Abstract
In the central nervous system, a constant microenvironment required for neuronal cell activity is maintained by the blood—brain barrier (BBB). The BBB is formed by the brain microvascular endothelial cells (BMEC), which are sealed by tight junctions (TJ). To identify genes that are differentially expressed in BMEC compared with peripheral endothelial cells, we constructed a subtractive cDNA library from porcine BMEC (pBMEC) and aortic endothelial cells (AOEC). Screening the library for differentially expressed genes yielded 26 BMEC-specific transcripts, such as solute carrier family 35 member F2 (SLC35F2), ADP-ribosylation factor-like 5B (ARL5B), TSC22 domain family member 1 (TSC22D1), integral membrane protein 2A (ITM2A), and epithelial membrane protein 1 (EMP1). In this study, we show that EMP1 transcript is enriched in pBMEC compared with brain tissue and that EMP1 protein colocalizes with the TJ protein occludin in mouse BMEC by coimmunoprecipitation and in rat brain vessels by immunohistochemistry. Epithelial membrane protein 1 expression was transiently induced in laser-capture microdissected rat brain vessels after a 20-min global cerebral ischemia, in parallel with the loss of occludin immunoreactivity. The study identifies EMP1 as a novel TJ-associated protein of the BBB and suggests its potential role in the regulation of the BBB function in cerebral ischemia.
Introduction
Despite significant advances made in understanding the structure and biology of the blood—brain barrier (BBB) since its first description by Paul Ehrlich in 1903, it continues to pose a major challenge for the treatment of central nervous system diseases. The restrictive nature of the BBB is conferred by the tight junctions (TJs), polarized transporter systems, and unique metabolic properties of the brain endothelial cells.
The major distinctive feature between brain and peripheral vascular beds are the TJs, complex multiprotein structures sealing adjoining endothelial cell membranes. Together with the adherens junctions, the TJs form the junctional complex that restricts the permeability of the BBB. Whereas adherens junctions are ubiquitous in the vasculature, TJs are selectively expressed in cellular layers forming barriers of the body, where they limit paracellular permeability by separating the apical and basolateral cell membranes. The TJs consist of several transmembrane proteins, most notably occludin and claudins, whose extracellular domains provide a cell—cell interface, whereas intracellular domains connect the cell membrane to the cytoskeleton by associating with the MAGUK family of proteins (i.e., ZO-1, ZO-2, and ZO-3) (reviewed in Bazzoni and Dejana, 2004).
The intracellular N- and C terminals of occludin are connected by two extracellular loops spanning the intercellular cleft; the second extracellular loop is necessary for establishing high BBB electrical resistance (Wong and Gumbiner, 1997). Occludin's C-terminal tail controls permeability of low-molecular-weight substances (Balda et al., 1996). Occludin expression is decreased in many disease states accompanied by increases in BBB permeability (Yang et al., 2007).
The claudin family of proteins includes 24 members described so far in mammals, among which claudin-1, -3, -5, and -12 are expressed at the BBB (Nitta et al., 2003). Claudin extracellular loops form the primary seals of the TJs by interacting in a homo- and heterophilic manner. The first and larger extracellular loops harbor a WWCC motif, which is conserved among all claudins (Katoh and Katoh, 2003), whereas the carboxy tail has an YV PDZ binding site. Loss of claudin-1, but not claudin-5, from cerebral vessels occurs under pathologic conditions associated with increased BBB permeability including stroke (Lippoldt et al., 2000).
Occludins and claudins assemble into heteropolymers and form intramembranous strands along endothelial cell contacts, which have been proposed to contain fluctuating channels regulating the selective diffusion of ions and hydrophilic molecules. Through their extracellular loops, they interact with the junctional adhesion molecules JAM-1, JAM-2, and JAM-3 to form the extracellular part of TJs (reviewed in Bazzoni and Dejana, 2004).
To identify novel structural or regulatory molecules that associate with TJs in brain microvascular endothelial cells (BMEC), we constructed a subtractive cDNA library from freshly isolated porcine BMEC (pBMEC; tester) and porcine aortic endothelial cells (AOEC; driver). Among the 26 pBMEC-specific genes isolated from this library, the epithelial membrane protein 1 (EMP1) exhibited both high-expression specificity for BMEC and structural similarity to the TJ molecules occludin and claudin. We show here that EMP1 colocalizes with TJs in both isolated mouse BMECs and
Materials and methods
Isolation and Culture of Endothelial Cells
Porcine BMECs were isolated as described (Franke et al., 2000) with some modifications. Briefly, brains were removed from freshly slaughtered pigs; meninges and large meningeal vessels were peeled off and choroids plexus was removed; gray matter was then dissected and cut into small pieces. Tissue was disaggregated using Dispase II (Roche Diagnostics, Mannheim, Germany) in Earle's M199 medium (PAA, Pasching, Austria) at 37°C for 2 h as described by Franke et al. (2000). Capillaries were then isolated by centrifugation in 7.5% (w/v) dextran (MW 100,000 to 200,000; Sigma, Taufkirchen, Germany) solution. To release BMEC from brain capillaries, sediments were digested using collagenase D (Roche Diagnostics) for 1 h at 37°C (similar to as described by Franke et al., 2000) in the presence of the protease inhibitor Na-
Primary cultures of mouse brain endothelial cells (MBECs) were generously provided by Dr Maria Moreno (National Research Council-Institute for Biological Sciences (NRC-IBS), Ottawa, ON, Canada). Briefly, MBECs were isolated from cortices of 19-day-old mice by sequential filtration through 112 and 20 μm Nitex meshes and subsequent digestion by Collagenase IV (Sigma, Oakville, ON, Canada). The endothelial cells were plated on 0.5% gelatin-coated plastic culture dishes and fed every 3 to 4 days with Dulbecco's modified Eagle's medium (high glucose—4,500 mg/L) (Invitrogen, Carlsbad, CA, USA) containing 10% fetal bovine serum, 5% human serum (HyClone, Logan, UT, USA), 20% murine melanoma cell-conditioned media, 25 μg/mL insulin, 25 μg/mL transferrin, 25 ng/mL selenium, 20 μg/mL endothelial cell growth supplement (Collaborative Biomedical Products, Bedford, MA, USA), and 10 U/mL Heparin (Sigma, Oakville, ON, Canada).
Porcine AOECs that were isolated as described (Lauth et al., 2000) were kindly provided by Professor Hecker (University of Goettingen, Germany). Isolated AOECs were cultured as described (Lauth et al., 2000). More than 99% of AOECs in culture were positive for Factor VIII. In all experiments, cultured AOECs were used as primary (P0) cells, because it was difficult to procure sufficient number of freshly isolated cells and to enable cell transport.
