Abstract
Optimizing the buffer conditions of the selection of nucleic acid binding species (aptamers), increases the likelihood of producing a target aptamer. Aptamers, with high target affinity and specificity, are often compared to antibodies, as aptamers emerge in the industry as diagnostic and therapeutic tools. The increased demand for aptamers encourages high-throughput aptamer generation. The selection buffer conditions may vary as widely as the selection targets, and therefore buffer optimization is helpful if not required for effective aptamer selections. Such optimization work is time consuming and repetitious, which bodes well for high-throughput applications. To accommodate this, an automated buffer testing protocol has been developed to test target-to-unselected RNA pool binding in the presence of 96 different buffer conditions. The dynamic program may vary the monovalent salt(s) identity, monovalent salt(s) concentration, divalent salt(s) identity, divalent salt concentration, buffer identity, buffer concentration, and pH. The optimized buffer conditions likely increase the probability of a successful selection and therefore promote higher ratios of successful aptamer selections against a variety of targets.
Preliminary results show trends with the buffer matrix solutions and lysozyme:unselected pool binding. In general, an inverse relationship between lysozyme binding and monovalent salt concentration is observed. (JALA 2004;9:117-22)
Keywords
Introduction
Aptamers, with high target affinity and specificity, are often compared to antibodies, as aptamers emerge in the industry as diagnostic and therapeutic tools. 1,2 The increased demand for aptamers encourages high-throughput aptamer generation. This has encouraged the generation of aptamers against a wide variety of targets, including targets that do not necessarily bode well to aptamer selection. These difficult selection targets include, for instance, targets that are negatively charged, which are, therefore, repelled by negatively charged RNA pools. Modifying and optimizing selection buffer conditions may counter-act or otherwise stabilize the interactions between the target and the unselected pool. For instance, as with the example of the negatively charged target, decreasing the pH below a target's isoelectric point (pI) will neutralize the negative charge and encourage the binding of the negatively charged pool. Hence, optimizing the buffer conditions is helpful if not required for effective aptamer selections.
Such optimization work is time consuming and repetitious, which bodes well for high-throughput applications. To accommodate this, an automated buffer testing protocol has been developed to test target-to-unselected RNA pool binding in the presence of 96 different buffer conditions. The dynamic program may vary the monovalent salt(s) identity, monovalent salt(s) concentration, divalent salt(s) identity, divalent salt concentration, buffer identity, buffer concentration, and pH. The optimized buffer conditions likely increase the probability of a successful selection and therefore promote higher ratios of successful aptamer selections against a variety of targets.
Purpose: automated buffer matrix test
Essentially, using 13 different stock solutions, the Automated Buffer Matrix Test prepares 96 different solutions and uses these solutions in 96 different target:pool (32P-labeled unselected pool) binding reactions. The assay utilizes the characteristic of nitrocellulose membranes to retain protein, and therefore any molecules bound to the protein target. 3 The percentage of bound pool contained on the nitrocellulose membrane, quantitated using an Amersham (Piscataway, NJ) phosphorimager, is compared to buffer solution conditions. The conditions providing some of the greatest percentage of pool:target binding are chosen for future automated aptamer selections.
