Abstract
We describe automated methods for purifying genomic DNA from plant tissue. The Wizard Magnetic 96 Plant System uses MagneSil paramagnetic particles (PMPs) in a 96-well format to purify sufficient DNA for polymerase chain reaction (PCR)-based plant genotyping. The system can be scaled up for high yield to provide genomic DNA for large numbers of PCR-based tests or archiving. In contrast, the MagneSil ONE Fixed Yield System uses MagneSil ONE PMPs to purify a fixed amount of ultraclean DNA for high sensitivity in Third Wave Technology Invader single nucleotide polymorphism (SNP) Genotyping assays. These methods may be implemented on Beckman Coulter Biomek automated workstations.
Introduction
Selection of the most appropriate DNA purification methods for industrial plant molecular biology research requires an understanding of the intended downstream applications for the DNA. The goal may be large-scale plant genomic research, seed quality control, or marker-assisted breeding. Increasingly, automation is being considered to improve throughput using limited resources. Purified DNA may be used in marker-assisted breeding, SNP (single nucleotide polymorphism) detection, and population studies involving the interaction of an array of gene variants. All of these methods require reliable DNA purification. Automation simplifies purification and improves consistency of yield and purity of DNA to be used in applications such as amplified fragment length polymorphism (AFLP), 1 random amplification of polymorphic DNA (RAPD), 2 inter simple sequence repeat (ISSR), 3 polymerase chain reaction (PCR), 4 as well as tests based on Third Wave Technology's Invader platform. 5 For small-scale studies, manual, moderate, or low-throughput robotic methods may be more practical. What is needed overall is a flexible purification system (robotic instrument and chemistry) that can be adapted to low throughput or high throughput, small and scaled-up sample size, and provide DNA suitable for downstream applications.
Traditionally, plant scientists have used home brew methods such as CTAB (hexadecyltrimethylammonium bromide) 6 or modified Dellaporta 7 extractions. These protocols are labor- and timeintensive, and often involve organic solvents such as chloroform. There are multiple incubations, and centrifugation steps as well. A common problem with these methods is that they work well with some sample types but cannot be used reliably with others. More importantly, they are not easily amenable to automation or high-throughput purification.
Promega has developed the Wizard Magnetic 96 DNA Plant System (“Wizard Plant”) for plant DNA isolation from a single leaf punch or seed, and from scaled-up sample sizes. In addition, the MagneSil ONE Fixed Yield System (“MagneSil ONE”) is available for the isolation of high-quality DNA with a fixed yield from a given sample type where normalized amounts of DNA can streamline applications that would otherwise require post purification DNA quantitation and normalization. For high throughput, 96-well robotic methods have been developed on the Biomek 2000 and Biomek FX (Beckman Coulter, Fullerton, CA) instruments and can be adapted to other liquid handling instruments. Medium throughput batches of up to 15 samples can be adapted to lower cost and lower throughput systems such as the Thermo Electron LabSystems KingFisher (Thermo Electron, Waltham, MA). For small sample batches, protocols may also be performed manually in 96-well plates. Methods are available from www.promega.com. In this paper, we present data generated using DNA isolated by these 96-well robotic methods.
DNA purification using the Promega MagneSil paramagnetic particles (“PMPs”) has been performed with blood samples; 8 the plant tissue protocols described here use similar technology to lyse and purify DNA. The faster binding kinetics, coupled with the high surface area available with MagneSil PMPs, provide an advantage over column-based 96-well methods. The particles maintain their robust physical integrity in a wide variety of chemical environments and rapidly respond to magnetic fields. Promega scientists and customers have focused their efforts on DNA purification from “high-value” agricultural crops such as corn, soy, cotton, tomato, canola, and other vegetables and seeds. A typical sample size for the Wizard Plant System is one 6-8 mm leaf punch or 1-5 seeds. We have also developed a scaled-up 96-well protocol for customers who require larger amounts of DNA for large-scale marker analysis or for archiving. 9 The Wizard Plant System offers scalability and circumvents the limitations of extensive centrifugation or vacuum filtration methods.
One of the variables in the purification protocol is the efficiency of the sample-grinding step. Whether using a SPEX CertiPrep Geno/Grinder 2000 (Metuchen, NJ), a Retsch Mixer Mill 300 (Haan, Germany), or a homemade grinding apparatus, variations in the amount of sample and the nature of the metal ball being shaken with the plant sample increases variability in the lysis procedure. Some samples may be lysed to a greater or lesser extent than a similar sample in an adjacent well. The results described here were obtained using the SPEX Geno/Grinder. Leaf punches were dispensed into deep (1.2 ml) 96-well plates. One stainless steel grinding ball was added to all the wells simultaneously, using a 96-well plate dispenser available from SPEX. Lysis buffer was added with a multichannel pipettor, and the plate was sealed with aluminum tape. The plate was placed on the grinder, which rapidly and vertically shakes the plate and homogenizes the plant tissue. Times and speeds may be adjusted for each individual plant type. For very fibrous tissue, two balls may be needed. After grinding, the plate is centrifuged briefly to pellet any plant debris and excess liquid from the sealer, avoiding cross-contamination with removal. The plate is then ready to be placed on the robot for automated processing. The whole process may be completed within 60 to 90 min for most samples.
For many applications, there is considerable tolerance for a wide range of DNA concentrations without variability in the final results. However, for those applications where it is desirable to isolate DNA in a more narrow concentration range, Promega offers an alternative: MagneSil ONE Fixed Yield System, based on patented Promega technology. 10 Sample concentrations will fall into a range of +/− 50% of the average value as measured by PicoGreen (Invitrogen, Carlsbad, CA).
The MagneSil ONE System is an intelligent way to normalize sample-to-sample variability. The key feature of this method is the limiting amount of PMPs that bind a fixed amount of DNA. In the standard protocol, the amount of particles will only bind 200 ng of DNA from a sample, although the lysis procedure is designed to liberate greater amounts of DNA (e.g., 500 ng). The excess DNA is not bound to the particles and is removed in downstream steps in the protocol. With this system, all samples yield this predetermined amount of DNA. The predetermined amount of DNA can be raised or lowered by raising or lowering the amount of PMPs added per prep. The final concentration of DNA can be adjusted by raising or lowering the elution volume.
Materials and Methods
Automated protocols for DNA purification were performed using robotic methods developed for the Biomek 2000, Biomek FX, and Thermo Electron KingFisher instruments. Please contact Promega for more information about these methods, or visit our website www.promega.com/automethods. Manual DNA purifications were performed as described in the Wizard Magnetic 96 DNA Plant System technical bulletin 11 and the MagneSil ONE, Fixed Yield System technical bulletin. 12
Figure 1 outlines the DNA purification procedure using the Wizard Plant System. The system comprises two sequentially added lysis buffers, an optimized wash solution containing ethanol and isopropanol, and MagneSil PMPs. DNA can be eluted in nuclease-free water or Tris-EDTA (TE) buffer. After grinding and centrifugation, all steps can be performed robotically in standard 96-well plates, or for the scaled-up methods, deep well (1.2 ml) 96-well plates.
