Abstract
We describe a reagent system and robotic method for purifying plasmid DNA for restriction digestion, PCR, and fluorescent sequencing applications. The method uses two types of Wizard® MagneSil™ paramagnetic particles. Following lysis and neutralization procedures, the first particle type binds and removes cell debris; the second type is then used to bind plasmid DNA from the cleared lysate. The particles are then washed to eliminate unwanted contaminants. Purified plasmid DNA is then eluted from the particles with nuclease free water. When using a cell mass of approximately 4 O.D.600, the yield is 10–12μg of DNA when using high copy number plasmid. When used in BigDye® terminator sequencing, these DNA templates typically yield read lengths greater than 700 bases and Phred 20 scores of 600 to 750 bases. This purification method has been adapted for use on several robotic platforms in a 96-well format.
Introduction
MagneSil™ paramagnetic particles are coherently clad silica micro-particles with a magnetite core. This proprietary matrix 1 2 combines the advantages of silica/DNA chemistry 3 magnetic capture, and solution kinetics. 4 There are numerous purification applications based on the MagneSil™ chemistry; PCR fragment clean up, 5 sequencing extension products clean up 6 and genomic DNA from food product, 7 plant material, 8 and blood. 9 This chemistry has been adapted for the purification of plasmid DNA from bacterial cells. 10
Purifying plasmid DNA is a valuable procedure for investigating and analyzing cloned genes. While several purification systems are commercially available, most use a solid phase, fixed-membrane extraction system, which requires a vacuum system for both filtration and concentration of the plasmid DNA. These systems require specialized labware and large reagent volumes, particularly for wash procedures, before eluting the plasmid target.
To address the increased interest of the scientific community for both higher throughput and lower cost per preparation, Promega developed a system that incorporates our MagneSil™ paramagnetic particle technology for purifying plasmid DNA from bacterial culture. Historically, either centrifugation or vacuum filtration has been used to clear genomic DNA and cell debris from the bacterial lysate. The plasmid DNA is then reversibly immobilized, washed, and eluted.
Promega has developed two formulations of MagneSil™ paramagnetic particles to facilitate purification of plasmid DNA without centrifugation or filtration steps. 11 MagneSil™ BLUE removes genomic DNA and cell debris from the bacterial lysate, leaving plasmid DNA behind in the aqueous phase. MagneSil™ RED is then used to selectively bind the plasmid DNA from the cleared lysate.
Separation of the MagneSil™ plus bound material complex for both of these steps is accomplished by using a strong external magnetic field. The MagnaBot® 96 Magnetic Separation Device is designed to perform simultaneous separation of 96 samples in a multiwell plate format. The combination of the two MagneSil™ particle formulations and the magnetic separation device produce an easily automated purification system based entirely on solution chemistry. The system is compatible with a variety of robotic platforms with 96-well plate footprint workstations.
Materials and Methods
Reagents used for purification were from the Wizard® MagneSil™ Plasmid Purification System (Cat. # A1630, Promega Corporation, Madison, WI). Plasmid DNA was extracted from Escherichia coli JM109 transformed with pGL3 or E. coli DH5a transformed with pGEM®-3Zf(+). Cells were grown in 2ml, 96-well plates using Terrific Broth media (1.2% tryptone, 2.4% yeast extract, 0.4% glycerol in 89mM potassium phosphate buffer) containing 100μg/ml ampicillin to maintain selective pressure. Each well, containing 1ml of broth, was inoculated with a single isolated colony from overnight agar plates (LB + Amp100). The plates were loosely covered with aluminum foil to allow gas exchange then placed on an orbital shaker inside a 37°C incubator. Plates were shaken at 800rpm for 16–18hrs. Using these conditions, an average total cell mass of approximately 4 O.D.600 was obtained. The cells were harvested by centrifuging in a table-top centrifuge at 300rpm (1500 μg) for 10 minutes.
