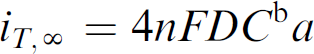Abstract
The development of highly sensitive biosensors for the detection of biomolecules, such as the biomarker interleukin-8 for the early detection of oral cancer, requires optimization of sensor design. To augment the performance of an electrochemical sensor, this study used a microscale, aptamer-based electrochemical sensor for detecting botulinum neurotoxin aptamer hybridization. We first used top—down lithographic processing to define the pattern of the electrodes and then used bottom—up manufacturing to modify the surface molecular properties for reducing nonspecific binding. We systemically examined the effects of the design parameters of an aptamer-based electrochemical sensor. Specifically, five key design parameters were examined: the area of the working electrode (WE), the area of the counter electrode (CE), the separation distance between the WE and CE, the overlap length between the WE and CE, and the aptamer concentration. Through an analysis of the signal and noise generated across variations of the different parameters, the significance of each parameter in sensor performance was determined. In particular, we found that the area of the WE was the only key parameter that influenced the performance of the sensor. The output signal level increased with the area of the WE and the signal-to-noise ratio was about constant in the tested range (i.e., from 0.02 to 4 mm2).
Introduction
Electrochemical biosensors synergize the biological recognition element of a biosensor with an electrochemical transducer and have been used to detect a variety of biological targets, such as uropathogenic bacteria, the hepatitis B virus, and the cholera toxin produced by Vibrio cholerae. 1 –4 Amenable to miniaturization, the development of microfabricated electrochemical biosensors has resulted in high sensitivity, portability, improved performance and spatial resolution, low power consumption, and the opportunity for integration with other technologies, such as microfluidics. 5,6 Such advantages enable Micro-Electrical-Mechanical Systems (MEMS)-based electrochemical biosensors to identify pathogens present in a variety of mediums and bodily fluids, such as blood, saliva, and air more quickly than previous, laboratory-based detection methods. 7 –10
Previous work has been done using an electrochemical biosensor in which a sensing paradigm was tested and developed. 7,11 –13 For example, Gau et al. 14 developed a MEMS-based electrochemical biosensor for the amperometric detection of Escherichia coli. The electrochemical detection system resulted from the synergy of microfabricated electrodes, self-assembled mono-layers via thiol chemistry, DNA hybridization, and enzyme-mediated signal amplification. 14 Gau's 14 electrochemical biosensor used three microfabricated gold electrodes: a working electrode (WE), a counter electrode (CE), and a reference electrode (RE). Using an identical electrode design, Wei et al. 11 and Wei and Ho 15 detected salivary interleukin-8 (IL-8) mRNA using a hairpin probe and botulinum neurotoxin (BoNT) with an aptamer. However, no study has been conducted to systematically determine the relationship between different sensor parameters and performance. To optimize and study the parameters and phenomena that operate as the foundation of the sensor, a parametric study was conducted in conjunction with an electrochemical detection method with enzymatic amplification, an aptamer probe, and a conducting polymer matrix for probe immobilization. 16 –20 The results of the study are reported herein. By manipulating the overall design of the electrochemical sensor according to hypotheses regarding possible sensing mechanisms, several parameters were studied to determine which design parameters are most critical to the function of the electrochemical sensor. Specifically, four design parameters in addition to the aptamer concentration were investigated: the area of the WE, the area of the CE, the separation distance between the WE and CE, and the overlap boundary shared between the WE and CE. These four parameters are illustrated in Figure 1A
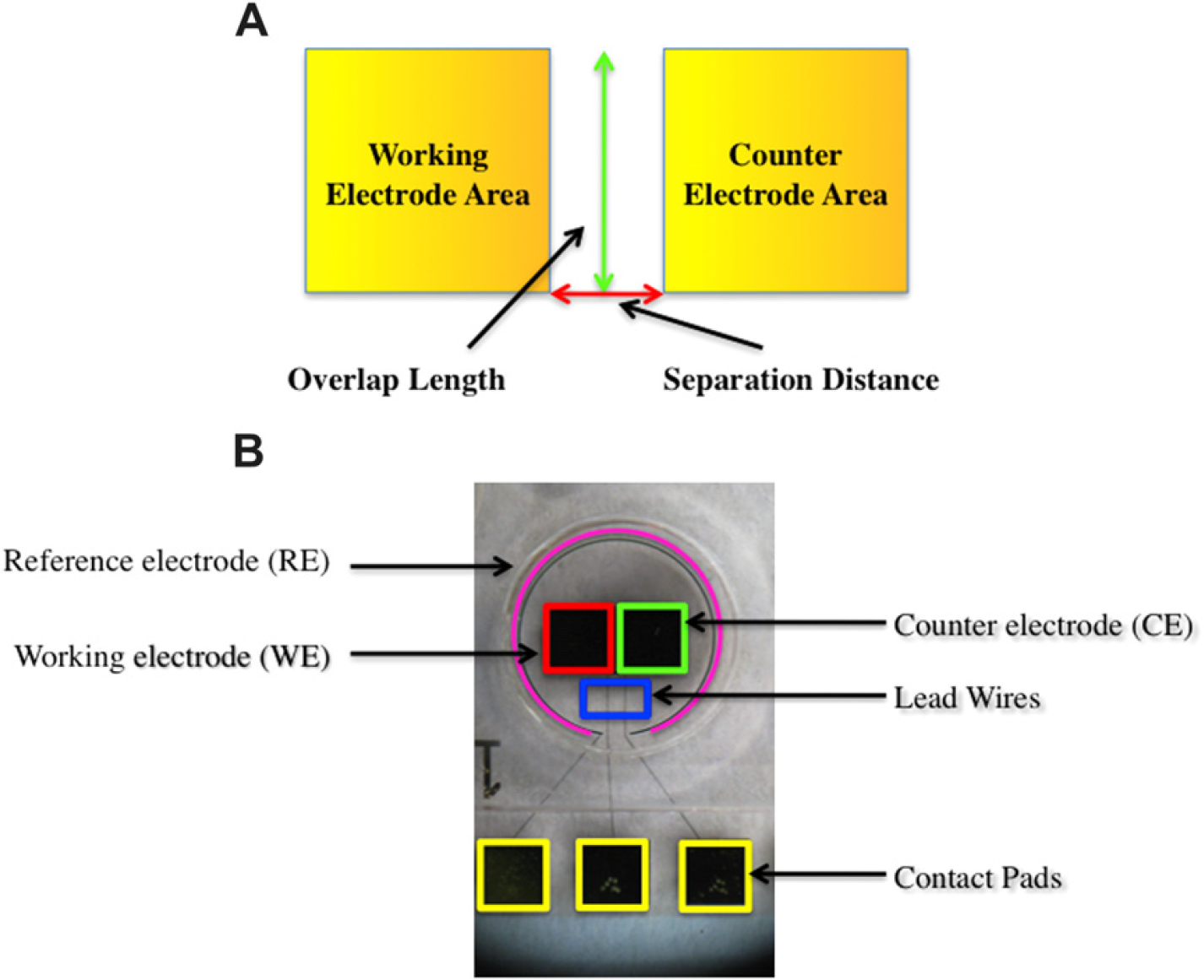
(A) Illustration of parameters for optimization of electrochemical sensor design. (B) Microfabricated electrochemical biosensor.