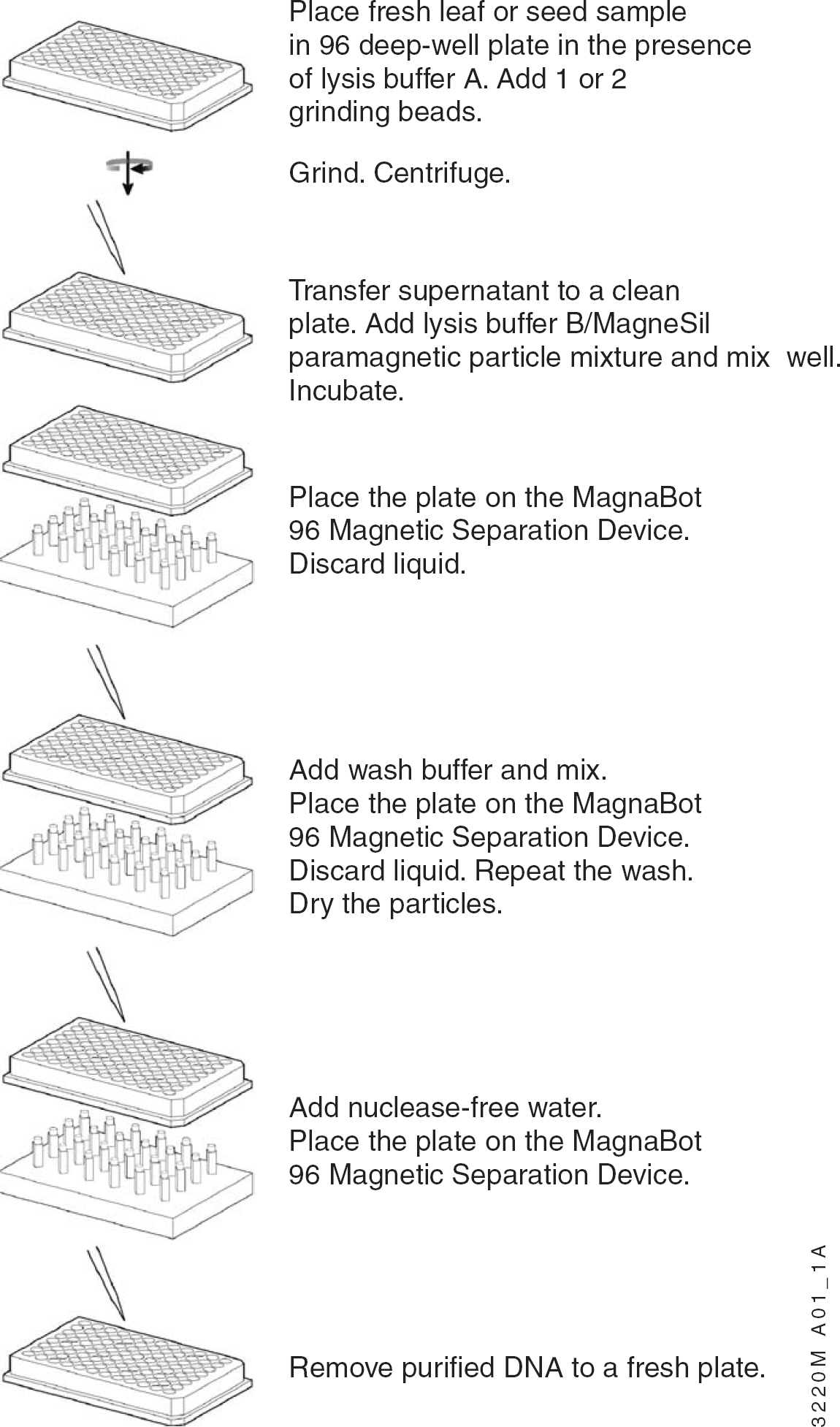
Schematic of DNA isolation using the Wizard Plant System.
The MagneSil ONE System is similar to the Wizard Plant System but uses only one lysis buffer that contains both guanidinium salts and detergent and uses lower-porosity MagneSil Fixed Yield PMPs, limiting the binding of DNA and decreasing the amount of contaminants that compete for binding. Reliably defined yields are obtained and may eliminate the need for DNA quantitation before downstream applications, providing obvious advantages for the highthroughput plant genomic laboratory testing multiple samples of the same plant species.
Figure 2 shows a schematic to assist the scientist in choosing the method that will meet his or her needs for throughput, quality, and quantity of DNA.
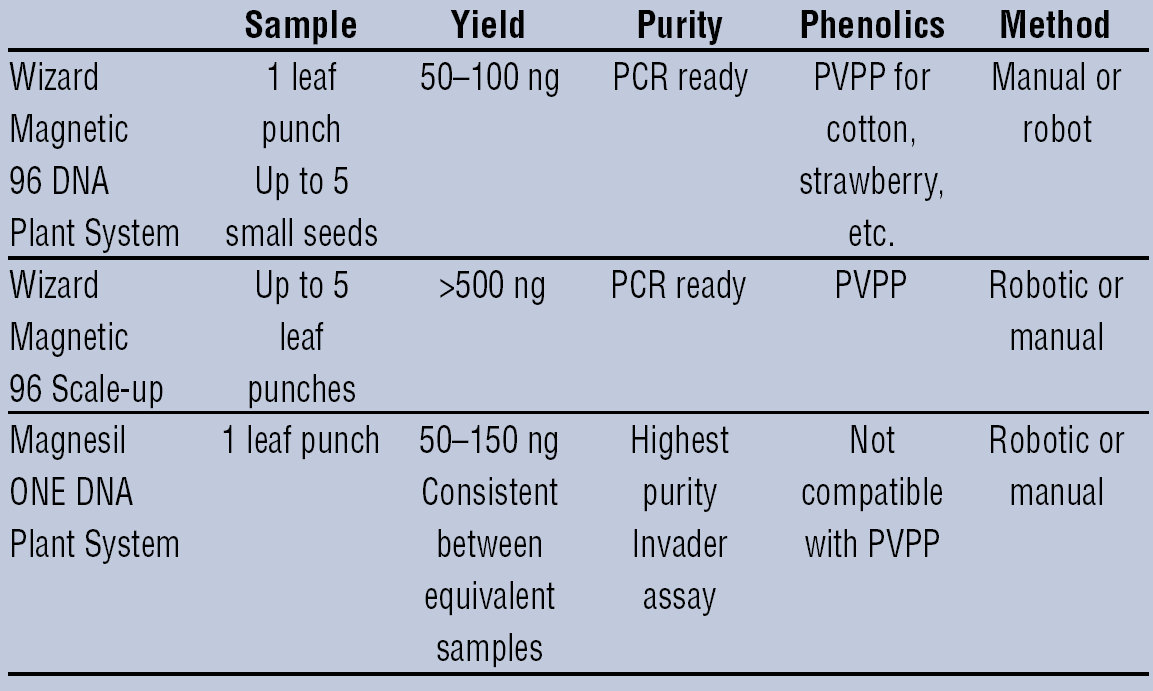
Method selection for plant DNA isolation.
A key element in this process is the grinding of plant samples in a 96-well format. Plant tissue may be fresh, frozen in liquid nitrogen, or lyophilized. All of these samples work well with both the Wizard Plant System and the MagneSil ONE System; however, data presented here was generated with fresh or frozen plant material ground at room temperature using a SPEX CertiPrep Model 2000 Geno/Grinder. A Retsch Mixer Mill provides comparable results under similar conditions. DNA from a wide variety of different plants has been purified using the Wizard Plant System. A number of these plants are described in Figure 3. For the scaled-up method, complete homogenization of the plant material and centrifugation to separate the debris from the lysate are critical. Suspended plant material may easily clog pipette tips and reduce yields. The aspiration height of the liquid handling device must be optimized for a given sample type so that lysate is aspirated above the plant debris level and transferred to a clean plate for the purification protocol.