Plasmid DNA purification was performed on a Biomek® FX Liquid Handling Workstation (Beckman Coulter, Inc., Fullerton, CA) equipped with a single multichannel pod and 96-well pipetting head (Figures 1). The deck setup for a single 96-well plasmid purification process used the following:
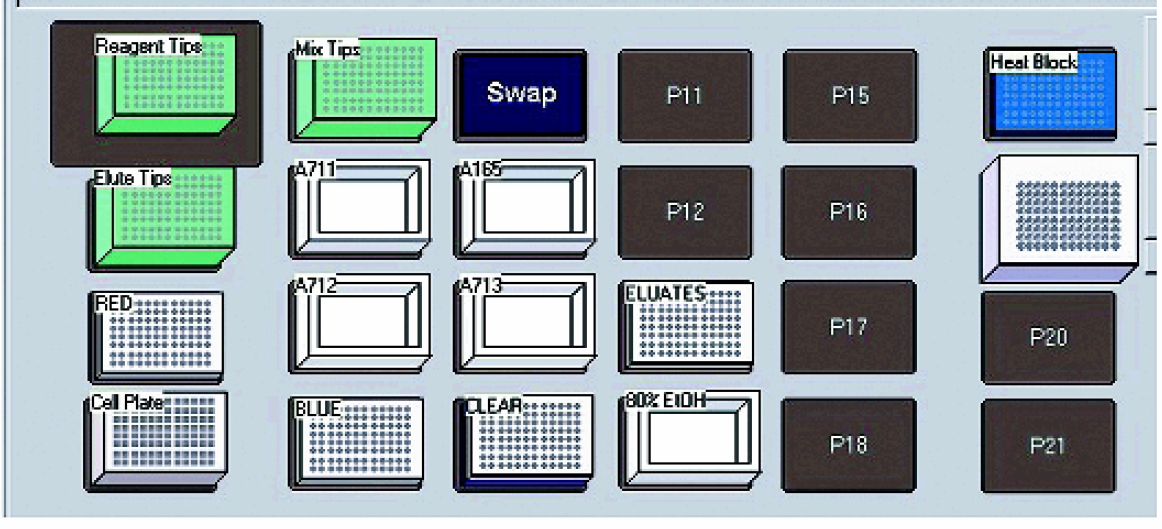
A diagram of the Biomek® FX deck layout.
One Tip Loader ALP (Beckman Coulter part # 719856) A 96-well tip wash ALP (Beckman Coulter part # 719804) Three single position labware ALP (Beckman Coulter part # 719360) One 16 position labware ALP (Beckman Coulter part # 719360) One heating/cooling ALP (Beckman Coulter part # 719361) An Orbital Shaker ALP (Beckman Coulter part # 379410) A Device Controller Unit (Beckman Coulter part # 719366) Three P250 Tip Rack Assembly (Beckman Coulter cat. # 717251)
Additional required equipment included the following:
A circulatory water bath capable of 65°C heating (for heating/cooling ALP) One Promega MagnaBot® 96 Magnetic Separation Device (Cat. # V8151) One Promega Heat Transfer Block (Cat. # Z3271)
Labware used for this method development is listed below:
Greiner 96-well U-bottom microplates (Cat. # 650101) Innovative Microplate reagent reservoirs (Cat. # S30014) Costar Deepwell Culture Plates (Cat. # 3960)
Many types of reservoirs and deep-well culture plates are available; however, the MagnaBot® 96 Separation Device was specifically designed for use with Greiner U-bottom microplates. Optional lids are available for the Innovative Microplate reservoirs (Cat. # A50014).
Cells were harvested by centrifugation, and the supernatants were discarded. The cell pellets were resuspended in cell resuspension solution containing 100μg/ml RNase; then cell lysis solution was added, followed by gentle mixing to lyse the cells. This short incubation also served to digest contaminating RNA, preventing co-purification with the plasmid DNA and subsequent interference with downstream applications. The addition of neutralization solution precipitated both genomic DNA and cell debris from the lysate. MagneSil™ BLUE was first mixed to homogeneity, and then added to each well of the 96-well plate. The magnetic particles bound precipitated genomic DNA and cell debris. The lysate was transferred to a round-bottomed, multi-well plate positioned atop a MagnaBot® 96 Separation Device (Cat. # V8151, Promega Corporation, Madison, WI), and the MagneSil™ bound debris was pulled to the sides of the wells. The cleared lysate containing the solubilized plasmid DNA was transferred to a second round-bottomed plate containing 50μl of MagneSil™ RED. Vigorous mixing allowed selective absorption of the plasmid DNA to the particle surface. A second magnetic separation collected the MagneSil™ particle-bound plasmid DNA, and the spent lysate was withdrawn and discarded. To remove unwanted contaminants (salts, RNA, etc.), the particles were resuspended and washed three times in 80% ethanol. Particles were magnetically separated and the wash solution was discarded after each wash. Following the final ethanol wash, the particles were dried to remove residual ethanol, which might interfere with downstream applications. Finally, the MagneSil™ particles were resuspended in elution buffer, and the plasmid DNA was released. A final magnetic separation removed the MagneSil™ particles, and clear eluates containing the purified plasmid DNA were transferred to a clean collection plate.
