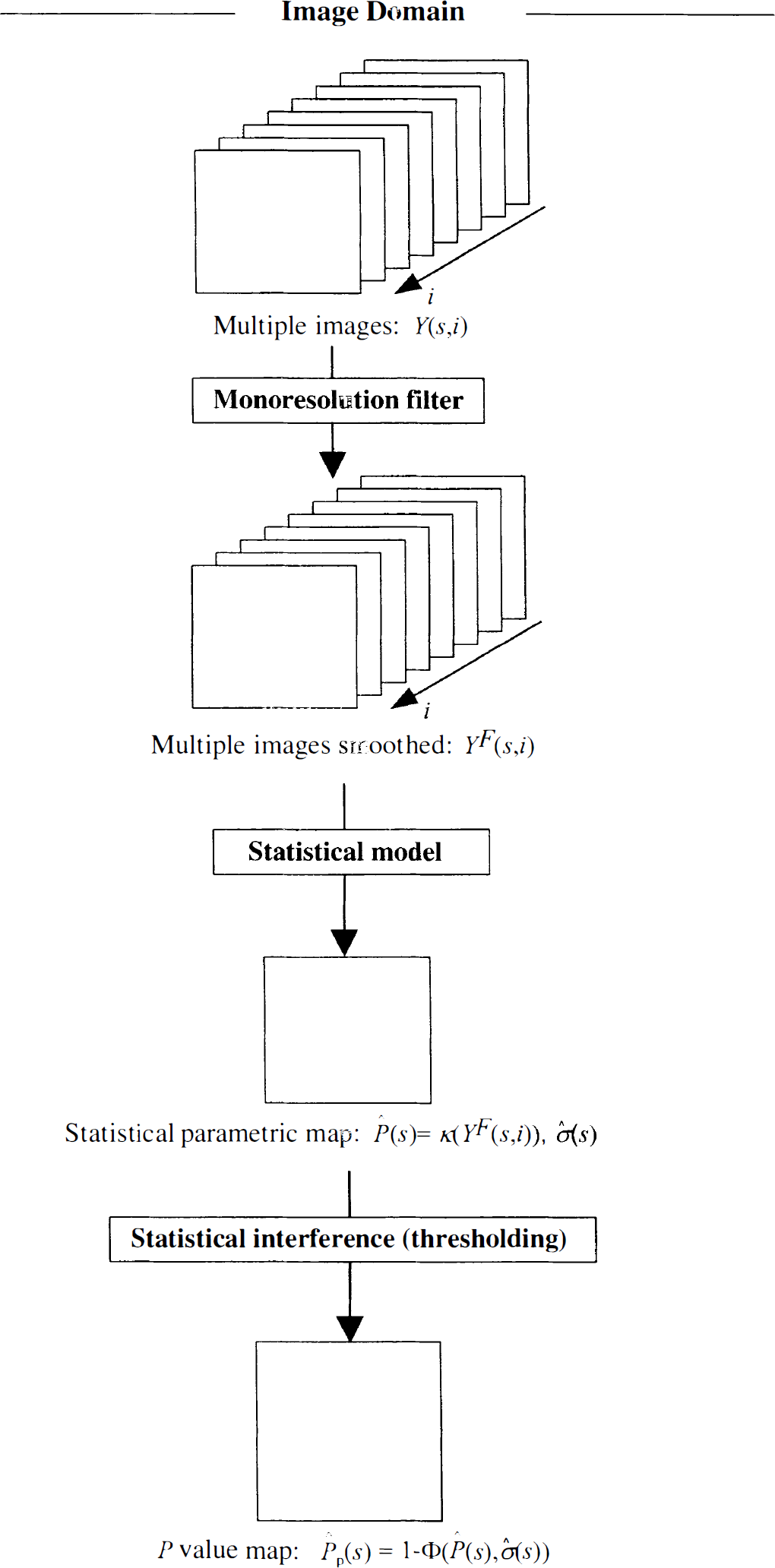
Scheme describes a common approach for the analysis of neuroimages. Y (s, i) (represented as bidimensional set of images indexed by i) represents a set of multiple images acquired in various conditions and all normalized to the same anatomic reference. Frames Y (s, i) are smoothed with a monoresolution filter to produce the filtered set YF(s, i). A functional κ() is then applied to produce the parametric pattern P̂ (s) and correspondent standard error σ̂(s). It is common to obtain P̂ (s) from the pixel-wise applications of ANOVA or ANCOVA linear models and to express it as a statistical map where each value of the parameter of interest is normalized by correspondent values of σ̂(s). These statistical scores are then inserted in an appropriate cumulative probability distribution function that allows the control of the rate of false positive in the volume of interest. The output is a probability map P̂p(s). Those locations where P̂p(s) is under a predefined threshold are then defined as those anatomic coordinates where pattern P̂ (s) has a statistically significant value.
