Abstract
The in vitro selection of aptamers that bind to low molecular weight targets is commonly a tedious, time-consuming project. We have expanded current automated selection protocols to include aptamer selections against small molecules including the aminoglycosides neomycin, kanamycin, and tobramycin. This modified procedure decreases both the frequency of manual handling of the selection reagents and the time required to perform the experiment, generating aptamers against the chosen target at a much greater rate. Using this process, we have selected aptamers of good affinity against all three aminoglycosides chosen. The method is suitable for integration with high-throughput technologies, greatly expanding the possibility of discovering useful aptamers against other low weight targets. (JALA 2004;9:150-4)
Introduction
Aptamers are short (∼80 nt) segments of nucleic acid that have been shown to mimic many properties of antibodies and which bind with high specificity and affinity to molecular targets. The process of selecting aptamers against chosen targets (in vitro selection or SELEX) includes several rounds of combining the target with a randomized pool of nucleic acid sequences, washing away the non-binding species, and amplifying the bound members. Subsequent iterations of this process narrow the nucleic acid pool from approximately −1015 unique members down to the strongest binding species.
The experiments outlined here suggest a method for the automated selection of aptamers against low molecular weight targets. The robotic protocols are an expansion of those outlined previously. 1 2 3 4 Modifications made include a longer RNA-target binding time and a less stringent wash cycle.
