Abstract
Robotic liquid-handling stations (RLHSs) are the mainstay of high-throughput biomedical/forensic DNA sample processing facilities. These liquid-handling systems can be alternatively tooled with either disposable or fixed-tip pipetting heads. The use of disposable tips is often perceived as the best tip configuration to eliminate cross-contamination between biological samples processed on liquid-handling stations. However, this suppression can be effectively achieved on instruments equipped with fixed tips with optimally designed tip wash station (WS) configurations. Fixed-tip instruments offer many significant sample processing advantages with respect to precision, pipetting of liquids that may contain aggregates, and operational cost. This report discusses how cross-contamination suppression was achieved for the reliable processing of forensic casework samples on specially configured fixed-tip TECAN Genesis RSP/Freedom EVO RLHSs. A critical analysis of the major components involved in tip washing, as well as the specifications of a redesigned tip-washing routine that increases wash effectiveness and significantly reduces processing time and cost is also presented.
Introduction
Full- or semiautomation of analytical processes generally provide increased throughput, improved quality/efficiency, and reduced operational costs of laboratory testing, 1 –18 attributes that are in high demand in biomedical, forensics, or environmental laboratory applications. Combined with integrated sophisticated sample-tracking systems, robotic liquid-handling stations (RLHSs) offer the potential to process samples under the highest standards of quality control. RLHSs can be programmed to closely replicate the manual version of virtually any liquid-pipetting process.
When configured with pipetting heads accepting plastic disposable tips, RLHSs generally operate under operational limitations similar to those of the original hand-operated manual process they mimic, with the potential for elimination of cross-contamination. However, some limitations associated with the use of disposable tips must be considered when planning their deployment in a forensic casework processing environment. The use of RLHSs equipped with disposable tips may present programming challenges with respect to their ability to: (1) pipet liquids that may contain aggregated materials (e.g., clumps during magnetic bead-based DNA capture of high DNA yield samples) without contaminating the pipet head or sealing tip aerosol barriers which can cause tips to drip and (2) prevent the production of aerosols when performing complex, multistep pipetting maneuvers with detergent-containing reagents. From a practical standpoint, compared with regular pipet tips, racked disposable tips for robotic use are more complex to fabricate (e.g., reinforced racks, stringent tolerances on center-to-center spacing for 96-tip racks, carbon fiber-containing plastics to support liquid-level detection capabilities) and, therefore, are more expensive. Also, significant deck space must be allocated to supply the procedure with the required racked tips, restricting the number of deck positions available for process containers, thereby reducing throughput.
Conversely, RLHSs configured with fixed tips carry little in the way of consumable costs and offer more deck space for process containers. Fixed-tip RLHSs pipet liquids containing aggregated material more readily (i.e., pipetting displacement is provided by a system fluid that is far less compressible/expandable than air), with no risk of splashing and sealing aerosol barriers. With smaller air gaps (i.e., the air between the solid/liquid providing momentum displacement [e.g., piston or system fluid] and the pipetted liquid), fixed-tip RLHSs can perform complex, multistep procedures without generating aerosols, and are capable of higher precision and accuracy at low dispensing volume when compared with disposable tips. 1,4,9,19,20 However, fixed-tip RLHSs require well-planned pipetting regimens to avoid diluting a pipetted reagent with system fluid. They may also require customized tip-washing routines if elimination of the potential for sample cross-contamination is required for a given application. 2 –4,9,12,17,19,21 –23 This last limitation can be effectively addressed.
The initial success experienced using fixed-tip RLHSs for the automated processing of convicted offender samples submitted to the Canadian National DNA Data Bank 3 was the impetus for us to consider the use of the same tip configuration for processing casework samples on RLHSs. Years ago, tip-washing steps using 2% sodium hypochlorite (stock from manufacturer at 10.8%) were incorporated strategically within all robotic pipetting scripts (i.e., DNA extraction and quantification, DNA stock transfer from plate to tubes, and pre- and post-PCR setup) of our automated sample processing platform to eliminate the potential for carryover from one pipetting step to the next. 23 We have since been using these bleach wash routines for more than 2 years on fixed-tip RLHSs dedicated for the processing of casework samples (e.g., high-volume, no-suspect crimes [break and enter] and general casework) with no evidence of carryover. However, with standard passive wash station (WS) configurations on TECAN Genesis and EVO platforms, the “no cross-contamination” status (as measured by real-time PCR quantification [Q-PCR]) is achieved only at the cost of lengthy, time-inefficient wash cycles.
This report presents a detailed review of a fixed-tip RLHS configuration for the TECAN Genesis/Freedom EVO that provides no cross-contamination of samples, no increase in process execution time and dramatically reduced operating costs for the processing of casework samples. This report may serve as a validation template for clinical or forensic laboratories faced with similar goals using any fixed-tip RLHS.
Materials and Methods
RLHS Configuration Redesign
The TECAN standard passive WS design was reviewed to reduce wash cycle time while still achieving as good a wash or better. Evaluated modifications included: (1) the reduction of the wash cuvettes' diameter and depth in the “shallow” WS to increase the laminar flow speed of the robot system's liquid (reverse osmosis [RO]-grade water) used as washing fluid over the tips' outer surfaces, (2) the active draining of the wash cuvettes after use to ensure that no liquid remains stagnant in the cuvettes for extended periods of time, and (3) the redirection of the flow of system liquid channeled from the RLHS pressure relief circuit (PRC) to the cuvettes of both “shallow” WSs from the waste to effect an active flush step. In addition, the removal of the bulk of the contaminants was confined to a first WS while a polishing wash was subsequently performed by a second, paralleled, and identical redesigned WS.
The custom design evaluated for the new WS was made of high-density polyethylene instead of polypropylene (to further reduce adherence of materials), and the connection originally designed by TECAN for use with an external vacuum source (to help in the active draining of the wash cuvettes) was used to connect to the PRC instead. During initial trials, the components for the external vacuum source proved cumbersome necessitating excessive space demands. The new WS design uses a Y-connector to concurrently direct secondary flow to the “dirty” and “clean” shallow WSs (see Figs. 1 and 2). Ball check valves are used to ensure unidirectional flow and prevent diffusion. The shallow cuvette volume for the redesigned WS has been reduced from 400 to 200 μL as a result of a decrease in depth (from 2.5 to 2.1 cm) and diameter (from 4 to 3 mm) to promote higher-speed laminar flow and wash-fluid turnover.
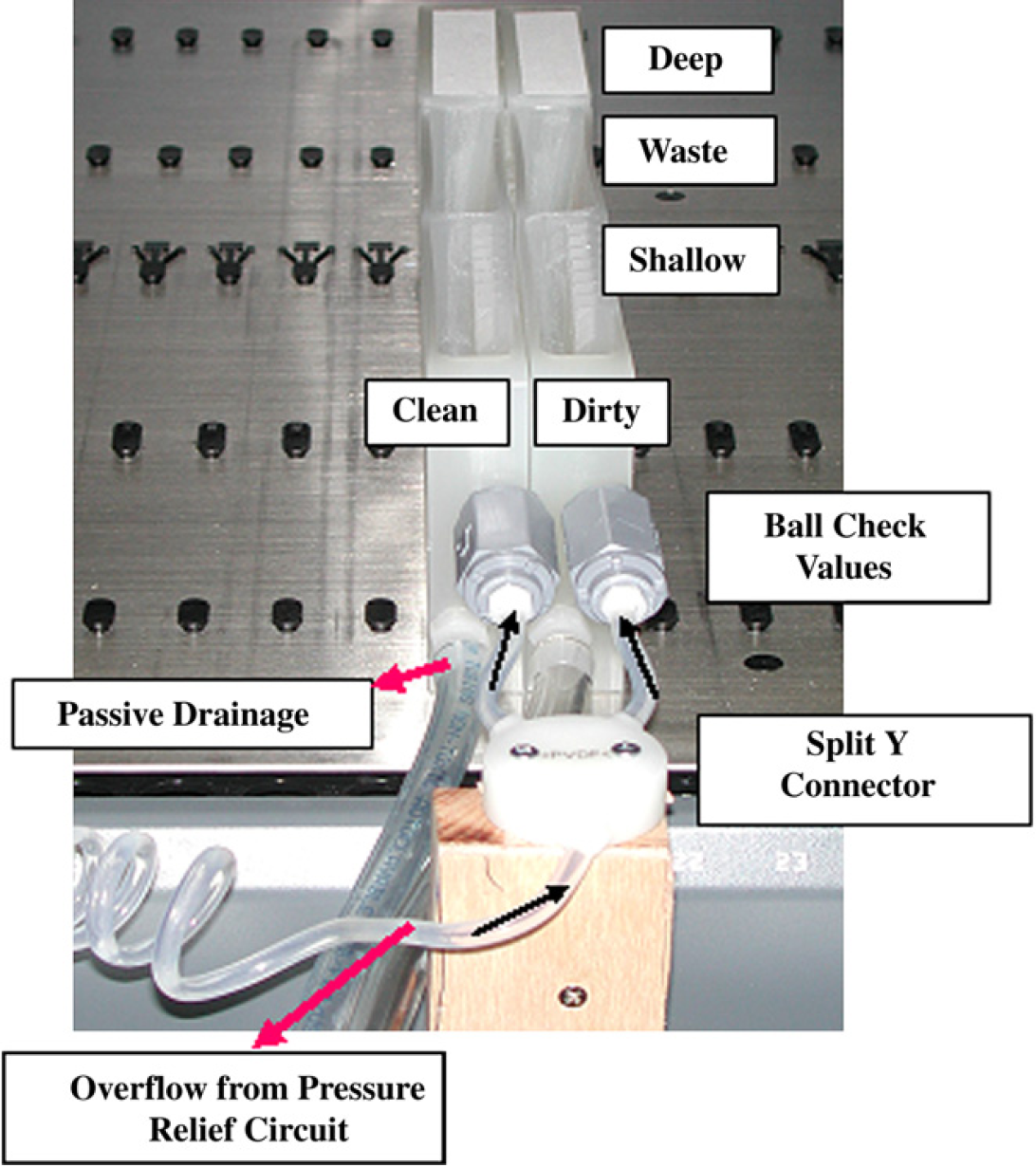
Redesigned WS configuration evaluated. See text for details.