Isolation of RNA and mRNA
Total RNA was isolated from cells or tissues using Trizol reagent (Invitrogen, Karlsruhe, Germany). The integrity of total RNA was analyzed using formaldehyde agarose gels. Dynabeads (Dynal, Oslo, Norway) were used for the enrichment of mRNA. Total RNA in water was mixed with an equal volume of binding buffer (20 mmol/L Tris-HCl, pH 7.5, 1.0 mol/L LiCl, 2 mmol/L EDTA) and heated to 65°C for 2 mins to disrupt secondary structures. The reaction was stopped by placing the sample on ice; the sample was then mixed with the Oligo dT(25) magnetic beads and allowed to anneal by rotating continuously on a mixer for 5 mins at room temperature. After placing the tube on a magnet for 1 to 2 mins to pellet the beads, the supernatant was carefully removed and mRNA captured (annealed) to the beads was washed twice before eluting with 10 mmol/L Tris-HCl (pH 7.5) at 80°C for 2 mins.
Construction of a Subtractive cDNA Library and Differential Screening
The preparation of the subtractive cDNA library was performed with the commercial PCR-Select cDNA subtraction kit (Clontech, Heidelberg, Germany) according to the manufacturer's specifications, using mRNA from either freshly isolated pBMEC (tester) or primary P0 porcine AOEC (driver). The efficacy of the subtraction was controlled using comparative PCR for the housekeeping gene glyceraldehyde-3-phosphate dehydrogenase (GAPDH).
Clones from the subtractive cDNA library were verified by differential hybridization of BMEC versus AOEC using the PCR-Select Differential Screening Kit (Clontech) applying forward- and reverse-subtracted probes, respectively, on two filters that were identically spotted with 92 subtractive clones, two negative controls, the housekeeping gene GAPDH, and the BHS marker apolipoprotein Al. Only those clones that had a ratio of at least 5 for the hybridization signal obtained with the subtracted tester cDNA compared with the subtracted driver cDNA were selected for DNA sequencing.
Both procedures are described in detail in Supplementary Methods.
Bioinformatical Analysis of Identified Blood—Brain Barrier Targets Using Litminer™
To analyze whether identified differentially expressed genes have been reported in association with the BBB, literature mining was performed using LitMiner software tool (http://www.litminer.ca) as described (Haqqani et al., 2007). This program performs searches of ‘knowledge terms’ (tissue, process, disease) and links them to ‘query genes’ (and their aliases) in titles and abstracts of published literature. LitMiner also enables easy generation of interactive tables (see Supplementary Table 2) and schematics of literature networks among selected genes and knowledge terms. All steps performed by LitMiner can be automatically replicated manually using Entrez Gene and HUGO Gene Nomenclature Committee (HGNC) to obtain synonyms/aliases for each gene, PubMed to find entries that describe each gene synonym and the selected knowledge term in the same document, and Gene Ontology to obtain molecular/biologic function of a gene.
Published literature was searched for entries containing knowledge terms ‘blood—brain barrier,’ ‘brain endothelial cells,’ ‘brain vessels,’ or ‘vessels’ in conjunction with each differentially expressed gene; molecular and biologic roles of identified genes were determined using a built-in Gene Ontology term function in LitMiner.
Northern Blot Analysis
A 15 μg portion of total RNA from the cells and tissues was separated on 1.5% formaldehyde agarose gels, transferred onto a Hybond-XL membrane (Amersham), and fixed onto the filter in a UV Crosslinker (Stratagene, Amsterdam, The Netherlands) at 70,000 μJ/cm2. The filter was then stained for 1 min in methylene blue solution to visualize the RNA and washed in water for 2 mins to de-stain the background. The stained filters were photographically documented. The hybridization was performed with radioactively labeled (Rediprime II; Amersham) cDNA probes, which were purified using ProbeQuant G-50 Columns (Amersham) and ExpressHyb solution (Clontech) according to the manufacturer's specifications. After verifying hybridization using phosphoimager (FLA-5000; Fujifilm, Duesseldorf, Germany), autoradiograms were prepared on Biomax MS films (Kodak, Taufkirchen, Germany).
Reverse Transcription-PCR and Quantitative PCR
For reverse transcription-PCR (RT-PCR) experiments, cDNA was prepared from total RNA after a pretreatment with DNase I (Invitrogen, Karlsruhe, Germany) by random priming using SuperScript II (Invitrogen, Karlsruhe, Germany) for reverse transcription.
Quantitative RT-PCR (qRT-PCR) was performed in porcine cell cultures or rat tissue samples using either the Opticon (MJ Research, Waltham, MA, USA) or ABI Prism 7700 Sequence Detector System (Applied Biosystems, Foster City, CA, USA) together with the QuantiTect SYBR Green PCR Kit (Qiagen) or SYBR-Green PCR Core Reagent Kit (Applied Biosystems). Primer sequences are listed in Supplementary Table 1.
The reference and target genes were amplified in duplicate during each experiment, and the amount of a given target gene was normalized to the reference (18S rRNA or β-actin). For each sample, the formation of PCR products was monitored fluorimetrically on the basis of the incorporation of SYBR Green. The PCR cycle number at which fluorescence reaches a threshold value of 10 times the standard deviation of baseline emission was used for quantitative measurements. This cycle number represents the cycle threshold (C
Western Blot and Immunoprecipitation
Porcine proteins were separated for Western blot analyses on 12.5% polyacrylamide gels and transferred onto nitrocellulose membranes (Whatman, Dassel, Germany). An immunochemical staining of transferred proteins was performed with a polyclonal EMP1 antiserum from rabbits (1:4,000; Eurogentec, Seraing, Belgium), which was raised against a peptide from the second extracellular loop of porcine EMP1 (which has no homology to second extracellular loops of occludin or claudin) and tested for specificity using peptide enzyme-linked immunosorbent assays. Visualization was achieved using a secondary antibody conjugated to alkaline phosphatase (anti-rabbit-IgG-antibody (goat), 1:5,000; Sigma, Taufkirchen, Germany), followed by color development with 0.016% (w/v) nitrotetrazolium blue chloride (Sigma, Taufkirchen, Germany) and 0.033% (w/v) 5-bromo-4-chloro-3-indolylphosphate-disodium salt (Sigma, Taufkirchen, Germany) in AP buffer (100 mmol/L Tris-HCl, 100 mmol/L NaCl, 5 mmol/L MgCl2, pH 9.5).
