Abstract
In bolus-tracking perfusion magnetic resonance imaging (MRI), temporal dispersion of the contrast bolus due to stenosis or collateral supply presents a significant problem for accurate perfusion quantification in stroke. One means to reduce the associated perfusion errors is to deconvolve the bolus concentration time-course data with local Arterial Input Functions (AIFs) measured close to the capillary bed and downstream of the arterial abnormalities causing dispersion. Because the MRI voxel resolution precludes direct local AIF measurements, they must be extrapolated from the surrounding data. To date, there have been no published studies directly validating these local AIFs. We assess the effectiveness of local AIFs in reducing dispersion-induced perfusion error by measuring the residual dispersion remaining in the local AIF deconvolved perfusion maps. Two approaches to locating the local AIF voxels are assessed and compared with a global AIF deconvolution across 19 bolus-tracking data sets from patients with stroke. The local AIF methods reduced dispersion in the majority of data sets, suggesting more accurate perfusion quantification. Importantly, the validation inherently identifies potential areas for perfusion underestimation. This is valuable information for the identification of at-risk tissue and management of stroke patients.
Introduction
Bolus-tracking magnetic resonance imaging (MRI) is frequently used to quantify perfusion in stroke patients via an intravenous injection of paramagnetic contrast followed by repeated acquisition of magnetic resonance images to track the bolus passage through the brain (Ostergaard et al, 1996a, 1996b). Commonly, a global Arterial Input Function (AIF) is measured by averaging the bolus concentration time-course (CTC) profiles in voxels selected near a major contralateral artery (Calamante et al, 1999). Perfusion maps are calculated by deconvolving this AIF from whole brain CTC data (Ostergaard et al, 1996b). In stroke patients, delay and temporal dispersion (D/D) of the bolus may be considerable in the ipsilateral hemisphere (IH) due to arterial stenosis, occlusions, or collateral flow (Calamante et al, 2003). Since D/D is not included in the measured global AIF (GAIF), the severity and extent of any perfusion deficit may be exaggerated (Calamante et al, 2000). Such misdiagnosis may lead to dangerous mistreatment, since tissues that are not at risk of infarction may appear to be so due to erroneously low perfusion estimates resulting from the D/D.
Although deconvolution algorithms can be made delay-insensitive, e.g., singular value decomposition with a block-circulant deconvolution matrix (oSVD) (Wu et al, 2003), dispersion cannot be easily accounted for in standard deconvolution methods. Dispersion error may be circumvented using multiple local AIFs—located downstream of any arterial abnormality—to deconvolve small branch arterial territories. Previously suggested methods to identify the local AIFs include: automated heuristic approaches to select CTC profiles appearing most arterial (Alsop et al, 2002; Lorenz et al, 2006b); and spatial Independent Component Analysis of the CTC map to isolate the arterial-like temporal components (Calamante et al, 2004). These methods assumed that a region of tissue is fed mainly by the local AIF to which it is
Although some studies have investigated the potential role of local AIFs in predicting stroke outcome (Christensen et al, 2009; Lorenz et al, 2006a), to date there are no published studies investigating the veracity of the estimated local AIFs. The validity of the local AIFs can be indirectly inferred through the minimization of D/D, since reduced D/D indicates a more accurate AIF, and consequently, a more accurate perfusion estimate. Here, we evaluate two local AIF methods (including one novel method that is introduced in this work), and compare them with the standard GAIF analysis by measuring the dispersion remaining after deconvolution.
Materials and methods
Local Arterial Input Function
In this study, the principles for identifying local AIFs and their territories were based on those presented by Christensen et al (2008), as follows: along the supply route and toward the distal small branches of the arterial tree, blood arrives later (increasing BAT) and flows more slowly (increasing intervoxel BAT differences). At typical temporal resolution repetition time (TR) (1.5 to 2.5 seconds), only the larger local BAT differences in the smaller distal arteries are detectable: the relatively low flow speeds of small pial arteries translate to a BAT difference of ∼150 ms between voxels spaced 2 mm apart; in contrast, at the higher velocities in large arteries this time difference would be only a few milliseconds, too small to be detected with the TRs of bolus-tracking MRI (see Christensen et al (2008) for more detail on this argument). Therefore, the small branch vascular territory is determined by following the lowest BAT gradient descent path from each tissue voxel toward a local minimum, inferred as the local origin of supply, i.e., the local AIF voxel. This approach assumes that voxels supplied by the same local AIF have similar BATs, and that the supply route has no large BAT discontinuities. Voxels belonging to the same local AIF territory are therefore both spatially close and temporally similar.
Given that the
In practice, the small BAT differences,
In this work, the BAT was estimated using two different approximations. First, BAT was estimated as the
Although

where
Following the identification of potential local AIF voxels and their associated territories using either LAIF1 or LAIF2, heuristic criteria similar to those in Mouridsen et al (2006) were applied to disregard the 10% of local AIFs with smallest area, and the 10% with largest roughness. Tissue voxels linked to an invalid local AIF were assigned the local AIF of the nearest neighboring territory. Finally, all local AIF curves were normalized to unit area. Automated methodologies for selecting the local AIFs of LAIF1 and LAIF2 were implemented in MATLAB (Mathworks, Natick, MA, USA).