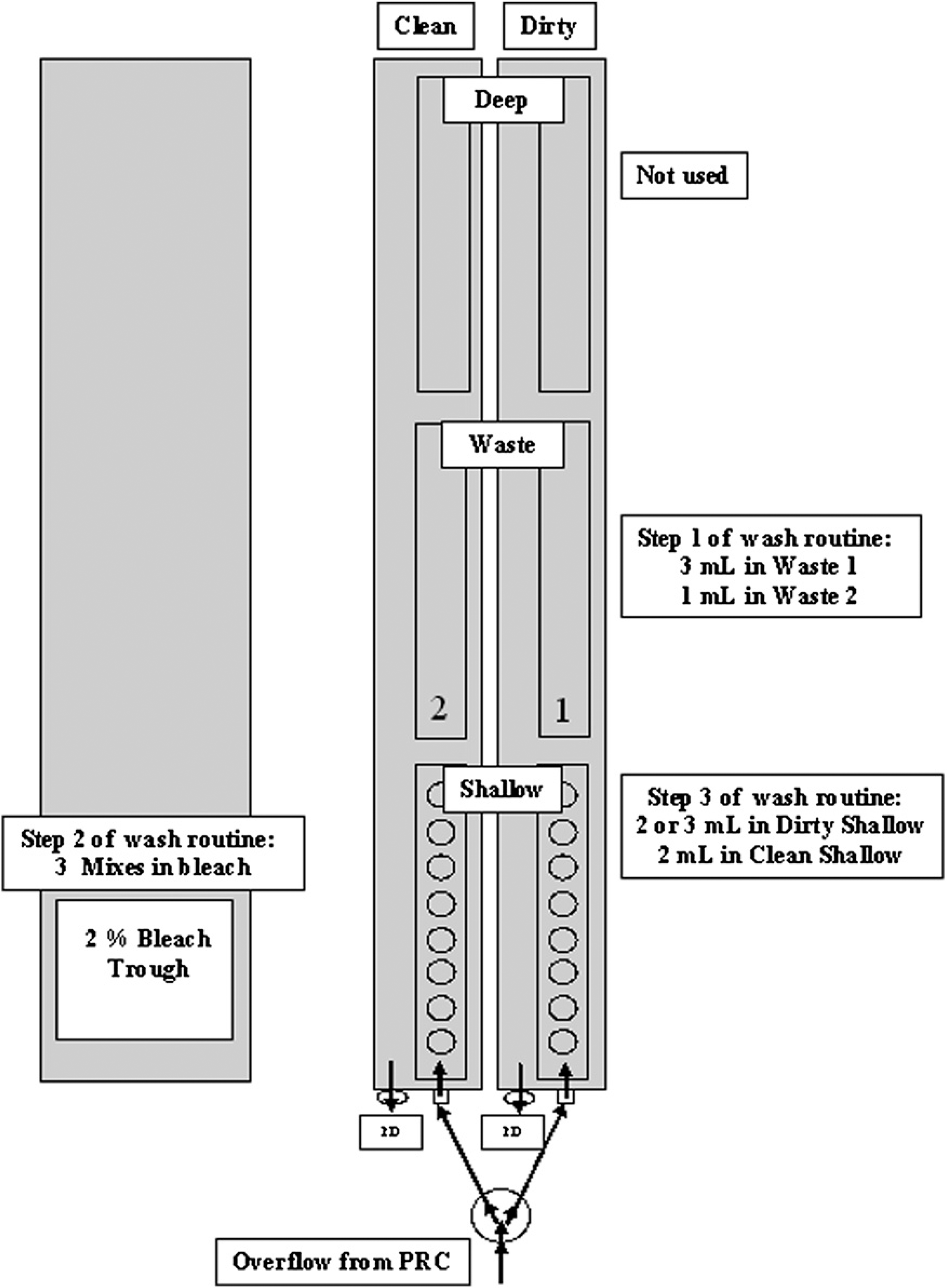
Diagram showing the redesigned WS and optimized final bleach wash routine adopted to eliminate biological sample cross-contamination. PD refers to passive drainage; PRC refers to pressure relief circuit. The volumes of system liquid shown for Step 3 of the wash routine represent the most frequently used volumes adopted during our automated DNA extraction process. Refer to Table 6 for other specific volumes adopted.
The principle of operation of the standard and redesigned WSs is shown in Figure 3. Both WSs share a standard washing path where the RLHS system fluid is expelled through the tips at high speed to wash both the tubing leading from the syringe to the tip as well as the inside of the tip, while creating turbulence in the wash cuvette to facilitate the removal of aggregate material and dilute liquid contaminants on the outside of the tip. The redesigned WS supplements this effect with a secondary flow of system liquid channeled from the RLHS PRC that is normally directed to waste on a standard WS. A 500-ms postwash delay (i.e., delay between pump stop and valve switch to syringe pumping) was used initially but increased to 750 ms to help ensure the absence of line pressure after the wash cycle completes.
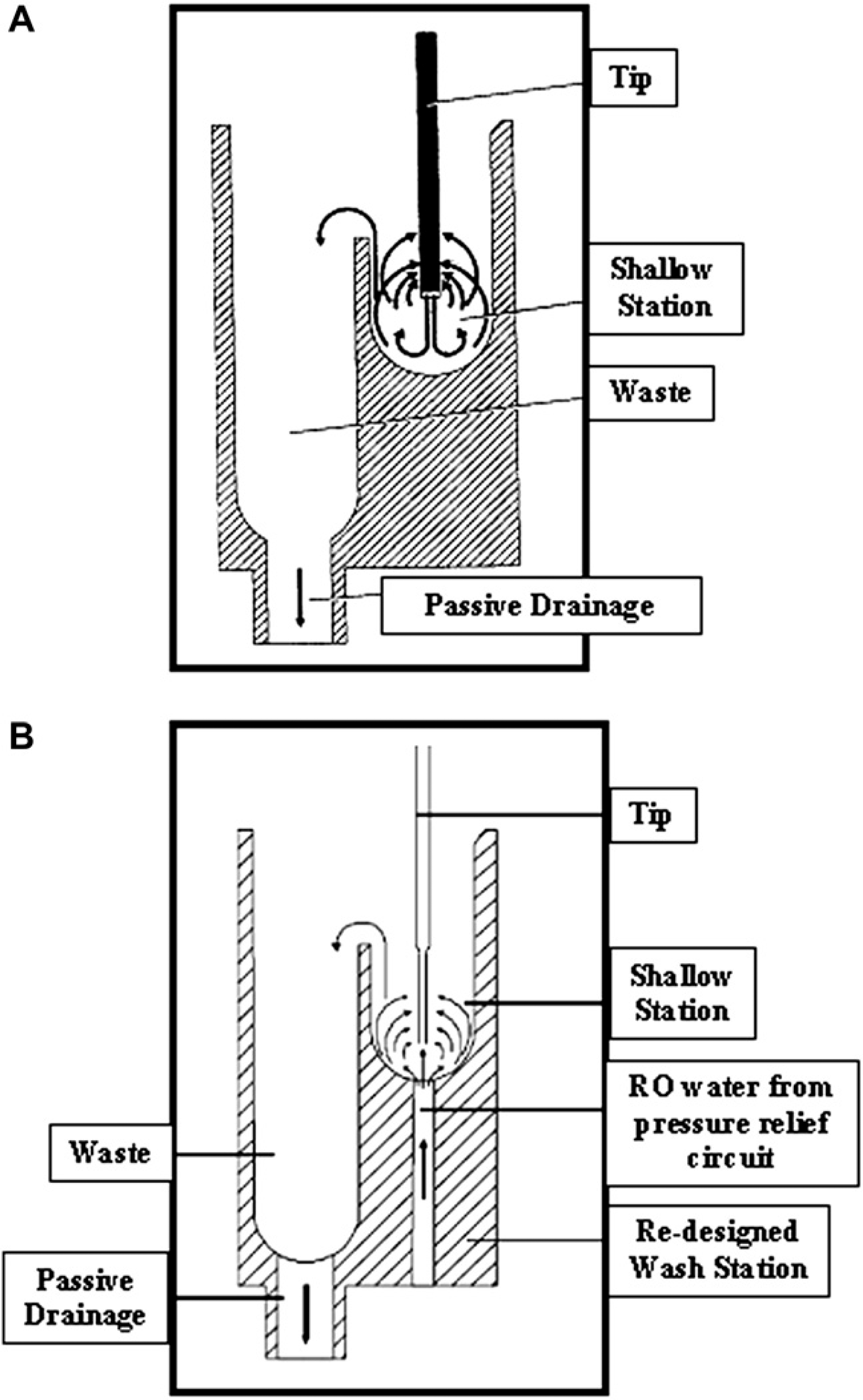
Diagram showing the principles of operation of a standard WS (Panel A) and the redesigned WS (Panel B) (see text for details). Diagrams adapted with permission from TECAN Genesis RSP and Genesis NPS Instruments Operating Manual—DOC ID 390783 V2.6, Oct 2001, pages 3–27 and 3–28, Tecan, Mannedorf, Switzerland.
Finally, the nondispensing, tapered end of the Teflon-coated stainless steel tips were redesigned. The tapered end of the tip that was intended to facilitate connection to the tubing leading to the syringe body was creating a void at the junction between the tip and tubing where stale liquid or beads could accumulate (Fig. 4). This void needed to be removed to eliminate every potential source of cross-contamination. The nondispensing end of the redesigned short, low volume adjustable tips are straight rather than tapered.
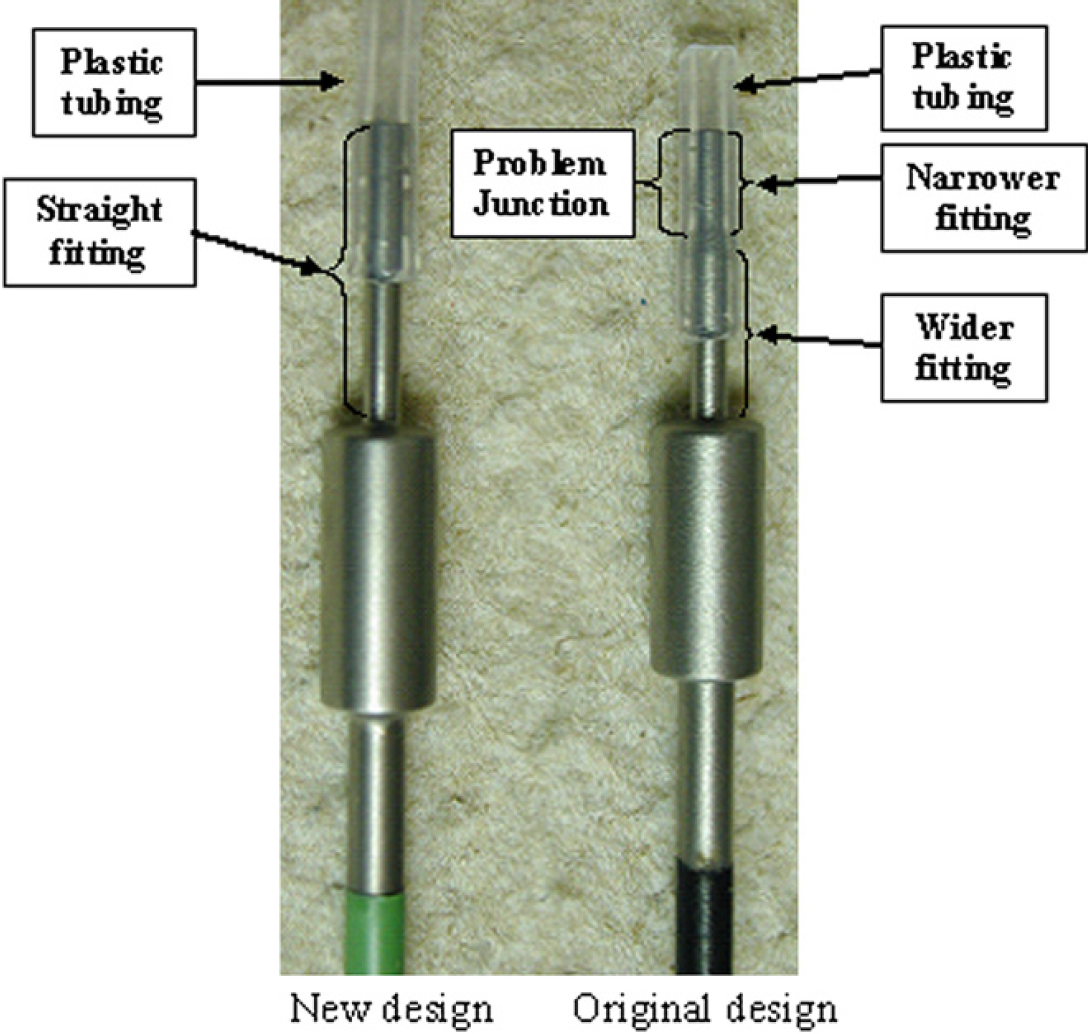

New versus old design for the low volume adjustable Teflon-coated stainless steel tips. See text for details.