The scheme of Fig. 1 was designed to obtain an estimate of the pattern P̂ (s) with a controlled statistical risk. However, the final output is a probability map and not the pattern itself. Such a map may be used to assess the location of statistically significant values of P (s), but not to obtain a numeric estimate of its spatial distribution (Worsley, 1997; Turkheimer et al., 1999). The reason for this lies in the incomplete modeling of the data: whereas the dimension i of Y (s, i) is modeled by κ(), no model is used for the spatial dimension s. The use of a monoresolution filter is suboptimal as it cannot adapt to the spatially varying features of the signal nor to the local signal-to-noise ratios.
One general approach to the modeling of images consists of the use of a mathematical transform that projects the data onto an appropriate functional space. Previous work (Unser et al., 1995; Ruttiman et al., 1996, Turkheimer et al., 1999) showed that bases of smooth wavelets are able to model efficiently the signal of images measured with positron emission tomography (PET) and single photon emission computed tomography (SPECT). Besides, the projection onto wavelet space is obtained through application of the dyadic wavelet transform (DWT;Mallat, 1989) that is a linear and orthogonal operator. Such properties therefore can be used for the spatial modeling of the dataset Y (s, i) by using the method shown in Fig. 2. In this scheme, every single frame of the dataset is transformed by application of the wavelet transform to obtain W (w, i) = WT(Y (s, i)). The application of the statistical model through the operator κ() produces the parametric image in wavelet space P̂wt(w) and correspondent estimates of the standard error σ̂wt(w). Both P̂wt(w) and σ̂wt(w) then are passed through a wavelet filter h () that thresholds each wavelet coefficient P̂wt(w) proportionally to its error σ̂wt(w) and accordingly to a certain statistical risk (Turkheimer et al., 1999). By application of the inverse WT one can obtain a final estimate of the parametric pattern P̂F(s) = WT−1(P̂Fwt(w)).
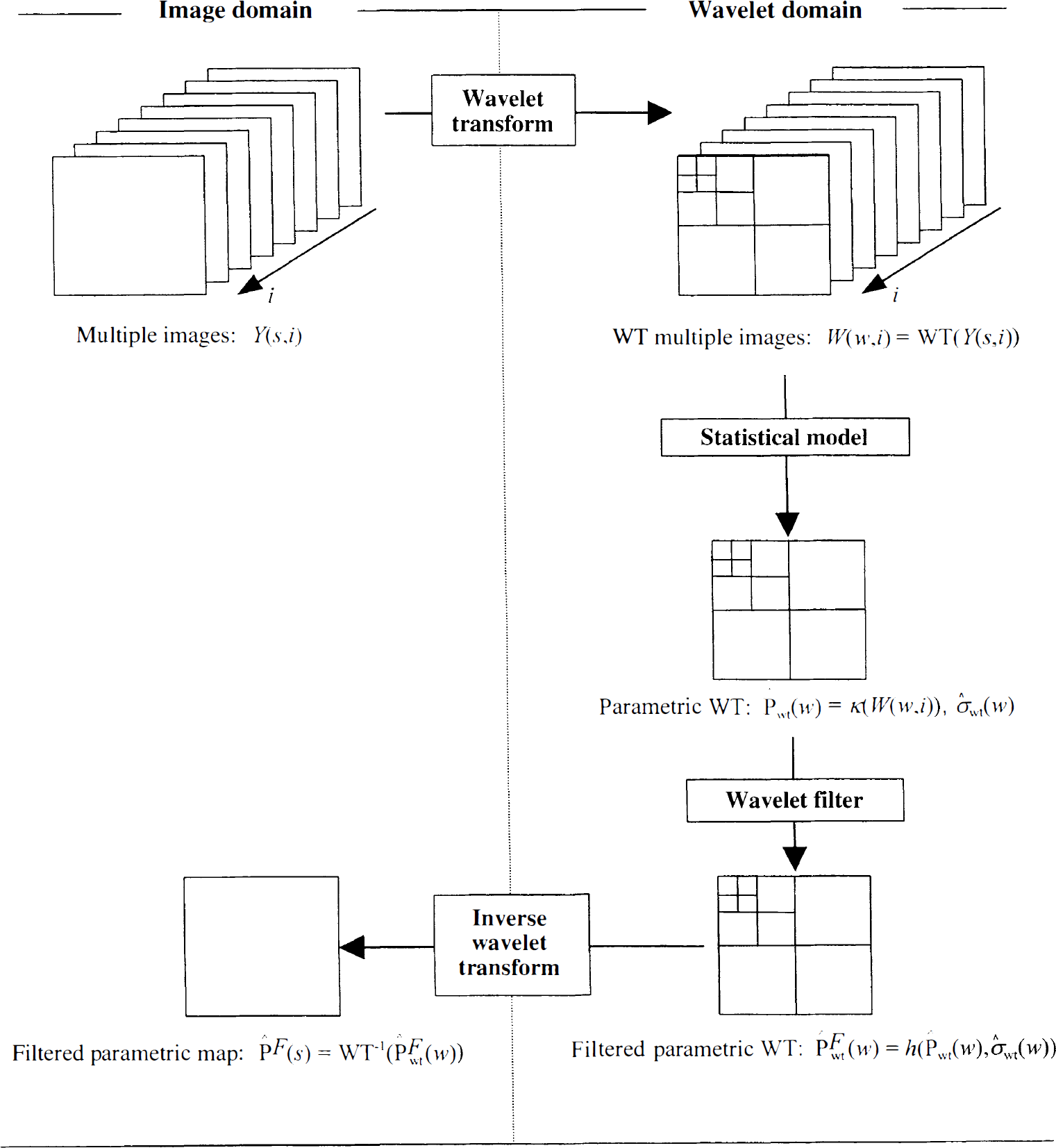
Solution to the problem of estimation of P̂ (s) with a controlled statistical risk. All frames Y (s, i) are projected in wavelet space through the dyadic wavelet transform. A pattern P̂wt(w) and correspondent error estimate σ̂wt(w) are obtained by application of an appropriate linear statistical model in wavelet space. A wavelet filter h () is then applied to P̂wt(w) that shrinks each wavelet coefficient proportionally to its error by use of a specified statistical risk. The filtered map P̂Fwt(w). is then inverted in image space to produce the final estimate P̂F(s).