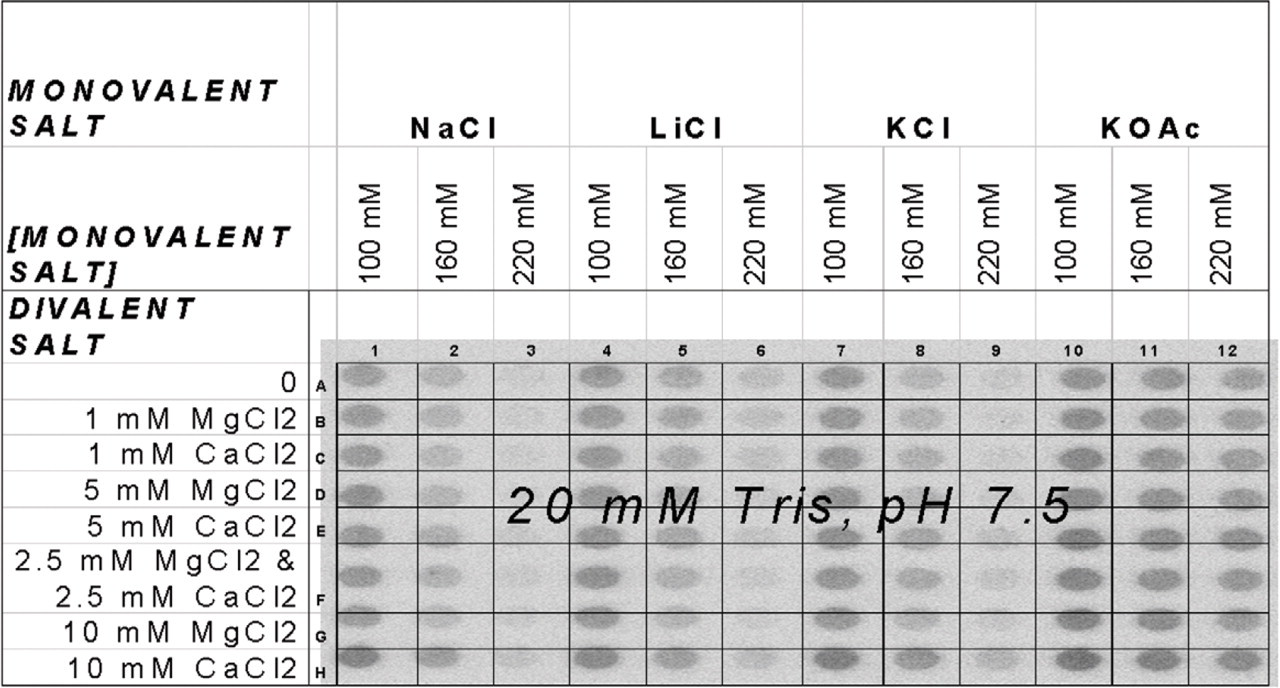
The Automated Buffer Matrix Test was performed against the target hen egg white lysozyme, obtained from Sigma (St. Louis, MO). Lysoyzme was originally chosen as an Automated Selection Buffer Matrix Test target because it has been successfully selected against in automated aptamer selections (scheme illustrated in Figure 1 4 ); it is positively charged, and it is relatively inexpensive. The automated program tested the lysozyme: unselected RNA pool binding in the presence of 96 different buffer solutions (see Figure 2), including the automated aptamer selection buffer (20 mM Tris, pH 7.5, 100 mM sodium chloride, and 5 mM magnesium chloride) 4 . The 96 different buffers were prepared on a Beckman (Fullerton, CA) Biomek 2000 Automated Liquid Handling Workstation using a stock buffer solution of Tris, pH 7.5, although other buffers may be used. Stock solutions of the monovalent salts, sodium chloride, lithium chloride, potassium chloride, and potassium acetate were used to prepare the matrix solutions at three different concentrations, 100, 160, and 220 mM, although other monovalent salts may be used. Additionally, stock solutions of the divalent salts magnesium chloride and calcium chloride (again, other solutions may be used) were available in an eight-row (horizontal) reservoir plate for the preparation of the buffer solutions (see Figure 3).