Coimmunoprecipitation of ZO-1 and EMP1 with occludin and claudin-5 antibodies was performed in cultured MBEC. Mouse brain endothelial cells (P2) grown to confluence in T-25 flasks were washed two times with physiologic saline (PBS); membrane proteins were extracted by adding 500 μL of Thesit (Sigma, Oakville, ON, Canada), a nonionic detergent used for the extraction of membrane-bound proteins, containing 0.5 μL protease inhibitors (Sigma, Oakville, ON, Canada) to each monolayer. The samples were incubated on ice for 30 mins with intermittent vortexing and subsequently sonicated two times for 10 secs. Cellular debris was pelleted by centrifugation, supernatant was collected, and protein concentration determined using BCA protein assay kit (Pierce/Biolynx, Brockville, ON, Canada). The supernatants (200 μL; 1 to 1.5 mg protein per mL) were precleared with 50% Protein A sepharose (Amersham, GE Healthcare, London, ON, Canada) for 1.5 h at 4°C and centrifuged at 500 ×
Cerebral Ischemia/Reperfusion
The global cerebral ischemia/reperfusion model was chosen for this study, as it produces a reproducible biphasic opening of the BBB and results in more uniform injury to brain vessels in the forebrain regions compared with focal ischemia models. The detailed experimental protocol was described earlier (Preston et al., 1993; Haqqani et al., 2005). Bilateral cerebral ischemia (20 mins) was induced in rats under sodium pentobarbital anesthesia (65 mg/kg, intraperitoneally) and mechanical ventilation with a 30:70 O2/N2 mixture by bilateral occlusion of carotid arteries (two-vessel occlusion) combined with blood withdrawal through a tail-artery cannula to maintain arterial pressure at 42 to 47 mm Hg. After blood reinfusion, clamp removal and wound closure, rats were taken off ventilation, were maintained normothermic (37.0°C to 37.5°C) until recovery from anesthetic and then returned to housing. Control animals were sham operated, but their arteries were not clamped and their blood pressure was not lowered. After the surgery, the animals were allowed to recover for a period of 1, 6, or 24 h and then killed by decapitation; their brains were rapidly removed, wrapped in aluminum foil, and placed on dry ice. Abdominal aorta and liver from sham-operated controls were also dissected and flash-frozen for immunochemistry. Experiments were performed at the Animal Care and Veterinary Services of the National Research Council (Ottawa, ON, Canada) in accordance with guidelines of the Canadian Council on Animal Care.
Tissue Sectioning and Laser Capture Microdissection
Experiments were performed as described elsewhere (Haqqani et al., 2005). Briefly, frozen rat brains were embedded in Tissue-Tec freezing medium (Miles Laboratories, Elkhart, IN, USA) and sectioned in a cryostat (Jung CM3000; Leica, Richmond Hill, ON, Canada) at 8 μm thickness. Sections were placed on Superfrost Plus microscope slides (Fisher Scientific, Nepean, ON, Canada) and kept at –80°C until use. The sections were fixed in 75% ethanol for 30 secs, rinsed in water, and stained with
Immunohistochemistry
12-μm-thick frozen (−80°C) sections of rat abdominal aorta, liver, or brain from saline-perfused sham-operated animals and brain sections from two-vessel occlusion-treated animals after 1, 6, or 24 h of reperfusion were used for immunohistochemistry. Slides were quickly thawed at room temperature for a few seconds, fixed in 75% ethanol in PBS for 30 mins at 4°C, and then rinsed in PBS and subsequently in Milli-Q water. Blocking was performed with 10% normal goat serum (NGS; Sigma, Oakville, ON, Canada) containing 0.01% Triton X-100 in PBS for 1 h at room temperature. Immunostaining was performed at room temperature using successively the following primary and secondary antibodies: (a) for abdominal aorta: mouse anti-occludin (1:100; Zymed, San Francisco, CA, USA) in 5% NGS in PBS for 1 h, goat anti-mouse IgG Alexa680 (1:300; Invitrogen, Burlington, ON, Canada) in PBS for 1 h, rabbit anti-EMP1 (1:100; Lee et al., 2005) in 5% NGS in PBS for 1 h, goat anti-rabbit IgG Alexa568 (1:300; Invitrogen) in PBS for 1 h, rat anti-mouse CD31 (1:300; BD Pharmingen) in 5% NGS in PBS for 1 h, and goat anti-rat IgG Alexa568 (1:500; Invitrogen) in PBS for 1 h; (b) for brain: anti-occludin (1:100; Zymed) in 5% NGS in PBS for 1 h, goat anti-mouse IgG Alexa680 (1:300; Invitrogen) in PBS for 1 h; rabbit anti-rat EMP1 (1:300; Lee et al., 2005) in 5% NGS in PBS for 1 h, and goat anti-rabbit IgG Alexa568 (1:800; Invitrogen) together with
Slides were coverslipped in the fluorescent mounting medium (Dako Diagnostics, Mississauga, ON, Canada) spiked with 4′,6-diamidino-2-phenylindole (2 μg/mL; Sigma, Oakville, ON, Canada). Images were obtained using Axiovert 200M Fluorescent Microscope (Carl Zeiss, Maple Groove, MN, USA) with × 20 and × 63 objectives and the following exposure times: FITC (green) detecting
Results
Identification of Blood—Brain Barrier-Specific Genes
Blood—brain barrier-specific transcripts were isolated from a cDNA library that was constructed by subtracting mRNAs from primary P0 porcine AOECs and freshly isolated pBMECs. The subtraction process led to a 50,000-fold enrichment of BMEC transcripts as assessed by GAPDH-specific qRT-PCR (data not shown). The enriched cDNA library was screened using forward- and reverse-subtracted probes, and clones showing at least five-fold enrichment in BMEC-specific signal were selected for sequencing. Supplementary Table 2 lists genes identified by this strategy, their molecular and biologic functions, and literature associations with selected ‘knowledge terms’ (i.e., blood—brain barrier, brain endothelial cells, brain vessels, and vessels) analyzed by LitMiner. Out of 26 BMEC-enriched transcripts identified, 8 (30%) had been directly linked to the BBB in the literature (Supplementary Table 2).
Expression Patterns of Selected Blood—Brain Barrier Targets
From the list of the BBB-enriched transcripts, we selected five genes for further validation; these genes either have not been associated with the BBB in the literature (Supplementary Table 2) or have predicted membrane localization and unknown molecular/biologic function: (a) EMP1, (b) the ADP-ribosylation factor-like 5B (ARL5B), (c) the TSC22 domain family member 1 (TSC22D1), (d) the integral membrane protein 2A (ITM2A), and (e) the solute carrier family 35 member F2 (SLC35F2). Tissue-specific expression of these genes was analyzed by Northern blot using RNA from pBMEC and various other porcine organs. EMP1, ARL5B, ITM2A, and SLC35F2 transcripts were highly enriched in BMEC compared with the whole brain tissue (Figure 1), suggesting that they are brain-endothelial cell-selective. Reverse transcription-PCR analyses confirmed at least approximately 10-fold enrichment of EMP1, ARL5B, ITM2A, and TSC22D1 in freshly isolated BMEC compared with AOEC (Supplementary Table 3).