This sequence is identical in all aspects to that used previously for the estimation of parametric maps from dynamic ET scans (Turkheimer et al., 2000). In fact, the application of a statistical model to a series of ET studies is in no way different from the application of any other model (for example, a kinetic one) to a sequence of tomographic images. In both cases, the final aim is the estimation of the pattern of the parameters of interest.
The kinetic modeling of a dynamic ET acquisition is used to recover the spatial distribution of physiologic indexes such as blood flow, metabolism, receptor density, and so on (Phelps et al., 1986). The statistical modeling of a series of PET images is used to investigate the pattern either of variables (factors and covariates) relating to the conditions under which the scans were made, or of linear combinations (contrasts) of such variables.
The only difference between the analysis of dynamic scans and the analysis of multiple scans is the statistical risk that characterizes the choice of filter h (). If P (s) is the pattern of a physiologic parameter one may be more interested in the mean squared error (MSE; the mean squared error is equal to the sum of the variance and squared bias) properties of the final estimate (Turkheimer et al., 2000). Instead, when P (s) is the pattern of a certain measurement related to a certain experimental condition then one is usually more interested in the control of the variance of P̂F(s).
A number of wavelet filters exist in the statistical literature for each risk of choice. For example, the SURE filter (Donoho and Johnstone, 1995) has excellent MSE properties. However, variance control can be exerted by minmax filters (Donoho and Johnstone, 1994; 1995) or by thresholding approaches that adopt multiple testing corrections like the Bonferroni (Turkheimer et el., 1999).
The scheme in Fig. 2 already has been discussed in detail (Turkheimer et el., 2000). The previous manuscripts (Turkheimer et al., 1999; 2000) also contain background material on wavelet bases, DWT, and filtering procedures in wavelet space. However, for the context of this article, it is relevant to review the following aspects of the methodology.
First, the scheme illustrated in Fig. 2 is a comprehensive modeling approach that uses appropriate functional support for dimensions s and i. In fact, the outcome P̂F(s) is a parametric map and not a map of probabilities. Second, if the functional κ() is linear then, given that the DWT is a linear and orthogonal operator, a solution P̂Fwt(w) that is optimal under a certain statistical risk in wavelet space will produce an inverse P̂F(s) that is also optimal under the same risk in image space (Turkheimer et al., 2000). Therefore, such procedures may have extensive use in the analysis of ET images where linear models, such as GLM or PCA, are common. Third, noise processes in wavelet space are spatially uncorrelated given the well-known whitening properties of the DWT (Turkheimer et al., 1999; 2000). In this particular context, this means that to control the variance of the solution any standard multiple hypothesis testing approach can be efficiently used (Ruttiman et al., 1996). Notably, in image space, the control of error rates is more difficult because the spatial correlation of the original images, or the one introduced by the filter, has to be taken into account; consequently, the methods rely on Gaussian random fields theory (Adler, 1981) or randomization procedures (Poline and Mazoyer, 1994; Holmes et al., 1996).
Finally, the scheme of Fig. 2 allows the point-wise estimation of the noise process in wavelet space σ̂wt(w) and is therefore able to handle the computational process in nonstationary gaussian noise conditions typical of this type of study (Turkheimer et al., 2000).
Filters that control the false positive rates were selected for the application section of this article to mirror common thresholding strategies in image space. The two properties of risk equivalence between image and wavelet space and of noncorrelation of statistical scores indicate the use of a simple filter in the form of hard-thresholding operator defined, for a wavelet coefficient d with error σ̂, as
where I () is the indicator function. The threshold τ is derived from the Bonferroni adjustment for multiple tests as (Turkheimer et al., 1999):
where α is the desired probability of at least one false positive in the set (for example, 0.5, 0.01, and so on), #() is the cardinality of the set (number of non-null coefficients). Given the assumption of normal noise, the inverse cumulative distribution function ϕ−1 can have the simple form of an inverse Student's t distribution with degrees of freedom equal to those of the estimated error σ̂.