Working Principles
DNA-Based Biosensors
Nucleic acids can be exploited for their specificity in the development of biosensors. A single strand of DNA, consisting of a string of nucleotides, will bind with a high degree of selectivity and specificity to its complement strand. Thus, single-stranded DNA molecules may be used to detect target DNA in a variety of media. For example, Wei et al. 11 recently used an oligonucleotide hairpin probe for the detection of target RNA in saliva. A hairpin probe consists of a loop component that is complementary to the target and a stem component that contains the fluorescein reporter element. In the absence of the target molecule, the hairpin probe remains closed and the reporter element remains near the surface of the sensor. The molecule responsible for signal amplification (i.e., antifluorescein horseradish peroxidase [HRP]) is unable to bind the reporter element on the hairpin because of steric hindrance and thus unable to produce a signal. 11,13 However, in the presence of the target molecule (i.e., salivary RNA), the oligonucleotide sequence that forms the “loop” portion of the hairpin probe binds the target molecule, causing the hairpin probe to open and make the fluorescein accessible to the antifluorescein HRP enzyme that is responsible for signal generation in combination with an electron transfer mediator.
Although proving useful in the detection of nucleic acid targets, oligonucleotide hairpin probes are unable to detect proteins. On the other hand, aptamers are ligand-binding nucleic acid sequences that are able to form secondary structures and bind specifically to both nucleotide sequences and proteins; they are isolated from combinatorial oligonucleotide libraries via in vitro selection. 21 –25 For example, aptamers have been used for the detection of BoNT. 15 Moreover, the use of aptamers as part of the detection scheme of a biosensor offers the advantages of high binding affinity, specificity, and stability. 15,26 –28 Like a hairpin probe, aptamers remain closed in the absence of the target molecule and may contain both an element that anchors it to the sensor surface (i.e., biotin) and a reporting element (i.e., fluorescein). Thus, target binding to aptamers eliminates steric hindrance and allows the reporter (i.e., HRP) to bind and amplify the signal. Although the study described herein used aptamers as the capture probe, the results are applicable to other types of electrochemical sensors, such as those with antibodies or hairpin loops as the mechanism to capture the target.
Electrochemical Biosensors
A typical electrochemical biosensor is designed based on principles used in electrochemical cells. 29 In the research described herein, a three-electrode system, consisting of a WE, a CE, and a RE, is used in which the electrochemical reaction occurs on the WE. The electroreduction current produced by this electrochemical reaction is ultimately sensed at a third electrode, a CE, and is directly proportional to the number of hybridization events occurring on the WE. Moreover, the current measured directly depends on the transfer of reactants to the surface of the WE and the transfer of electrons for the oxidation and reduction of the electrically active species; the CE operates to maintain the WE at a constant potential relative to the RE by gathering any current produced by the electrochemical reaction. 29,30 Because the current is measured, this is an example of an amperometric biosensor.
In this parametric study of an electrochemical sensor, an electrochemical detection method is combined with enzymatic amplification and an aptamer that experiences target-induced conformational changes. When an electrochemical sensor is used, the aptamer must be immobilized only on the WE. 6,14 A conducting polymer, specifically poly-pyrrole (PPy), mixed with the aptamer is immobilized on the surface of the WE through electrical polymerization. A variety of biosensors have used electrically conducting polymers for the immobilization of biomolecules, including an electrochemical biosensor that detected salivary biomarkers. 31 –36 The specific electrochemical reaction of biosensor used in this study is catalyzed by HRP and electrons are transferred from the WE to the mediator tetramethylbenzidine (TMB), reducing it. Subsequently, HRP couples the oxidation of TMB to the reduction of hydrogen peroxide (H2O2) to water (H2O). In doing so, HRP is regenerated and is thus able to amplify the signal produced by the oxidation-reduction reaction. This oxidation-reduction reaction is illustrated in Figure 3B.