Biological Sample and Blank Sample Preparation
Buccal swabs and blood swabs (2, 1, and 0.5 μL) from different volunteers as well as vaginal swabs from a few female donors spiked with neat semen (20 μL) were stored frozen at −20 °C until processed. For blood swabs, a 1:10 dilution in phosphate-buffered saline was prepared from the neat blood samples before applying 20, 10, and 5 μL on cotton swabs. Blank samples consisting of lysis buffer (LB) (10 mM Tris, pH 8.0, 10 mM EDTA, 100 mM NaCl, 0.5% sarkosyl, 40 mM dithiothreitol [DTT]) were included in all experiments to test the effectiveness of modifications to the wash routine in preventing the occurrence of carryover during automated DNA extraction/quantification processes.
Sample Lysis and Preparation for DNA Extraction
Samples destined for “direct” DNA extraction were lysed overnight at 56 °C in 350 μL of LB with 1.5 mg/mL of proteinase K (Prot. K). Samples were pulse-centrifuged to bring down condensation, and swabs were transferred to Spin-eZe baskets (Fitzco Inc., Spring Park, MN) using either sterile wooden sticks or sterile forceps. The baskets were reinserted in the tube where the contained swab originated and the tube/basket assemblies were centrifuged for 3 min at 13,200 rpm (16,300 g) to pellet debris. The baskets containing the swabs were discarded. Lysates were either processed immediately or stored frozen at −20 °C.
Samples from mock sexual assault cases and destined for “differential” DNA extraction were lysed for 2 h at 37 °C in 350 μL of LB without DTT with 0.2 mg/mL Prot. K. Samples were pulse-centrifuged and the swabs were transferred to Spin-eZe baskets as described above. The tubes were centrifuged for 10 min at 15,000 rpm (21,000 g) to pellet sperm cells, and the baskets containing the swabs were discarded. A volume of 315 μL was removed from the epithelial cell (EC) lysates to an intermediate tube for safekeeping as a backup. A volume of 315 μL of fresh LB without DTT with 0.2 mg/mL Prot. K was added to the EC lysates, and the tubes were vortexed. Lysates were either processed immediately or stored frozen at −20 °C.
Biological Sample and Blank Sample Layout for Contamination Checks
The zebra-stripe format (alternating columns of samples containing an abundant source of DNA and reagent blank samples) was used to assess cross-contamination of samples. Buccal swabs and vaginal swabs spiked with a large aliquot of semen were specifically evaluated in our study to present the worst case scenario for potential contamination because they normally yield large amounts of DNA and could challenge the RLHS tip-washing routine.
Automated DNA Extraction
Unless specified otherwise, all pipetting steps described below are performed by the RLHS.
With the 2-mL plate positioned on the activated TE-Shake unit, lysates (350 μL) were transferred from tubes to the deep-well plate with two additional volumes of Promega Lysis Buffer (PLB) to effect DNA capture by the beads. After 15 min on the shaker, the bead/DNA complexes were pelleted by quick centrifugation (ramping to 4000 rpm [2254 g], 20 s) then magnetized on a LifeSep 96F flat magnet (Dexter Magnetic Technologies, Chicago, IL). The spent liquid was removed from all wells to the waste station and the bead/DNA complexes were washed once with PLB and twice with Promega Wash buffer on the activated TE-Shake unit. Each washing step was carried out by placing the plate on the magnet between wash steps to aspirate off the solution. The bead/DNA complexes were air-dried for 5 min and the DNA eluted in 60 μL of low TE buffer (10 mM Tris, 0.1 mM EDTA, pH 9.0) twice at 65 °C for a total of 16 min (8 min each time) with vortexing for 10 s between elutions.
While DNA from the EC fraction was being captured by the magnetic beads, 50 μL of LB without DTT with 0.2 mg/mL Prot. K was added by the RLHS to each of the original sample tubes containing the sperm pellets. These samples were vortexed for 5–8 s and then centrifuged for 5 min/15,000 rpm in a microfuge before being returned to the RLHS. The wash supernatants were discarded, and then 350 μL of heated (65 °C) PLB with DTT (10 mM) was added to the sperm cell pellets. After a 5-min incubation, the sperm cell lysates were transferred in Columns 6–11 of the same deep-well plate already containing the EC lysate/PLB liquid columns (in Columns 1–5). Extraction was continued for both EC and sperm fractions according to the guidelines of the original “direct” protocol. Sperm cell DNA was captured on beads by agitating for a total of 5 min. The bead/DNA complexes were pelleted by quick centrifugation, the spent liquid was removed and samples were washed, beads were air-dried, and the DNA was eluted as per the “direct” protocol described above.
DNA Quantification
All extracted DNA samples and blank samples were quantified by Q-PCR with the AB Quantifiler Human DNA Quantitation assay (Applied Biosystems, Foster City, CA) on an ABI Prism 7000 Sequence Detection System as outlined in the AB draft protocol. A volume of 22.5 μL of master mix (10 μL of primer/probe solution and 12.5 μL reaction mix [containing the Taq Gold polymerase]) was dispensed with 2.5 μL of extracted DNA or standard DNA for a final PCR volume of 25 μL. The K562 cell line DNA was used as standard curve material, with data points between 32 and 0.0156 ng/μL. All PCR reactions were set up robotically. Amplification was carried out for 40 cycles.
Amplification Conditions for DNA Extracted from Biological Samples
A spreadsheet template was written in-house to feed the RLHS with correct volumes of stock DNA and diluent required to perform normalization to a target concentration suitable for PCR. Low yield samples (<1 ng total DNA) were filtered through Montage PCRμ96 Filter Plates (Millipore, Mississauga, ON, Canada) as detailed in the next subsection for blanks. AmpF/STR Profiler Plus PCR reactions were set up by the RLHS in 96-well plates with 1 ng of DNA in a final PCR volume of 15 μL. Plates were sealed with a Simport Secure Seal membrane (Simport, Beloeil, QC, Canada) in combination with a Corning Thermowell Sealing Mat (Corning, Acton, MA). This sealing mat was used as a cushion between the sealed plate and the thermal cycler heat bonnet by positioning it upside down (i.e., bumps in an upward position; different from manufacturer's instructions) on the sealed PCR plate; it proved very effective in preventing evaporation by uniformly pressing around the rim of each well of the sealed plate. PCR plates were cycled in an MJ Research DNA Engine Peltier Thermal Cycler (PTC-200) according to the following run parameters: 95°C/11 min, once; 94 °C/60 s, 59 °C/90 s, and 72 °C/90 s, for 28 cycles; 60 °C/45 min, once; and room temperature/overnight.
Processing Conditions for Blank Samples
Blank eluates (50 μL of low TE) were filtered through Montage PCRμ96 Filter Plates on the RLHS configured with a TECAN Vacuum Separation Module (Te-VacS) as follows: (1) wells of filter plates were prewetted with 100 μL of a 0.01-mg/mL bovine serum albumin as a blocking agent to prevent nonspecific binding of DNA to the filter, (2) vacuum was applied on the filter plate for 4–6 min at which point the blank eluates were transferred to the filter plate, (3) vacuum was applied until all liquid had drained from the wells (3–6 min), (4) vacuum was broken off and 16 μL of AmpF/STR Profiler Plus PCR cocktail were added to each well, (5) the filter plate was moved to the TECAN TE-Shake unit and shaken for 80 s/800 rpm to promote recovery of any material captured on the filter, and (6) the recovered material was transferred to a PCR plate and amplified as detailed in the previous section.
Analysis of Fluorescently Labeled Amplification Products
Amplicons were analyzed by capillary electrophoresis on ABI Prism 3100 Genetic Analyzers. Aliquots (1 μL) of the amplified material were combined by the RLHS with 0.5 μL GeneScan-500 and 20 μL of HiDi formamide (Applied Biosystems), heated for 2 min at 90 °C and snap-cooled at 4 °C for 3 min using the robot cool block. Samples were electrokinetically injected for 10 s at 3 kV onto 36 cm capillary arrays, and electrophoresed through POP-4 (Applied Biosystems) at 15 kV and 60 °C.
Chromatograms were generated with the GeneScan Analysis 3.7 and Genotyper 3.7 programs (Applied Biosystems). The peak-detection threshold used during AmpF/STR Profiler Plus profile analysis and interpretation was set at 20 relative fluorescence units (RFUs) to maximize the detection of peaks in all samples including blanks.
Results
Efficient washing protocols for fixed tips and integrated robotic platform design are important aspects of the development of cross-contamination containment and effective high-throughput contamination control.
Evaluating Redesigned WS Configurations with New Settings for the Fast Wash Pump (FWP)
The system liquid's flow distribution at different wash volumes and FWP settings was assessed for the new WS configuration in reference to the 15% “control” FWP setting implemented initially for casework processing and 500 ms postwash delay conditions selected for the development of the bleach wash routine. Table 1 shows that partial diversion of system fluid through the PRC could be measured at FWP setting of 30% and over. At lower flow rates, the system fluid was expelled strictly through the tips. With a setting of 50%, equivalent volumes of system fluid enter the wash cuvettes through the tips and PRC. At that setting, the total wash volume (tip + pressure relief) is at or near maximum (74 mL for a 10-mL wash volume per tip) and the tip flow rate is also at or near maximum (0.51 mL/s or 188.9% compared with “control”), thus representing optimal conditions for effective tip washing for this tip diameter. Further increase in FWP settings did not provide significant additional time saving. Similar results were obtained with a 750-ms postwash delay, so this setting was used in subsequent experiments (see Table 1).