The following steps detail the addition of reagents and sample handling:
Resuspend cell pellets in 90μl of cell resuspension solution. Add 120μl of cell lysis solution and gently mix for three minutes. Add 120μl of neutralization solution and vigorously mix for 30 seconds. Mix the MagneSil™ BLUE slurry to homogeneity, then add 25μl to each sample. Mix vigorously for 30 seconds. Transfer the lysates to a 96-well U-bottom plate positioned atop a MagnaBot® 96 Separation Device. Allow 1–2 minutes for separation. Transfer the clear lysate to a second 96-well U-bottom plate containing 50μl per well of MagneSilTM RED. Mix for 15–20 cycles. Move the plate onto the MagnaBot® 96 Separation Device and allow 60 seconds for separation. Remove and discard the spent lysate. Wash by resuspending the particles in 100μl of 80% ethanol. Move the plate onto the MagnaBot® 96 Separation Device and allow 30 seconds for separation. Remove and discard the spent wash solution. Repeat step 8 two additional times. Dry the particles by incubating 10 minutes at room temperature or alternatively with gentle heat for 5 minutes. Resuspend the particle pellets in 100μl Elution Buffer, mixing gently for 2 minutes. Move the plate to the MagnaBot® 96 Separation Device and allow 60 seconds for separation. Transfer eluates containing the purified plasmid DNA to a clean 96-well U-bottom plate.
Purified plasmid DNA was analyzed to gel electrophoresis in 1.2% TBE-buffer agarose gels followed by densitometry analysis using the Alpha Innotech Digital Imaging System (San Leadro, CA). Pure pGEM®-3Zf(+) plasmid DNA (Promega Corporation, Cat. # P2271) was used to produce a concentration standard curve. The yield of purified plasmid DNA was estimated by comparison to the standard curve. Restriction digests of the purified plasmid DNAs were performed using 0.5μg of plasmid DNA in an appropriately buffered 50μl reaction containing 5u/μg of either XbaI (Promega Corporation, Cat. # R418A) or 5u/μg of Sca I/Sph I (Promega Corporation, Cat.# R6211, R6261 respectively) for 60 minutes at 37°C. Samples were analyzed on 1.2% TBE-buffer agarose gels.
Sequencing was performed using the Applied Biosystems BigDye® Terminator Cycle Sequencing Ready Reaction Kit V2.0 using either 2-fold or 4-fold diluted mix. Sequencing reactions were analyzed on an ABI Prism® 3700 DNA Sequencer by a commercial DNA sequencing facility (SeqWright, Houston, TX).
Cross-contamination assays were performed using a PCR 1 assay. 50μl reactions contained pUC/M13 Forward and Reverse primers (Promega Corporation, Cat. # Q5391 and Q5401, respectively) at a final concentration of 0.2μmol and 2X PCR Master Mix (Promega Corporation, Cat. # M7505). A Perkin Elmer PE 9600 Thermocycler® was used for amplification. The thermal cycle profile consisted of: 95°C for 1 minute, 55°C for 1 minute, for 30 cycles. The final extension step consisted of a 7-minute hold at 68°C. Ten microliter aliquots of each reaction were loaded onto 2% TBE-buffered agarose gels for analysis.
Results and Discussion
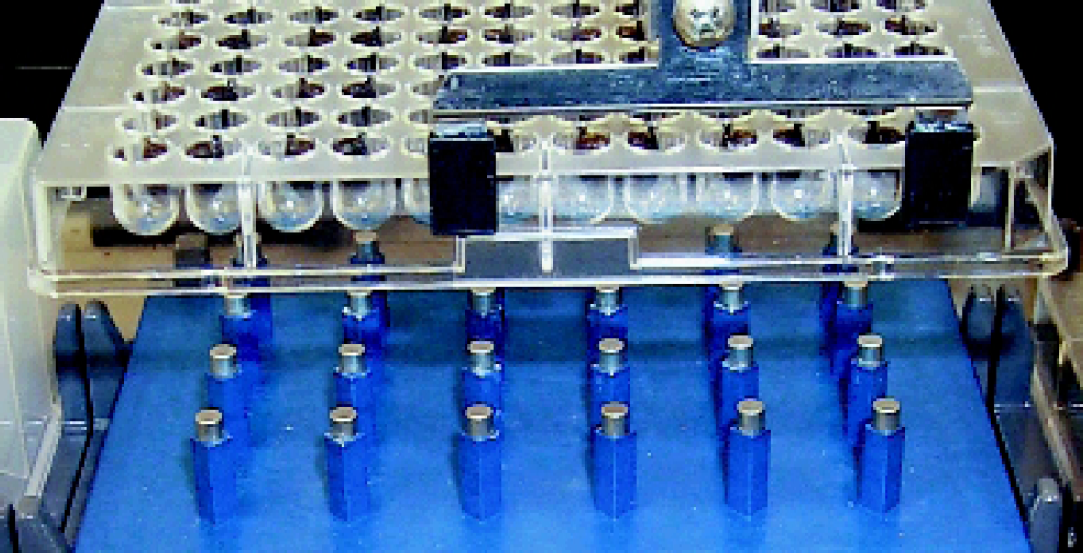
To address high-throughput applications, we developed a custom magnet for 96-well separations. This device, the MagnaBot® 96 Magnetic Separation Device (Cat.# V8151, Promega Corporation, Madison, WI), was designed with a standard SBS footprint to be amenable to a variety of robotic platforms. The MagnaBot® device uses magnets oriented such that each captures particles from four wells, allowing waste material to be easily removed without disturbing the magnetic particles (Figures 2). When used in conjunction with the method described, 96 samples were purified in a total elapsed time of approximately 40 minutes, after the plate is placed on the Biomek® FX.