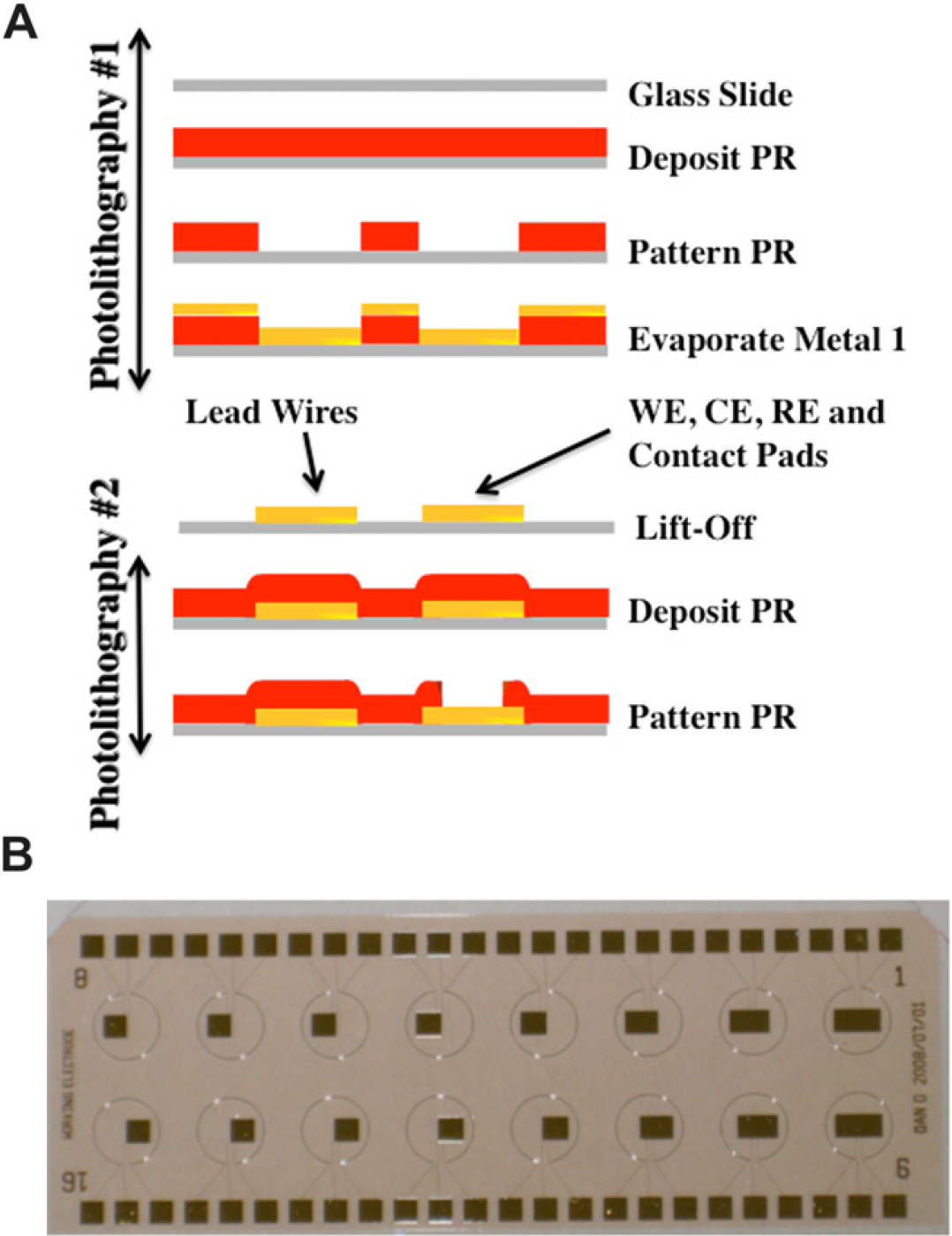
(A) Fabrication of microelectrode amperometric system. (B) 16-sensor array of microelectrode electrochemical sensors.
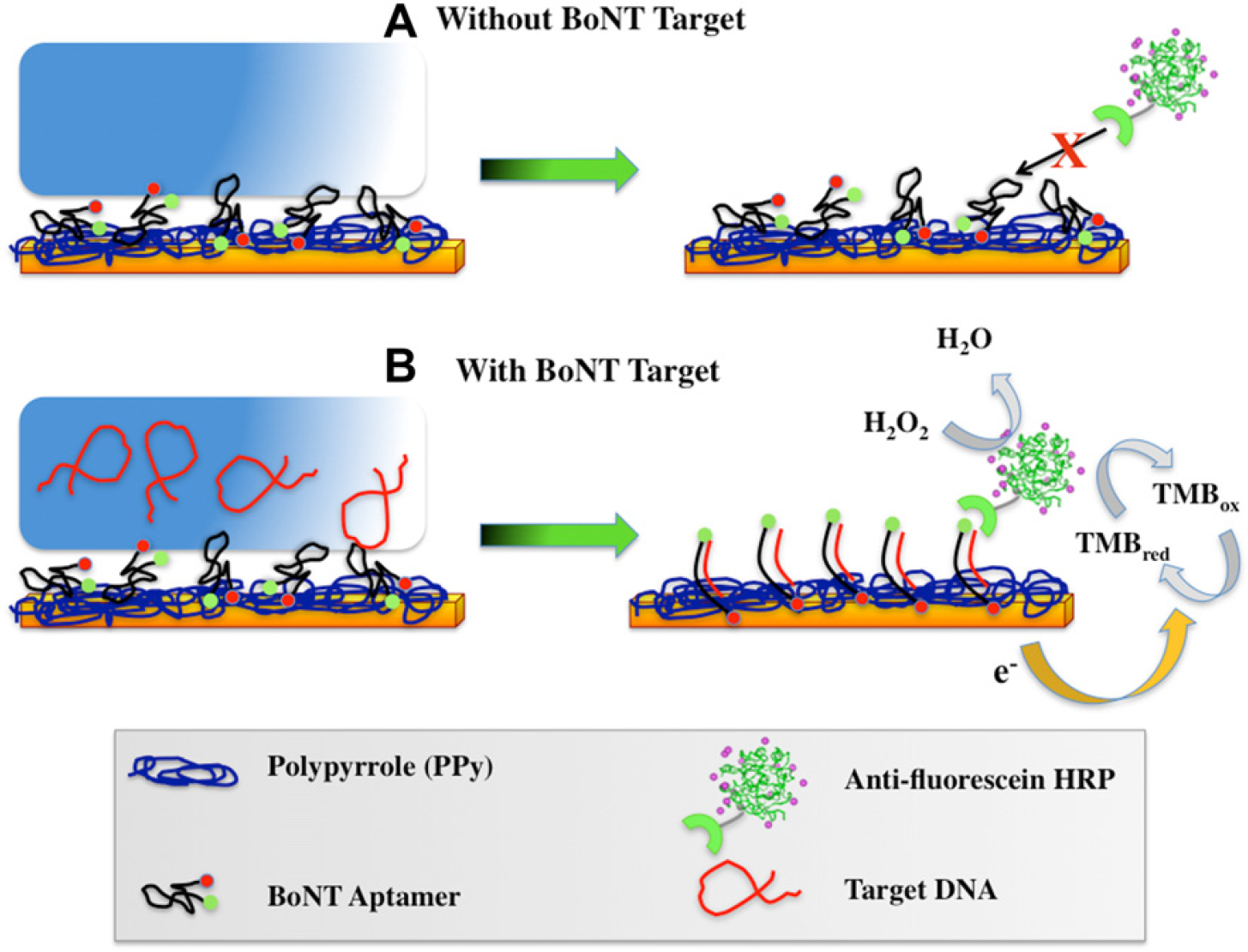
Aptamer-based electrochemical detection scheme. (A) Without Target DNA. (B) With Target DNA.