Tip and PRC volume output using various FWP settings and wash volumes with the redesigned WS configuration a
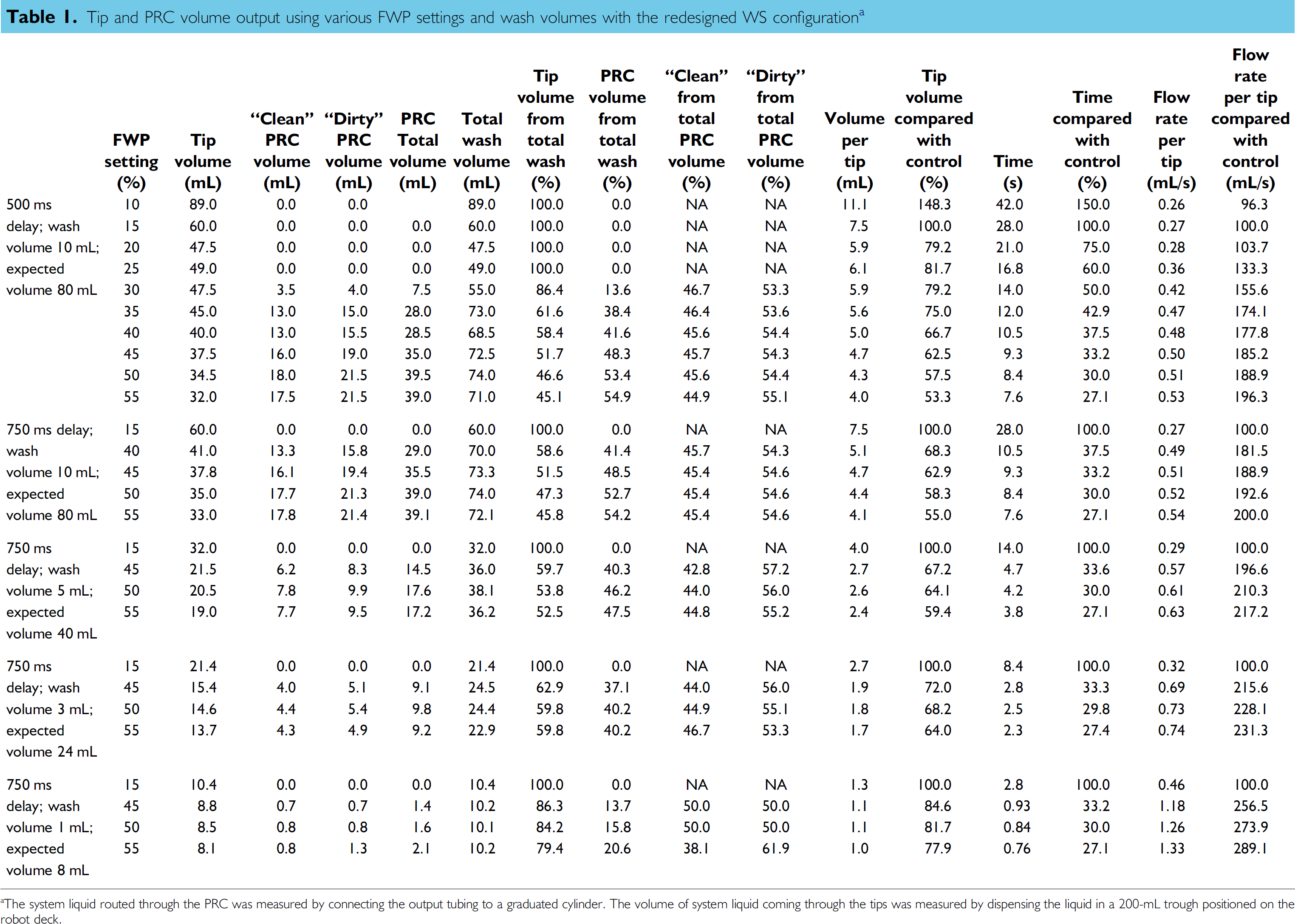
The system liquid routed through the PRC was measured by connecting the output tubing to a graduated cylinder. The volume of system liquid coming through the tips was measured by dispensing the liquid in a 200-mL trough positioned on the robot deck.
Determining Time Savings Using the New WS Configuration and New FWP Setting
A series of “blank runs” were performed to estimate process time saving with the use of the new WS configuration with the FWP set at 50%. The automated DNA extraction protocol was run incorporating the wash conditions as defined in Table 2 using RO water in place of biological samples. As shown in Table 2, approximately 1 h is dedicated to washing the fixed tips using the wash routine developed to process casework in our operational unit (FWP = 15%; 4 mL [waste], 10 mL [“dirty” shallow], and 10 mL [“clean” shallow]) when comparing fixed tip with simulated disposable tip processes. Increasing the FWP to 50% while retaining the same wash volumes results in an overall time gain of 36 min. More significant time saving was realized on following the reduction of the volumes of system fluid used in the washing steps (see below).
Process time for extracting a full batch of samples (N = 88) with and without washing steps
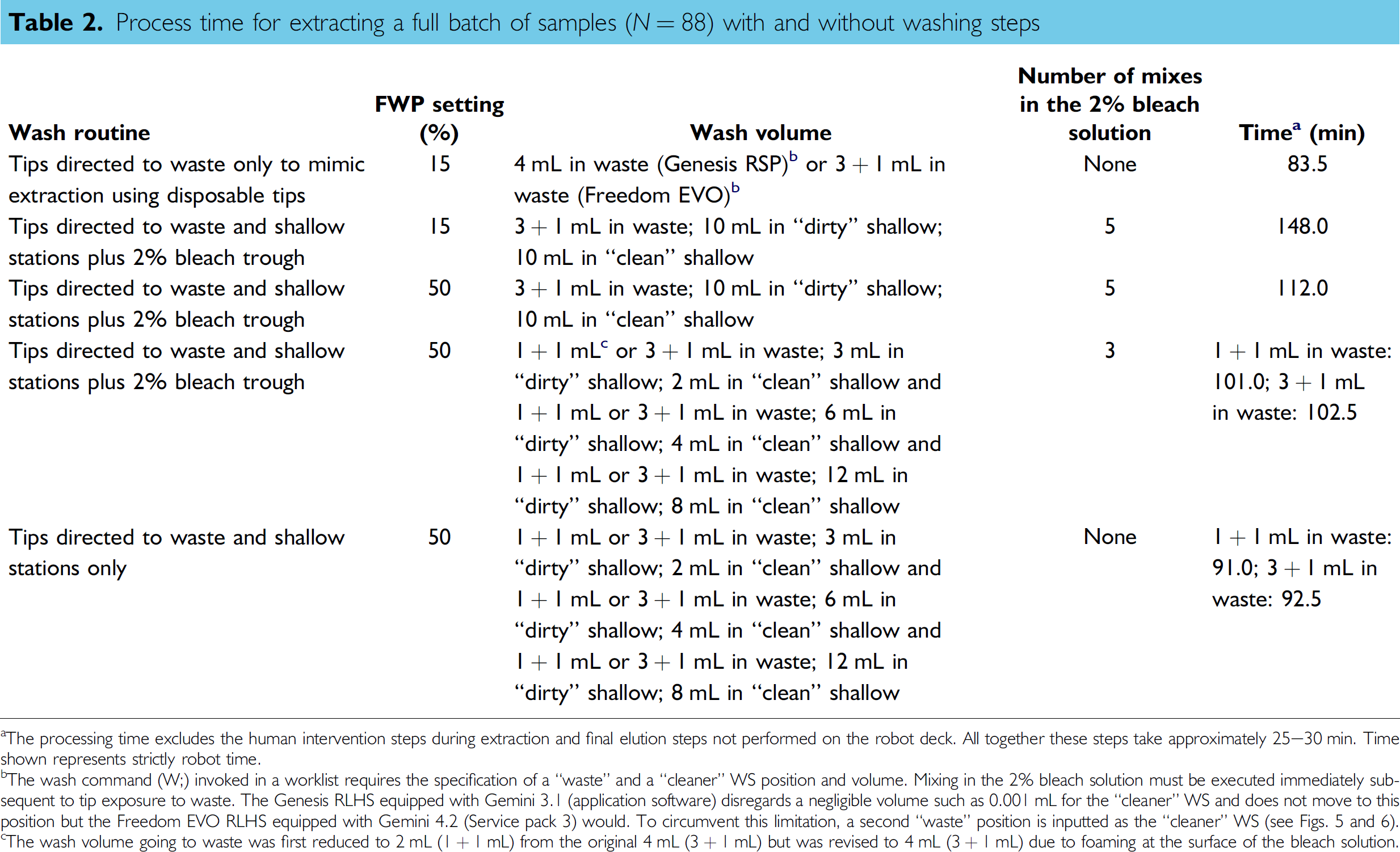
The processing time excludes the human intervention steps during extraction and final elution steps not performed on the robot deck. All together these steps take approximately 25–30 min. Time shown represents strictly robot time.
The wash command (W;) invoked in a worklist requires the specification of a “waste” and a “cleaner” WS position and volume. Mixing in the 2% bleach solution must be executed immediately subsequent to tip exposure to waste. The Genesis RLHS equipped with Gemini 3.1 (application software) disregards a negligible volume such as 0.001 mL for the “cleaner” WS and does not move to this position but the Freedom EVO RLHS equipped with Gemini 4.2 (Service pack 3) would. To circumvent this limitation, a second “waste” position is inputted as the “cleaner” WS (see Figs. 5 and 6).
The wash volume going to waste was first reduced to 2 mL (1 + 1 mL) from the original 4 mL (3 + 1 mL) but was revised to 4 mL (3 + 1 mL) due to foaming at the surface of the bleach solution.
Reducing Wash Volumes Enhances Process Efficiencies
Table 3 presents the original wash volumes that were used to prevent cross-contamination during the automated DNA extraction using tapered fixed tips, the original WS configuration and a FWP setting of 15%. The “tip interior rinse factor” and “tip exterior rinse volume” are included. The lysate remaining in the tips/lines after lysate transfer was diluted 58 times and the lysis column remaining in the tips/lines after lysate column aspiration was diluted 18 times. Depending on the amount of bleach wash aspirated into the line, the 2% bleach wash solution was diluted 15, 38, or 60 times during the DNA extraction process. Although these wash volumes were shown to be effective at preventing contamination, they were potentially beyond what is necessary to achieve the goal of no cross-contamination. Table 4 summarizes the new reduced wash volumes that were adopted as an initial attempt to gain efficiencies by decreasing the time required to perform the automated DNA extraction process. A dilution factor of 8 and a volume of 4 mL were arbitrarily selected as the minimum “tip interior rinse factor” and “tip exterior rinse volume,” respectively. Establishing this reference allowed for a measured, proportional alteration of wash conditions if required. For example, at lysate transfer followed by a wash step in our automated protocol, 330 μL of lysate and 400 μL of bleach are aspirated. The volume of system fluid used in the washing steps specific for this portion of the automated protocol was set at 1 + 1 mL (waste), 3 mL (“dirty” shallow), and 2 mL (“clean” shallow) which corresponded to a “tip interior rinse factor” of 16.7 for the lysate-transfer step and 8.3 for the bleach-aspiration step and a “tip exterior rinse volume” of 4.3 mL for both steps (see Table 4). Each step of the protocol was carefully evaluated to minimize the wash volumes while maintaining the minimum established dilution factor and exterior rinse volume for the DNA extraction and DNA quantification protocols. The wash steps' commands were adjusted so that 60% of the wash volume was directed to the “dirty” station and 40% to the “clean” station. The intent was to maximize cleaning in the “dirty” cuvettes such that the “clean” cuvettes were exposed to minimal contaminants and served primarily as a final pristine rinse. The number of mixes in the bleach solution was reduced from five to three for additional time saving. As indicated in Table 2, the reduction in wash volumes and number of mixes in the bleach trough decreased processing time by an additional 9.5–11 min for a full batch of samples. The elimination of the bleach step from the wash routine while conserving the flushing steps with system fluid in the “waste,” “dirty shallow,” and “clean shallow” stations could result in an extra 10 min saving in the overall process. However, in the stepwise development of any automated DNA protocol, gains in efficiency must be evaluated with respect to achieving a valid result without enhancement of contamination. As noted in the next section, elimination of specific bleach mixes during the wash steps could promote sample carryover.