The Results section of this article is devoted to validation and practical illustration of the method. Although extensive work has been performed for validation of WT methods in tomography (Turkheimer et al., 1999, 2000), it is of interest to verify the assumptions behind the filters of Eqs. 1 and 2 that are the ones of choice for the present context. In particular, the use of standard Student's t distributions in Eq. 2 and relative Bonferroni adjustment are appropriate only if the noise processes of the set W (w, i) strictly match the independent normal noise conditions in wavelet space described above. By analogy with the approach of Worsley et al. (1993), such correspondence was verified by comparing the randomization distribution of studentized wavelet coefficients generated from a group of real images with the expected probability distribution function.
Practical application of the method considered two PET datasets in which the statistical model consisted of a GLM that is a common choice for data analysis in neuroimaging. For an example on the use of PCA refer to the previous article (Turkheimer et al., 2000).
The first dataset is a parametric study of CBF response to word recognition. In this case, the pattern of interest P (s) is the map of the rate of change of CBF toward the specified rate of words. The second study considers the measurement of [carbonyl-11C]WAY-100635 binding in brain in a group of normal controls and in a group of depressed subjects. In this case, it is of interest to estimate the distribution of the difference in serotonin receptors density between the two populations.
Details on the modeling of the experimental conditions are given in the following sections. The software used was written in MATLAB (Mathworks, Nadick, MA, U.S.A.) and run on an Ultra-Sparc 10 Sun Workstation (SunSystems, Mountain View, CA, U.S.A.). The DWT was implemented as detailed in Turkheimer et al. (1999) using Battle-Lemarie wavelets (Battle, 1987; Lemarie, 1988). The three-dimensional DWT was used in all studies. Because DWT requires a data size that has to be a power of 2, all original images were imbedded in a 128 × 128 × 128 volume with 2 × 2 × 2 mm pixel size. The translation invariant DWT used previously (Turkheimer et al., 1999, 2000) could not be extended to the third dimension because such an approach requires storage space and computational power that is not provided by the hardware available. However, the use of information from all three axes compensated for the translation invariant approach as the amount of artifacts in the parametric maps was negligible (see Results section). In all studies, the hard-threshold filter of Eq. 1 was used and the threshold was derived from Eq. 2 by using a standard t-distribution with appropriate degrees of freedom derived from the GLM used. The overall error rate in Eq. 2 α = 0.05 for both studies. Wavelet coefficients outside the brain were suppressed.
It is of interest to verify the ability of the wavelet filter of Eq. 2 to control error rate in wavelet space. This means that under the null hypothesis of all frames Y (s, i) being acquired in the same condition, the Bonferroni adjusted threshold of Eq. 2 must be able to allow a proportion of studentized wavelet coefficients d/σ̂ equal to the desired error rate α. Such ability of the procedure may be verified by constructing from the null dataset Y (s, i) a number of simulated datasets consisting in two groups of scans made by random assignment of the original ones (Worsley et al., 1992). Statistical analysis is then performed on each simulated dataset, and in each occasion the occurrence of at least one wavelet coefficient over the threshold defined in Eq. 2 is recorded as a positive event. After all possible permutations of Y (s, i) in the two groups have been performed, the number of positive events divided by the total number of simulations performed is compared with the error rate α.
The null dataset was composed by the scans in activation state of a previously published protocol (Brett et al., 1998). Such protocol consisted in two conditions: activation and rest. In all conditions, subjects looked at a computer monitor displaying a series of stimuli, consisting of abstract designs and video clips of four abstract hand gestures. In the activation condition, subjects performed one of the four abstract hand gestures with their right hand. In the rest condition, they were told to observe the stimuli without moving, and to let their minds go blank. Subjects were scanned with an ECAT 953B PET camera (CTI/Siemens, Knoxville, TN, U.S.A.) operating in three-dimensional mode. Approximately 9.2 mCi H2O15 were injected by the bolus method for each scan. For each subject, reconstructed images were realigned to the first scan in the session, and then coregistered to the Montreal Neurological Institute standard brain (Evans et al., 1993) using SPM99 software (Functional Imaging Laboratory, London, U.K.). The null set Y (s, i) consisted of data from 8 subjects, 5 frames each.
The statistical model consisted of a GLM with 8 subjects factors plus 8 subject specific covariates to normalize each scan to its global counts. Single frames from all subjects were randomly assigned to 2 groups of 20 scans each, then the statistical analysis in wavelet space was performed. Error estimates were obtained by pooling residuals from all 40 scans. Because the set of all possible permutations was too large (40!/(20!20!) = 1.3e + 11), analysis was repeated for a subset of 1000 random assignments, and the number of times that studentized wavelet coefficients were over prefixed thresholds (0.1, 0.05, 0.01) was recorded.