Materials and Methods
Aptamer and Target DNA Sequence
Custom-designed and HPLC-purified aptamer oligonucleotides were obtained (Operon Inc., Huntsville, AL). The aptamer oligonucleotides were labeled with biotin on the 5' end and with fluorescein on the 3' end. The 76-bp aptamer sequence was 5'-ATACCAGCTTATTCAATTGACATGA CTGGGATTTTTGGCGAAATCGAAGGA AGCGGAG AGATAGTAAGTGCAATCT-3'. 37 The target oligonucleotide used in the experiments was the complement of this DNA sequence.
Fabrication of Amperometric Microelectrode Biosensor
The fabrication of the amperometric microelectrode system, illustrated in Figure 2A consisted of two photolithographies. The first photolithography defined the microelectrodes using a positive polarity photoresist AZ5214 (AZ Electronic Materials USA Corp., Branchburg, NJ) in conjunction with lift-off after metal deposition via electron beam evaporation. The end result of the first photolithography was a 16-sensor array, an example of which is shown in Figure 2B. Lead wires function to connect the different electrodes to the contact pads, as shown in Figure 1B. To cover the lead wires, a second photolithography was performed in which the photoresist, AZ4620 (AZ Electronic Materials USA Corp., Branchburg, NJ), was patterned such that it covered the lead lines. As a consequence of the second photolithography, only the WE, CE, and RE were exposed during the electrochemical detection process. A detailed description of the fabrication process is described in the Supplementary Materials section I.
Electrochemical Detection
The electrochemical microelectrode biosensor system consisted of a 16-sensor array, as shown in Figure 2B. After the fabrication, a prefabricated plastic well manifold (GeneFluidics, Monterey Park, CA) was bonded to the sensor array. As shown in Figure 1B, each sensor unit consisted of three electrodes: the WE, CE, and RE. By having a 16-sensor array, the signal readout for all 16 electrodes was able to be determined simultaneously, providing information about both the signal and noise.
The aptamer-based electrochemical detection scheme used in this study is illustrated in Figure 3. To embed the fluorescein-labeled BoNT aptamer within the conductive polymer PPy on the surface of the WE via electropolymerization, 0.5μL of the BoNT aptamer (stock concentration = 100μM) was added to 890μL of 1 × phosphate-buffered saline (PBS) (Invitrogen, Carlsbad, CA), 100μL of potassium chloride (Mettler-Toledo, Columbus, OH), and 10μL of pyrrole (Sigma-Aldrich, St. Louis, MO). The mixture was mixed via vortexing and then 60μL was aliquoted onto each sensor immediately before electropolymerization. The electric waveform used for the electropolymerization step was pulse amper-ometry. 38 Each pulse consisted of a pulse for 9 s at a potential of +350 mV and 1 s at +950 mV; a total of 30 cycles were applied. After the electropolymerization step, the sensor array was rinsed with deionized H2O (18.3 Mω cm) and then dried with a purified stream of N2.
During the recognition phase, solutions with and without target were applied to the surface of the electrochemical sensor. In the case of the target solution, this step facilitated binding of the target BoNT DNA oligonucleotide with the BoNT aptamer that was deposited on the WE during the electropolymerization step. In this step, the target BoNT DNA is diluted 1:100 in the recognition buffer, which consisted of a solution of 100 mM MgCl2/1 × Tris-HCl; 60μL of the mixture is added to each of eight sensor wells. Likewise, a solution of 100 mM MgCl2/1 × Tris-HCl without the target BoNT DNA is added to the other eight sensor wells. To promote hybridization between the target sequence and the BoNT, pulse amperometry was used as the electric waveform. Each pulse consisted of a pulse for 9 s at −300 mV and 1s at 200 mV; a total of 30 cycles were applied. A 5-min passive hybridization step was done after the pulse amperometry. After the recognition step, the sensor array was rinsed with deionized H2O(18.3 Mω cm) and then dried with a purified stream of N2.