Original wash volumes (system fluid) used for effective cross-contamination prevention during automated DNA extraction using the original tapered fixed tips and the standard WS configuration with the FWP at 15% on TECAN Genesis RLHSs
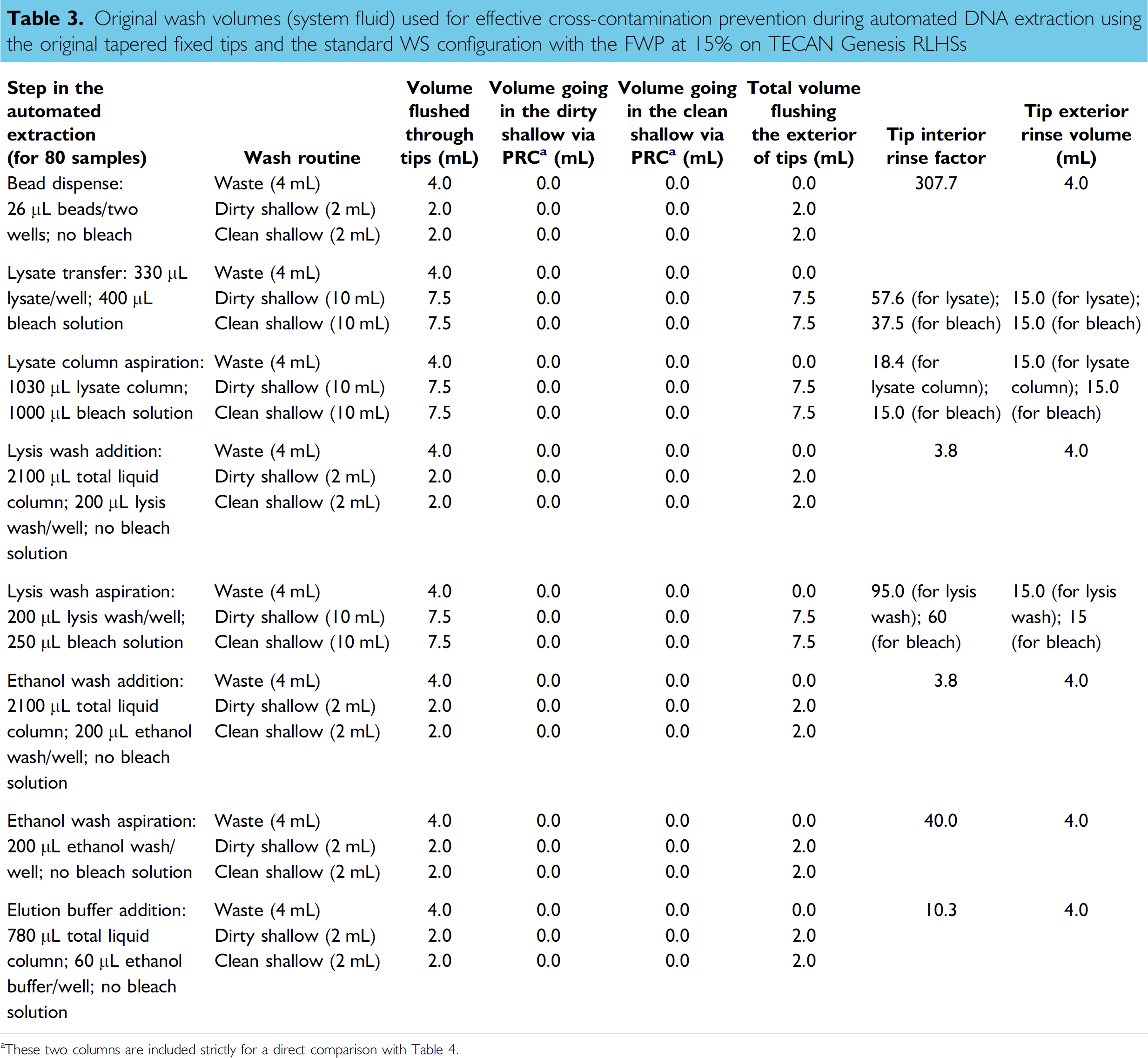
These two columns are included strictly for a direct comparison with Table 4.
Reduced wash volumes (system fluid) used for effective cross-contamination prevention during automated DNA extraction using nontapered fixed tips and the new WS configuration with the FWP at 50% on TECAN Freedom EVO RLHSs
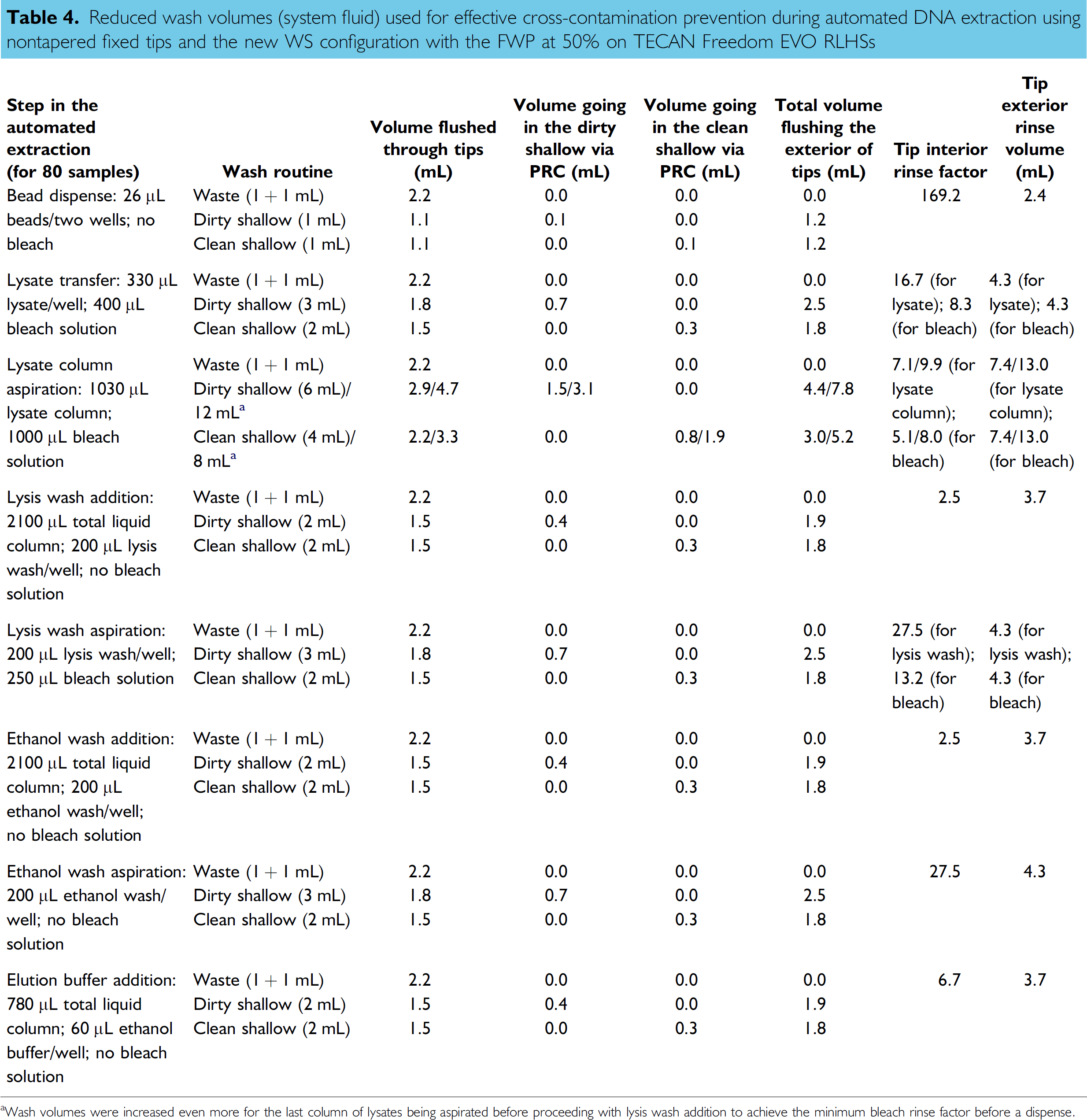
Wash volumes were increased even more for the last column of lysates being aspirated before proceeding with lysis wash addition to achieve the minimum bleach rinse factor before a dispense.
A similar review of the wash volumes implemented during automated DNA quantification was also carried out and volumes were reduced according to our reference minimum “tip interior rinse factor” and “tip exterior rinse volume” (data not shown).
Assessing the Robustness of the New Wash Routine Using Highly Concentrated Biological Samples
The new wash routine incorporating the reduced wash volumes shown in Table 4, redesigned tips, and WS configuration was evaluated for prevention of cross-contamination while enhancing biological sample processing efficiencies. Buccal swabs (yielding between 500 and 1000 ng total DNA) were used to determine the robustness of this new wash routine using our automated “direct” extraction protocol. Table 5 summarizes the DNA quantification results for blanks extracted using this new wash routine as well as the outcome of the AmpF/STR Profiler Plus amplifications of those samples. As shown, 72 blanks were extracted in the first three experiments performed. Three of those blanks showed marginal Q-PCR values following quantification, two had C T values greater than 36 cycles and one had a C T lower than 36. Based on our internal validation of the AB Quantifiler Assay, blank samples with a C T < 36 usually showed either complete or partial short tandem repeat (STR) profiles, whereas blank samples with C T > 36 did not produce any STR signals. Marginal and spurious Q-PCR values could be the result of the instability of the fluorochrome marker noted following many cycles of amplification (i.e., 36 cycles to 40 cycles; Applied Biosystems, personal communication). None of the blanks processed generated STR products when subjecting the entire concentrated blank eluate for amplification using 28 cycles, 1 μL of amplified material for analysis on the ABI Prism 3100 Genetic Analyzer and using a peak-detection threshold of 20 RFUs. This data indicated that blanks were not contaminated with DNA when evaluated under our operational experimental conditions.
Cross-contamination checks performed using the new nontapered fixed tips, the new WS configuration and the reduced wash volumes
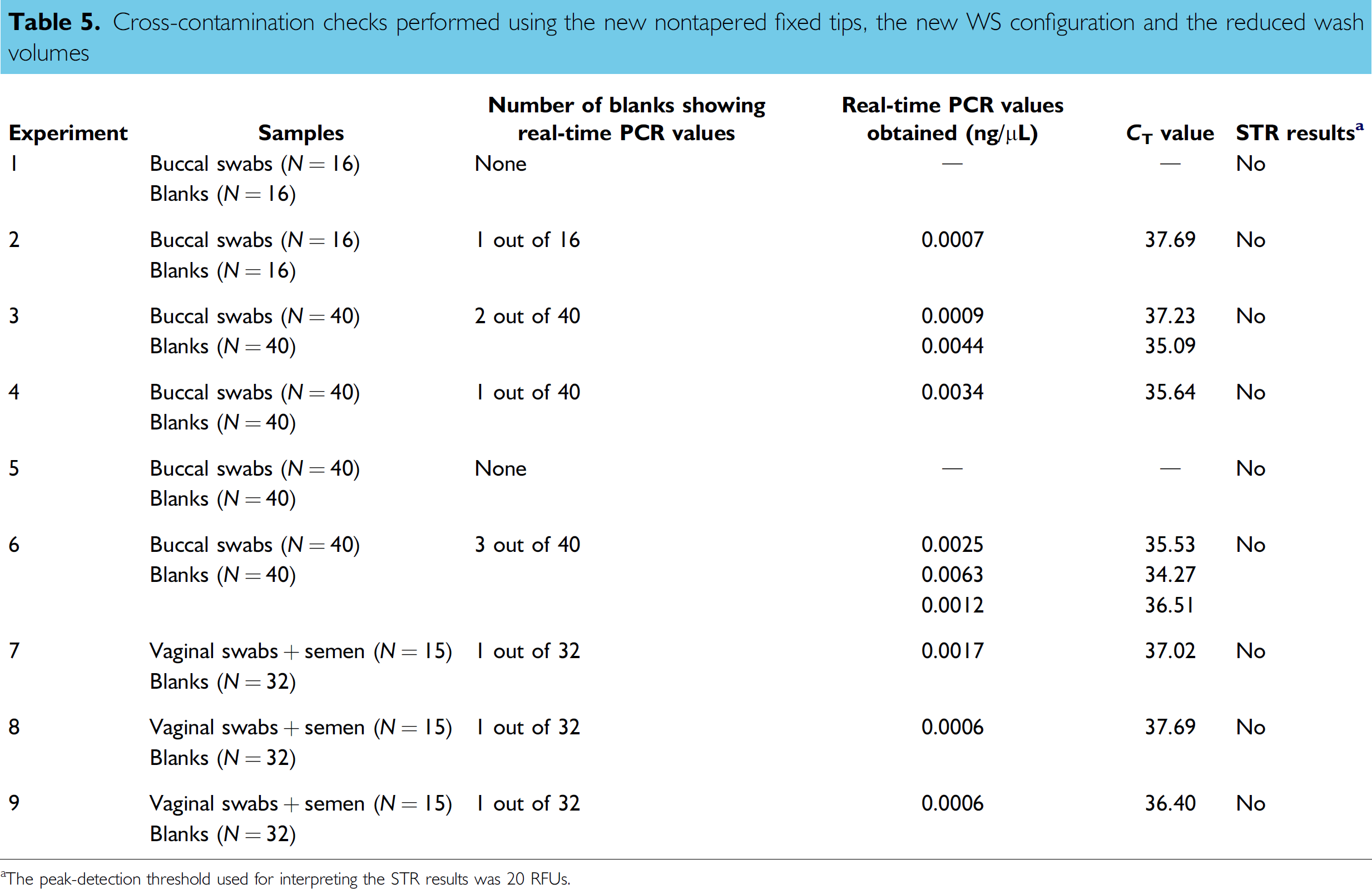
The peak-detection threshold used for interpreting the STR results was 20 RFUs.