This example illustrates the application of the WT method to a typical parametric study of CBF response to an experimental condition. By “parametric” it is usually meant that the CBF is related to stimulus intensity by a linear relation and is therefore of interest to estimate the pattern of such linear rate determined by the experimental setting. In this particular instance, data are selected from a previously published study that considered a word recognition task (Leff et al., 2000). Five right-handed, normal subjects participated in the study. All subjects claimed English as their first language. Subjects viewed single words, presented centrally, at different rates: 5, 20, 40, 60, and 80 per minute. Stimuli were displayed for 500 milliseconds and were proceeded by a central cross-hair. The cross-hair remained for the entire interstimulus interval. There was also a “baseline” condition—a single word appeared just before the scan started and was then replaced by a cross for the rest of the scan—to ensure that subjects attended to the screen at all times during the scan. All stimuli were single syllable, English words (nouns and verbs). Subjects were not given any explicit task related to the stimuli; instead, they were instructed to “read and understand” the words. Each participant underwent 8 scans, the order of which was randomized within and across subjects. Images were acquired using an ECAT 953B camera (CTI, Knoxville, TN, U.S.A.) operated in three-dimensional mode. Arterial samples of the input function were not available, therefore data represent normalized tissue concentrations after injection of H215O. For each subject, reconstructed images were realigned and then coregistered to the MNI standard brain (Evans et al., 1993) using SPM99 software.
Scans were entered into a design matrix for a GLM with stimulus rate as the covariate of interest. Subjects effects and subject-specific covariates for global normalization completed the matrix.
[carbonyl-11C]WAY-100635 is a selective 5-HT1A receptor antagonist that can be used in conjunction with PET to assess 5-HT1A receptor binding in vivo (Pike et al., 1996). In this section the authors reanalyze a previously published dataset (Sargent et al., 2000) in which this tracer was used to measure changes in 5-HT1A receptors in depression. [11C]WAY-100635 scans were performed with an ECAT 953B camera (CTI) on 15 patients with major depressive disorder (unmedicated) and 18 healthy volunteer subjects. Binding potential images were generated from the dynamic scans using a reference tissue model (Gunn et al., 1998). Images were spatially normalized with SPM-96 software (Functional Imaging Laboratory, London, U.K.) into Montreal Neurological Institute stereotactic space using a [11C]WAY-100635 template as previously described (Meyer et al., 1999). Statistical modeling was performed as in the original analysis to simplify the comparison between the results of the WT method and those previously obtained with statistical parametric mapping. The GLM contained only one factor (group) in which error was estimated by pooling residuals estimates from both populations.
Proportions of false positives at each of the α = 0.1, 0.05, 0.01 are shown in Table 1. All proportions are close to nominal levels, which suggests that the underlying noise processes in wavelet space are reasonably well approximated by normal, independent distributions. Table 1 also reports the error of simulations. This can be estimated by noting that the rejection of the null hypothesis is a binomial event. Using the normal approximation to the binomial distribution, the approximate 90% confidence interval surrounding each rejection frequency F may be easily computed as F ± 1.645[(F)(1−F)/N]½ where N is the number of simulated datasets (Noreen, 1989).
Specificity of wavelet filter with Bonferroni correction (± 90% confidence interval)
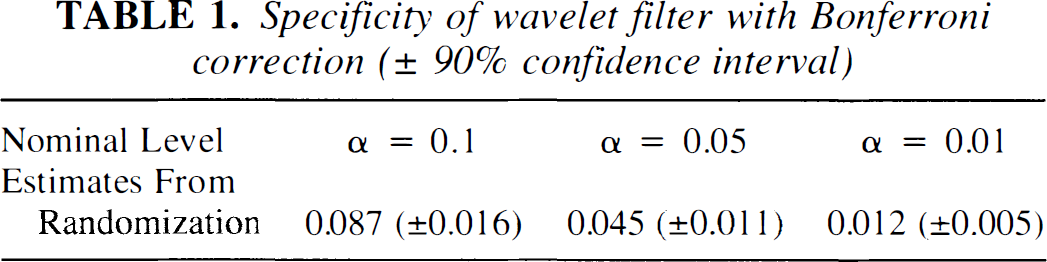
Empirically, one can appreciate the concordance of the empirical probability distribution function of wavelet coefficients with the expected one by looking at Fig. 3 where such distribution from one of the simulated datasets is compared with the Student's t-distribution with 23 degrees of freedom.