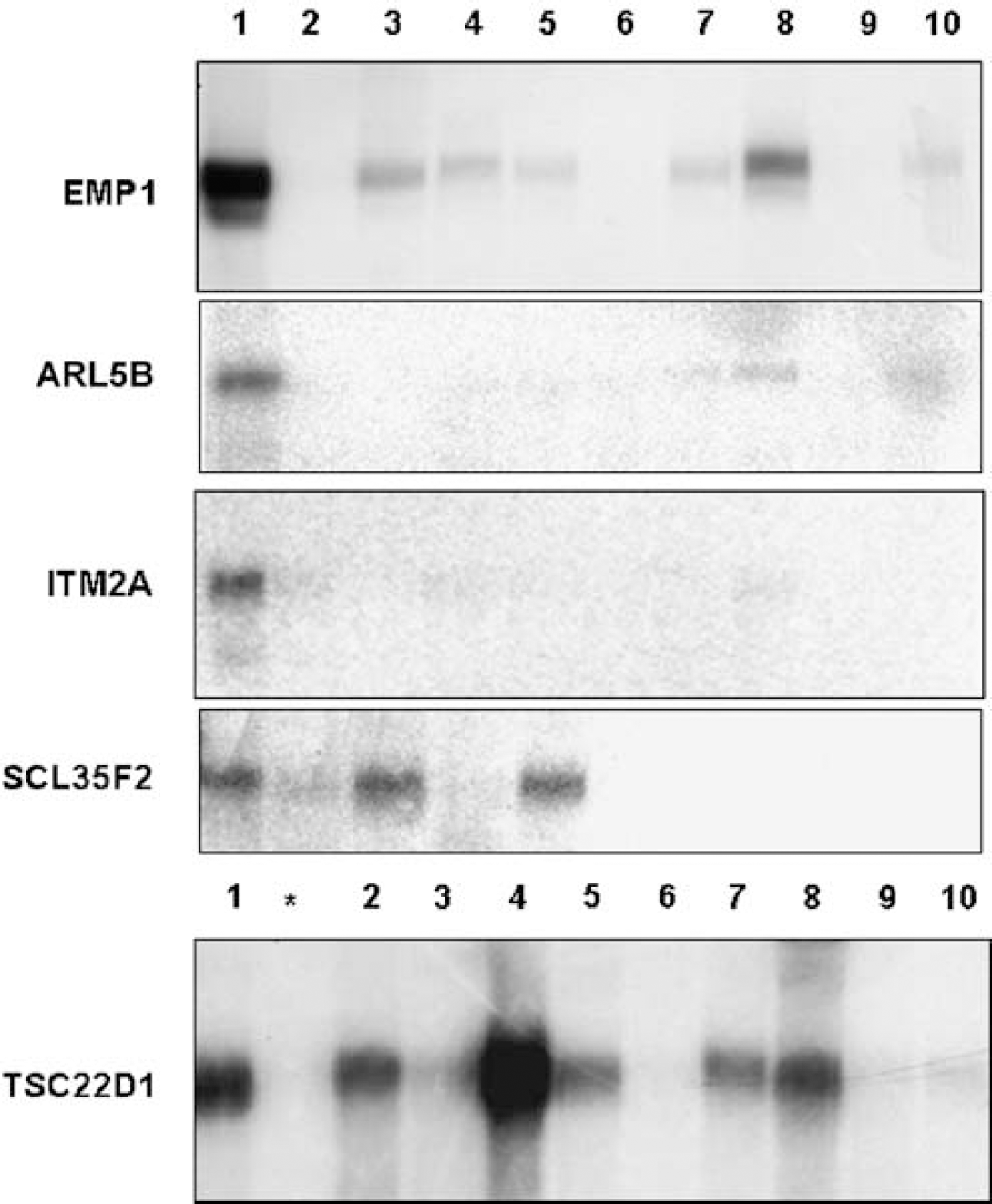
Validation of genes selectively enriched in brain microvascular endothelial cells (BMEC) using Northern blot analyses. Validation of selected pBMEC-enriched transcripts was performed by Northern blot analysis as described in Materials and methods. Subtractive clones for EMP1, ARL5B, ITM2A, SCL35F2, and TSC22D1 were used as specific hybridization probes with total RNA from porcine: BMEC (lane 1), whole brain tissue (lane 2), colon (lane 3), heart (lane 4), kidney (lane 5), liver (lane 6), lung (lane 7), spleen (lane 8), choroid plexus (lane 9), and testis (lane 10). Asterisk (∗) indicates additional hybridization with porcine aortic endothelial cells (AOEC) for TSC22D1.
With the exception of SLC35F2, four transcripts (EMP1, ARL5B, ITM2A, and TSC22D1) were repressed in primary and propagated pBMEC cultures (Figures 2A and 2B) compared with freshly isolated BMEC. This observation is consistent with reported loss of phenotypic properties of BMEC in culture, including downregulation of BBB-specific genes/proteins such as apolipoprotein A1 (Weiler-Güttler et al., 1990).
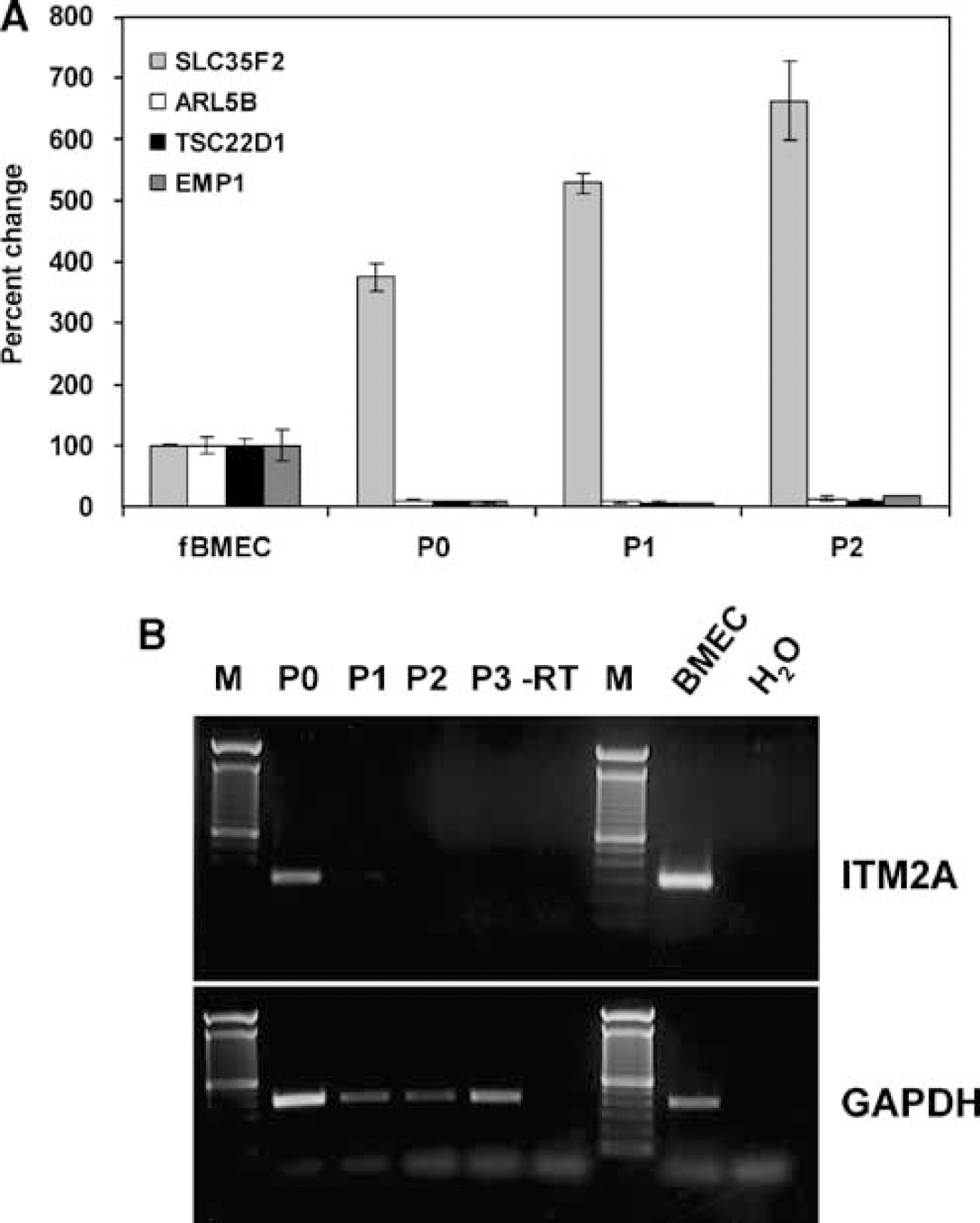
The expression of BMEC-enriched genes in cultured cells. The expression of selected BMEC-enriched genes was determined by quantitative RT-PCR (
Collectively, the data suggest that four out of the five validated transcripts are highly enriched in or, in some cases, even selectively expressed by BMEC and may participate in establishing a unique brain endothelial phenotype.