Plasmid DNA purified with this automated procedure consistently yields high quality sequence data. Sequence analysis was performed on plasmid templates using ABI BigDye® Ready Reaction Mix Version 2 and analyzes with an ABI 3700 DNA Sequencer. The Phred 20 value is shown for 48 samples sequenced with a two-fold or four-fold dilution of the BigDye® Ready Reaction Mix (Figures 3). Read lengths were consistently greater than 600 bases. The Wizard® MagneSil™ Plasmid Purification System is quality control tested to ensure reliable and reproducible sequencing results of greater than 600 bases at a minimum 98% accuracy.
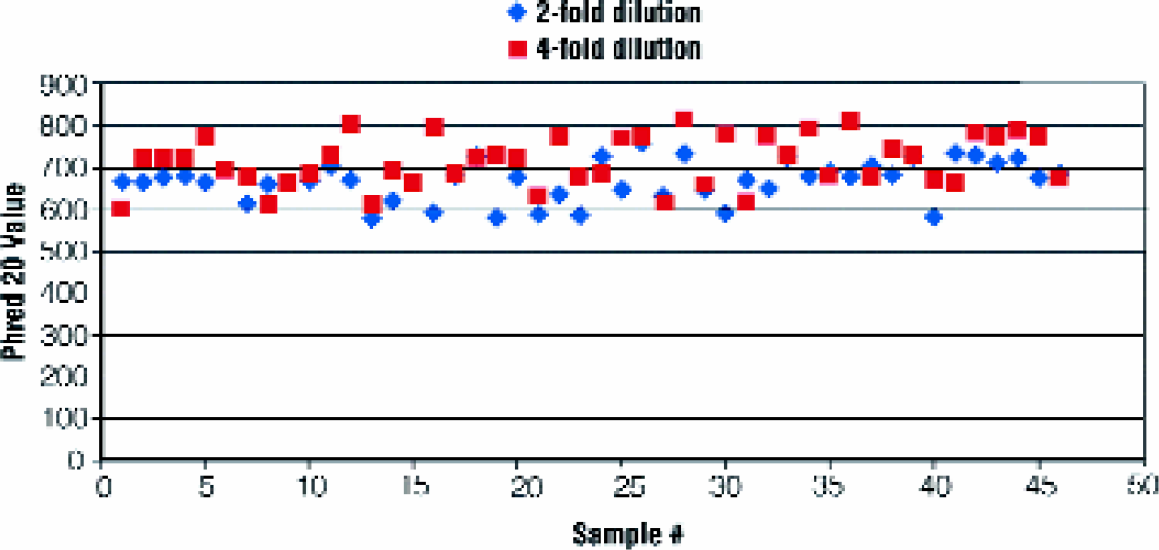
ABI3700 sequencing results. Samples of plasmid DNA purified from a 96-well plate were sequenced using a two-fold or four- fold dilution of the Big Dye® Ready Reaction Mix Version 2. The Phred 20 values were plotted for each sample.
Purified plasmid DNA was subjected to agarose gel electrophoresis to visually assess the quality of the DNA samples (Figures 4). The preparations contain minimal levels of contaminating RNA. A small amount of chromosomal DNA was observed in the plasmid preparations. The low level of contaminating chromosomal DNA does not negatively affect DNA sequence analysis. It is also apparent from visual inspection that the procedure yields a consistent amount of plasmid DNA. This observation was confirmed by densitometry analysis of the gel (Figures 5). In this experiment, the preparations yield approximately 12μg of plasmid pGL3, with minimal variation across the 96-well plate. Plasmid yield was estimated by densitometry, as opposed to absorbance or fluorescence assays, to minimize the potential contribution from contaminating chromosomal DNA.