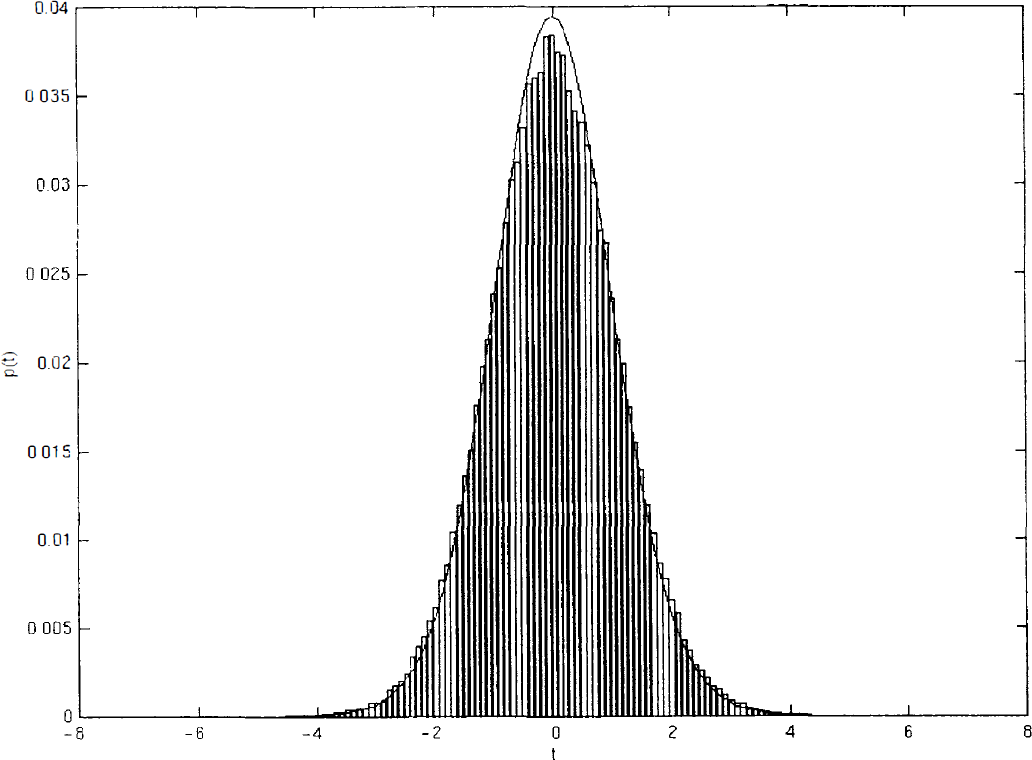
Distribution of wavelet scores computed from 1 of the null-datasets obtained by random assignment of 40 scans acquired in the same experimental conditions in 2 simulated experimental groups of 20. Figure displays the empirical distribution of studentized wavelet coefficients superimposed with the expected distribution, a standard Student's t-distribution with 23 degrees of freedom.
The estimated pattern of CBF response to stimulus rate is shown in Fig. 4 superimposed to the correspondent MNI space. The main area of activity was in the posterior part of primary visual cortex (BA17 or V1), and prestriate cortex (BA18 and 19), bilaterally. From prestriate cortex, activity ran anteriorly, medially, and ventrally through the occipito-temporal junction and into the fusiform gyrus (BA 37). This effect was greater on the left. The other two main brain regions, which also responded bilaterally, were in the prefrontal cortex: the supplementary motor area (SMA) and premotor cortex, and medial and lateral BA6, respectively.
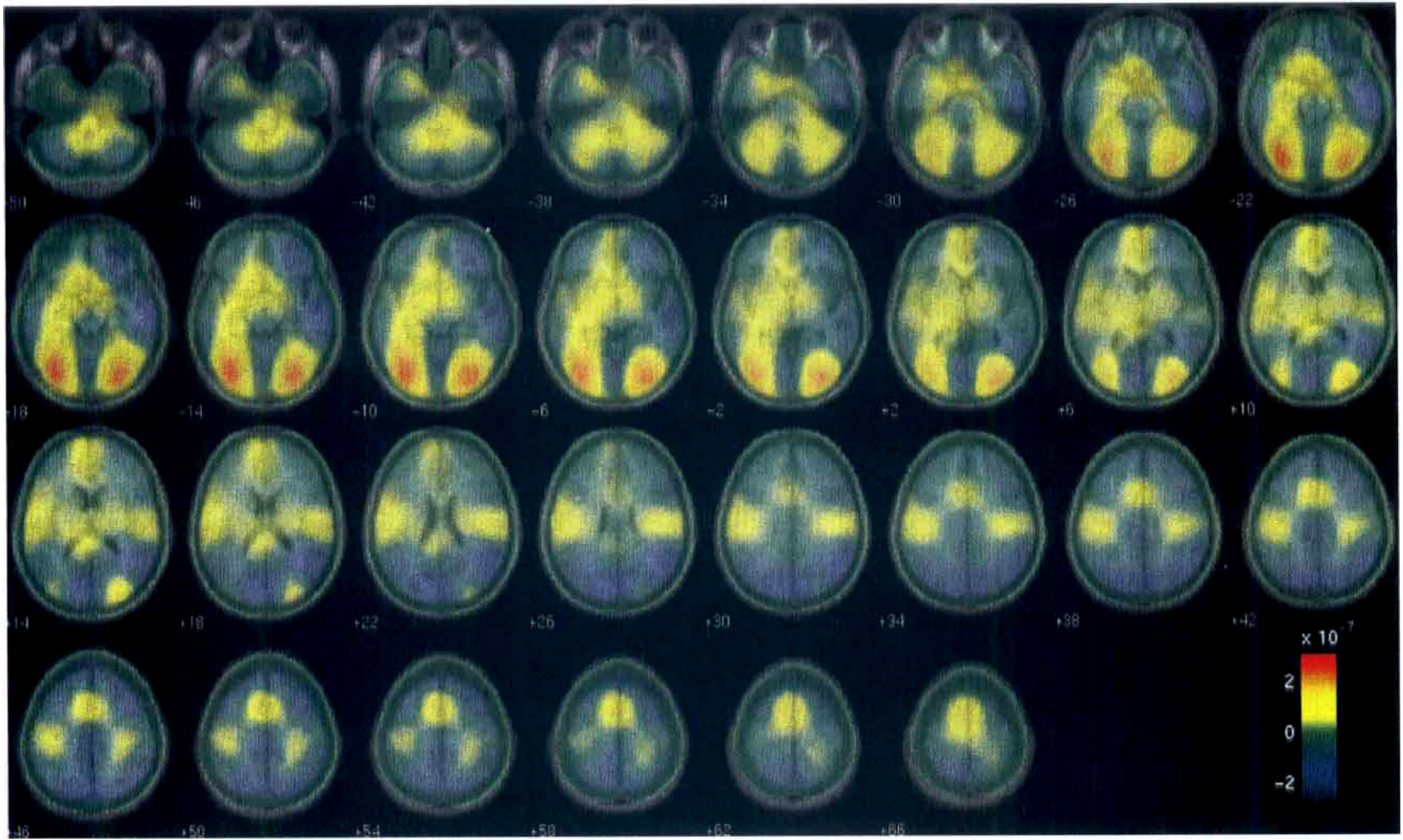
Results of the wavelet procedure when applied to the analysis of a water activation study. The activation study was designed to activate areas that respond to word recognition. The filtered activation image is displayed in color overlaid on a gray scale magnetic resonance imaging of the MNI standard brain. Values represent the pattern of the change of blood flow toward the rate of words estimated with an error rate α = 0.05 over all slices. The left side of the brain is displayed at the left of the slices. Slices are labeled with their distance in millimeters from the transverse plane containing the anterior and posterior commissures. Units are in normalized counts for unit word rate.
Previous processing in SPM99 (Leff et al., 2000) identified the same areas with smaller clusters. Bilateral primary and secondary visual cortex and SMA were activated, but only the left premotor (lateral BA6) survived correction at the overall error rate α = 0.05 (one resolution only analyzed using a gaussian filter with 10 mm full width at half maximum); activity on the right was present but only at a lower threshold (P = 0.001 uncorrected). This was also true for the bilateral fusiform area.