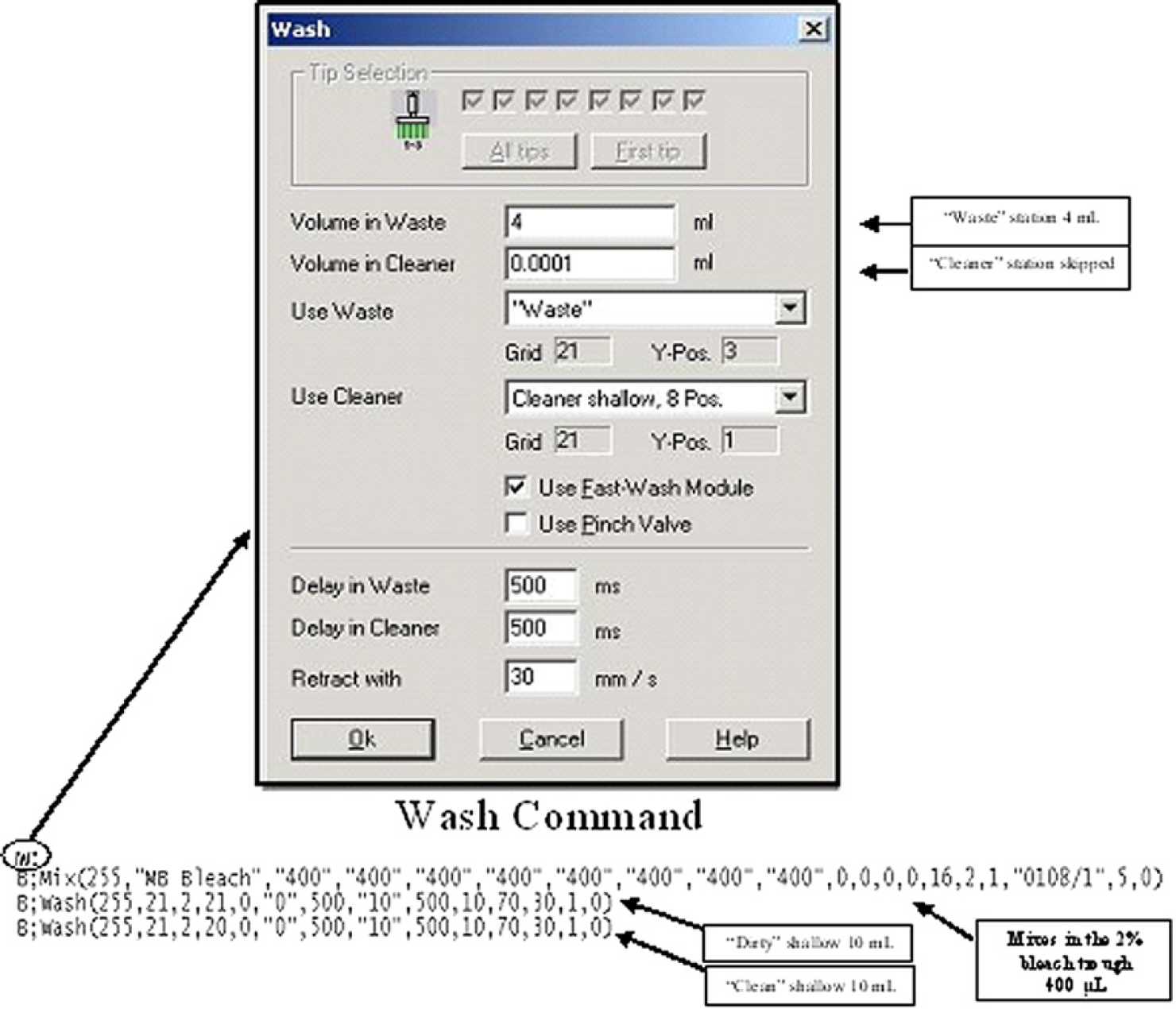
Details of the wash command executed during the bleach wash routine for fixed tips on the TECAN Genesis RSP 150/8 work-station using Gemini 3.1. The minor differences to the definition of the wash command (W;) invoked in a worklist required to maintain a uniform bleach routine between the Genesis and Freedom EVO RLHS models are shown. Mixing in the 2% bleach solution in a trough must be executed immediately subsequent to tip exposure to the waste station. The wash command requires the specification of a “waste” and a “cleaner” station position and volume. RLHSs equipped with Gemini 3.1 disregard a negligible volume such as 0.001 mL for the “cleaner” station and do not move to this position.
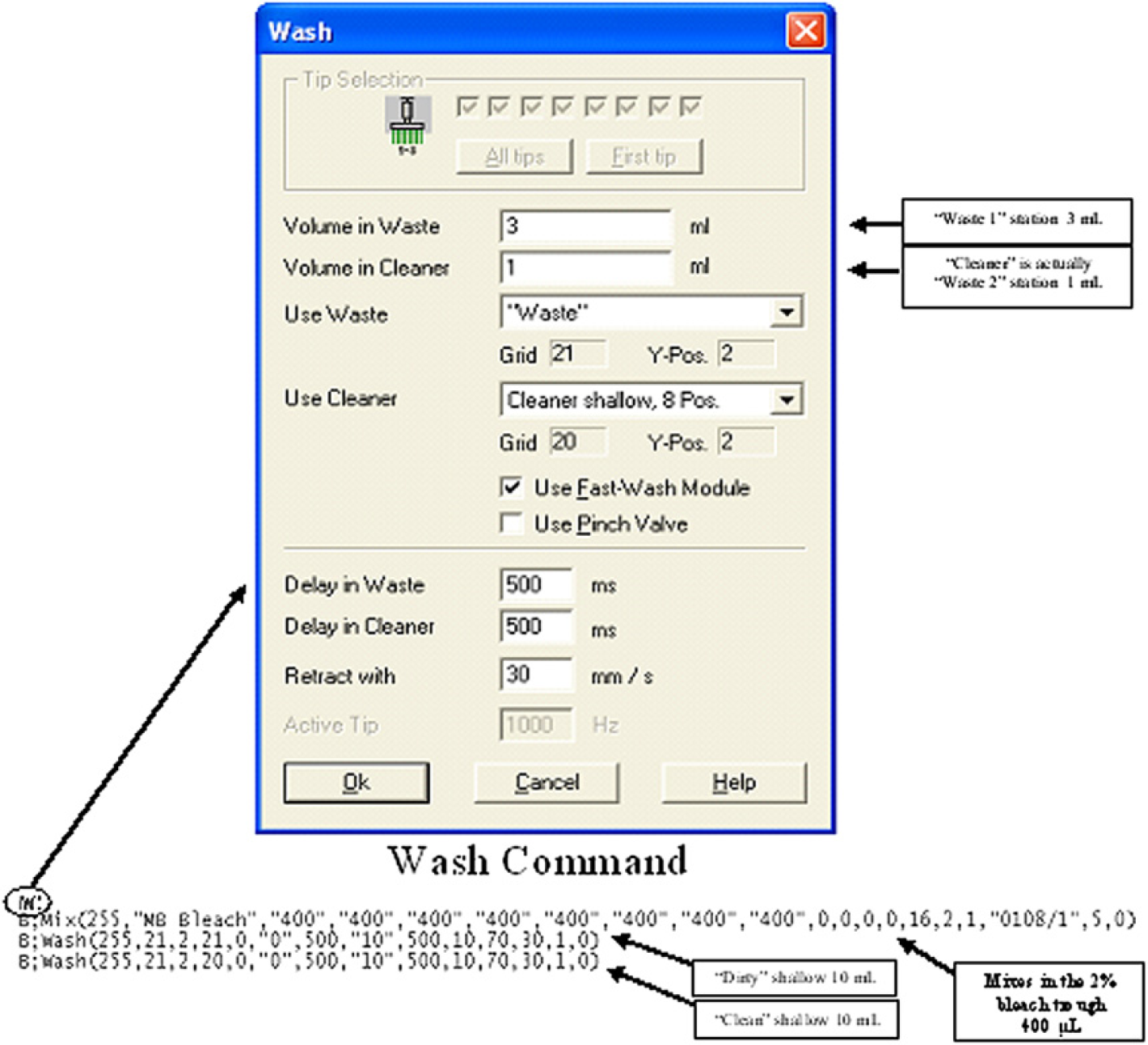
Details of the wash command executed during the bleach wash routine for fixed tips on the TECAN Freedom EVO workstation equipped with Gemini 4.2 (Service pack 3). Refer to legend of Figure 5 for further explanation. RLHSs equipped with a higher version of Gemini (v. 4.2) cannot disregard a negligible volume such as 0.001 mL for the “cleaner” station and will move to this position. To circumvent this limitation, a second “waste” station position is inputted as the “cleaner” station.
Based on this first series of experiments, the reduced wash volumes implemented during DNA extraction and quantification (process thoroughly reviewed) appeared effective at preventing cross-contamination. However, foam was noted at the surface of the bleach trough during the wash steps. This phenomenon was noted previously while developing the bleach routine 23 and was likely the result of the presence of lysis solution (high guanidium isothiocyanate content) remaining in and around the tips. During the wash routine, tips are first directed to waste to expulse remaining liquid and then to the bleach trough. The use of a 1 + 1 mL wash command to flush the lines and tips in the waste appeared insufficient to eliminate remaining lysis solution inside and around the tips. To prevent foaming, a 3 + 1 mL waste-volume command was required while keeping all other wash-volume commands to their minimum (see Table 6). Three additional experiments were carried out using the adjusted wash volume for the waste station to confirm that no contamination was detected using high concentrated DNA samples such as buccal scrapings. As indicated in Table 5, 120 blanks were extracted in the second set of three experiments. Four of those blanks showed marginal Q-PCR values following quantification and all four had C T values very close to 36 cycles. No STR products were detected when using the entire blank eluate for amplification. These results confirmed that the reduced wash volumes implemented during DNA extraction and quantification appeared effective at preventing cross-contamination. A total of 45.5 min of processing time was saved following the adoption of this new wash routine. Further reduction of wash volumes was not pursued as the potential time saving was minimal with only 10 min of wash time remaining in the protocol. The new wash routine was also tested during the automated “differential” DNA extraction of vaginal swabs spiked with semen and similar results were obtained.