Simulated Data
To demonstrate the limitations of the LAIF1 method and investigate the approximations involved in the design of the LAIF2 method, the dependence of their respective BAT estimates,
In Vivo Data and Analysis
Nineteen bolus-tracking MRI data sets from 17 patients were retrospectively selected from clinical scans acquired between 2001 and 2006 from anterior circulation stroke patients. Consent was obtained from all participants and the study protocol was approved by the local (Austin Health) Ethics Committee. The selection criteria were minimal patient motion during the bolus passage, and an MTT abnormality based on conventional analysis of bolus-tracking data. For the purpose of this study, conventional analysis corresponded to using the delay-insensitive oSVD deconvolution with a GAIF automatically measured in a branch of the contralateral middle cerebral artery. This analysis was carried out using the PENGUIN software (PErfusioN Graphical User INterface; http://www.cfin.au.dk/software/penguin), and provided a preliminary indication of the perfusion abnormality, though possibly exaggerated by dispersion (but not delay). For all the analyses (local and global AIF methodologies), the CTC data were calculated assuming a linear relationship between the transverse relaxation rate change and contrast concentration (Calamante et al, 1999). The perfusion MRI data were acquired on a 1.5 T General Electric Signa Horizontal SR 120 (General Electric, Milwaukee, WI, USA) using a gradient-echo echo planar imaging and 0.2 mmol/kg bolus of a Gd-based contrast agent injected by a power injector and followed by 15 mL saline. The sequence parameters were echo time/TR=60/1,740 ms and 10 slices, echo time/TR=70/2,240 ms and 12 slices, or echo time/TR=60/1,999 ms and 12 slices; 256 × 256 matrix; slice thickness=6 mm; slice spacing=7 mm; voxel size 1.56 × 1.56 mm. For the corresponding diffusion images, echo time/TR=100/12,000 ms, 19 slices, 256 × 256 matrix, slice thickness=5 mm, slice spacing=6.7 mm, voxel size 1.56 × 1.56 mm, and
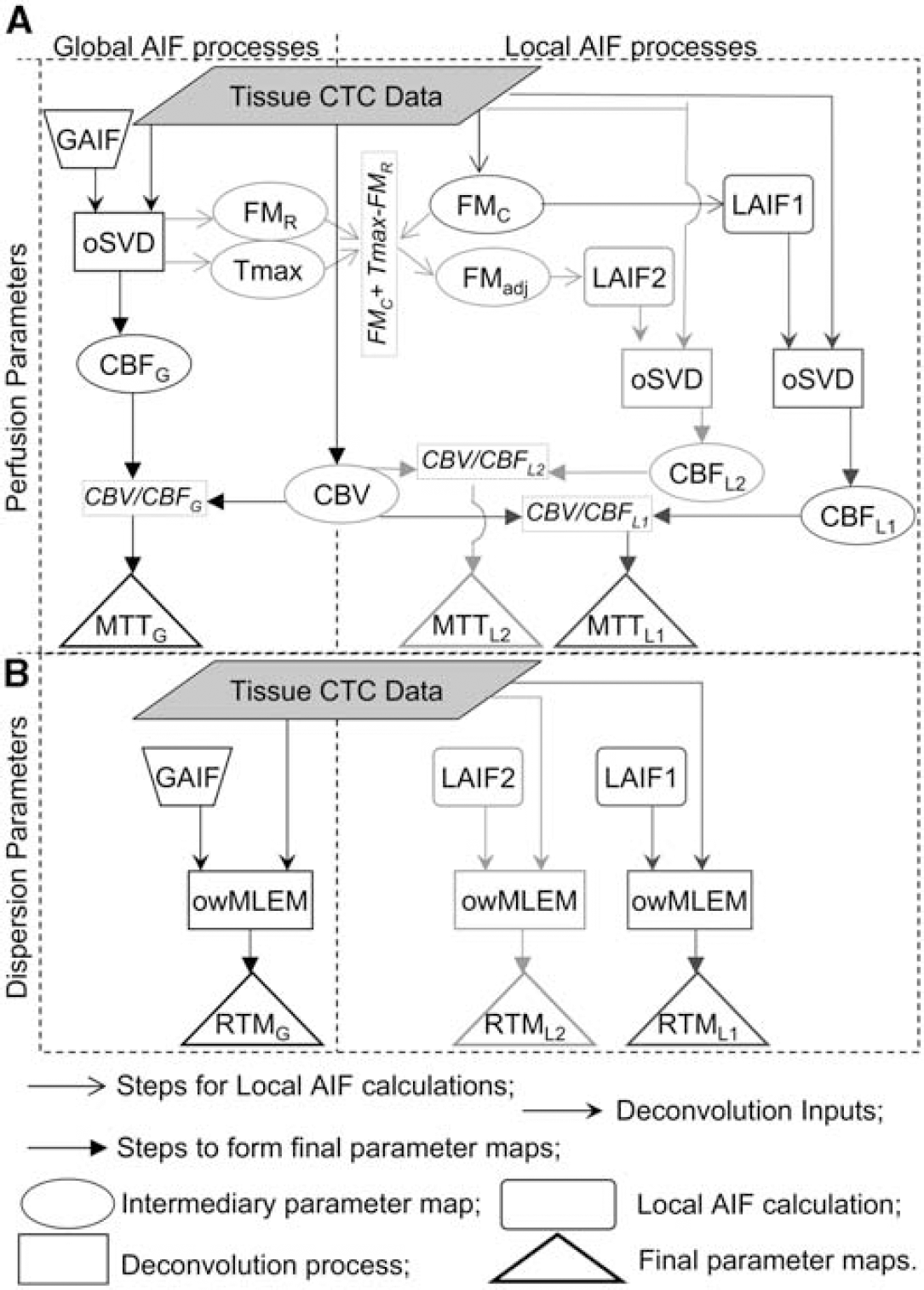
Flow chart outlining the steps involved in forming the various parameter maps, and their relationship to each other. The chart is divided into four quadrants. The top two quadrants (
A reference MTT value was estimated for each method (GAIF, LAIF1, and LAIF2) in each data set by manually segmenting the contralateral grey matter (GM) and averaging the MTT values, MTTcontraGM. Abnormal MTT areas were defined by highlighting ipsilateral voxels with MTT>Thld, where Thld=MTTcontraGM+1.78 seconds (Christensen et al, 2009). This threshold was shown in Christensen et al (2009) to have the greatest sensitivity and specificity for predicting future infarction from oSVD MTT maps. From herein, MTT above this threshold is referred to as abnormally (abn) perfused, or hypoperfused, and the hypoperfusion maps for GAIF, LAIF1, and LAIF2 are labelled as abnMTTGAIF, abnMTTLAIF1, and abnMTTLAIF2, respectively. These are in fact apparent (rather than true) hypoperfusion maps since their MTT values may be biased by dispersion.