Results are presented in Fig. 5 as binding potential values superimposed on the MNI brain. In the depressed population, binding potential values were reduced across frontal, temporal, and limbic cortex (maximum difference 0.6 ∼ 10% difference from normal values). Anatomic location of such differences matches the one reported previously using SPM96 (Sargent et al., 2000). However, such a pattern could not be obviously estimated by SPM and the detection of the areas of significant change of the parameter pattern required a much lower threshold (P < 0.01 uncorrected, images smoothed with a gaussian filter with 8 mm full width at half maximum) with equivalent increase in the error rate α. Fig. 5 also shows a greater reduction of binding in the right orbitofrontal cortex compared with the left side that could not be revealed in the previous analysis by SPM but only by region of interest analysis (Sargent et al., 2000).
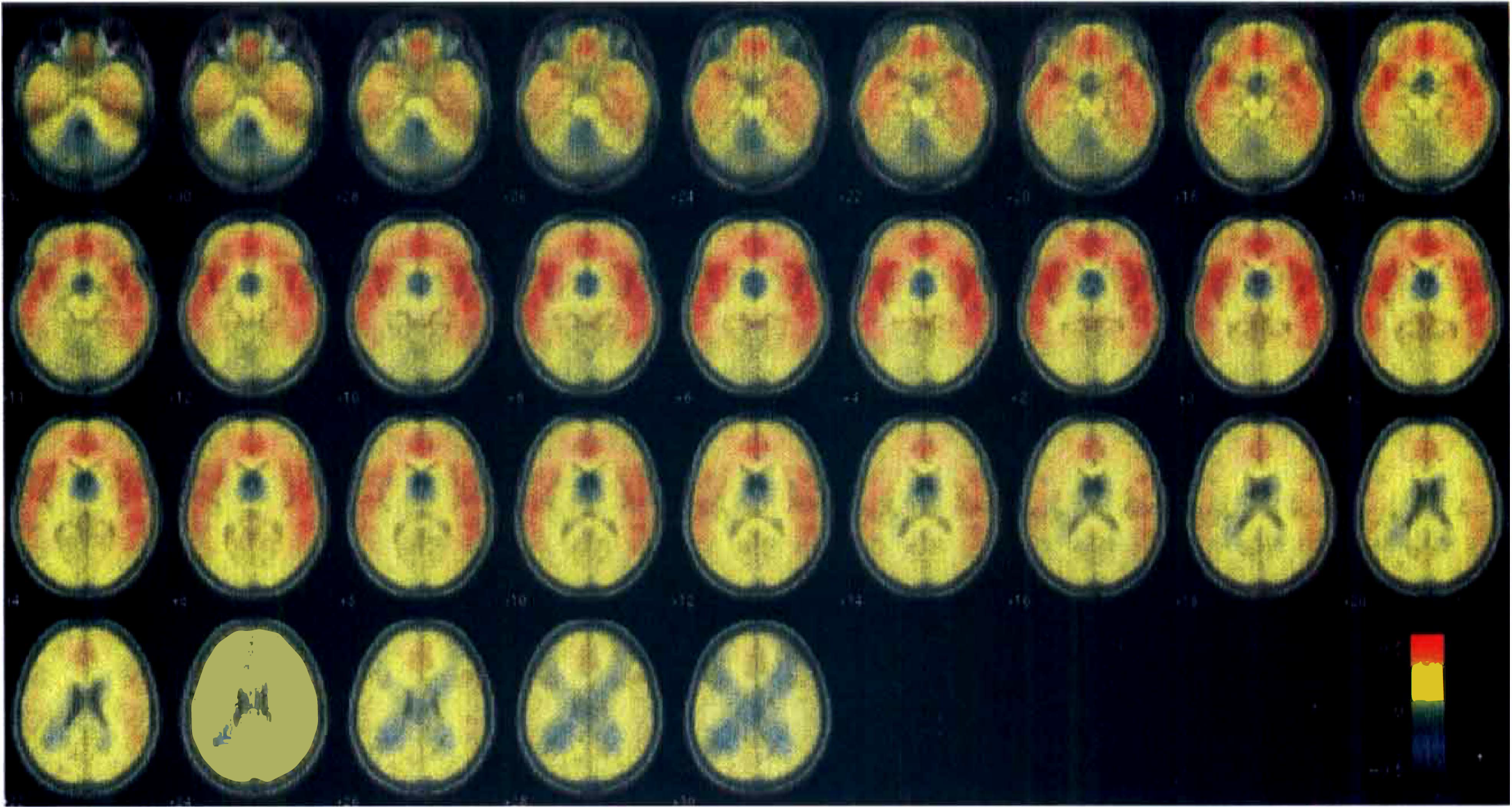
Results of the wavelet procedure for the [11C]WAY-100635. The colored pattern, overlaid on the MNI standard brain, represents the difference in binding potential (BP units are pure numbers) between the controls and the depressed group. Overall error rate for the estimation was α = 0.05. The left side of the brain is displayed at the left of the slices. Labels indicate the distance in millimeters from the transverse plane containing the anterior and posterior commissures.
A new method for the statistical analysis of sets of PET images was introduced. The method adopts current statistical models for the description of the set but is novel in the use of a mathematical transform, the wavelet transform, to model the spatial dimension of images. Use of a transform allows a complete modeling approach that produces spatial estimates of the parameters of interest under the desired statistical risk.
This property distinguishes this approach from current methods of analysis. These methods combine mono-resolution filters with thresholding procedures that control the rate of false positives; their output are probability maps and no proper estimation of effects is allowed.
In short, whereas SPM methods are meant to obtain statistical significance, the wavelet approach targets the estimation problem and this difference is substantial, as illustrated by the words of S.C. Pearce (1993): “Also, there should be no suggestion that statistical analyzes exist to find significant differences. Experiments are conducted in order to find answers to the questions being asked. Possibly no one doubts that a difference exists. If so, the task is to estimate its size and not to test for its existence. Further, a difference of means may be significant but not important or vice versa….”