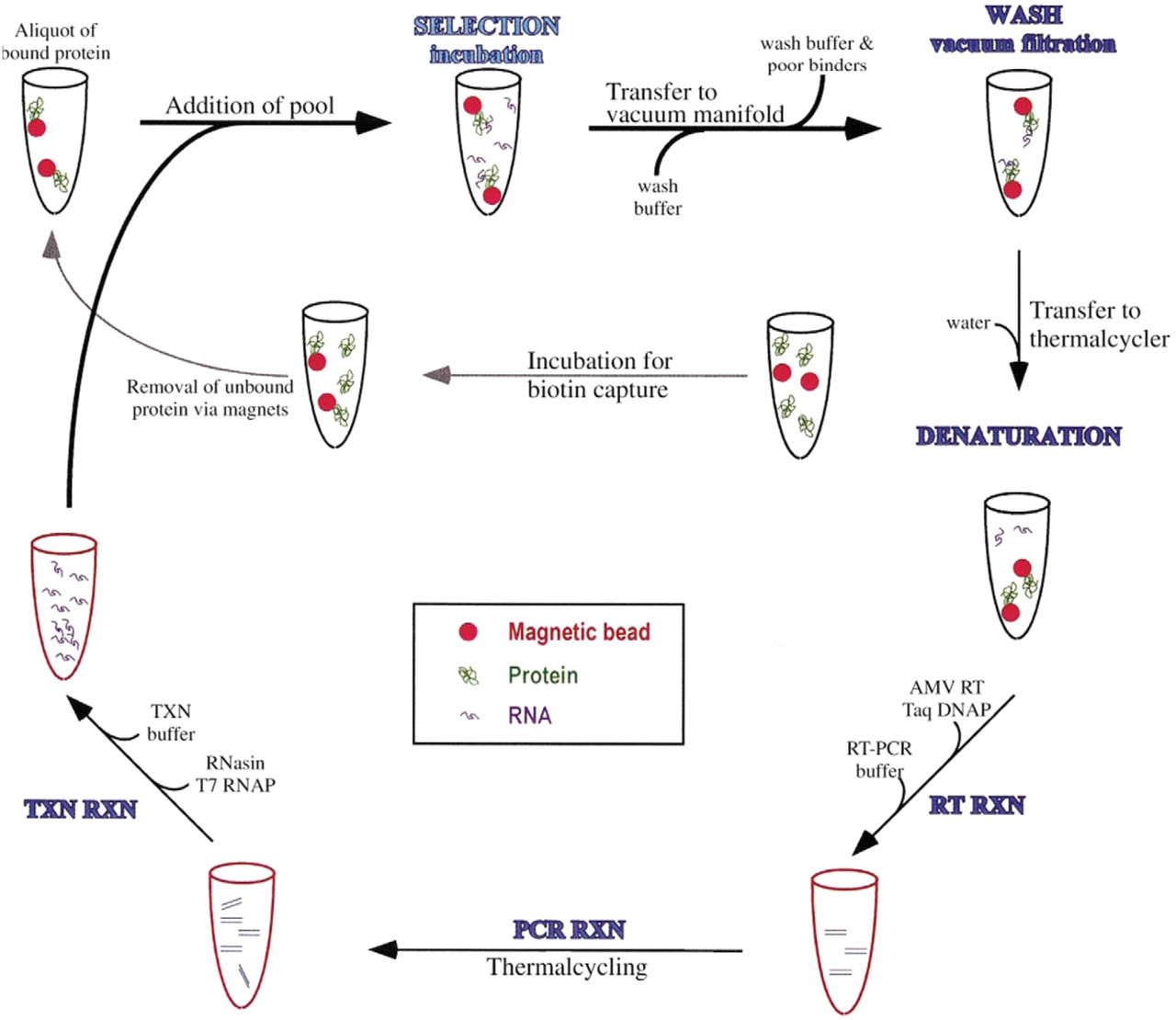
Automated in vitro aptamer selection scheme. 4
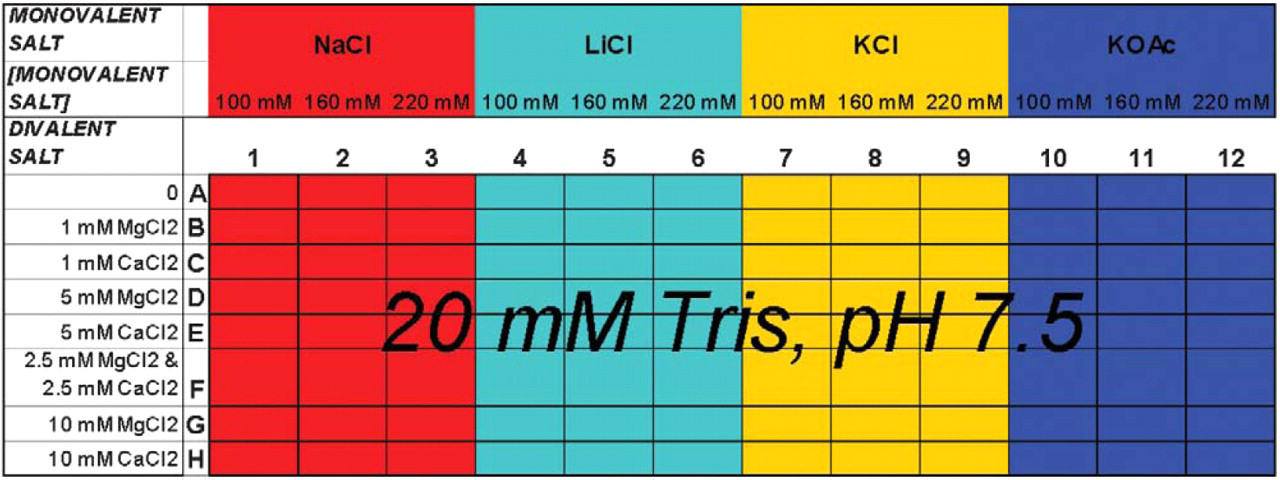