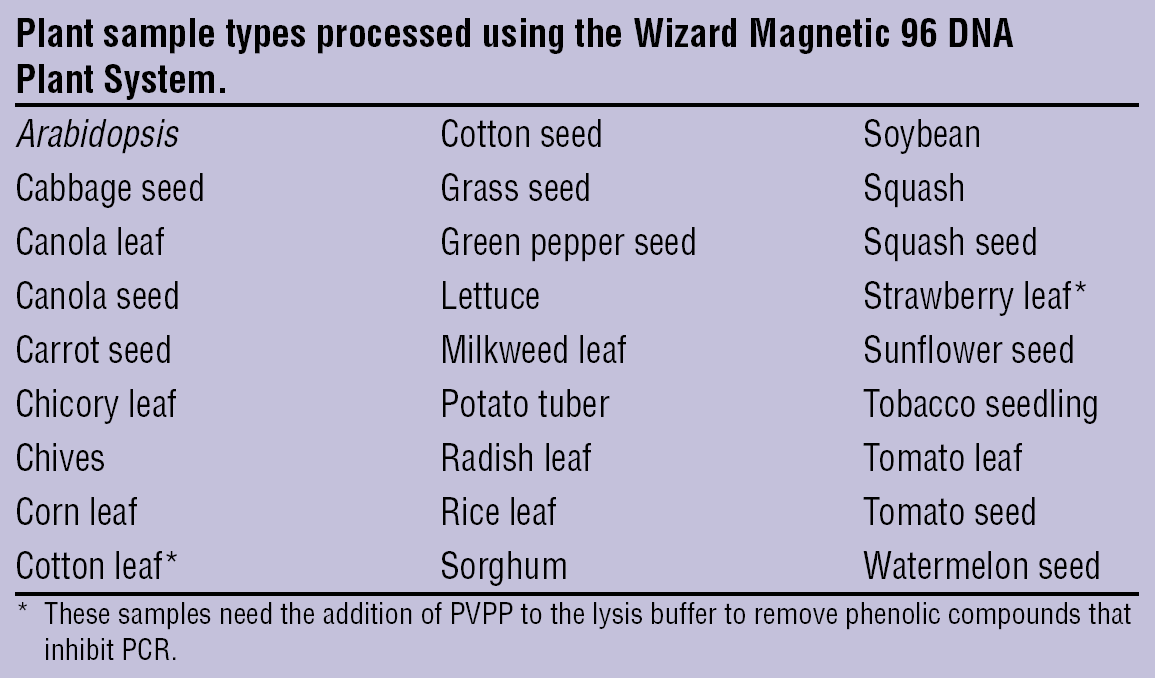

Plant sample varieties tested.
Results
Figure 4 shows DNA isolated from a variety of plants using the Wizard Plant System, and the automated method for the Biomek 2000 Laboratory Automation Workstation. Consistent recovery of high molecular weight DNA is evident in the gel photo.
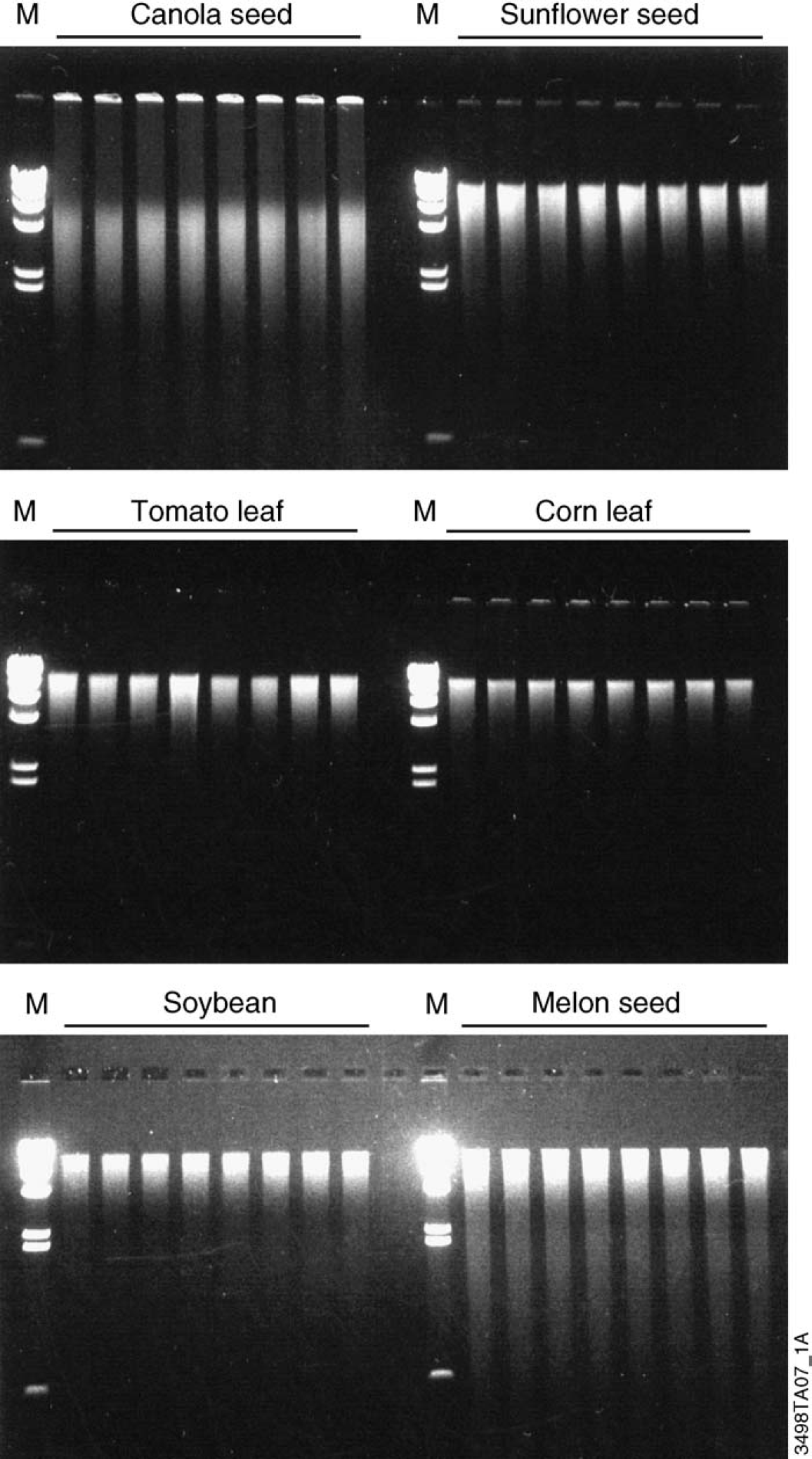
DNA isolated from different plant species using the Wizard Plant System. DNA was isolated using the automated protocol available for the Biomek 2000 instrument. The starting material for each lane was either five canola seeds, one sunflower seed, eight tomato leaf punches (8 mm in diameter), eight corn leaf punches (8 mm in diameter), 20 mg of soybean, or one watermelon seed. Samples were eluted in 50 μl nuclease-free water, and 15 μl of eluate was run on a Latitude (Cambrex Corporation, East Rutherford, NJ) 1% agarose Tris-Borate-EDTA buffer (TBE) gel stained with ethidium bromide. Eight samples were shown for each plant tissue to indicate reproducibility of the method. DNA from the canola seed appears partially sheared due to milling. Lanes M, Lambda DNA/Hind III Markers (Promega Cat.# G1711).