Epithelial Membrane Protein 1 is a Potential Blood—Brain Barrier Tight Junction Protein
The subsequent studies were focused on EMP1, because of its membrane localization and topological similarities to known TJ molecules.
The subtractive clone S231 showed significant homology to transcripts from other species encoding EMP1. The coding region of porcine EMP1 was obtained by screening a cDNA library from BMEC (accession number EF577081). The sequence was then translated
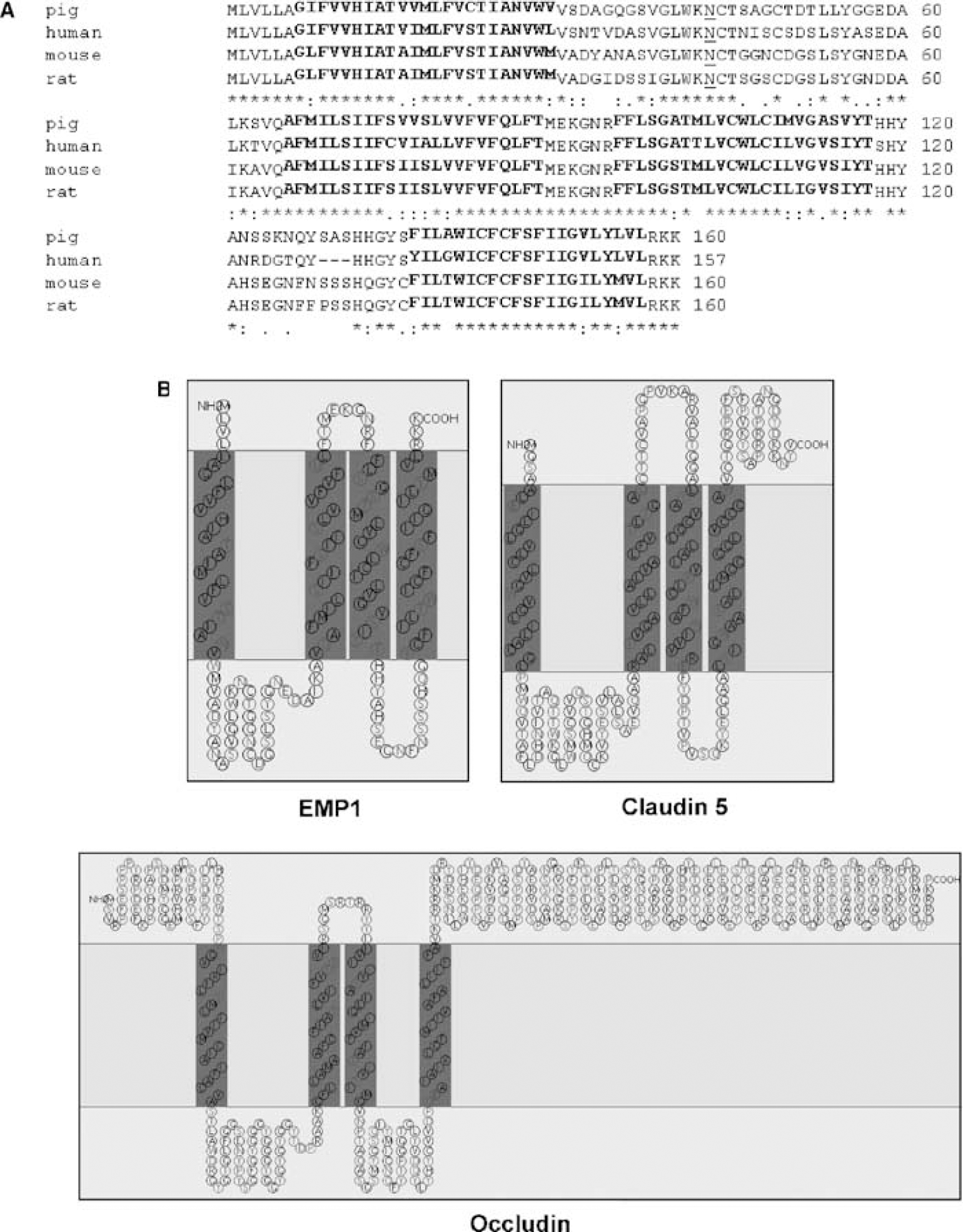
CLUSTAL alignment of EMP1 from different species and its topological similarity to tight junction molecules. (
On the basis of similar topology with the BBB TJ proteins occludin and claudin (Figure 3B), we hypothesized that EMP1 might be a hitherto unknown TJ protein. To investigate this further, polyclonal antibodies against porcine EMP1s were raised using a peptide from the second extracellular loop as antigen. Western blot showed a double band recognized by EMP1 anti-serum in freshly isolated pBMEC (Figure 4A), but not in primary P0 porcine AOEC (Figure 4A), corresponding to previously described migration patterns of EMP1 (∼20 kDa) and EMP1 glycosylated at a conserved
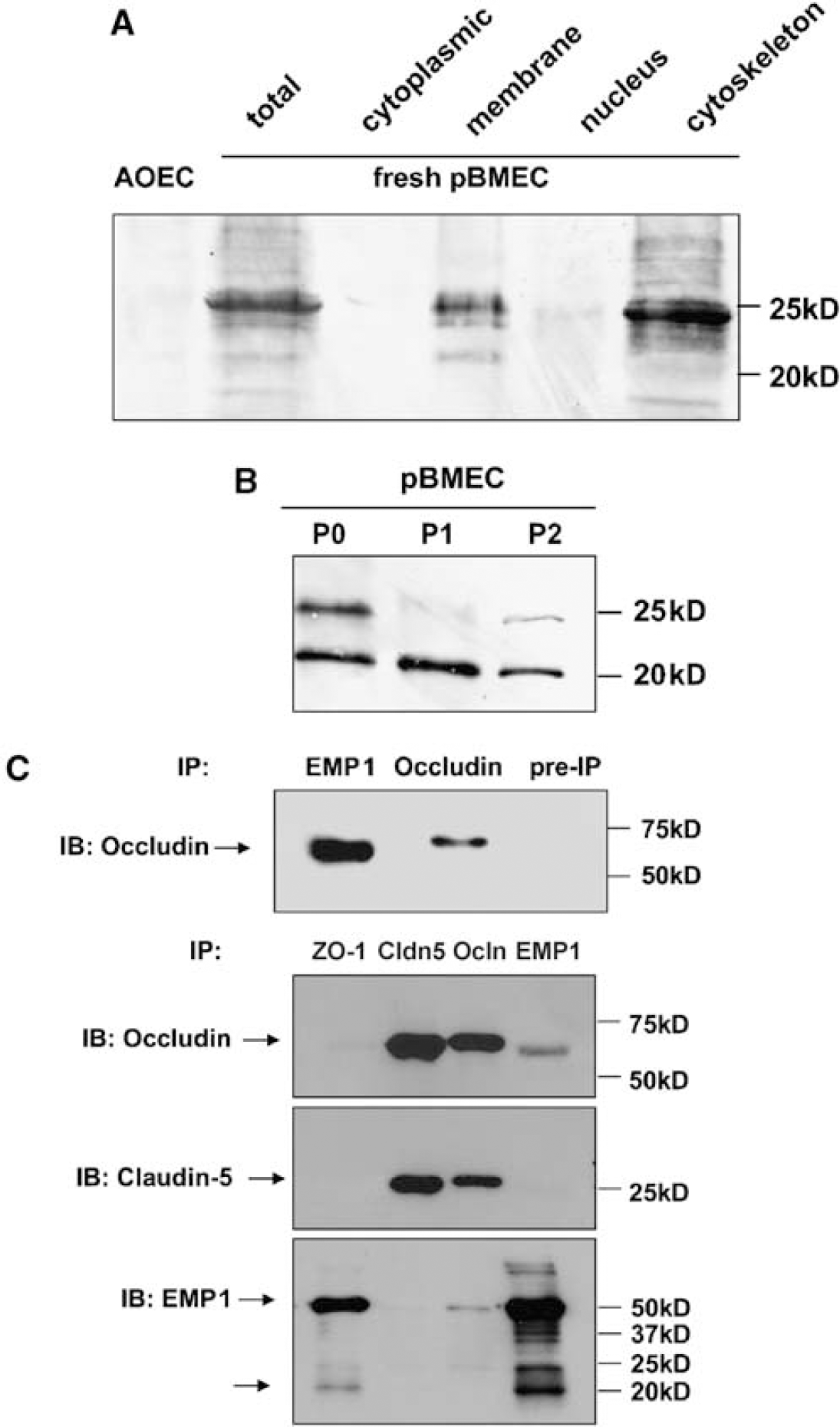
Western blot analyses of EMP1 protein expression in brain microvascular endothelial cells (BMEC) and its interaction with occludin and ZO-1. Total cell extract from porcine aortic endothelial cells (AOECs), freshly isolated porcine BMEC (pBMEC), various cellular fractions of freshly isolated pBMEC (
To determine whether EMP1 interacts with other TJ proteins, we analyzed coimmunoprecipitation of TJ protein complexes from extracts of cultured MBEC (P2). In this set of experiments, EMP1 was detected using an antiserum obtained against the second extracellular loop from rat EMP1 (Lee et al., 2005) and tested for EMP1 immunoreactivity in solid-phase enzyme-linked immunosorbent assay. The EMP1 antibody coimmunoprecipitated occludin and ZO-1, but not