Aminoglycosides are a class of low molecular weight compounds capable of inhibiting microbial growth by binding to the 30S subunit of ribosomes. They are good candidates for selection protocols as each molecule contains several primary amines creating a substantial positive charge on each molecule. The negatively charged pool can therefore bind more easily. The amines also provide a site for biotin attachment to the antibiotic.
Materials and methods
Aminoglycosides (Figure 1) were biotinylated via NHS ester linkage as per Pierce (Rockford, IL) protocols. The resulting product was added to streptavidin coated ferric beads. A 10x molar excess of biotinylated target to streptavidin was used and it was assumed that all binding sites on the streptavidin were filled by target.
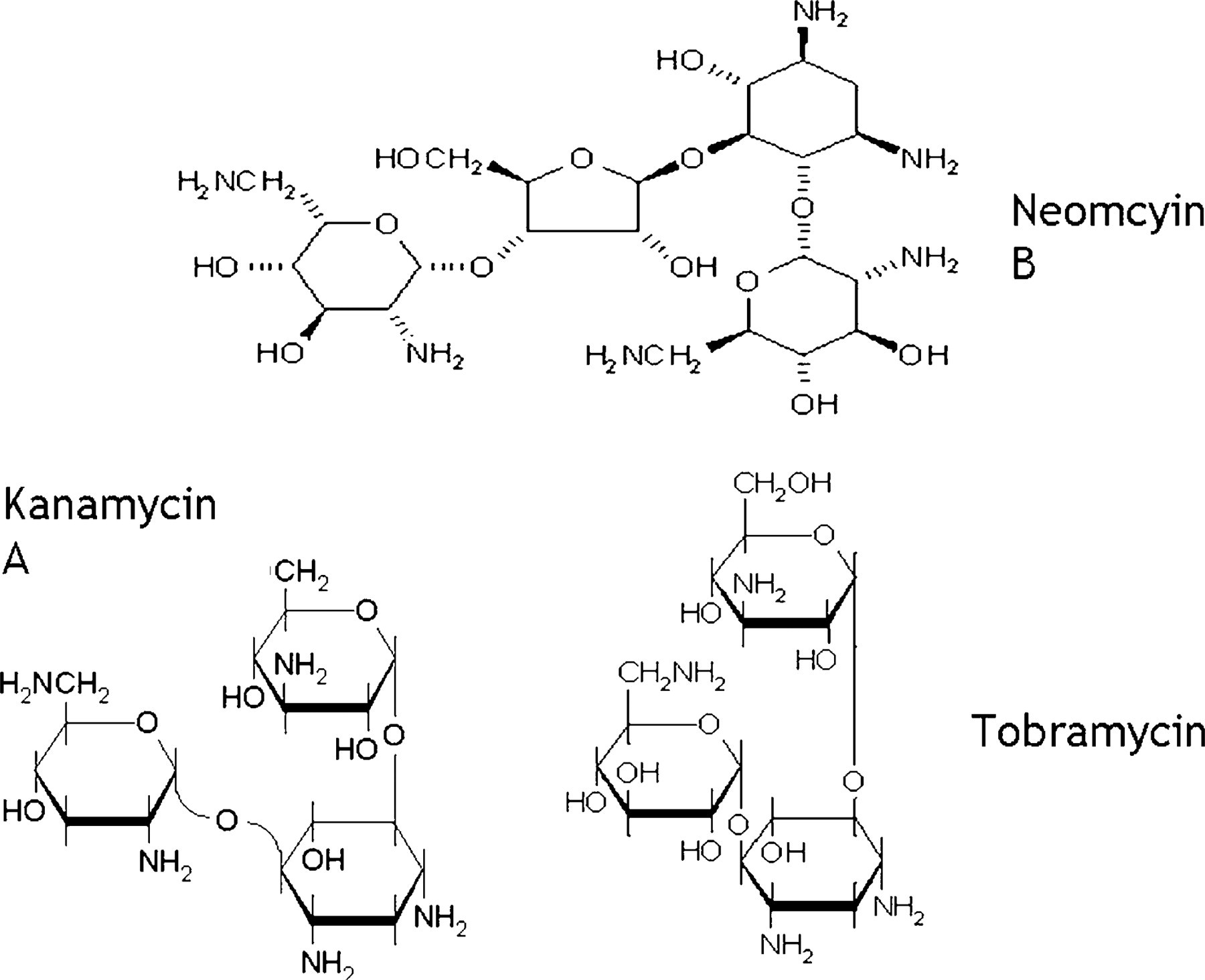
Molecular structures of three aminoglycosides.
All steps in the selection process (Figure 2) were performed by a modified Biomek 2000 workstation, shown in Figure 3 (Beckman Coulter, Fullerton, CA). The target was allowed to bind with the randomized RNA pool for ten minutes. The binding reaction was then transferred to a filter for a wash to remove non-specific binding RNA molecules. The binding species underwent RT-PCR to amplify the sequences. Transcription followed and the process repeated from the beginning. This continued for eleven more cycles. Samples of DNA and RNA were taken every cycle to ensure that reactions were proceeding as expected.
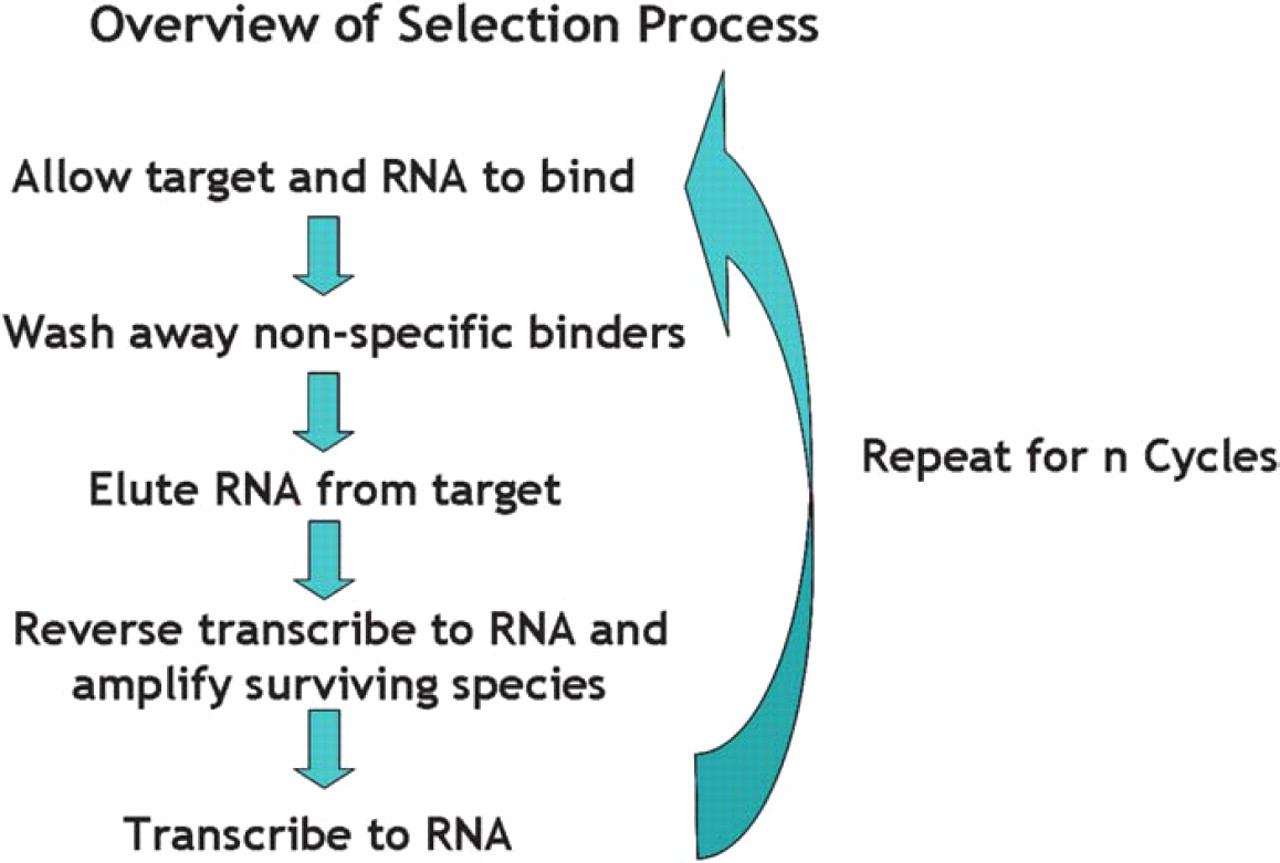
Schematic outline of selection process.