Figure 5 lists the plant sample types and yields obtained in our laboratory using the Wizard Plant System with the scaled-up method. To allow more plant lysate in the purification process, the method must be carried out in 96-well deep well plates. We have developed a deep well 96-well plate method for the Biomek FX instrument. 13 Figure 6 shows typical DNA yields from different plants using the Wizard Plant System. DNA was quantitated using PicoGreen fluorescence.
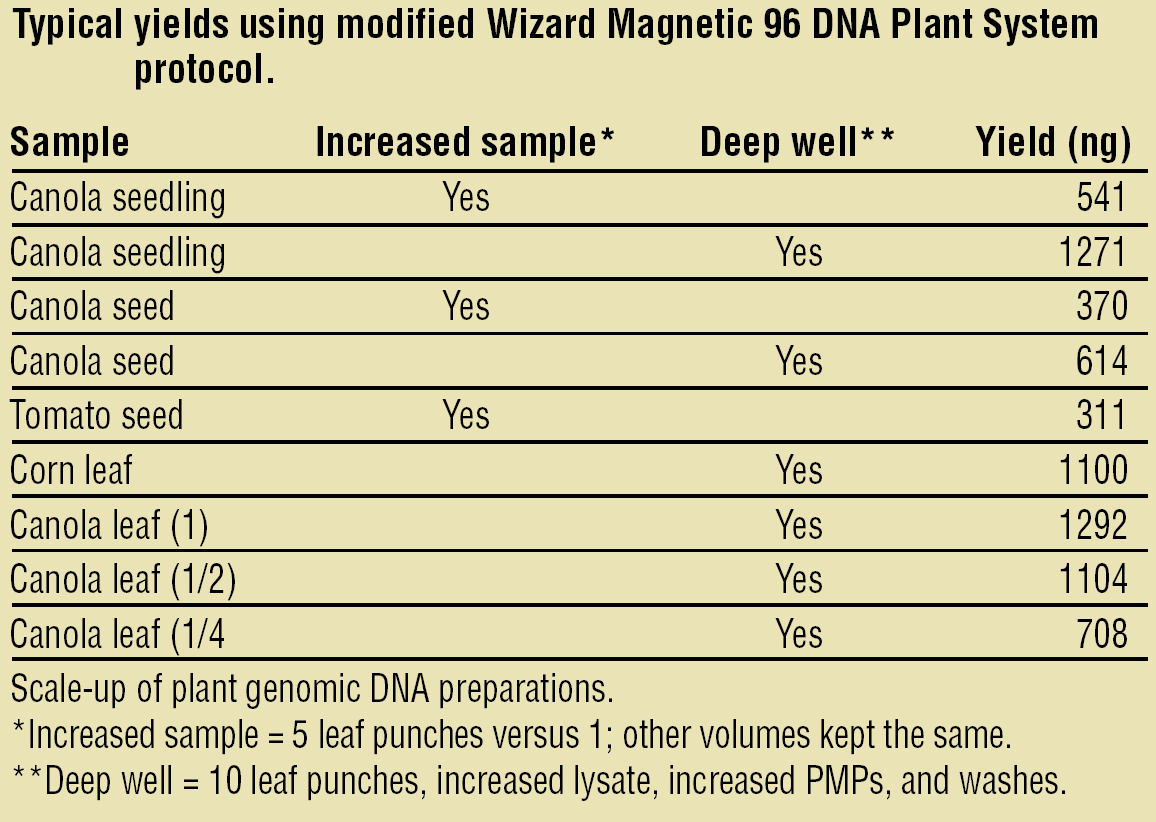

Sample DNA yields.
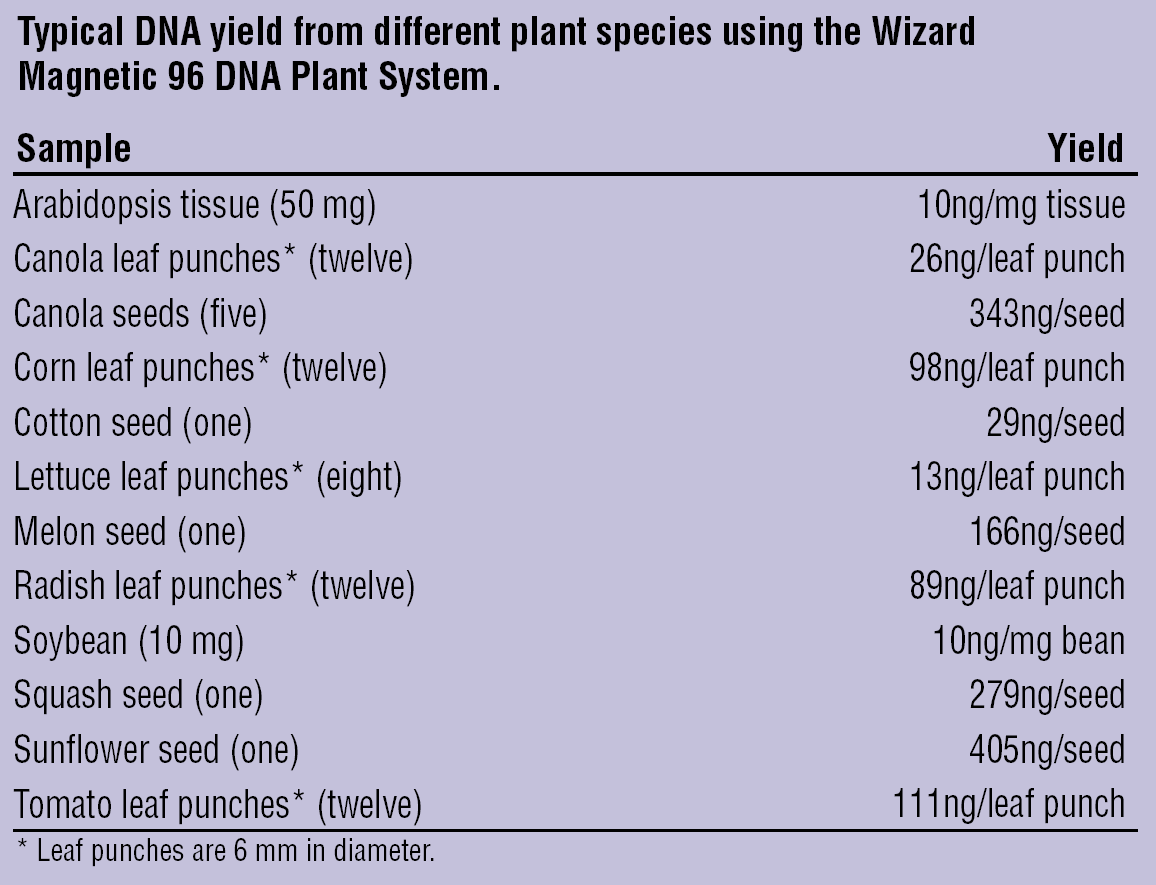

Plant species DNA yield.