claudin-5 (Figure 4C). Epithelial membrane protein 1 could be coimmunoprecipitated using antibodies against ZO-1 and occludin. The EMP1 antibody detected multiple bands pointing to the existence of EMP1 dimers (37 and 50 kDa) in addition to the described monomeric bands of 20 and 24 kDa. A double band specific for EMP1 in samples coimmunoprecipitated with ZO-1, occludin, and EMP1 antibodies (Figure 4C) suggests that EMP1 physically interacts with occludin and ZO-1, but not with claudin-5, in TJ complexes in cultured MBEC. As the efficacy of extraction of TJ proteins using described protocols was not determined, the data presented are qualitative in nature and are suggestive of EMP1 interacting with at least some TJ proteins.
Consistent with these results, immunohistochemistry with antibody against rat EMP1 (Lee et al., 2005) showed lack of both EMP1 and occludin immunoreactivities in sections of rat abdominal aorta (Figure 5Aa to Ad), in contrast to a brain vessel-specific expression of EMP1 and its close association/colocalization with both occludin immunoreactivity and endothelial ligand to lectin RCA-1 (Figure 5Ba to Be) in sections of the rat brain. In sections of rat liver, used as a positive control, a colocalization of occludin and EMP1 along hepatocyte borders as well as nonspecific staining of nuclei with EMP1 antibody was observed (Figure 5Ca and Cb), similar to what was reported by Lee et al., 2005. Collectively, the results suggest that EMP1 is likely a component of TJs in brain vessels
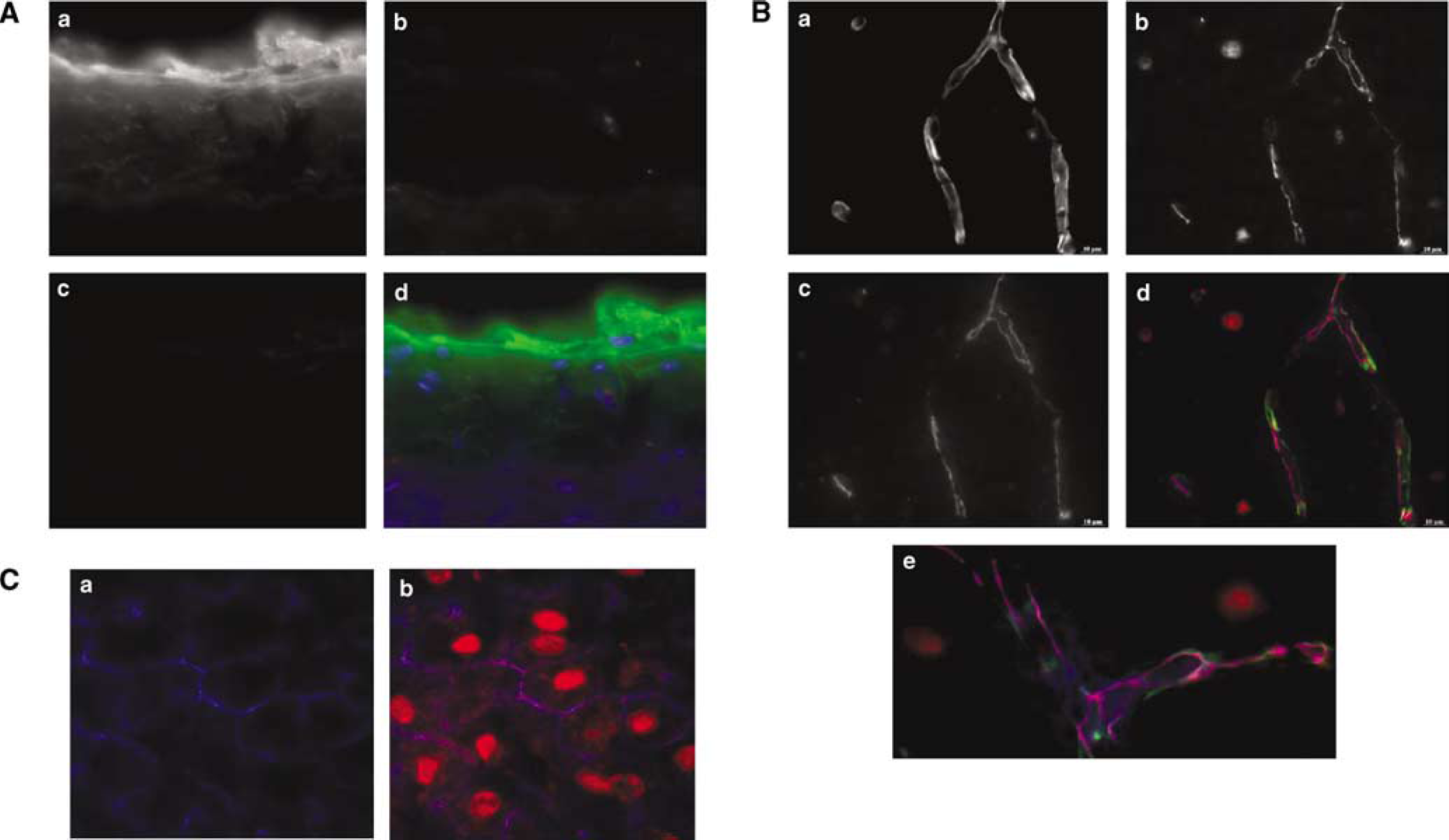
Localization of EMP1 in rat aorta, brain, and liver by immunohistochemistry. Cryosections of rat aorta (Aa to Ad), brain (Ba to Be), and liver (Ca and Cb) were immunostained for EMP1 and occludin, the endothelial-selective antigen CD31 (aorta), or with vessel-binding lectin RCA-1 (brain vessels) as described in Materials and methods. (
Epithelial Membrane Protein 1 in Brain Ischemia