Robotic workstation for automated selection. Binding reactions took place in the target plate. Buffer plate held selection buffer and water for washing on the filter plate. RT-PCR and transcription reactions ran in the thermocycler with reagents and enzymes pipetted from their respective plates. Enzymes cooler maintained a temperature of −20 °C throughout the run.
Selection Buffer 5 : Neomycin—50 mM Hepes pH 7.6, 5 mM MgCl2, 200 mM NaCl and Tobramycin/Kanamycin—50 mM Hepes pH 7.6, 5 mM MgCl2, 100 mM NaCl.
At the end of the selection process, the round twelve DNA was PCR amplified and cloned for analysis. Sequencing reactions were performed with DCTS Quick Start kits (Beckman Coulter) and capillary electrophoresis sequenced with the CEQ 2000XL (Beckman Coulter).
Binding reactions for gel shift assays 6 were performed in 1x selection buffer, 10% glycerol, and a variable concentration of target connected through a biotin link to free streptavidin. The radiolabled RNA concentration was held at a constant value under 1 nM. The RNA was allowed to bind to the target for 30 minutes and then was run on a 10% 100:1 acrylimide:bisacrylamide gel at 200 V for 1 hour. The gel was then developed on a Molecular Devices (Sunnyvale, CA) Phosphourimager.
Sequence Analysis
The sequences of aptamers obtained from the selections show a high degree of conservation (Figure 4). This is most striking in the case of the anti-neomycin aptamer. All sequences viewed contained the same thirty base pair random region. In the case of the other two targets, much of the intra-family sequence was identical. It is interesting to note that the selected sequences display no significant similarity to motifs seen in previously performed selections by other researchers. 5,7,8

Aptamer sequences. Static regions appear in gray. Random regions are black. Conserved random regions are shown in similar colors.