In the absence of delay and dispersion, the initial value of the tissue response functions,
Dispersion maps for GAIF, LAIF1, and LAIF2 were created by highlighting ‘abnormal’ voxels with RTM≥TR (maps labelled abnRTMGAIF, abnRTMLAIF1 and abnRTMLAIF2, respectively). These maps indicate areas affected by residual vascular dispersion, where the MTT estimates are thought to be unreliable (Willats et al, 2008).
The sizes of the highlighted (abnormal) regions in the abnMTT and abnRTM maps described above were expressed as a percentage of the IH excluding ventricles (%abnMTT and %abnRTM, respectively), for a single slice with significant diffusion abnormality. The sizes of these abnormal regions were pairwise compared between each method (GAIF versus LAIF1, GAIF versus LAIF2, and LAIF1 versus LAIF2). Comparisons were done using a signed Wilcoxon's test. To aid discussion, the data sets were ordered according to the difference in dispersion area between %abnRTMGAIF and %abnRTMLAIF2, and labelled from A to S accordingly. A summary of information for each data set is given in Table 1.
Summary information for data sets A–S
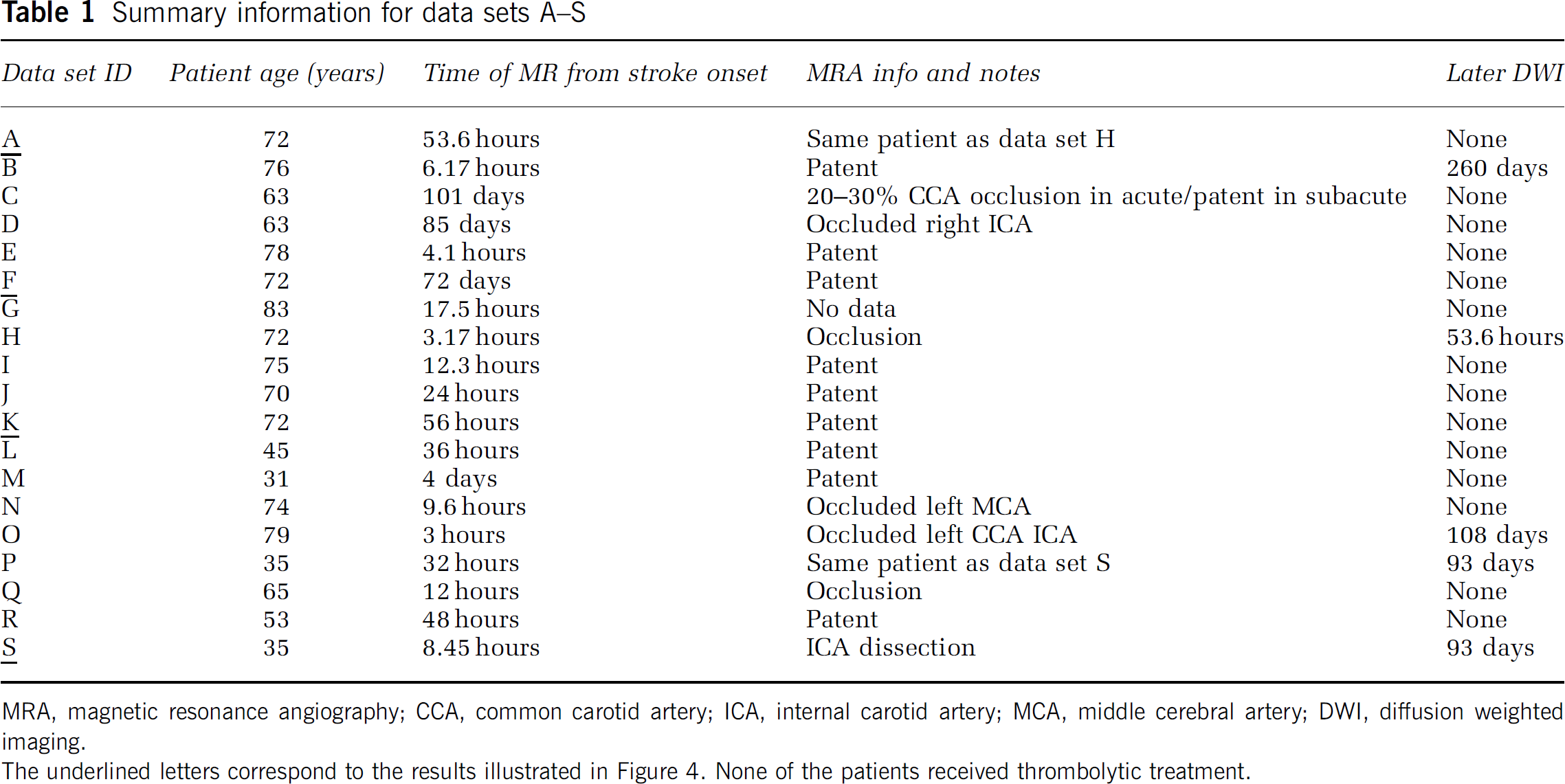
MRA, magnetic resonance angiography; CCA, common carotid artery; ICA, internal carotid artery; MCA, middle cerebral artery; DWI, diffusion weighted imaging. The underlined letters correspond to the results illustrated in Figure 4. None of the patients received thrombolytic treatment.