Changes in EMP1 expression were analyzed in brain vessels after a transient 20-min global brain ischemia/reperfusion in rats, induced by two-vessel occlusion and hypotension as described (Preston et al., 1993). This model is known to induce a biphasic BBB disruption at 1 and 24 h, with transient recovery at 6 h after ischemia (Haqqani et al., 2005). Forebrain small vessels were LCM captured from brain sections 1, 6, and 24 h after a 20-min global ischemia. Laser capture microdissection capturing of cells or tissues from organ sections enables preservation of their molecular characteristics
Immunohistochemistry showed EMP1 and occludin colocalization at 1 and 6 h after ischemia (Figures 6B and 6C) as well as an increased intensity of EMP1 staining compared with sham-operated controls (Figures 6A to C). Quantitative RT-PCR using LCM-captured vessels from sections adjacent to those used for immunohistochemistry revealed a moderate and reversible upregulation of EMP1 mRNA expression in the brain vascular fraction 6 h after reperfusion (Figure 6E). At 24 h after ischemia EMP1 immunoreactivity remained strong and tightly associated with cellular contacts (Figure 6D) despite almost complete loss of occludin immunoreactivity from brain vessels (Figure 6D), suggesting that the two proteins are differentially regulated in ischemic brain vessels and may participate in the regulation of the BBB permeability.
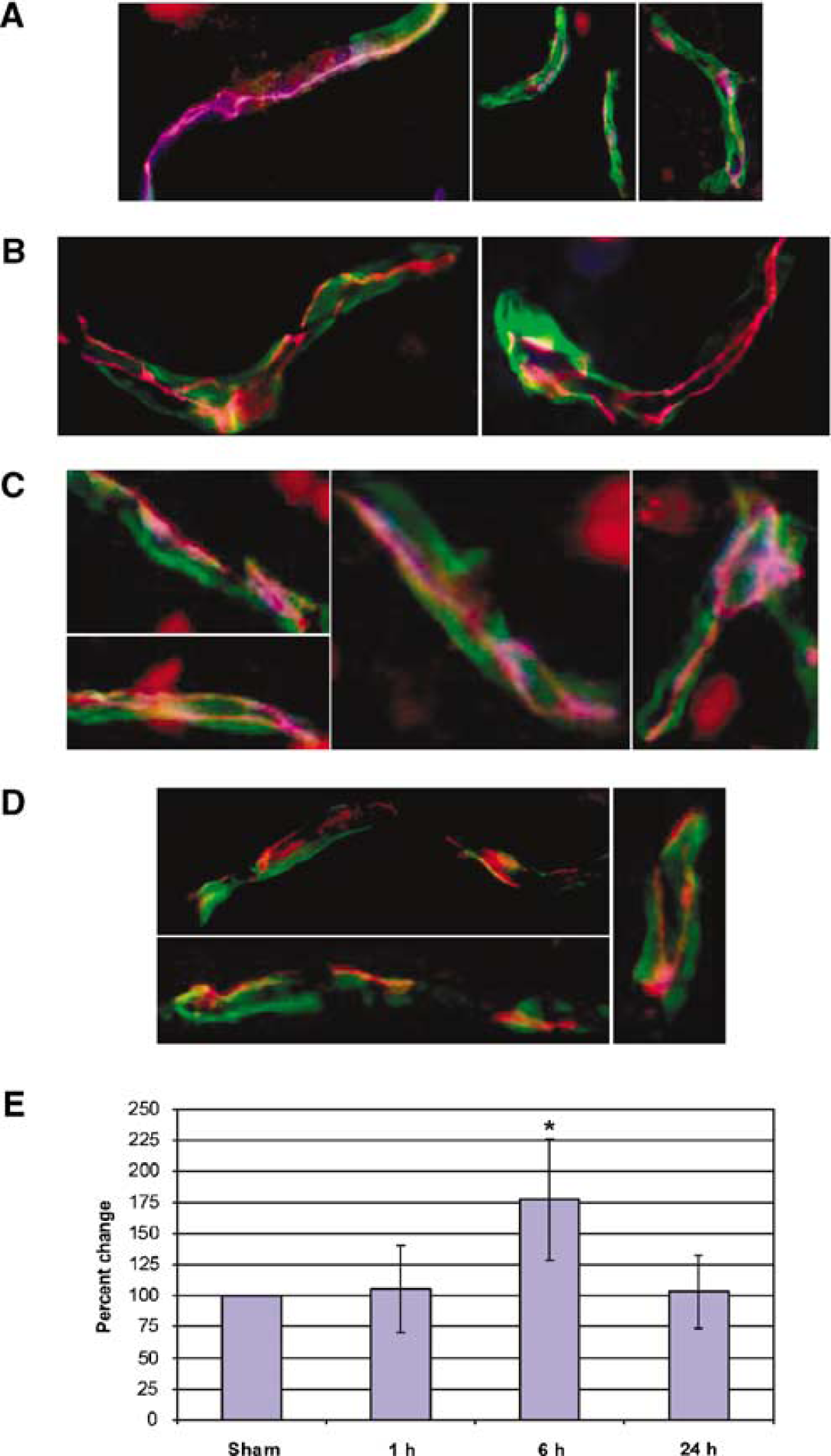
Changes in EMP1 and occludin expression in rat brain vessels after a transient global cerebral ischemia. (
Discussion
Despite recent advances in understanding the biology and molecular characteristics of the BBB, the pharmacological modulation of TJs and the BBB permeability remains a major challenge. The discovery of novel pharmacological targets to modulate the BBB was facilitated by screening platforms, including genomic (Kallmann et al., 2002; Shusta, 2005), proteomic (Haqqani et al., 2005, 2007), and phage display (Muruganandam et al., 2002; Wang et al., 2007).