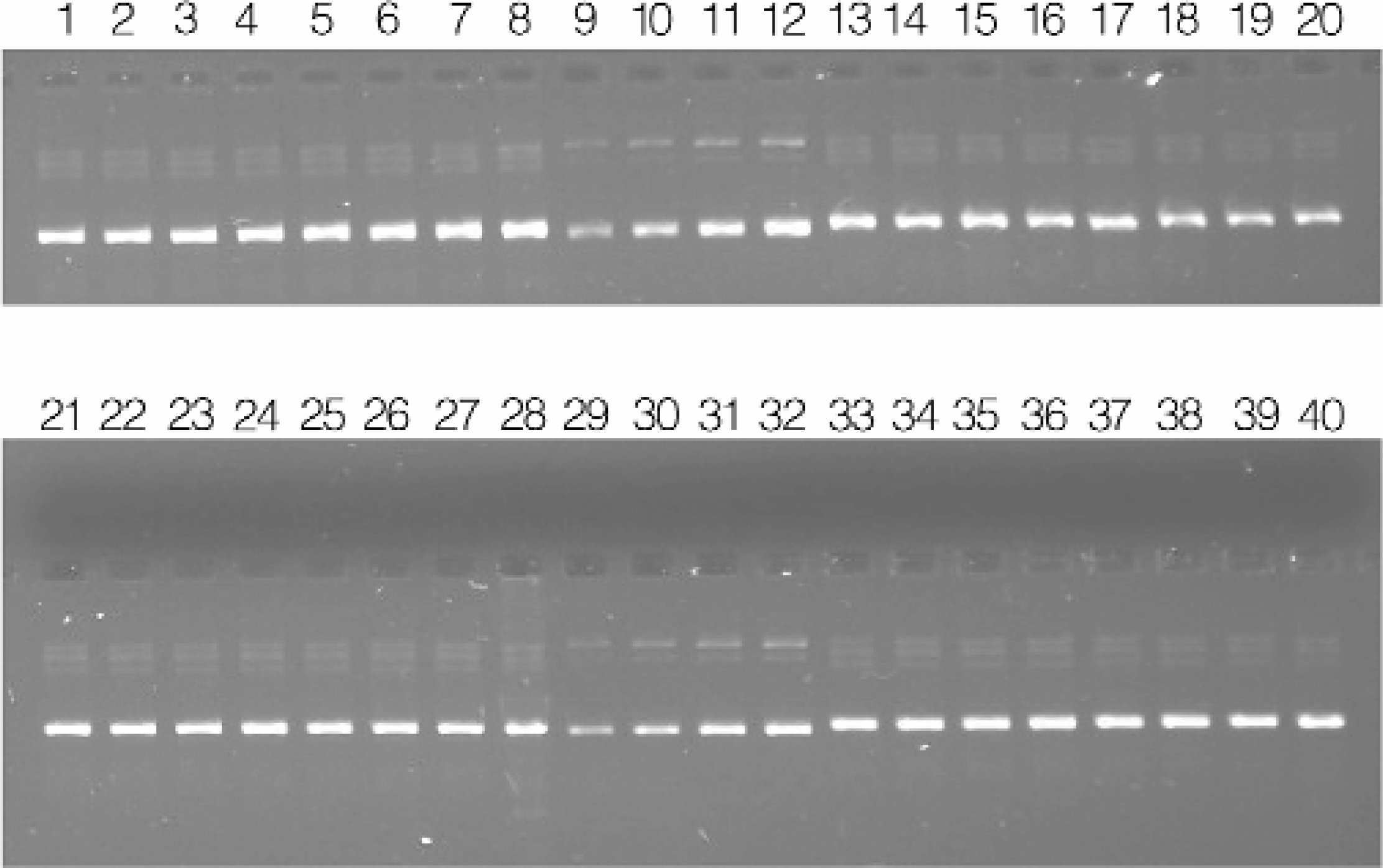
Agarose gel electrophoresis of purified plasmid DNA. Purified plasmid DNA resulting from 96-well cultures of E. coli transformed with pGL3. Lanes 1–8, 13–20, 21–28, and 33–40: 0.5μl of eluate representing wells 1A through 4H. Lanes 9–12 and 29–32: 25, 50, 75, and 100ng, respectively of pGL3 standard.