Possible secondary structures of the sequences were determined using the Zucker Mfold web server. 9 Three representative structures can be seen in Figure 5.
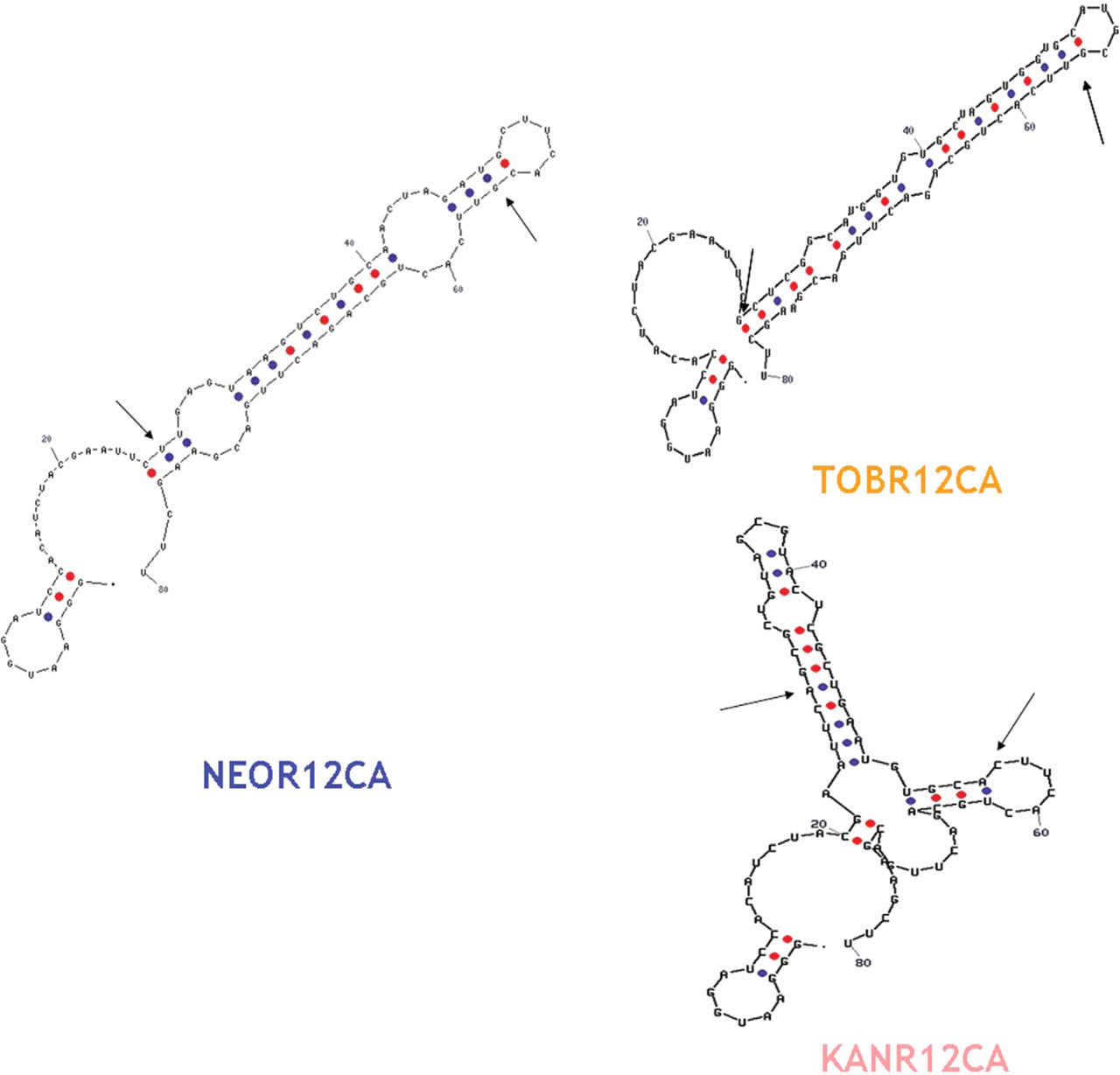
Representative RNA aptamer secondary structures motifs from Mfold server. Blue dots are U-A bonds and U-G wobbles. Red dots are G-C bonds. Arrows denote the start and end of the random region.