This DNA can be used for many downstream applications. Figure 7 shows amplification of an intron from the TrnL chloroplast gene 14 using DNA samples isolated from a number of plant DNA samples as described in Figure 1. DNA yield from a single leaf punch is generally sufficient to perform at least 100 standard monoplex PCR amplifications. All samples showed strong amplification of the TrnL sequence. The size differences for the amplification products are due to polymorphism within these loci between plant species.
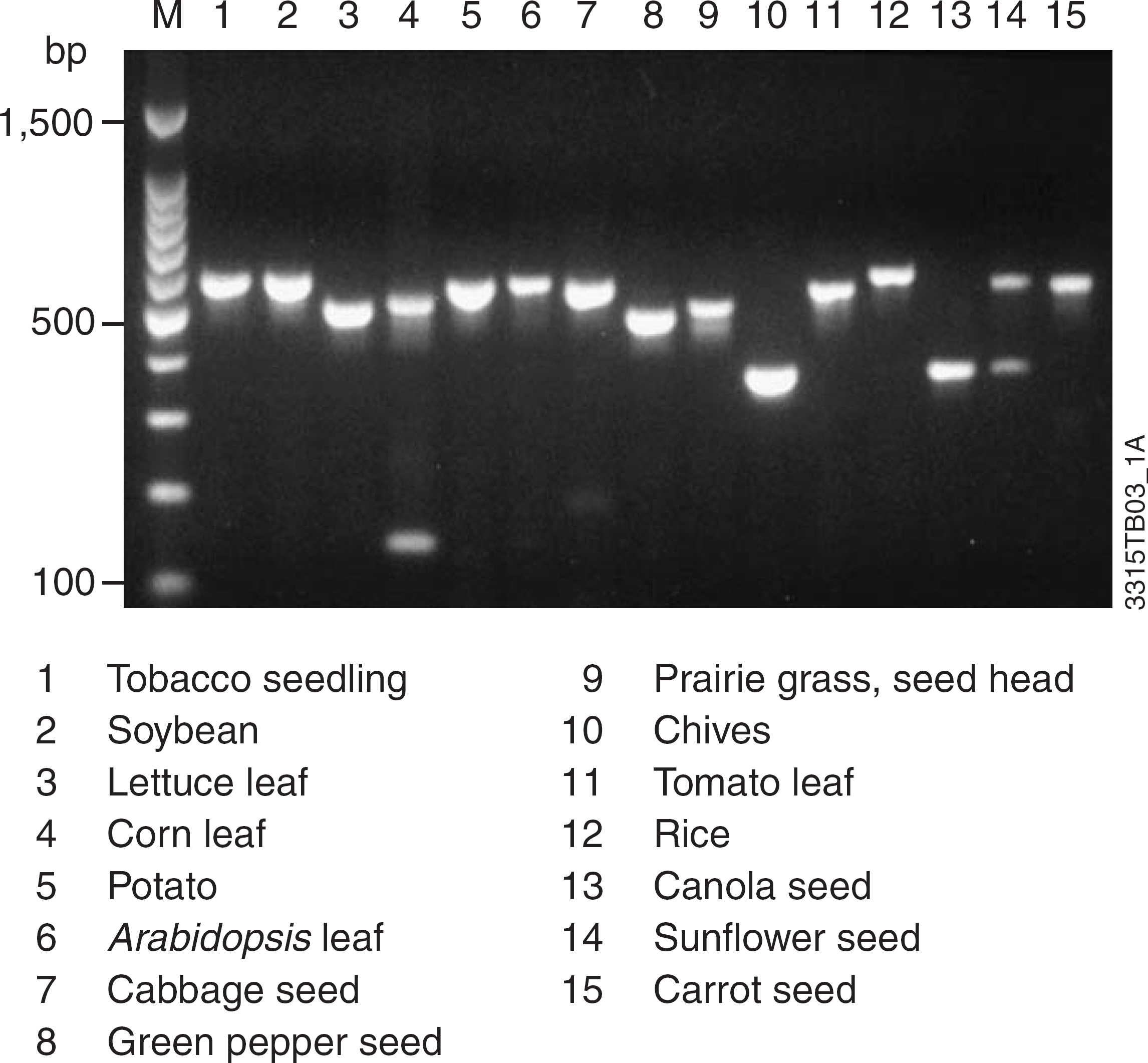
DNA was isolated from the indicated tissues using the Wizard Plant System. An aliquot (1 μl) of the eluted DNA was used to amplify the intron of the TrnL chloroplast gene using a universal primer pair. One or more bands are produced depending on the plant species.
A second confirmation of the quality of purified DNA is RAPD analysis (GE Healthcare/Amersham, Ready to Go RAPD Analysis Beads Cat. #27-9502-01; Piscataway, NJ). RAPD amplification was performed on purified DNA from several plants, and the products were electrophoresed on an ethidium bromide stained gel (Fig. 8). All amplified products showed the expected molecular weight bands.
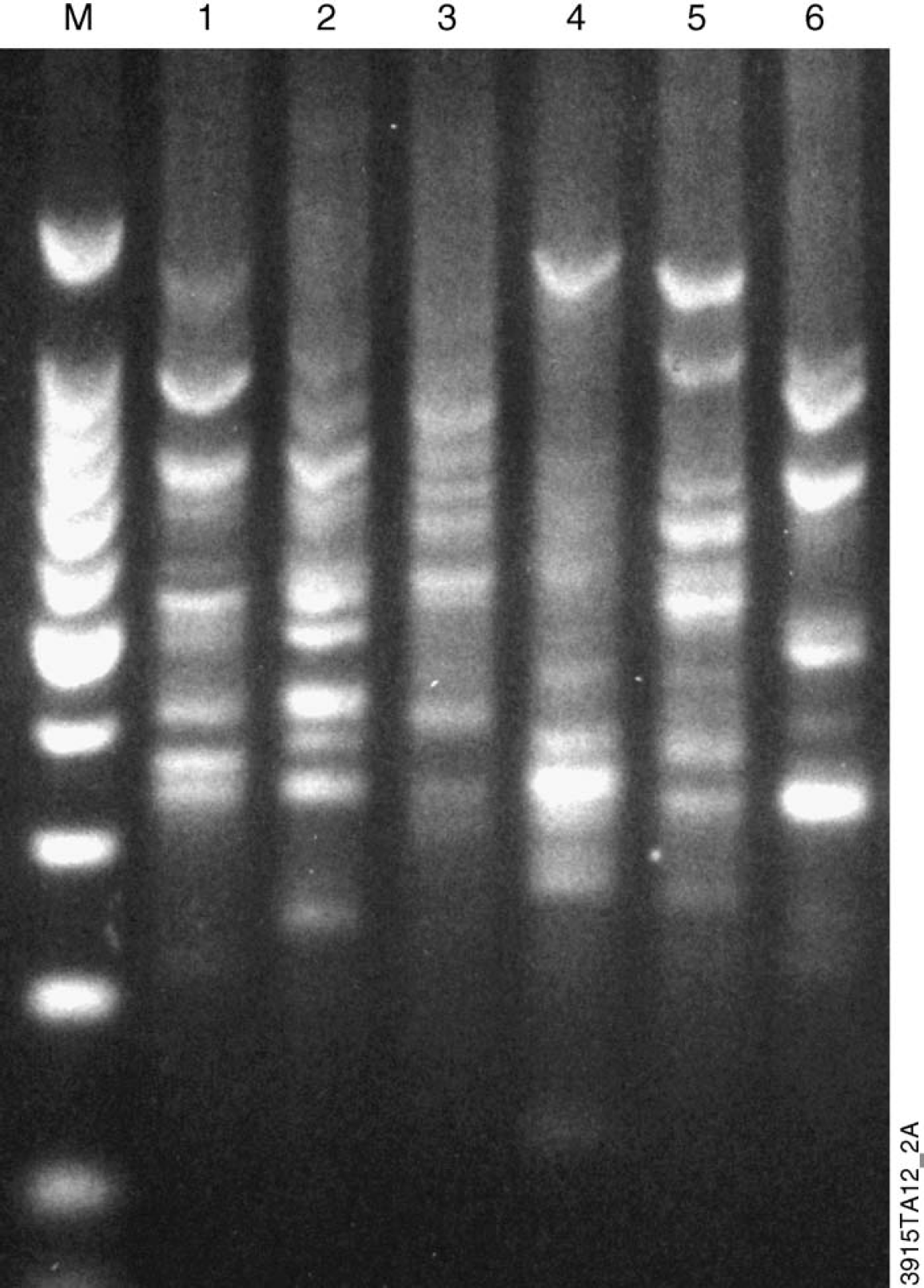
Multilocus PCR amplification of random sequences using DNA isolated from canola leaf tissue using the deep well scaled-up method and the Wizard Plant System. Samples were analyzed using six different RAPD primers from the Ready-to-Go RAPD Analysis Kit. One microliter of a 1:10 dilution of the DNA was used as a template. M = 100 bp DNA Ladder (Promega Cat.# G2101). Lanes 1-6: Canola leaf DNA amplified with RAPD primers 1-6, respectively.