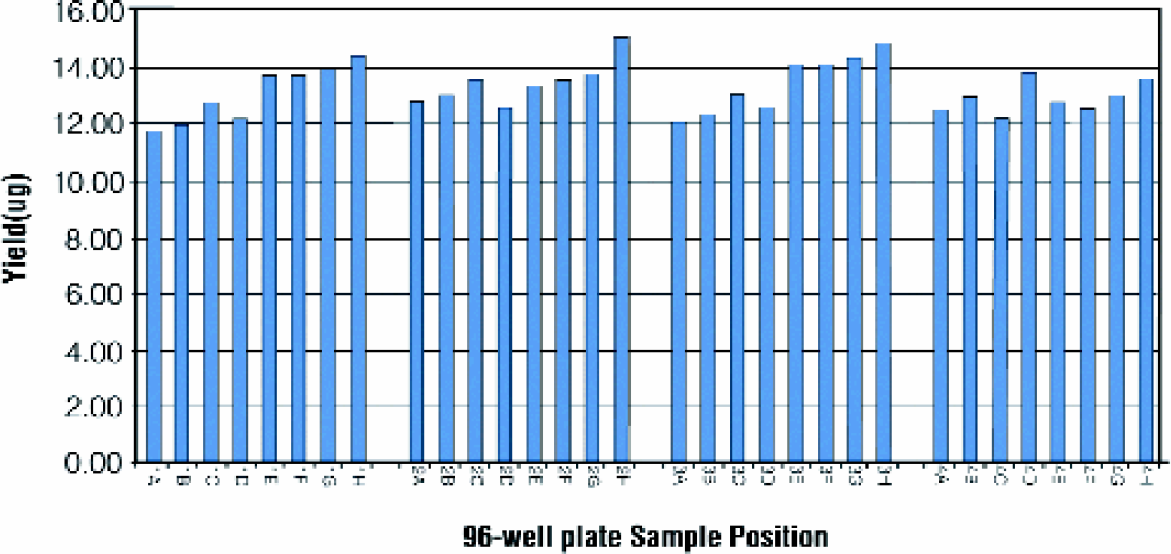
Plasmid DNA yield. Yield estimated by gel densitometry of the 32 samples separated on the agarose gel shown in Figures 4.
DNA purified with this automated procedure is intended for use as a template for DNA sequence analysis. Nonetheless, we tested the efficacy of the purified plasmid DNA in other applications (restriction digestion and PCR amplification) to further assess the quality of the DNA preparation. Purified plasmid DNA was subjected to restriction digestion with Xba1, or double digestion with a mixture of Sca1 and Sph1 (Figures 6). Complete digestion was observed for all the preparations.
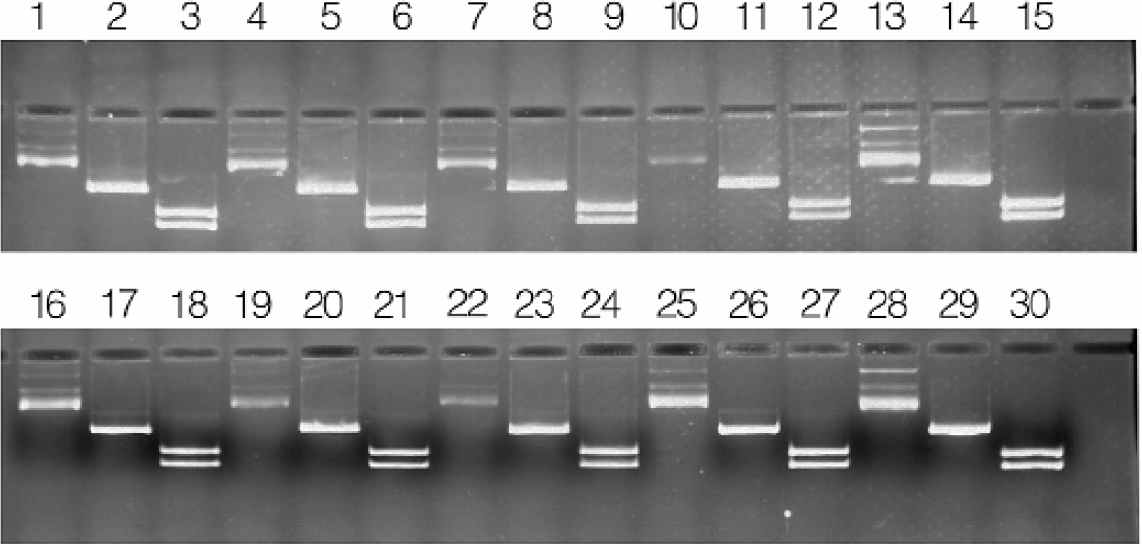
Restriction digestion of MagneSil™ purified plasmid DNA. Purified pGEM®−3Zf(+) plasmid DNA was digested with Xba I, or double-digested with Sca I and Sph I. Lanes 1–3, 4–6, 7–9, 10–12, 16–18, 19–21, 22–24, and 25–27: uncut, Xba I, and Sca I/Sph I, respectively, representing eight randomly selected preps. Lanes 13–15, and 28–30:100ng pGEM∀ −3Zf(+) control DNA, Xba I-digested, and Sca I/Sph I-digested plasmid DNA, respectively.