After the hybridization step, an oxidoreductase enzyme, HRP was coupled to the aptamer-target complex for signal amplification. To accomplish this, 2μL of 150 U/mL antifluorescein HRP Fab fragments (Roche Diagnostics, Indianaopolis, IN), diluted 1:500 in PBS with 0.5% casein blocking buffer (Blocker Casein in PBS; pH 7.4 Pierce, Rockford, IL). The mixture (60μL) was added to each sensor of the 16-sensor array. To promote binding of the antifluorescein HRP to the fluorescein-labeled BoNT aptamer, pulse amperometry was used as the electric waveform. Each pulse consisted of a pulse for 9 s at −300 mV and 1 s at 200 mV; a total of 30 cycles were applied. Subsequently, the sensor array was rinsed with deionized H2O (18.3 Mω cm) and then dried with a purified stream of N2.
Next, 80μL of 3,3',5, 5'-tetramethylbenzidine—H2O2 solution (K-Blue Low Activity TMB Substrate; Neogen, Lexington, KY) was added to each sensor and then electrochemical measurements were immediately and simultaneously taken for all 16 sensors. Amperometric detection was performed by applying a −200 mV potential (relative to the RE) to each sensor unit and the amperometric current versus time was measured via a multichannel potentiostat (GeneFluidics, Monterey Park, CA).
Results
To better understand the mechanisms involved in the operation of the electrochemical sensor and to optimize its performance, the design of the electrochemical sensor was varied to study several key parameters, as illustrated in Figure 1A. Specifically, four parameters were studied: the area of the WE, the area of the CE, the separation distance between the WE and CE, and the overlap length shared between the WE and CE. The same detection process was used in the study of all of the parameters.
Parametric Study: Area of the WE
Because the electroreduction current that produces the amperometric signal occurs at the WE, the WE area was the first parameter to be investigated during the parametric studies of the electrochemical sensor. During the polymerization process, pyrrole polymerizes on the WE and incorporates the BoNT aptamer within its polymeric structure. By altering the area of the WE, different amounts of pyrrole will polymerize on the surface of the WE and thus different amounts of BoNT aptamer will be integrated. By varying the incorporation of the BoNT aptamer at the WE, it was hypothesized that the signal could be varied because the electro-reduction current generated during the detection step is correlated to the number of hybridization events.
To accomplish this, the area of the CE was fixed by maintaining the dimensions at 2 × 2 mm. Furthermore, the separation distance between the WE and CE was also fixed at 20μm. However, the area of the WE was varied from 0.02 to 4 mm2. The detailed dimensions that were used in the design are described in Supplementary Information Section II; Figure 4B shows a few images of the devices used to test the effect of WE area on sensor performance.
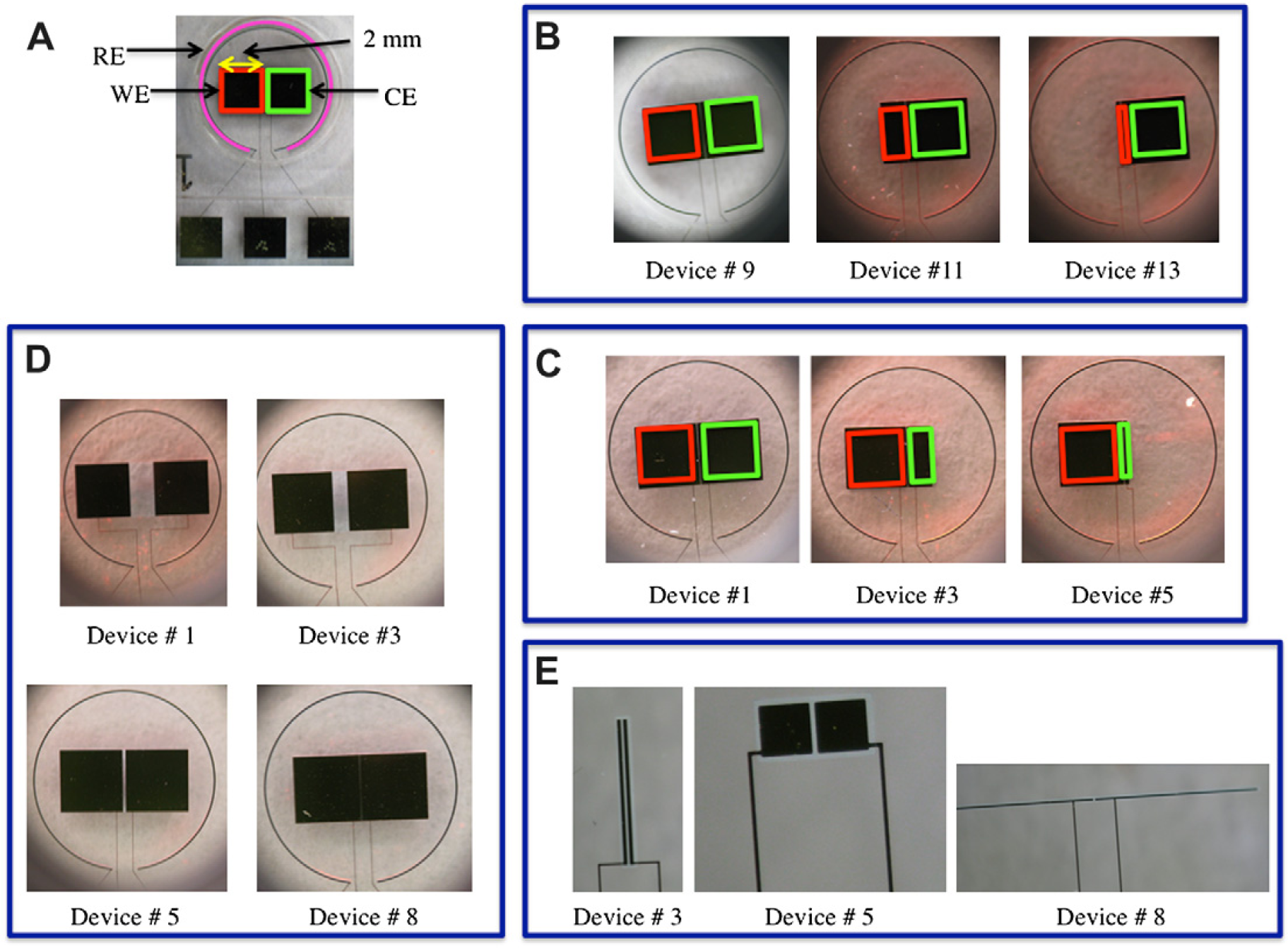

Images illustrating devices used to test the effect of variations in WE area. (A) The WE is highlighted with a red box, the CE is highlighted with a green box, and the RE is highlighted with a pink circle. (B) Devices used in parametric study of WE area. (C) Devices used in parametric study of CE area. (D) Devices used in parametric study of separation distance. (E) Devices used in parametric study of overlap length.