Results
Simulated Data
Figure 2 shows the dependence of
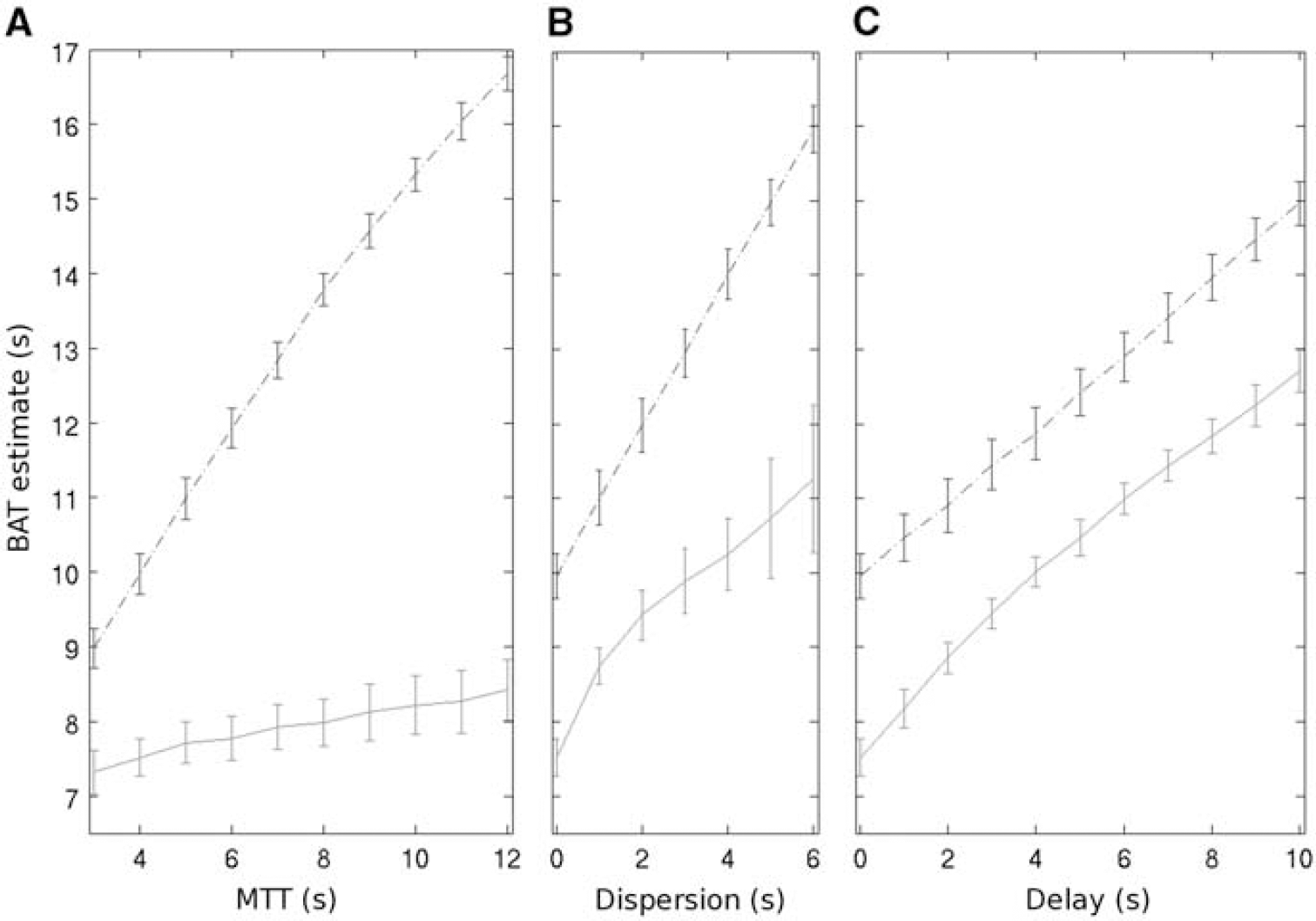
Example simulation results at TR=2 seconds and signal-to-noise ratio=50 showing the BAT estimate (
In theory,
In Vivo Data
While there was an overall good agreement between the results using LAIF1 and LAIF2, LAIF2 was marginally better in reducing dispersion in all but one data set (data set C, where the %abnRTMLAIF2 (=82%) was ∼4%IH larger than abnRTMLAIF1 (=78%), data not shown). Accordingly, %abnRTMLAIF2 was significantly smaller than %abnRTMLAIF1 (
Figure 3 indicates the size of the abnormal dispersion and apparent abnormal perfusion areas using GAIF and LAIF2. In all the data sets except A, B, and C, LAIF2 reduces the size of the dispersion area (%abnRTMLAIF2<%abnRTMGAIF), suggesting that the correspondingly smaller apparent hypoperfusion areas (%abnMTTLAIF2<%abnMTTGAIF) are less affected by dispersion error and therefore more likely to