MagneSil™ purified pGEM®-3Zf(+) plasmid DNA was serially diluted and aliquots were tested in a PCR amplification reaction (Figures 7). We observed efficient amplification of the multiple cloning site for all dilutions tested. Consequently, we concluded there were no significant inhibitors of Taq DNA polymerase in the purified plasmid DNA samples.
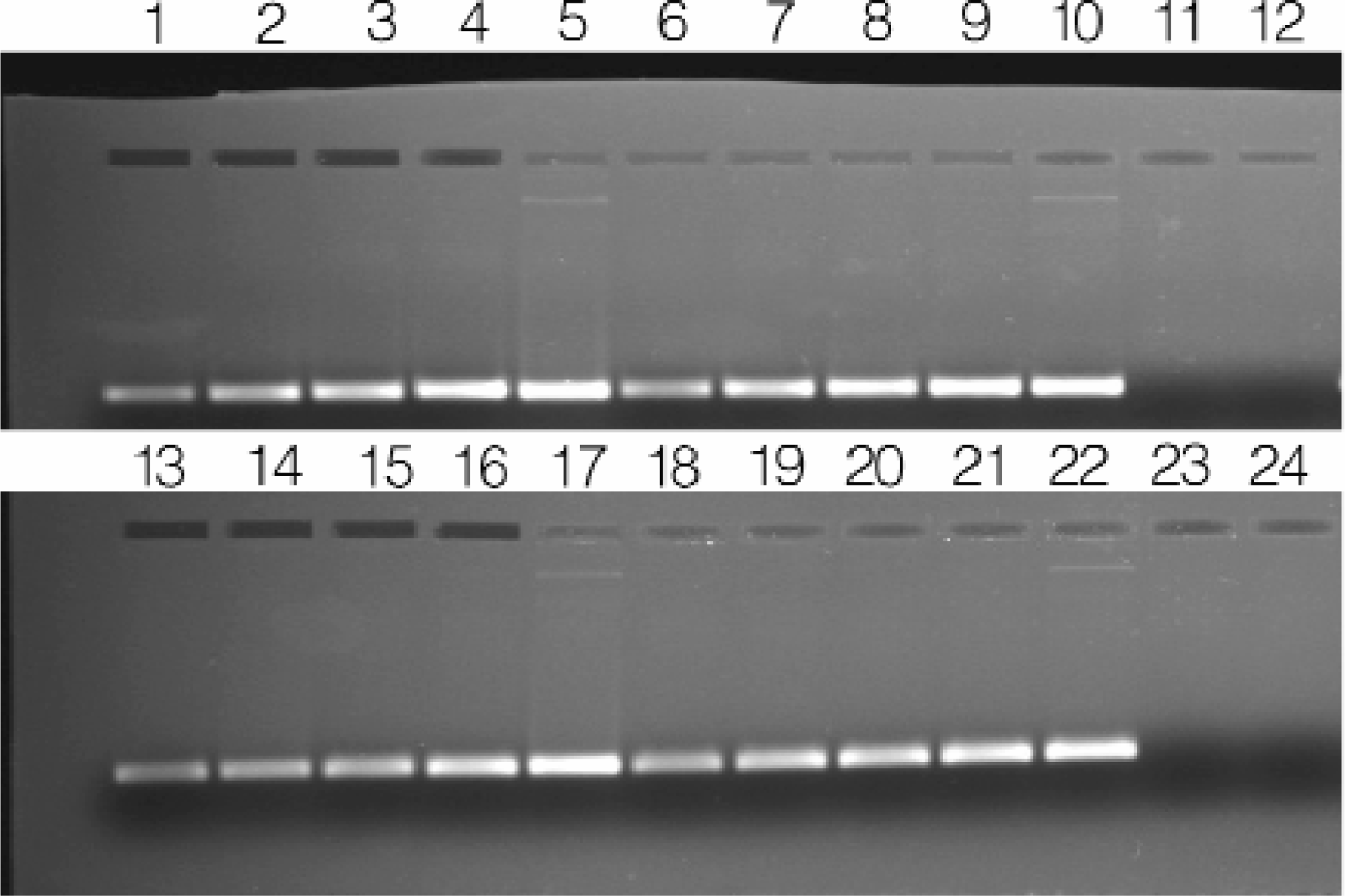
PCR amplification of the purified plasmid DNA samples. PCR amplification was performed on four randomly selected MagneSil™ particle purified pGEM®-3Zf(+) samples following serial dilution. Primers were M13/pUC forward and reverse 17-mers. Using this strategy, a 181 base pair fragment results from amplification. Lanes 1–5, 6–10, 13–17, and 18–22: 3pg, 30pg, 300pg, 3ng and 30ng of template, respectively. Lanes 11, 12, 23 and 24: No template controls.