To further illustrate the importance of high-quality DNA for amplification, we analyzed DNA purified from cotton leaf. Cotton leaf contains large amounts of secondary metabolites such as polyphenolics that irreversibly bind to DNA during purification. This can lower yield, as well as inhibit downstream analysis of purified DNA. 15 Polyvinyl-polypyrrolidone (PVPP, Cat # P6755, Sigma, St. Louis, MO) reduces the interaction of phenolic compounds with DNA that occurs during the initial crushing or grinding steps of purification by forming insoluble PVPP-phenolic complexes, allowing the DNA to remain free in solution. 16
Figure 9 shows RAPD analysis using DNA isolated from cotton leaf with the Wizard Plant System. PVPP was added to half of the sample wells prior to tissue grinding. Cotton leaf DNA purified without PVPP showed limited amplification. Inhibition of the RAPD reaction increased as increasing amounts of DNA sample were used, suggesting the presence of inhibitors in the purified DNA. In contrast, the addition of PVPP resulted in strong amplification. The use of PVPP with the Wizard Plant System is a simple, cost-effective method that can be used in an automated system or manually to isolate high-quality DNA from tissue containing high levels of phenolic compounds.
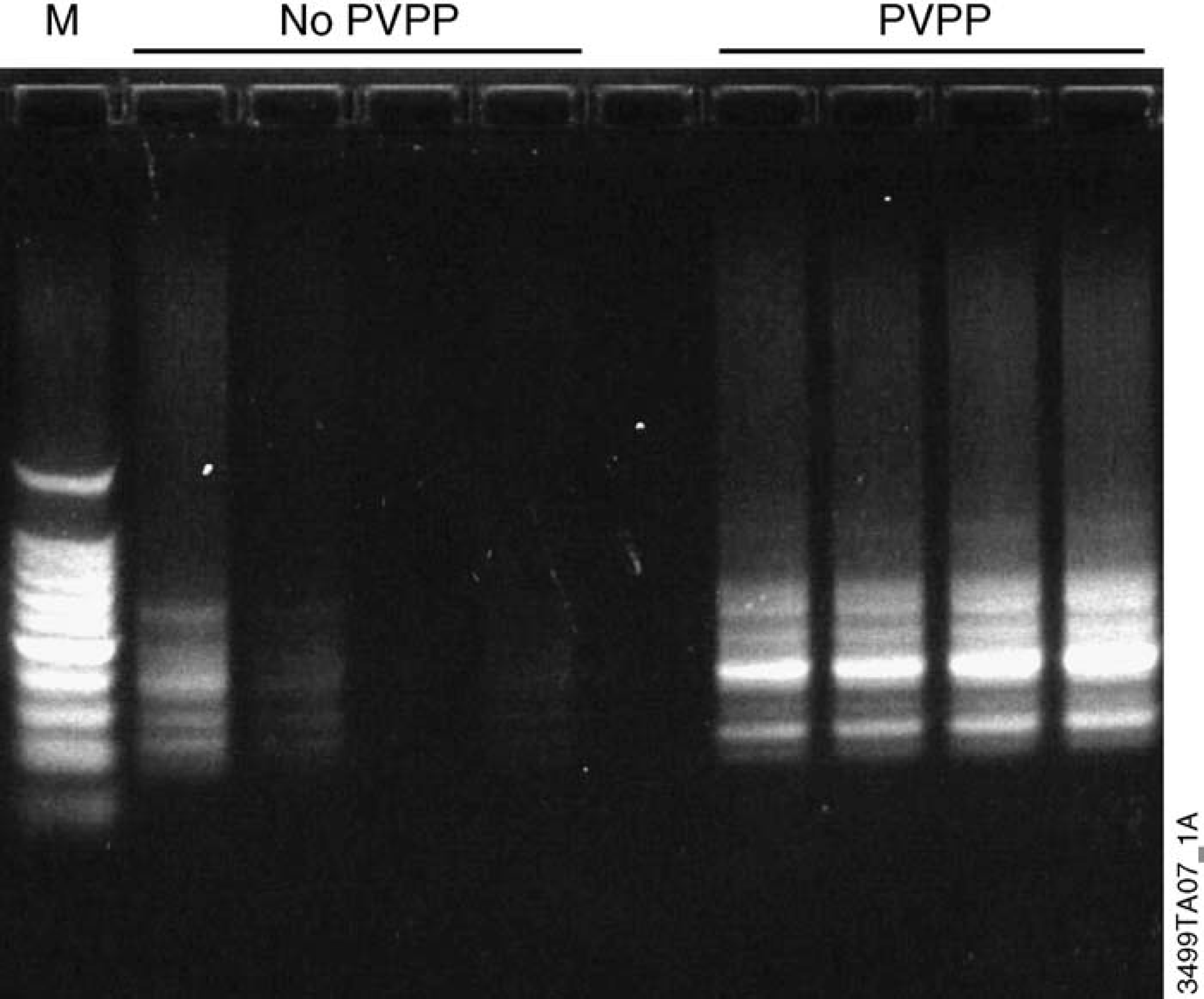
RAPD analysis of cotton leaf DNA isolated using the Wizard Plant System in the presence or absence of PVPP. A cotton leaf was homogenized using a SPEX CertiPrep Geno/Grinder mill in the presence or absence of 10 mg PVPP/300 μl lysis buffer A. DNA was isolated manually from each leaf as described. RAPD analysis was performed on cotton leaf DNA using the Ready-To-Go RAPD Analysis Kit. Samples were diluted 1:10 in water, and I, 2, 3 or 4 μl were used in a RAPD reaction with primer #3. Five microliters of each sample were analyzed on a 1.2% E-Gel agarose gel (Invitrogen).