represent the true extent of the hypoperfusion. In data sets A, B, and C, LAIF2 slightly increased the dispersion area (∼10%IH). Closer inspection of the RTM images shows that the additional dispersion occurs primarily in areas of deep white matter where it is difficult to find a local AIF (e.g., data set A, Figure 4i). While this increase is not desirable, there is no corresponding increase in the area of apparent hypoperfusion in data sets A and B (%abnMTTLAIF2≈%abnMTTGAIF), indicating that any errors introduced by the additional dispersion are too small to recategorize this tissue as hypoperfused. In the remaining 16 data sets, LAIF2 reduces both the dispersion and apparent hypoperfusion areas to various extents as indicated in Figure 3. In the paired comparison across all data sets, %abnRTMLAIF2 and %abnMTTLAIF2 were significantly smaller than %abnRTMGAIF and %abnMTTGAIF, respectively (
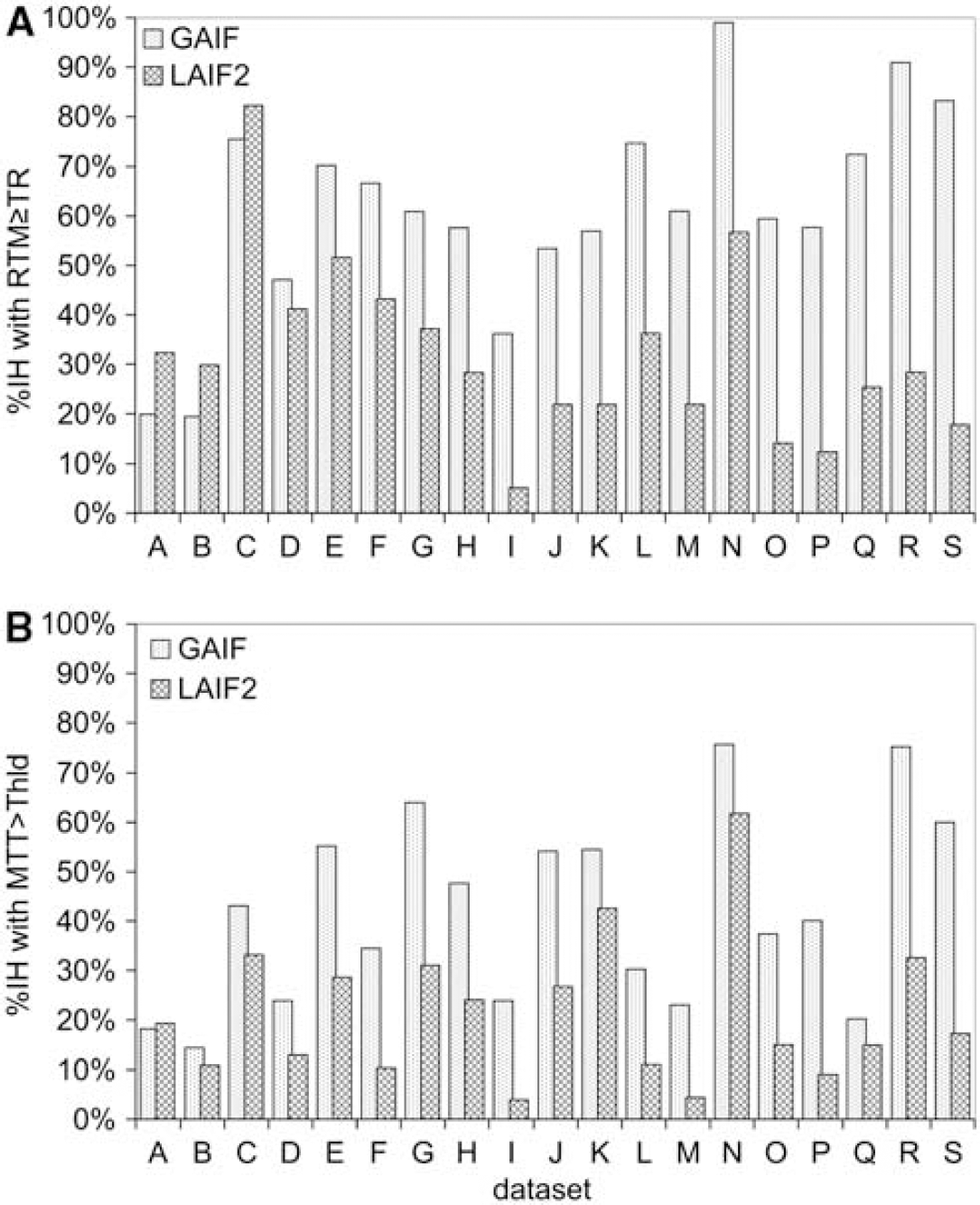