A log-log graph illustrating the results of the experiment examining the relationship between WE area and sensor performance is shown in Figure 5A. As indicated by the graph, both the signal and noise were directly proportional to the area of the WE; there is a linear relationship between the WE area and the signal and noise. Furthermore, it was shown that the lines representing the signal and noise had a nearly equal slope, where the slope of signal curve was (0.60 ± 0.063) nA/μm2, whereas that of the noise curve was (0.64 ± 0.040) nA/μm2. A consequence of this, the signal-to-noise ratio was nearly independent of the area of the WE, which is show in Figure 5B. Thus, although a larger WE area produced a larger signal, it did not offer an improvement in overall performance of the sensor.
When considering the dynamics of the detection method used for this electrochemical sensor in light of the data shown in Figure 5A, one may conclude that the primary agent responsible for affecting the signal and noise was the number of BoNT aptamer molecules. In other words, an increased number of BoNT aptamer molecules on the WE surface produced an increased signal because more target could bind and thus more current was generated by the electrochemical reaction. The “opening” of the BoNT aptamer and interaction with the antifluorescein conjugated HRP produced noise when it occurred in the absence of target. As the area of the WE was increased, such events occurred more frequently. However, because the BoNT aptamer was at the foundation of the signal or noise and because the area of the WE affected its number, there was a linear relationship between the WE area and both the signal and the noise.
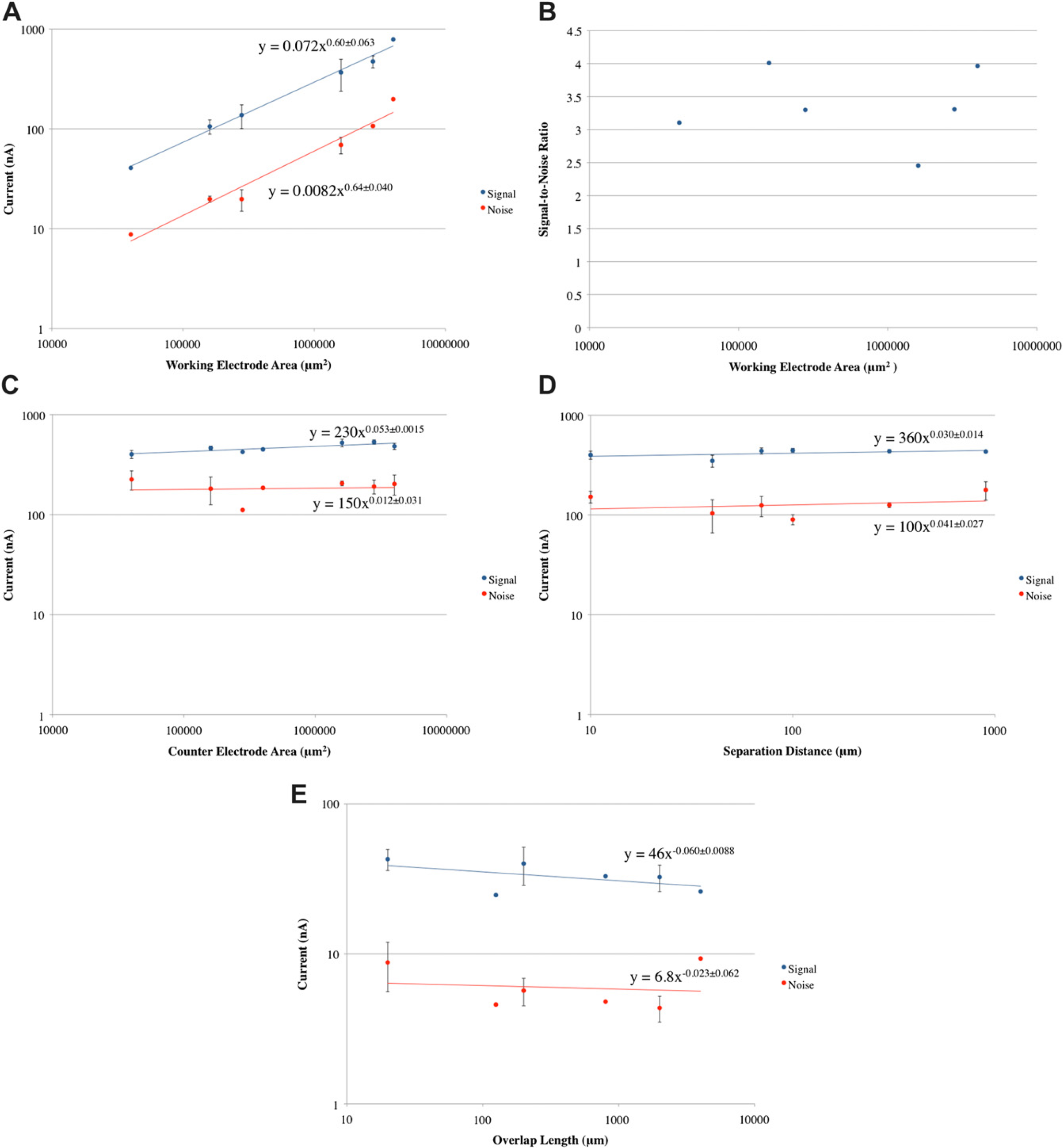
(A) Log—log graph of results of parametric study of WE area. (B) Graph illustrating relationship between WE area and signal-to-noise ratio. (C) Log-log graph of results of parametric study of CE area. (D) Log-log graph of results of parametric study of separation distance. (E) Log—log graph of results of parametric study of overlap length. (Error bars are indicated when device and electropolymerization yield enabled collection of triplicate data.)