Final wash volumes (system fluid) used for effective cross-contamination prevention during automated DNA extraction using nontapered fixed tips and the new WS configuration with the FWP at 50% on TECAN Freedom EVO RLHSs
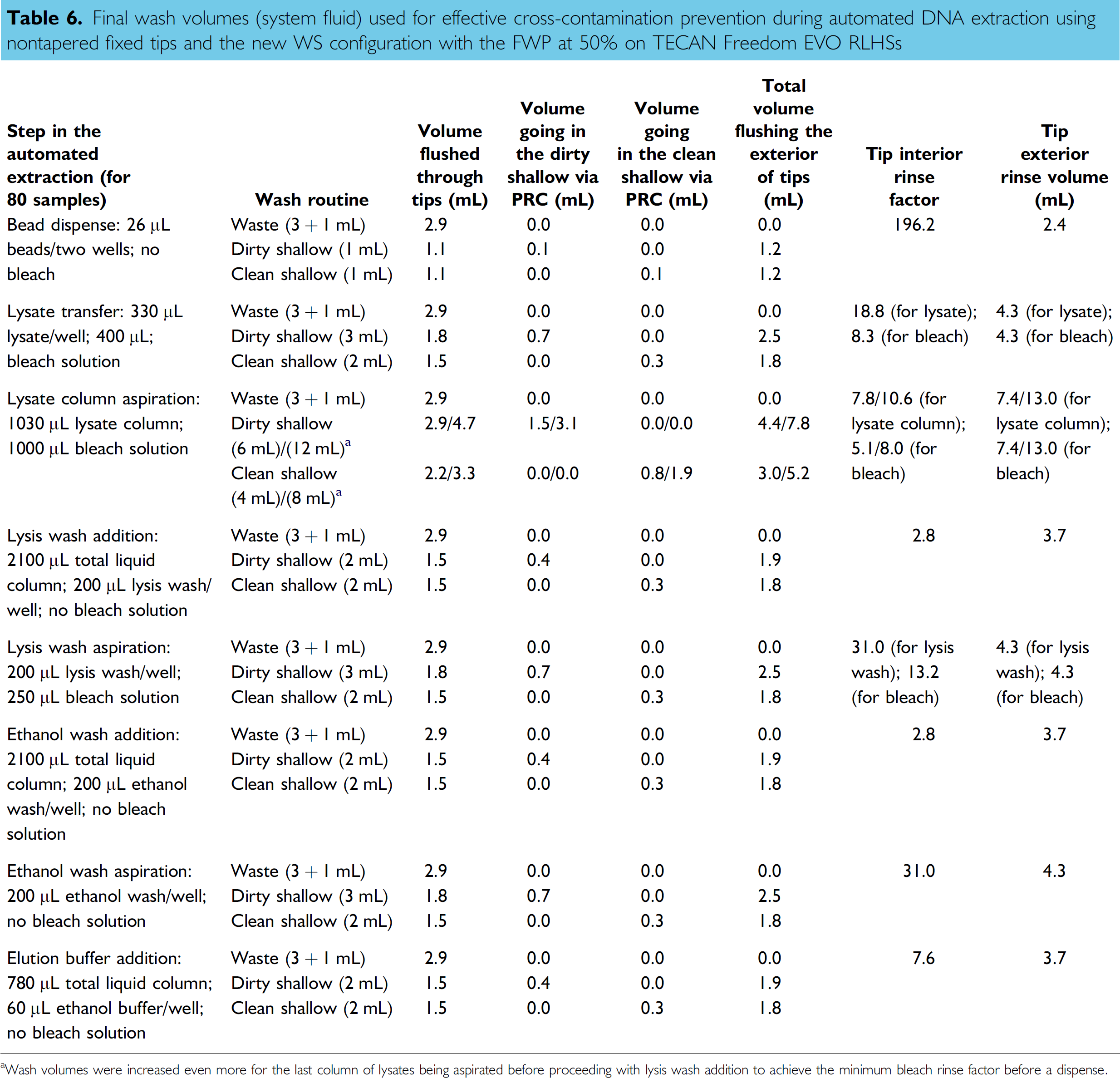
Wash volumes were increased even more for the last column of lysates being aspirated before proceeding with lysis wash addition to achieve the minimum bleach rinse factor before a dispense.
In an attempt to gain even more time, the bleach mixes were eliminated all together from the wash routine. Buccal samples from two individuals (N = 16) were processed alongside blank samples (LB buffer; N = 16) using our “direct” extraction protocol. Samples were set in a zebra layout in a deep-well plate. Only five blanks out of 16 had no Q-PCR values. Eleven blanks out of 16 blanks extracted showed marginal Q-PCR values following quantification. Six had C T values below 36 cycles and five had C T values above 36 cycles. All blanks processed produced either partial or complete STR profiles when using the entire eluate for amplification. In all instances, the profiles corresponded to the previous set of samples pipetted by the robot. Representative AmpF/STR Profiler Plus profiles obtained are shown in Figure 7.
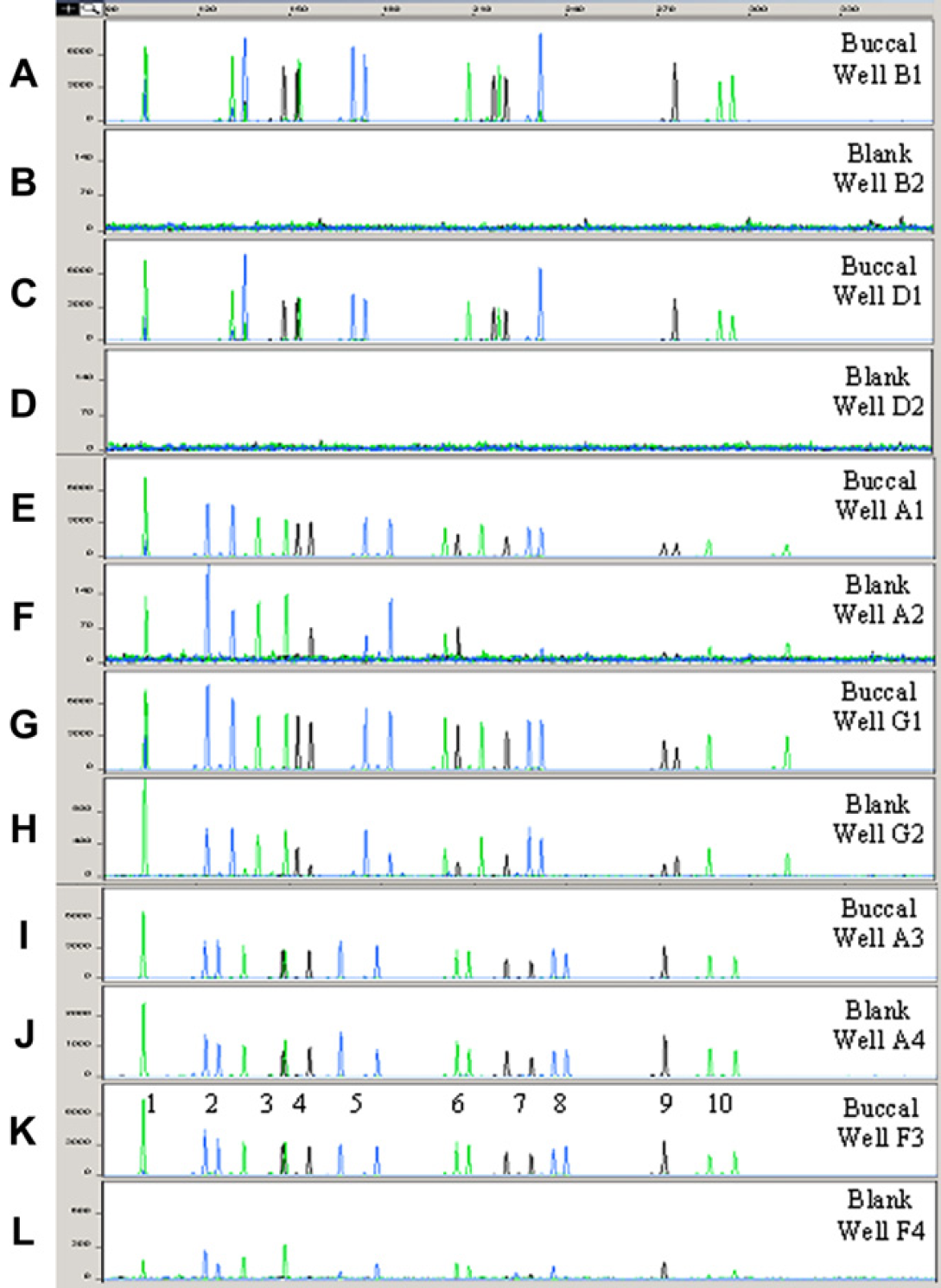
STR results from blank samples processed alongside buccal samples on RLHSs with bleach (Panels A–D) or without bleach (Panels E–L). Buccal and blank samples were extracted using the “direct” DNA extraction protocol under the new wash conditions (new WS configuration, FWP at 50%, reduced wash volumes, and new fixed tips). For amplification, 1 ng of template DNA was used for buccal swabs while the entire filtered eluate was subjected to PCR for blanks. For Panels E–L, only the 2% bleach step was omitted from the wash cycle, that is, tips were still directed to waste and to the dirty and clean shallow stations. Each panel depicts the relative fluorescence intensity (RFU, Y-axis) and the size estimate in bases (X-axis) derived from the internal lane standard GeneScan-500 [ROX] using the ABI GeneScan Analysis version 3.7 software. The genetic markers observed from left to right, labeled 1–10, are Amelogenin, D3S1358, D8S1179, D5S818, HumvWA, D21S11, D13S317, HumFGA, D7S820, and D18S51.