The Thermo Electron KingFisher Instrument is a medium-throughput instrument that reduces liquid handling and liquid-transfer steps between plates. Purification is straightforward: lysed samples, particles, and reagents are placed in a multi-well plate that is contoured at the bottom to eliminate crevices in which materials and particles can collect. To mix the MagneSil PMPs, the mechanical arm raises and lowers a row of plastic sleeves covering a magnetic probe, like a plunger, mixing the samples and PMPs. The sleeves bind and release the PMPs by applying and removing a magnetic field, respectively. This allows the PMPs to be mixed and transferred from well-to-well for washing and elution. Up to 24 samples may be processed per instrument run, 12 samples per plate and two plates per run. The standard run takes approximately 15 min.
Swaminanthan Applied Genomics, India has used the KingFisher instrument to isolate DNA from five crops: sorghum, pearl millet, chickpea, groundnut, and pigeon pea. 17 The quality of the extracted DNA was assessed on the agarose gel shown in Figure 10 and the DNA used to perform sequence-tagged microsatellite (SSR) 18 PCR using optimized primers. Amplification was compared to samples isolated using a cetyltrimethylammonium bromide (CTAB) protocol, and the results were equivalent (Fig. 11).
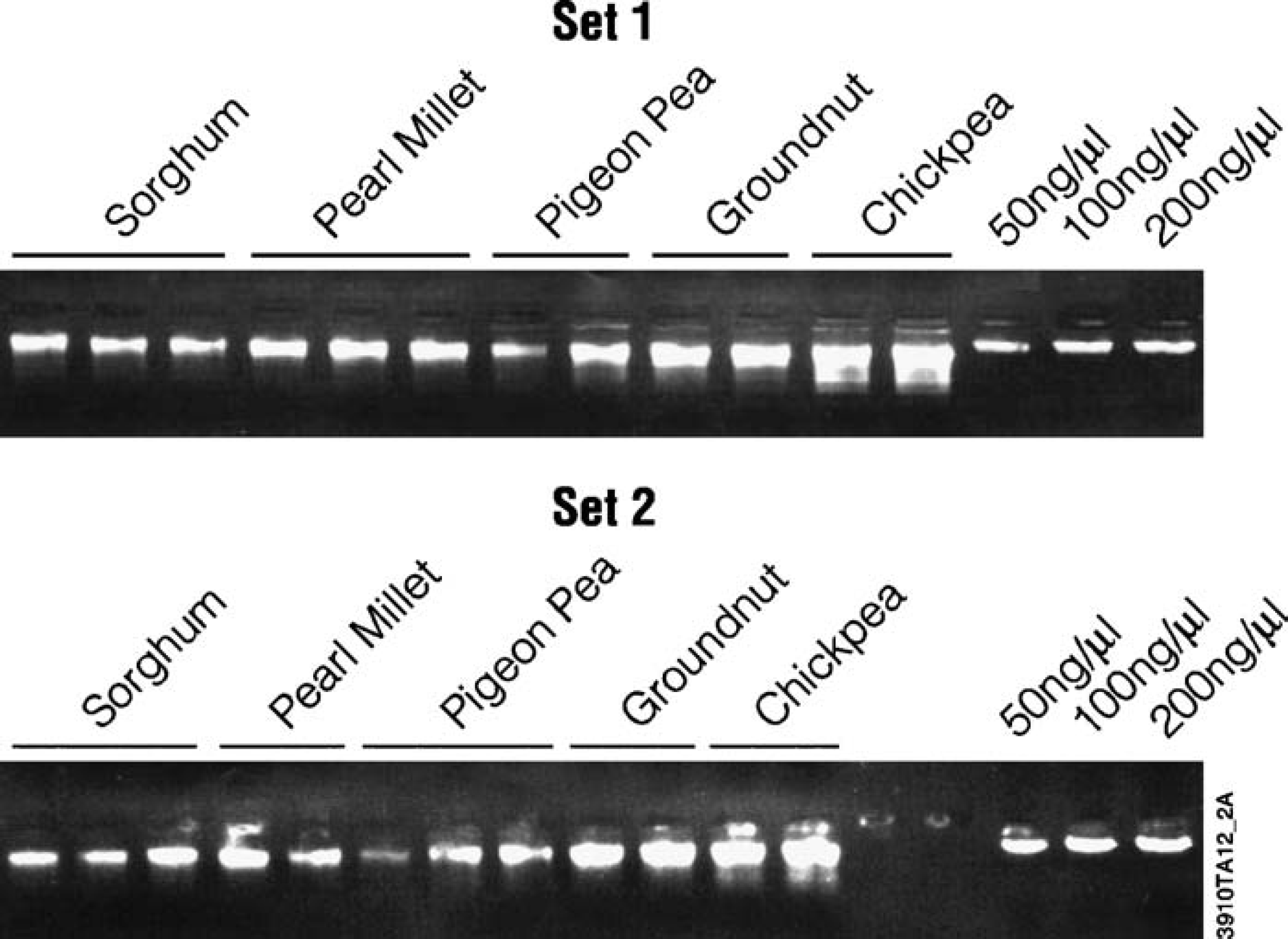
DNA extracted from International Crops Research Institute for the Semi-Arid Tropics (ICRISAT)-mandated crops. DNA was isolated from 30 mg (sorghum, pearl millet) or 20 mg (chickpea, groundnut, pigeon pea) of leaf tissue using the Wizard Plant System as described. 17 Five microliters of the purified DNA were separated on a 0.8% agarose gel, stained with ethidium bromide, and visualized with UV light. Aliquots of 50 ng/μl, 100 ng/μl, and 200 ng/μl of DNA purified with an optimized CTAB method were used as DNA standards.