Graph showing the %IH with abnormality using GAIF (dotted bars) and LAIF2 (hashed bars). The top graph (
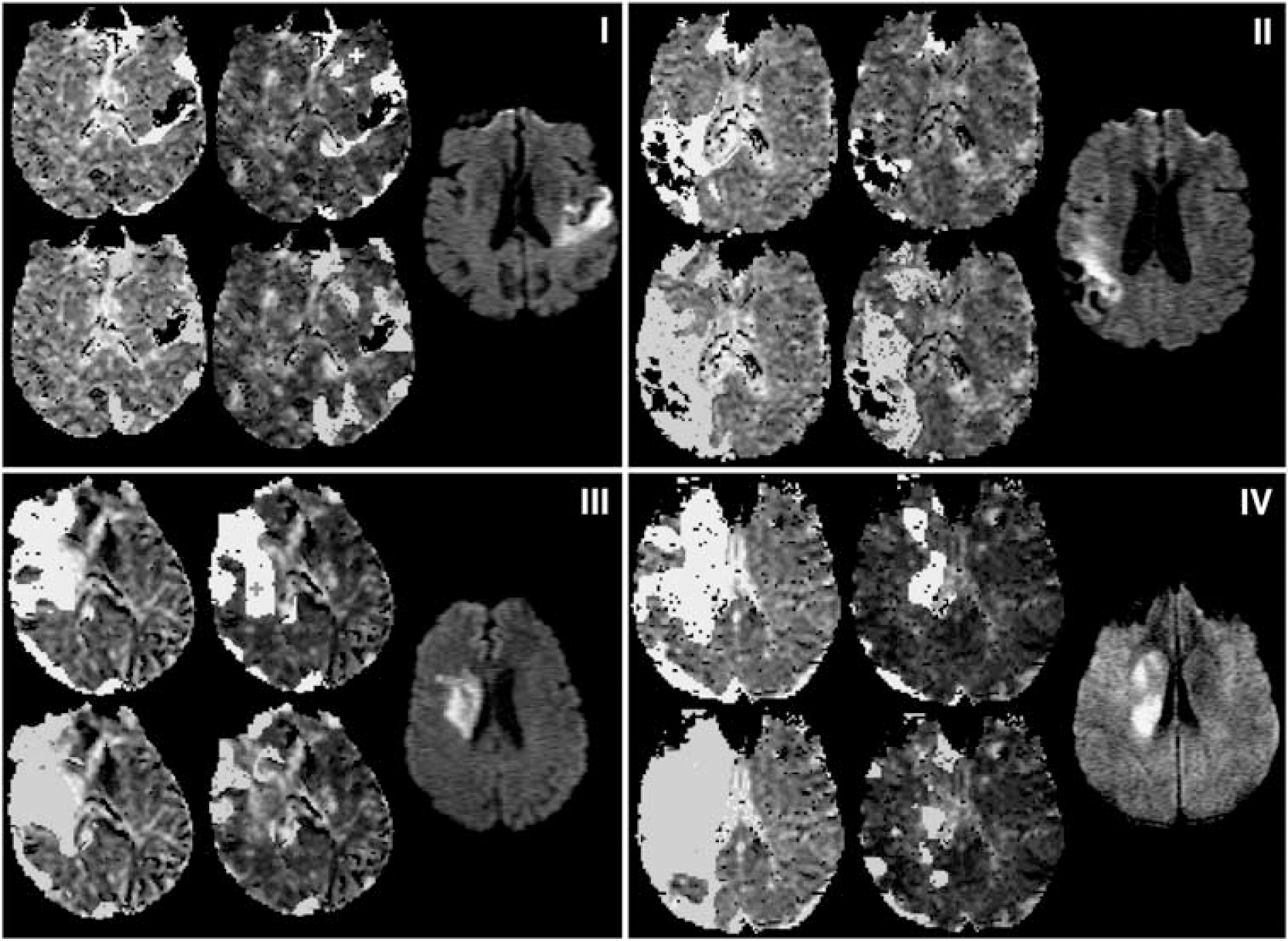
Abnormal perfusion and dispersion areas from example slices in data sets A (i), F (ii), K (iii), and S (iv). In each subfigure (i–iv), the diffusion weighted image slice most closely corresponding to the perfusion slice is shown to the right. The remaining four maps are MTT, with areas of MTT>Thld overlaid in yellow (top) and RTM≥TR overlaid in blue (bottom); the left-most maps are the GAIF analysis and the central maps are the LAIF2 analysis. The crosses marking the LAIF2 perfusion maps of (i) and (iii) are the locations from which the LAIF2 time-courses in Figure 5 are taken. GAIF, global Arterial Input Function; LAIF2, local Arterial Input Function (new method); RTM, time to rise-to-maximum value of the owMLEM residue function; MTT, mean transit time; Thld=MTTcontraGM+1.78 seconds (see text).