Assessing the Quality of the STR Profiles from Low Concentrated Samples Subjected to the New Wash Routine
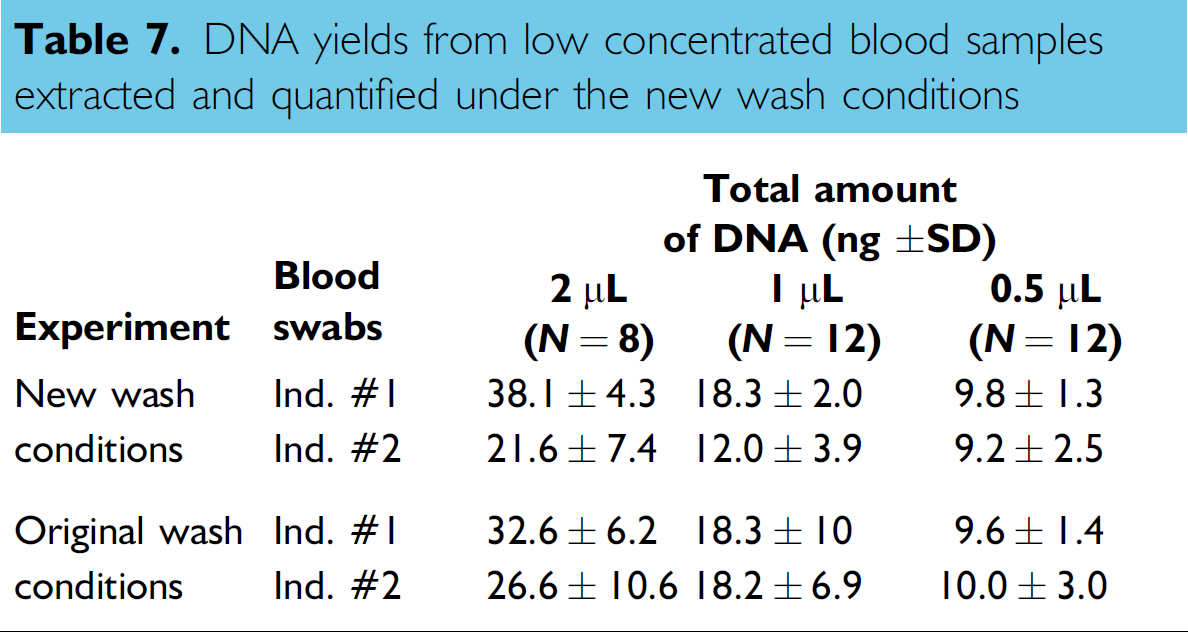
To rule out the possibility that traces of bleach could remain following the new wash conditions, additional studies were carried out using samples with low DNA concentrations. The experimental design ensured that DNA degradation was not masked by the high-concentration samples used in the previous experiments. The examination of the DNA yields and quality of the AmpF/STR Profiler Plus profiles produced from small aliquots of blood (≥2 μL) applied on cotton swabs should provide an indication that the wash routines effected sufficient dilution of the bleach to prevent bleach-induced PCR inhibition and DNA degradation. Table 7 and Figure 8 summarize the results obtained. All yields derived from blood swabs processed under the new wash conditions were comparable with those obtained using the old wash routine. This was especially true for Donor #1. More variation was noted for Donor #2 compared with Donor #1 for which a perfect doubling factor was observed between the various aliquots of blood tested. The quality of the AmpF/STR Profiler Plus profiles generated using 1 ng of target DNA under the new wash conditions was excellent as depicted in Figure 8. The profiles showed excellent balance in the allele peak height intensity across all STR loci amplified and peaks of heterozygous loci were equally balanced. These results confirmed the effectiveness of the new wash conditions in diluting the 2% sodium hypochlorite used during the automated DNA extraction and quantification processes.
DNA yields from low concentrated blood samples extracted and quantified under the new wash conditions
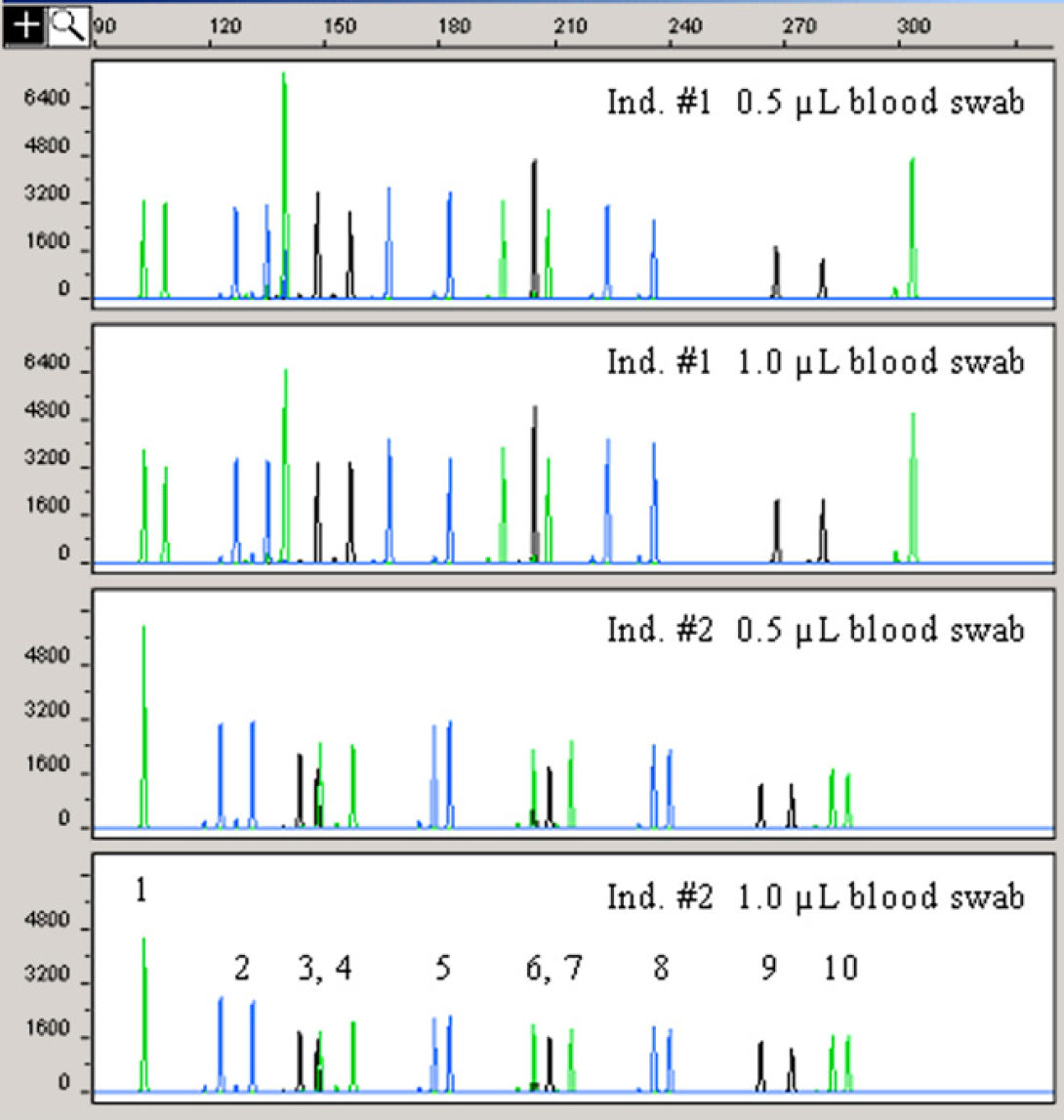
AmpF/STR Profiler Plus profiles from diluted blood samples processed using the new wash conditions. PCR amplifications were performed using 1 ng of template DNA. Refer to legend of Figure 7 for details on the genetic markers detected.
Discussion
Critical to the implementation of efficient DNA automation is the necessity to achieve an acceptable balance between a “no cross-contamination status” and a rapid washing protocol to economize on the length of time required to process a large batch of samples. During the developmental work that led to the design of our 2% bleach wash routine, it was recognized that the existing WSs, fixed tips, and FWP configuration could be improved to achieve better efficiencies and reduce processing time. A series of modifications were incorporated which allowed us to successfully reach our goal of processing a large number of samples in the shortest period of time while maintaining a “no cross-contamination status.”
During the execution of the original 2% sodium hypochlorite routine on all our robotic workstations, the fixed tips were directed consecutively to the “waste” station, a bleach trough, the “dirty” shallow and the “clean” shallow stations of the standard TECAN WS(s). 23 The incorporation of the bleach washing steps within the automated DNA extraction added 64 min to the overall process for a full batch of 88 samples.
In this paper, a simple modification to the FWP setting to 50% was shown to reduce the wash process by 36 min by increasing flow rate. A new WS configuration with narrower and shallower wells connected to the PRC created increased wash turbulence and force which, in conjunction with a new tip design, was more effective at dislodging and purging any contaminants present inside or around the tips. These modifications resulted in our capacity to further significantly reduce the wash volumes to gain an additional 9.5–11 min in the overall process. This reduction in the wash volumes did not impact our ability to achieve a “no-contamination” status based on the lack of STR profile development from all blanks tested. The new and enhanced wash routine proved very effective in preventing contamination for both the “direct” and “differential” automated DNA extraction protocols.
DNA automation is typically used to gain efficiencies in speed and capability in processing many samples with enhanced quality assurance. As our studies have shown, wash routines using select volumes and bleach decontamination can prove to be most effective in suppressing crossover contamination. However, each of these measures should be carefully introduced into the automated protocol and be subject to constant monitoring. Automation must be assessed with appropriate controls and involve routine maintenance to promote optimal delivery of service. We would recommend that users establish the “pump profile” of each of their RLHSs, that is, the relationship between the volume specified in the wash command and actual output volume and distribution. Variations in the FWP output between our TECAN RLHSs have been observed with associated variations in tip flow rate. The total pump output volume was observed to vary by as much as 25% for a 10-mL wash command at a 15% FWP flow rate. For a 5-mL wash command at a 50% FWP flow rate, the total pump output volume was observed to vary by as much as 10%. Therefore, instrument-specific wash commands and/or FWP percentage settings for each RLHS may prove necessary. The volume output could be greater or equal to those cited in this report. The inner diameter size of the fixed tips selected for use will also affect the volume output. Users should set the wash-volume commands such that the wash volume is greater or equal to those that have been validated at a greater or equal flow rate to maintain the minimum established dilution factor and exterior rinse volume across the protocols.
Continued monitoring of blanks should provide confidence in each laboratory's respective carryover preventive measures. We opted to assess sample-to-sample contamination by subjecting the entire blank eluate to STR amplification. Other investigators currently processing casework using RLHSs equipped with disposable tips evaluate contamination by using the maximal volume of bead eluate that can be easily accommodated in the PCR reaction as a representative aliquot of the entire eluate 7,15 or the same size aliquot as that used for regular samples is subjected to amplification. 13 These approaches were adopted based on the genotyping systems, amplification conditions, and sensitivity level of the detection instruments used for DNA analysis and validated in these respective laboratories. In the clinical field, users of automation assess potential sample carryover by treating the negative samples as regular samples in their respective assays. 2,4,19,24 –26
The elimination of the 2% sodium hypochlorite mixes from the wash routine, although permitting an additional gain of 10 min, is not recommended. We showed in a previous report that the bleach step was essential to prevent cross-contamination using RLHSs equipped solely with fixed tips. 23 These results were obtained using standard WSs in combination with tapered tips, larger wash volumes, and the FWP set at 15%. In the current study, under the new WS configuration and using nontapered tips, reduced wash volumes and the FWP at 50% for a more efficient wash, bleach was also proven essential to prevent sample carryover. Reverting back to greater volumes of system fluid using the FWP at 50%, in an attempt to bypass the bleach step and wash away DNA contaminants, may not be sufficient and would result in lengthier wash cycles. The 2% sodium hypochlorite mixes serve as a reassurance of contamination prevention and add essentially 10 min to our overall automated process for a full batch of 88 samples. Our automated process using fixed tips and including bleach and associated wash steps takes less than 20 min more than the homologous process using disposable tips and the savings in operational cost are significant. 23
Conclusion
The new and enhanced bleach routine for Teflon-coated stainless steel tips was proven very effective in eliminating contamination in both the “direct” and “differential” automated processes. An overall gain of 45.5–47 min was noted in processing time for a batch of 88 samples following an increase in the FWP output to 50% and a reduction in the overall wash volumes for DNA extraction and quantification. Under the new wash parameters, the quality of the AmpF/STR Profiler Plus profiles produced remained excellent. The combination of a new WS configuration, reduced wash volumes, and new nontapered fixed tips allow biological samples to be processed almost as quickly as under a disposable tip configuration. Overall, the elimination of cross-contamination, pipetting precision, capability to handle aggregated materials, and lower processing costs make this fixed-tip RLHS configuration a sound platform choice for the processing of casework material.
Acknowledgments
The authors would like to thank Michael Sahl, Susan Walker, Christine McEachern, and Pamela Stewart from the Biology Section of the RCMP Forensic Laboratory Services—Ottawa for their participation in the development of the automated filtration step for DNA eluates using Montage PCRμ96 Filter Plates and Dr. Benoît Leclair for his critical review of the manuscript.