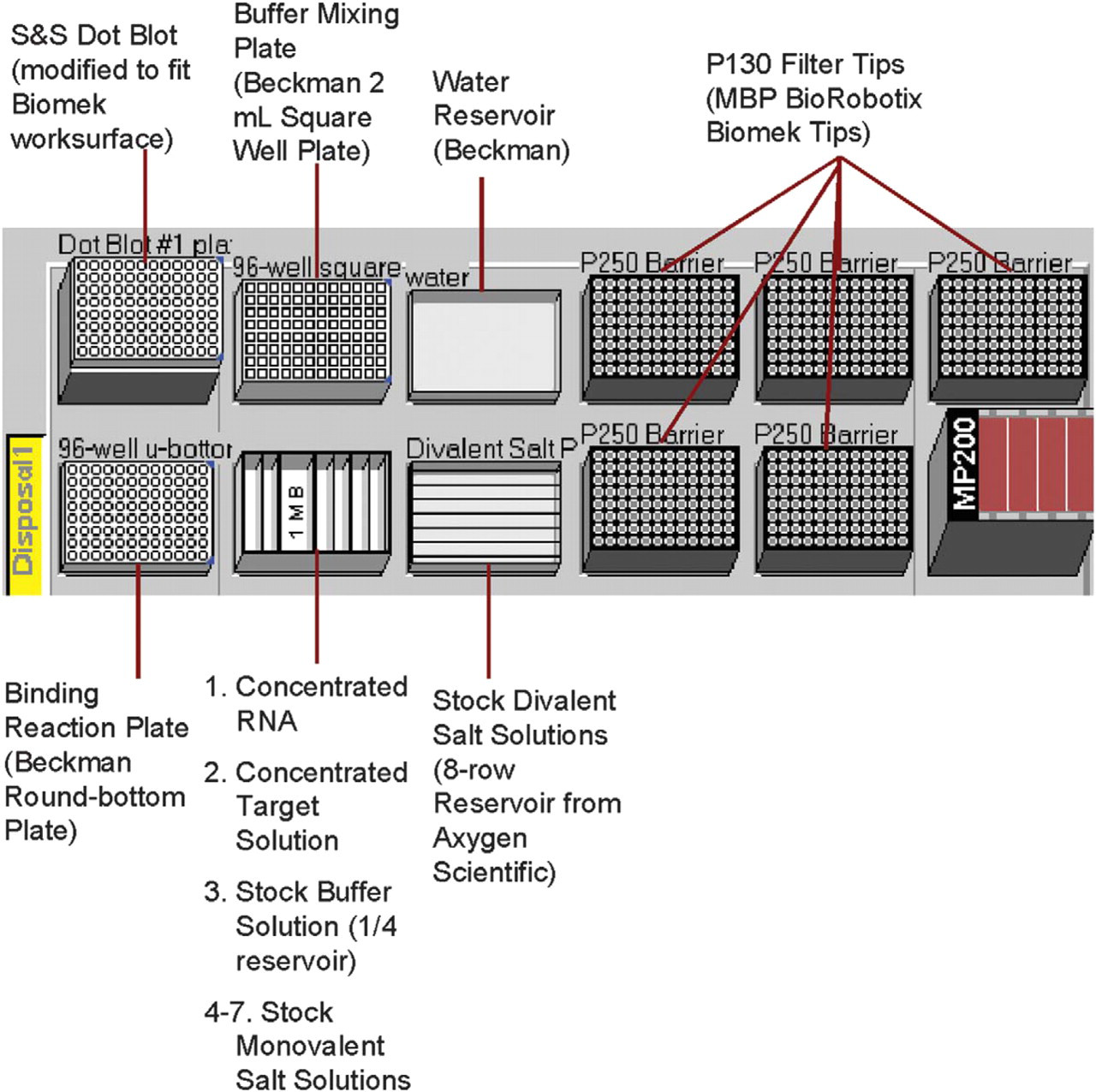
Automated buffer matrix test worksurface on Biomek 2000.