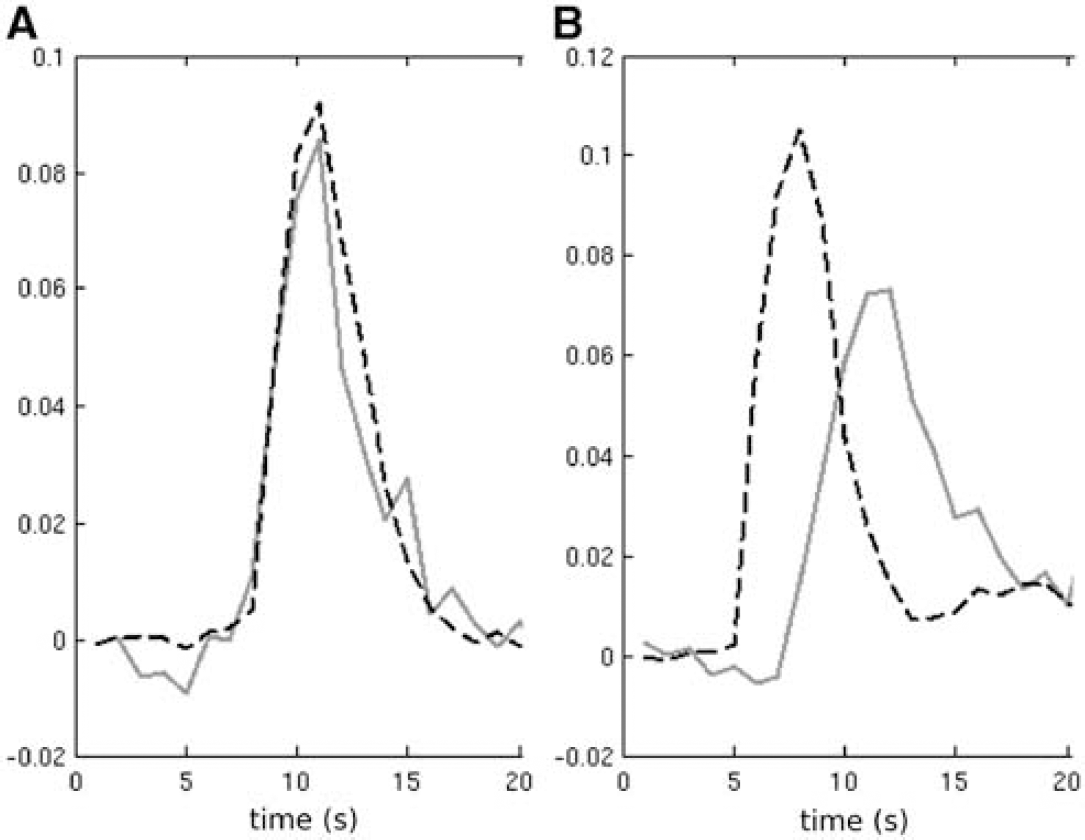
Example LAIF2 time-courses (red) from patients A and K (panels
To illustrate four different scenarios, data sets A, F, K, and S are shown in Figure 4. Data set A (Figure 4i) illustrates a case where LAIF2 slightly increases the dispersion area compared with the GAIF result (see previous paragraph). The similar perfusion maps may also be inferred from the comparison of the LAIF2 time-courses with the GAIF. In the example shown in Figure 5A, the LAIF2 and GAIF curves show similar characteristics, suggesting the dispersion of blood flow in the smaller feeding arteries is minimal in this case. In data sets F, K, and S, LAIF2 decreased the dispersion area compared with the GAIF result (
Discussion
In this study, we have evaluated two local AIF methods LAIF1 and LAIF2, whose source locations are determined as the local minima of
The
It should be noted that the parameter and threshold used for dispersion assessment, RTM≥TR, has good specificity but limited sensitivity in detecting dispersion. For example, a voxel with RTM<TR may still have a small dispersion (and therefore contain perfusion error), but it cannot be distinguished from delay (where perfusion is unbiased using oSVD). A more optimized protocol with shorter TR (Wintermark et al, 2008) would minimize this effect. Furthermore, the abnormal perfusion threshold MTT>MTTcontraGM+1.78 seconds was taken from work looking at infarct prediction in acute data (Christensen et al, 2009), and from a GAIF analysis. Because of dispersion, the MTT values obtained with a GAIF analysis may have been overestimated. It follows that for a LAIF analysis where dispersion is minimized, the abnormal MTT threshold may be lower. However, since the duration of ischemia is a significant factor in determining tissue infarction, the threshold with greatest sensitivity and specificity for infarct prediction will probably change with time of scan; and therefore, a time-dependent threshold could be more appropriate for our data. Further work is needed to determine the optimal threshold both for local AIF methods and for different scan times. Nevertheless, the threshold used in this study illustrates the issues of defining the perfusion abnormality using global and local AIF methods.