We confirmed that the robotic method does not introduce significant cross contamination between the wells. Alternating wells of a 96-well plate were inoculated with E. coli transformed with pGEM®-3Zf(+). After overnight incubation, the plate was processed as described in the Materials and Methods section. Aliquots from each well (1A through 4H) were separated on an agarose gel. No contaminating plasmid or chromosomal DNA was observed in the uninoculated wells (Figures 8).
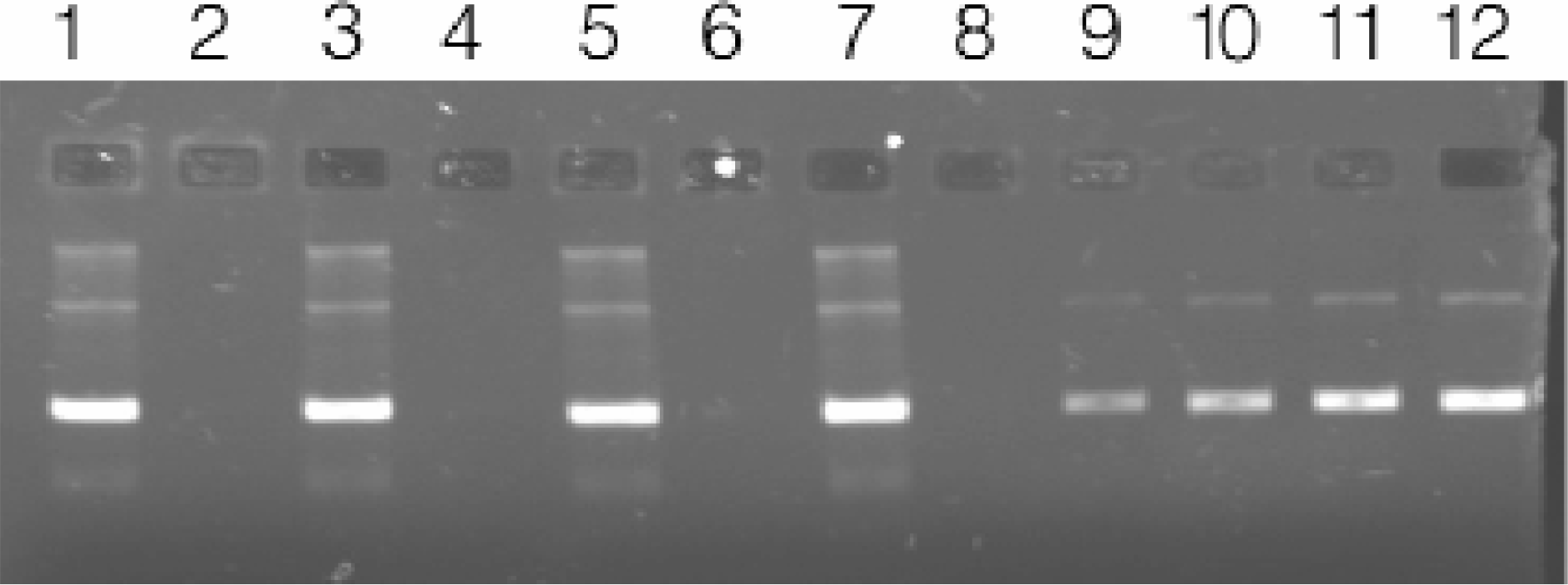
Testing for the presence of cross-contamination. Plasmid DNA purified by automated methods from 96-well cultures of E. coli DH5α transformed with pGEM∀ −3Zf(+); alternating wells inoculated with bacteria (Average yield = 5.24μg, CV=1.3%). Lanes 1–8: 1μl of eluate representing wells 1A through 1H of a 96-well purification. Lanes 9–12: 25, 50, 75, and 100ng, respectively of pGEM®-3Zf(+) standard.
Conclusion
The Wizard® MagneSil™ Plasmid Purification System has been adapted for automated plasmid DNA purification. Robotic methods have been developed for a number of automated platforms including the Biomek® FX workstation. These methods allow for simultaneous purification of plasmid DNA a 96-well format in less than 40 minutes. Plasmid DNA resulting from these purifications performs well in a multitude of downstream applications including restriction digestion, fluorescent sequencing, and PCR. Plasmid quality is reflected in sequencing results showing consistently high Phred 20 scores and PCR amplification from as little as 3 picograms of purified plasmid. The system is QC tested to ensure reliable and reproducible sequencing results of greater than 600 bases with a minimum 98% accuracy. Read lengths routinely are in the 700–750 base range.