Parametric Study: Area of the CE
In the three-electrode system, the RE is used to maintain the WE at a constant potential and, thus, no current is allowed to pass through the RE during the reaction. To facilitate this, the CE completes the circuit with the electroreduction current produced at the WE and gathers any current that is generated, thereby maintaining a constant voltage at the WE relative to the RE in the presence of the electrochemical reaction. Because the current generated results from the flux (moles/area x time) of the TMB molecules that travel to the CE, the hypothesis was made that the current could be adjusted because total moles/time = molar flux × (area of CE). To study this, the area of the CE was varied from 0.02 to 4 mm2 while everything else was maintained at a constant, as illustrated in Figure 4C. Detailed dimensions of the devices used in this parametric study are shown in Supplementary Information Section III.
As shown in Figure 5C, both the signal and noise were independent of the CE area. Specifically, the slope of the signal curve was found to be (0.053 ± 0.0015) nA/μm2,whereas that of the noise curve was calculated at (0.012 ± 0.031) nA/μm2. Because the amperometric signal measured results from the oxidation and reduction of the mediator TMB, the results would suggest that in the range studied, the CE was of a size sufficient to complete the circuit by facilitating the oxidation of TMB. This result is supported by the size of the TMB molecule (diameter ∼ 1 nm) compared with the dimensions of the CEs used in the study (height = 2000μm, width = 10–2000 mm). In other words, the signal and noise currents remained constant as the CE area was varied because all CE areas tested were significantly larger than the TMB molecule and thus did not affect the current transfer efficiency. The significance of this finding is that future sensor designs can minimize the area of the CE without affecting the performance of the sensor.
Parametric Study: Separation Distance between WE and CE
The strength of an electric field is proportional to the separation distance between two charged plates, so the separation distance between the WE and CE was varied to determine if this was an important parameter for the performance of the sensor. Specifically, it was hypothesized that by varying the strength of electric field, the TMB molecules could be accelerated toward the CE at different rates and thus affect the current that was generated.
To study the parameter of separation distance, the separation distance was varied from 10 to 900μm, whereas the other parameters were kept constant. This is illustrated with a few examples in Figure 4D a table showing the detailed dimensions is in Supplementary Information Section IV. The results of the experiment are shown in Figure 5D. As shown by the graph, the signal and noise was shown to be independent of the separation distance, where the slope of signal curve equaled (0.030 ± 0.014) nA/μm and that of the noise curve equaled (0.041 ± 0.027) nA/μm. These results suggest that electrophoresis and electric field are not dominant force in transferring species from the WE to the CE in the system studied.
Parametric Study: Overlap Length
The last parameter that was investigated was the parameter of overlap length, which is illustrated in Figure 1. The investigation of this parameter was based on the relationship between the electron transfer efficiency and the interface between the WE and CE. This is reflected by the following equation for the steady-state current between a circular electrode and a conductive substrate in the presence of an oxidation-reduction reaction: 39
where n = number of electrons transferred per molecule in the redox reaction, F = Faraday's constant, D = diffusion coefficient, C b = is the bulk concentration of the electrically active species (e.g., TMB), and a = radius of a circular electrode. The concept of electron transfer efficiency between a circular electrode and a substrate proves applicable to an electrochemical sensor and thus, in this study, the length of the interface or “overlap length,” was varied to determine if any observable change in the performance of the electrochemical sensor was observed. To accomplish this, only the overlap length was varied while the other parameters were kept constant. The dimensions are shown in Supplementary Information Section V, and some examples of the devices are shown in Figure 4E.
The results of the experiments are shown in Figure 5E. As shown by the graph, both the signal and noise were relatively independent of the overlap length, where the slope of the signal curve equaled (-0.060 ± 0.0088) nA/μm and that of the noise curve equaled (-0.023 ± 0.062) nA/μm. These results suggest that the flux of the TMB from the WE to the CE occurs all across both the area of the WE and CE and not just along the interface between the two electrodes.
Parametric Study: Effect of Aptamer Concentration on Sensor Performance
Following the results of the parametric study of the relationship between WE area and sensor performance, a study was performed to examine the relationship between the current generated and aptamer concentration. The same 16-sensor chip used in the parametric study of WE was used in these studies so to examine the effect of varying the WE area at different aptamer concentrations. Figure 6A shows a log-log graph of the results of an experiment using an aptamer that was diluted 1:1000 from the stock concentration of 100μM, whereas Figure 6B shows the results of an experiment in which the aptamer was diluted 1:500. As can be seen by the graphs, the signal and noise curves are parallel as they were in the previous experiments in which the aptamer was diluted 1:2000. Because the lines remain parallel, it is suggested that the signal-generation scheme and the noise-generation scheme were affected equally by the alteration in aptamer concentration. As a consequence of this, it may be concluded that one of the mechanisms for noise generation is the opening of the aptamer in the absence of target hybridization in addition to the nonspecific binding of the antifluorescein HRP to components on the surface of the WE. This conclusion is supported by an experiment previously done by Wei and Ho15 in which the specificity of the aptamer binding was investigated by comparing the signal obtained with BoNT/A toxoid, the target; four nontarget proteins (i.e., bovine serum albumin, casein, IL-8, and IL-1b); and PBS, a blank. The implication of this is that to reduce the noise, a protocol should be developed to enhance the likelihood that the aptamer only opens when in the presence of the target molecule and to reduce the nonspecific binding of antifluorescein HRP. Furthermore, the data also show that the signal-to-noise ratio remains constant for different aptamer concentrations in addition to the tested range of different WE areas.