Method: automated buffer matrix test
Materials
Beckman (Fullerton, CA) Biomek 2000 Liquid Handling Workstation; S&S (Keene, NH) Dot Blot, modified for Biomek 2000; S&S (Keene, NH) Nitrocellulose Paper (0.45 μm pore size); Amersham (Piscataway, NJ) Hybond-N+ Nylon Membranes; Beckman (Fullerton, CA) Reservoirs and Plates; 8-row reservoir plates—Axygen Scientific (Union City, CA) plates, purchased from VWR (West Chester, PA); Molecular BioProducts (San Diego, CA) Biomek Filter Pipette Tips; and Molecular Biology Grade Reagents.
Procedure
96 different solutions are prepared on the Biomek 2000 work surface using aliquoted stock solutions of monovalent salt(s), divalent salt(s), buffer(s), and any additional solutions (e.g. Tween, BSA, etc.). 2x concentrated buffer solutions are prepared by the Biomek 2000. 2x concentrated buffer solutions, 4x concentrated RNA, 4x concentrated target solutions, and water are added to the binding reaction plate. The RNA: target binding reactions incubate at room temperature for ∼30 minutes. The remaining 2x concentrated selection buffer solutions are diluted to 1 × solutions. The 96 buffer solutions pre-wet the nitrocellulose and nylon membranes sandwiched in an S&S dot blot on the Biomek 2000 worksurface. The RNA: target binding reactions are filtered through the dot blot and washed with 1 × selection buffer solutions. The target and any binding RNA molecules remain on the nitrocellulose membrane, whereas the unbound RNA molecules flow through the nitrocellulose membrane and become trapped on the nylon membrane underneath. The filtered RNA: target binding reactions are washed with 1 × buffer solutions. The radiation counts are measured. The percentage of bound unselected pool is calculated and compared to the buffer solution.
Conclusions: Automated Buffer Matrix Test
Preliminary results show lysozyme:unselected pool binding trends (Figure 4). An inverse relationship between lysozyme binding and monovalent salt concentration is observed (Figure 5). The binding trends are generally reproducible, but individual binding values may vary greatly. Preliminary results from automated buffer matrix tests against additional targets suggest the inverse monovalent salt concentration and target-binding trend does not necessarily occur with all targets.