The technique was built upon previous work with attention to the noise model; in particular, the application of the statistical model in wavelet space allowed the method to adapt to the local noise characteristics of tomographic images. Such an ability combines with the energy preserving and whitening properties of the transform to yield a theoretical setting that controls effectively the chosen risk by use of standard statistical procedures. In the framework of multiple hypothesis testing, which is common in neuroimaging, the control of false positives may be obtained by a simple Bonferroni correction.
Theoretical expectations were validated in the application section of the paper. In the first instance, the correspondence between the empirical distributions of data and the predicted ones for a null dataset (all frames in the same condition) was shown. Note that the verification of assumptions on a single dataset, however randomly selected and representative of the general experimental conditions that data set may be, does not imply that these assumptions would hold on any other set. Deviations from independence and normality can be either predicted or easily detected using routine diagnostics. In this case, one may divert from the use of parametric approaches and turn toward nonparametric approaches. In this particular instance, permutation methods applied to wavelet coefficients may be appropriate. Permutation methods for multiple testing are reviewed by Westfall and Young (1993). Also see Holmes et al., (1996) for an application to neuroimaging.
The same considerations regarding the robustness of parametric assumptions in neuroimaging also apply to current statistical parametric methods with the important difference that the methods in wavelet space do not require the additional constraints imposed by gaussian random field theory such as large number of degrees of freedom and upper and inferior bounds on the smoothness of the field (Holmes et al., 1996).
Finally, the method was also applied to two other datasets to clarify its use in typical instances of analysis of neuroimages. Comparison of these results with those obtained previously with statistical parametric mapping showed that the use of wavelet bases, which are optimal or near-optimal for PET signals (Turkheimer et al., 1999), combined with an adequate noise model, not only estimates correctly the parametric patterns but also produces a valuable increase in the detection power.