Although GAIF identified larger areas of hypoperfusion (due to dispersion-related perfusion error), it is necessary to determine whether a different (larger) MTT threshold would identify the same hypoperfused area as identified using LAIF2. If this was the case, the GAIF could still provide the same clinical information. For the data sets in Figure 4ii–iv, we found that a threshold of MTT>MTTcontraGM+
Interestingly, however, a visual comparison of the limited available follow-up data with the initial perfusion abnormality revealed that for all cases the LAIF2 methodology delineates an abnormal perfusion volume that is more closely aligned with the final infarct volume (an example is shown in Figure 6), compared with the much larger GAIF area. This underlines the caution that should be taken when using dispersion-contaminated GAIF perfusion maps to assess for tissue at risk of infarction. However, we acknowledge the very preliminary nature of these findings due to the limited amount of follow-up data available.
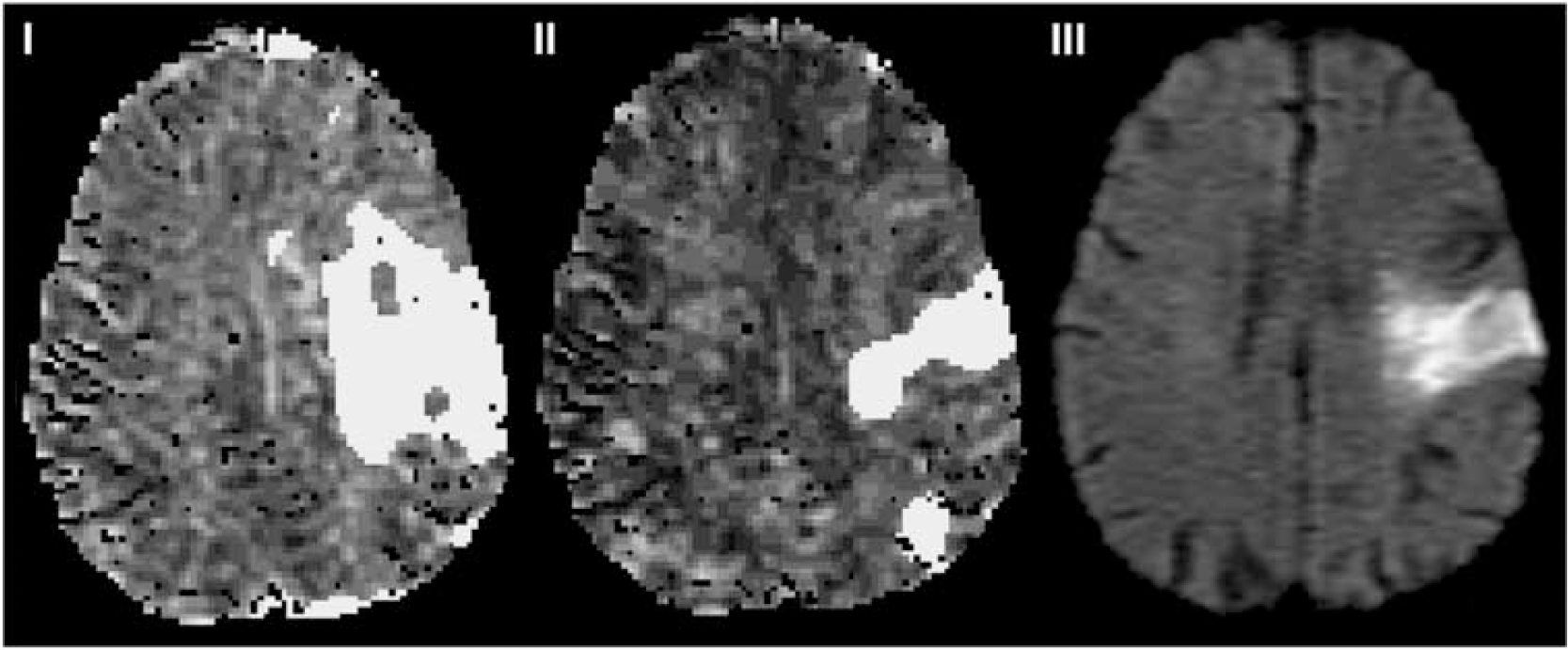
Example of how local AIF methodology may delineate final infarct volume more closely than the standard GAIF method. The left and central maps are MTT with areas of MTT>Thld overlaid in yellow for (i) GAIF and (ii) LAIF2 at acute scan time of 3 hours (data set O, Table 1). The right hand image (iii) is the follow-up 108 day diffusion scan. AIF, Arterial Input Function; GAIF, global AIF; LAIF2, local AIF (new method); MTT, mean transit time; Thld=MTTcontraGM+1.78 seconds (see text).
Several major clinical trials have used the
Due to the long computational time, the owMLEM validation methodology (to measure the residual dispersion) is not practical for use in acute clinical stroke analysis. For this reasons, we used the more conventional and fast oSVD method to calculate
To conclude, both of the local AIF methods presented here were found to reduce the amount of residual dispersion in the perfusion maps compared with the conventional GAIF analysis. The corresponding reduction in dispersion-related perfusion error suggests that the local AIF methods can better characterize the severity and extent of any perfusion deficit. Of the two local AIF methods, the novel method (LAIF2) was found to be marginally better in reducing dispersion.
Footnotes
The authors declare no conflict of interest.
Appendix
This section explains the derivation and calculation of
The perfusion convolution equation is (Ostergaard et al, 1996b):
where
FMs are additive under convolution, therefore:
(Note that the scaling factor
Using a global AIF (GAIF), equation (A1) becomes
where
and
Taking FMs of equation (A4),
where
where