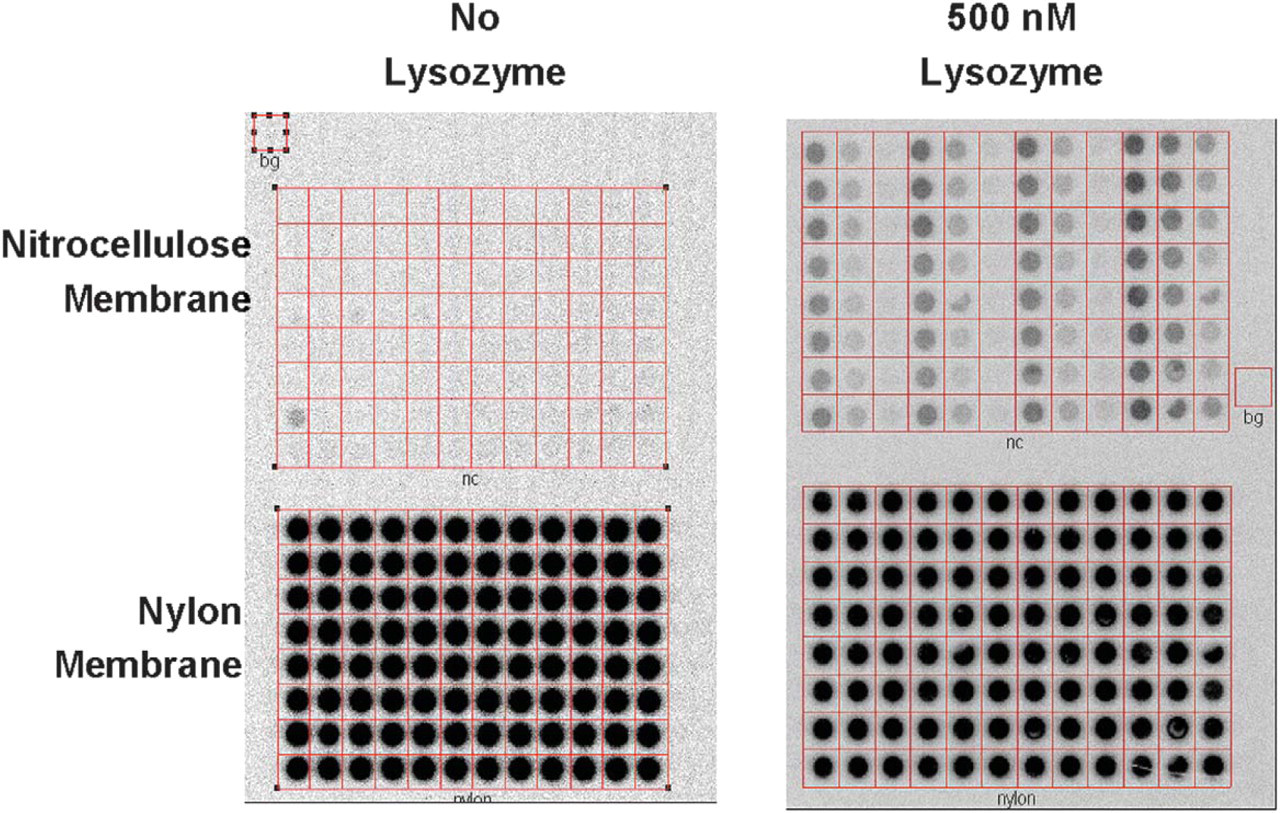
Results from Automated Buffer Matrix Test against hen egg white lysozyme protein.

Results of Automated Buffer Matrix Test against lysozyme (500 nM). Note: preliminary results show the inverse relationship of lysozyme binding:monovalent salt concentration does not necessarily occur with all targets.
Based on the results above (see Figure 6) buffer solutions #25–27, 49–56, or 73–80 would be buffer candidates for an automated lysozyme aptamer selection because of the great percentage of lysozyme binding. However, one may elect to use buffer solution #56 (20 mM Tris, pH 7.5, 100 mM KCl, and 10 mM CaCl2) in an automated lysozyme aptamer selection because of the reproducibility of the automated buffer matrix results (see Figure 7).