Binding Strength
Preliminary results from gel shift assays suggest that the Kd values for the selected aptamers lie under 1 μM (Figure 6). The aptamer against neomycin appears to have a Kd value in the lower range of the nanomolar scale. The KANR12CA clone has a dissociation constant in the range of 10–30 nM. Three species of aptamer against tobramycin have been tested for binding affinity. TOBR12CA has an apparent Kd of 30–100 nM. TOBR12CB and TOBR12CC seem to possess a Kd value of around 100 nM. Further planned tests will more precisely locate the dissociation constants of these as well as the as yet untested aptamers. We also intend to examine the cross reactivity of the aptamers when introduced to similar targets.
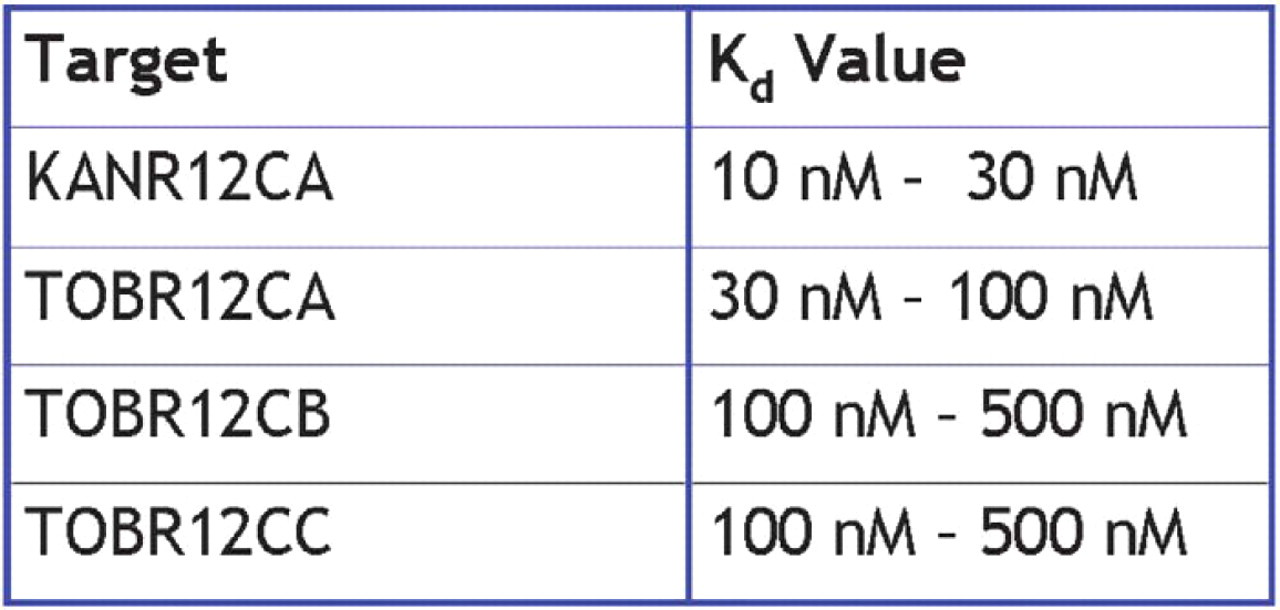
Selected aptamer dissociation constants.
CONCLUSIONS
We have developed an in vitro selection method for discovering aptamers against low molecular weight targets that in some cases selects stronger binders than published selections against identical targets.
We are continuing to examine the binding character of the selected aptamers. We plan next to select aptamers against molecules of higher interest, including diagnostically important molecules such as neurotransmitters and narcotic drugs.
Acknowledgments
This work was supported by National Science Foundation EIA-0218447.