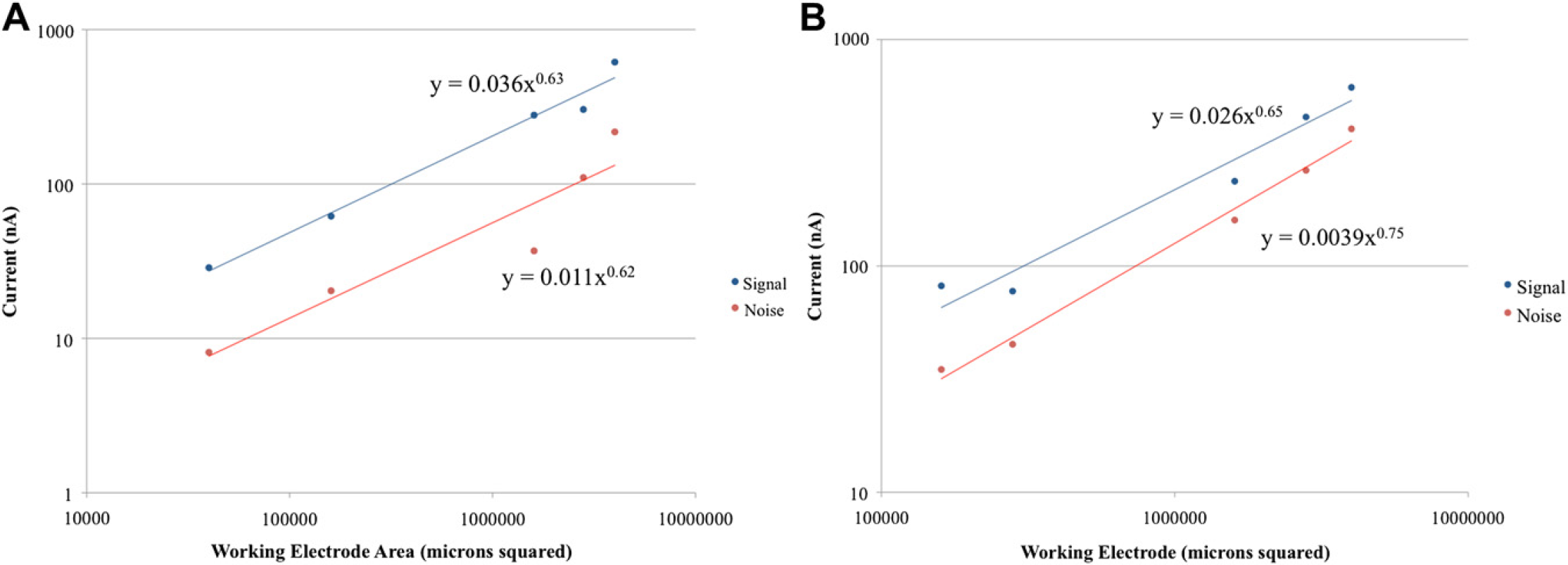
(A) Log—log graph of results of experiment in which aptamer was diluted 1:1000. (B) Log—log graph of results of experiment in which aptamer was diluted 1:500.
Conclusion
To better understand the operation and potential for optimization of electrochemical biosensors, an amperometric microelectrode electrochemical sensor was fabricated according to a variety of designs. In particular, the designs were created to examine the effect of five parameters on sensor performance: the area of the WE, the area of the CE, the separation distance between the WE and CE, the overlap boundary shared between the two electrodes, and the aptamer concentration. In addition, during the examination of these parameters, a protocol was developed to use an aptamer and the conductive polymer PPy in the electrochemical sensor detection scheme. As a result of this process, a few critical findings were presented.
First, a systematic characterization of the parameters for designing an electrochemical sensor was presented, which to the best knowledge of the authors, has yet to have been done. Second, it was determined that while the signal-to-noise ratio was independent of the WE area, the absolute value of signal and noise did vary linearly with the area of the WE and thus the signal and noise did exhibit dependence. This indicates that the area of the WE may be decreased without affecting the signal-to-noise ratio, but that the absolute values of the signal and noise will decrease. This is in contrast with the studies of the other parameters, in which the signal and noise were constant. All three values (i.e., signal, noise, and signal-to-noise ratio) are important in evaluating an electrochemical sensor. In addition, the absolute value of the signal and noise is important in determining the sensitivity of a sensor. For example, with the same signal/noise ratio, a higher signal will lead to a higher sensitivity. These findings will enable the design of high-density sensor arrays by fabricating electrochemical sensors in which the WE area is maximized when space permits, without considering the other parameters, such as areas of other electrodes and their shapes, thus greatly simplifying the design principle in the future.
In addition to the demonstrated significance of the area of the WE, other parameters will need to be investigated in the future for their contribution to the performance of the electrochemical sensor. For instance, the BoNT/aptamer hybridization dynamics may also prove important. Typically, the electrochemical reaction is the rate-limiting step among the ensemble of reactions involved in an electrochemical sensor that also includes the ion/charge transfer and mass transport. However, if the BoNT hybridization is very fast, and thus not the rate-limiting step, the final signal generated may only depend on the area of the WE area. Such a conclusion can only be ascertained through further experimentation.
Acknowledgments
This work was supported by the NSF funded SINAM (DMI-0327077) center grant.
Competing Interests Statement: The authors certify that they have no relevant financial interests in this manuscript
Appendix
Supplementary Information
Supplementary data associated with this article can be found, in the online version, at 10.1016/j.jala.2010.01.007