In this study, the cDNA library enriched for BBB-specific transcripts was constructed by subtracting mRNAs from porcine primary P0 aortic and freshly isolated brain endothelial cells. The subtractive hybridization identified 26 transcripts enriched in pBMEC; among these transcripts were several proteins with known BBB functions, including apolipoprotein A1 (Weiler-Güttler et al., 1990), calcitonin receptor-like (Kis et al., 2002), and vinculin (Schulze and Firth, 1993), as well as some rat and human brain microvessel-enriched transcripts (e.g. secreted protein, acidic, cysteine-rich (SPARC)-like 1, ITM2A, and ferritin) recently cataloged using either serial analysis of gene expression (SAGE) (Emerson and Drewes, 2006) or subtractive hybridization (Li et al., 2002; Shusta et al., 2002) approaches. Other known BBB-enriched genes (e.g. mdr-1, transferrin receptor) might have been represented in the library, but they were not identified because the subtractive profiling was not comprehensive (i.e. not every clone in the subtracted libraries was evaluated).
Integral transmembrane protein 2a and several novel BMEC-enriched genes present in the library, including SLC35F2, ARL5B, TSC22D1, and EMP1, were further analyzed. Integral transmembrane protein 2a, recently identified as brain microvessel-enriched transcript by subtractive hybridization of rat (Li et al., 2002) or human (Shusta et al., 2002) brain microvessels against mixed liver and kidney driver cDNA, is a single-pass type II membrane protein with roles in T-cell activation, thymocyte development, and chondrogenic differentiation (van den Plas and Merregaert, 2004 and references therein). Solute carrier family 35 member F2 belongs to the solute carrier family SLC35, which consists of at least 17 molecular species in humans (Ishida and Kawakita, 2004); the characterized family members encode nucleotide sugar transporters localizing in the Golgi apparatus and/or the endoplasmic reticulum. ADP-ribosylation factor-like 5B belongs to the family of ADP-ribosylation factor regulatory GTPases (Kahn et al., 2005) involved in a multitude of processes, such as cell growth, signal transduction, organization of the cytoskeleton, and regulation of membrane trafficking. TSC22 domain family member 1, a member of the TSC-22/Dip/Bun family, is a potential tumor suppressor gene (Uchida et al., 2000) and a transcriptional repressor on C-type natriuretic peptide promoter (Ohta et al., 1996). Epithelial membrane protein 1 was originally identified in brain tumors as a c-myc target that plays a role in cell proliferation and differentiation (Ben-Porath and Benvenisty, 1996). EMP1 is widely expressed in the adult mouse (Lobsiger et al., 1996) with high expression found in the developing central nervous system and peripheral nervous system at the onset of neurogenesis (Wulf and Suter, 1999). In the liver, EMP1 has been described as a putative intercellular junction protein (Lee et al., 2005). The functional roles of these genes in brain vasculature remain unknown.
Interestingly, with the exception of pronounced upregulation of SLC35F2, all identified novel BMEC-enriched genes were dramatically downregulated in cultured pBMEC. A recent study by Calabria and Shusta (2008) identified 25 transcripts with different abundance between isolated brain microvessels and cultured rat BMEC; among these, the BBB control genes, including glucose transporter, P-glycoprotein, and transferrin receptor, were significantly downregulated on cell culture. Both studies reinforce previous observations that cultured BMECs undergo a significant de-differentiation/adaptation reflected in the expressed transcriptome.
We focused further study on EMP1 because of its structural similarity to the known TJ molecules occludin and claudin. Epithelial membrane protein 1, together with EMP2, EMP3, and PMP22, belongs to the peripheral myelin protein PMP22 gene family (Taylor and Suter, 1996). In epithelia, PMP22 is coexpressed with occludin and ZO-1 at or near TJs (Notterpek et al., 2001). Another member of the family, EMP2, has been shown to regulate surface expression of αvβ3 integrin, caveolins, and glycosylphosphatidyl inositol-linked proteins (Wadehra et al., 2005), thus modulating cell—matrix and cell—cell interactions.
In this study, EMP1 was detected in membrane and cytoskeletal protein fractions of pBMEC; in mouse BMEC, EMP1 coimmunoprecipitated with TJ-specific proteins, occludin and ZO-1, but not with claudin-5, suggesting that it interacts or forms complexes with some proteins assembled in TJs. Within brain tissue
The protein sequence of porcine EMP1, when compared with its mammalian homologes, confirms EMP1 as a tetraspan membrane protein (Ben-Porath and Benvenisty, 1996) that harbors PMP22-claudin protein domains. The W-GLW-C-C motif (Katoh and Katoh, 2003) within the first extracellular loop, which is characteristic for the members of the claudin family, was also found in EMP1, although with different spacing between the conserved residues. Although this resemblance may suggest a functional overlap between EMP1 and claudins, there are also major differences between the proteins. For example, the YV PDZ binding motif, found at the C termini of claudins, is missing in EMP1. Epithelial membrane protein 1 has very short cytoplasmic domains, suggesting that it may lack the ability to bind other cytoplasmic proteins, in contrast to claudins and occludin, which are linked to the cytoskeleton via ZO-1. The analogy with the C-terminal truncated occludin, which results in a several-fold increase in paracellular flux of small molecular weight tracers (Balda et al., 1996), suggests that increased expression of EMP1 may lead to disturbed barrier function of the TJ. Another difference between EMP1 and claudins is the range of their extracellular loop isoelectric points (p
The BBB breakdown is a hallmark of several neurologic diseases including stroke. In the global cerebral ischemia model used in this study, the BBB disruption occurs 1 and 24 h after reperfusion and has been associated with changes in vascular expression of proteins involved in ion and nutrient transport, inflammatory responses, and vascular remodeling (Haqqani et al., 2005). Curiously, analyses of the TJ molecules in this model are sparse. In other models of cerebral ischemia, disappearance of occludin immunoreactivity from brain vessels was observed in the penumbra region 3 days after a permanent middle cerebral artery occlusion in mice (Li et al., 2007), whereas both occludin and claudin-5 were reduced 2 h after a transient 90-min middle cerebral artery occlusion in rats (Yang et al., 2007). This study is the first to document changes in occludin expression
In summary, this study shows that EMP1, which displays structural similarities with occludin and claudins, associates with TJs in brain endothelial cells
Footnotes
Acknowledgements
We thank Ms Ally Pen for performing LitMiner analyses and Dr Hsuan-Shu Lee for providing anti-EMP1 antibody. We also thank all members of the former Esplora GmbH for their help in isolating porcine BMEC.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