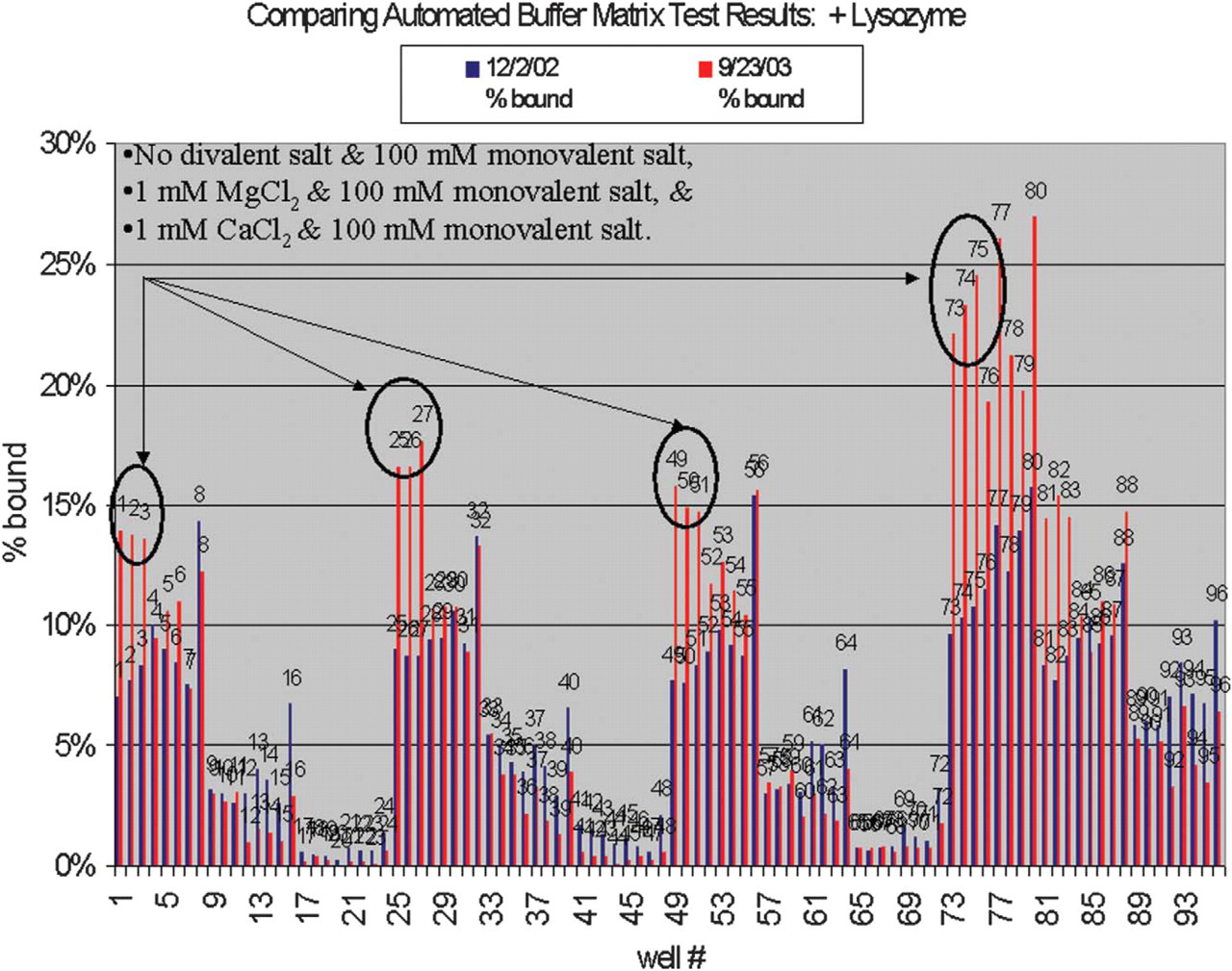
Reproducibility test results for Automated Buffer Matrix Tests against lysozyme (500 nM).
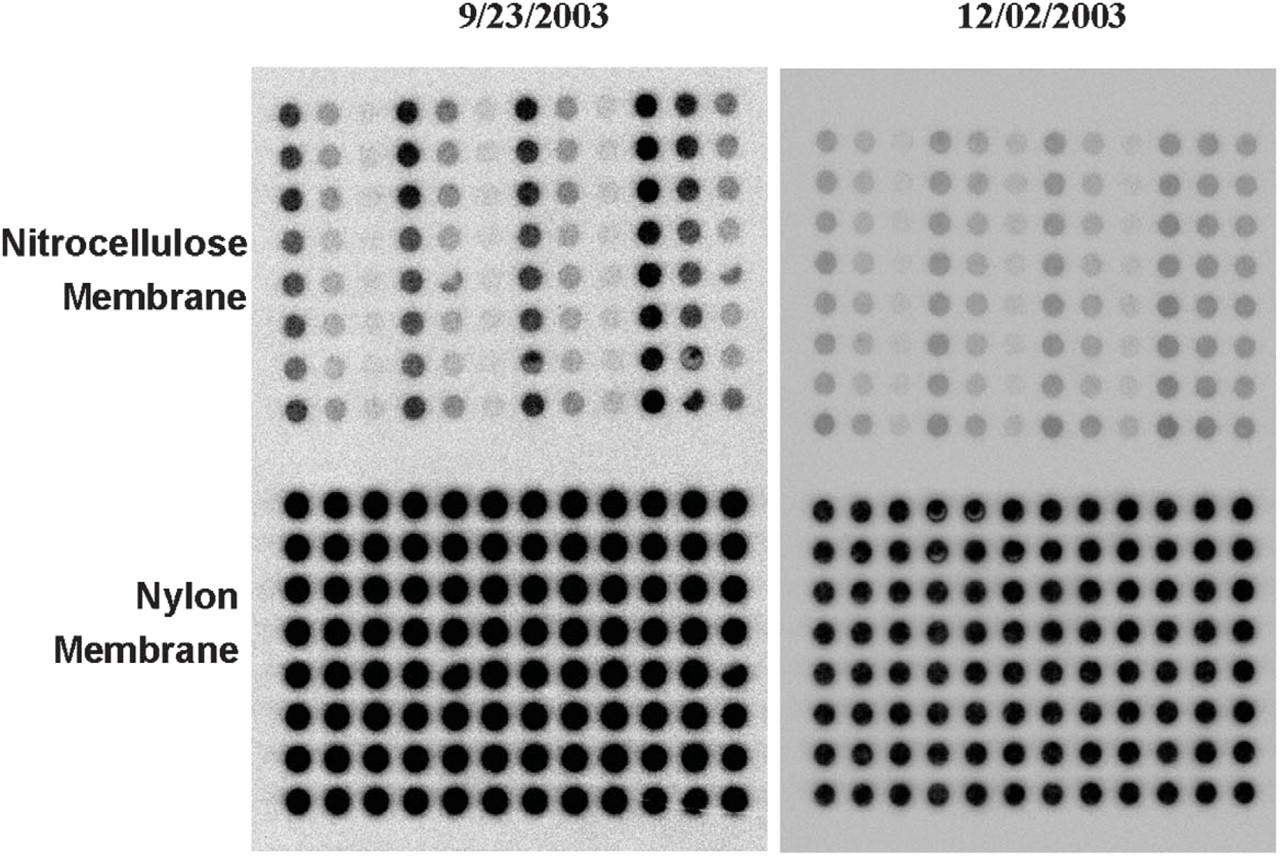

Reproducibility studies—comparison of Automated Buffer Matrix Tests against lysozyme (500 nM). Tests were performed on 9/23/2003 and 12/02/2003. Note: reproducibility is dependent on the quality of the target.
Additional reproducibility studies are planned. The Automated Buffer Matrix results of buffer solutions #1–3, 25–27, 49–51, and 73–75, which contain 100 mM monovalent salt, 20 mM Tris, pH 7.5, and 0 mM divalent salt, 1 mM MgCl2, or 1 mM CaCl2, varied greatly. These buffers, specifically, will be further studied and potentially eliminated from the matrix.
Additionally, we will perform aptamer selections using the conditions based on the Automated Buffer Matrix Test results. Last, we will correlate the Automated Buffer Matrix results with the Automated Aptamer Selection results.
Acknowledgments
The authors would like to thank Bruce Sullenger at Duke University (Durham, NC) for his helpful discussions and suggestions. This work was supported by a grant from the Welch Foundation (Grant No. F-1393).