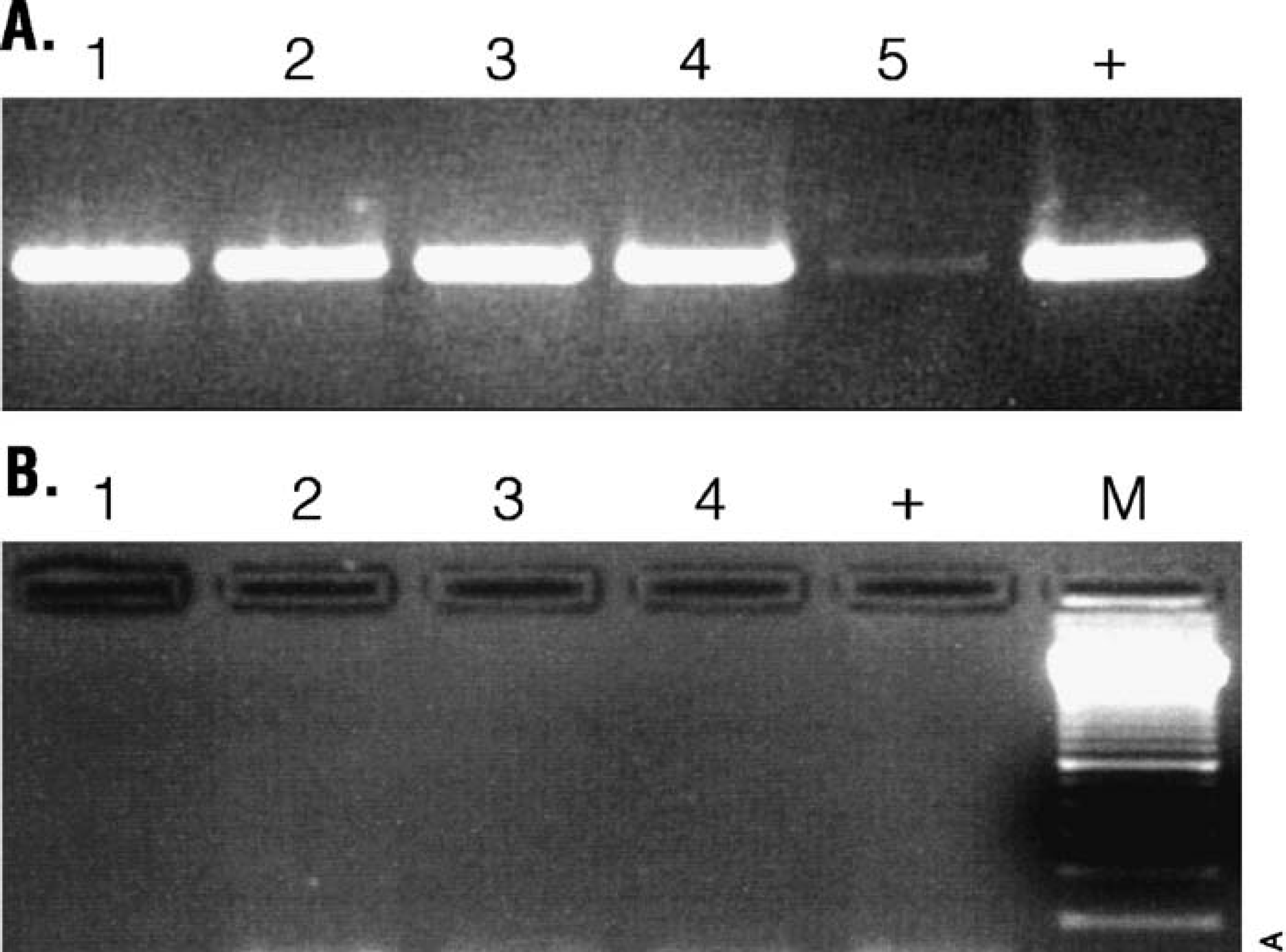
Amplification of purified DNA with SSR-PCR. DNA was purified using the method described. The purified DNA was amplified using SSR-PCR, and the amplification products were separated on a 1.5% agarose gel, stained with ethidium bromide and visualized with UV light. For reference, a positive control (+) of previously characterized DNA was included. (A) Five pearl millet samples amplified using SSR primer Psm2086 (3), and (B) four chickpea samples amplified using SSR primer Trl (4). Lane M contains a 100 bp DNA size marker (Promega Cat.# G2101).
Using the MagneSil ONE system, varying volumes of lysate from canola seeds were processed with 6μl and 8μl of MagneSil ONE Fixed Yield PMPs (40 μl of lysate corresponds to approximately one seed equivalent). As shown in Figure 12, lower amounts of lysate do not saturate the PMPs, but using 80-120 μl lysate had approximately equivalent yields. The yield with a fixed amount of PMPs needs to be evaluated for each plant material but can be expected to remain within a narrow range. Figure 13 shows yield data obtained from three consecutive runs of n = 32 samples of corn leaf punch DNA isolation.
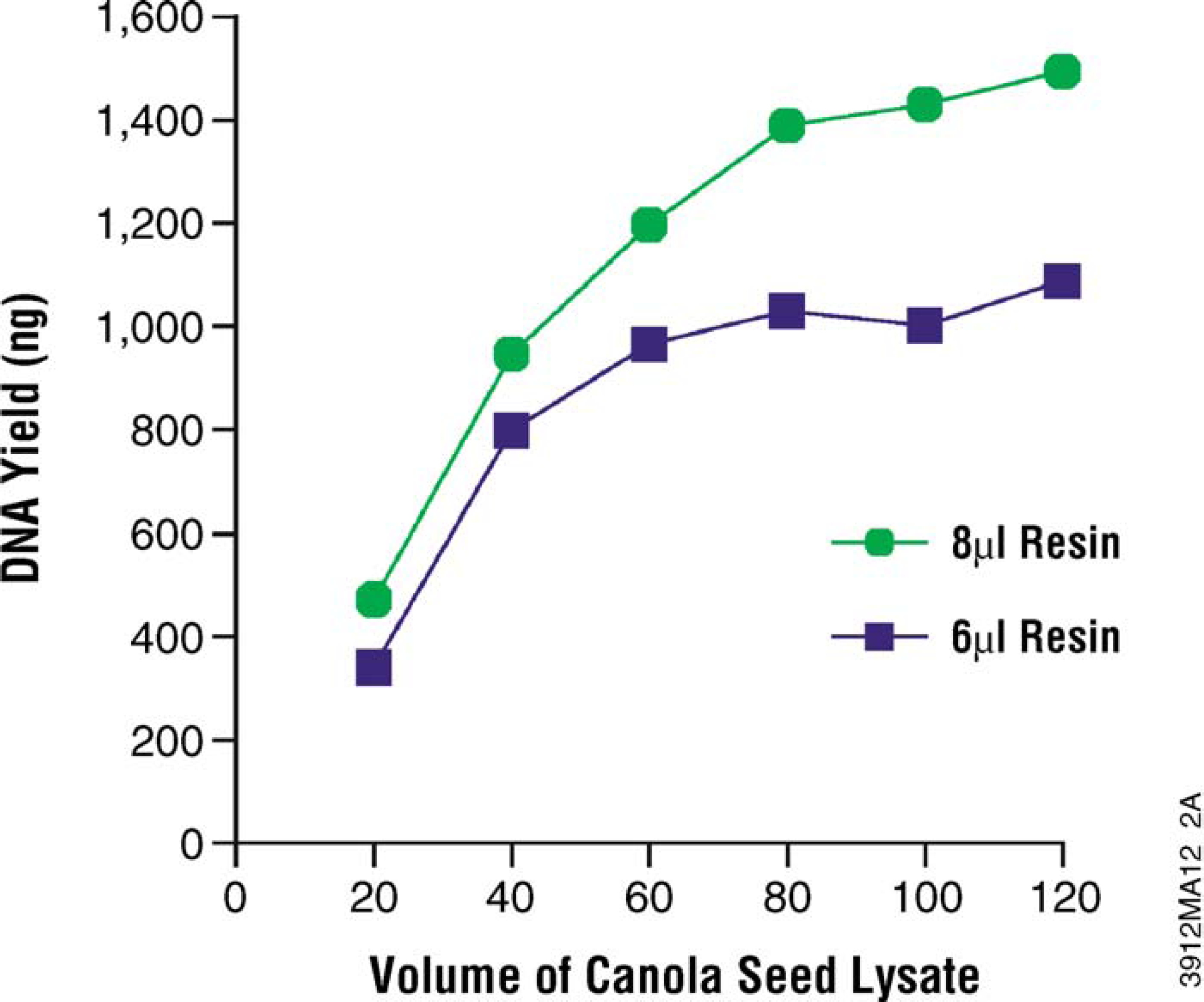
The MagneSil ONE System allows fixed-yield genomic DNA isolation from plant material. Plant genomic DNA was isolated as described. Yields were determined by PicoGreen fluorescence.
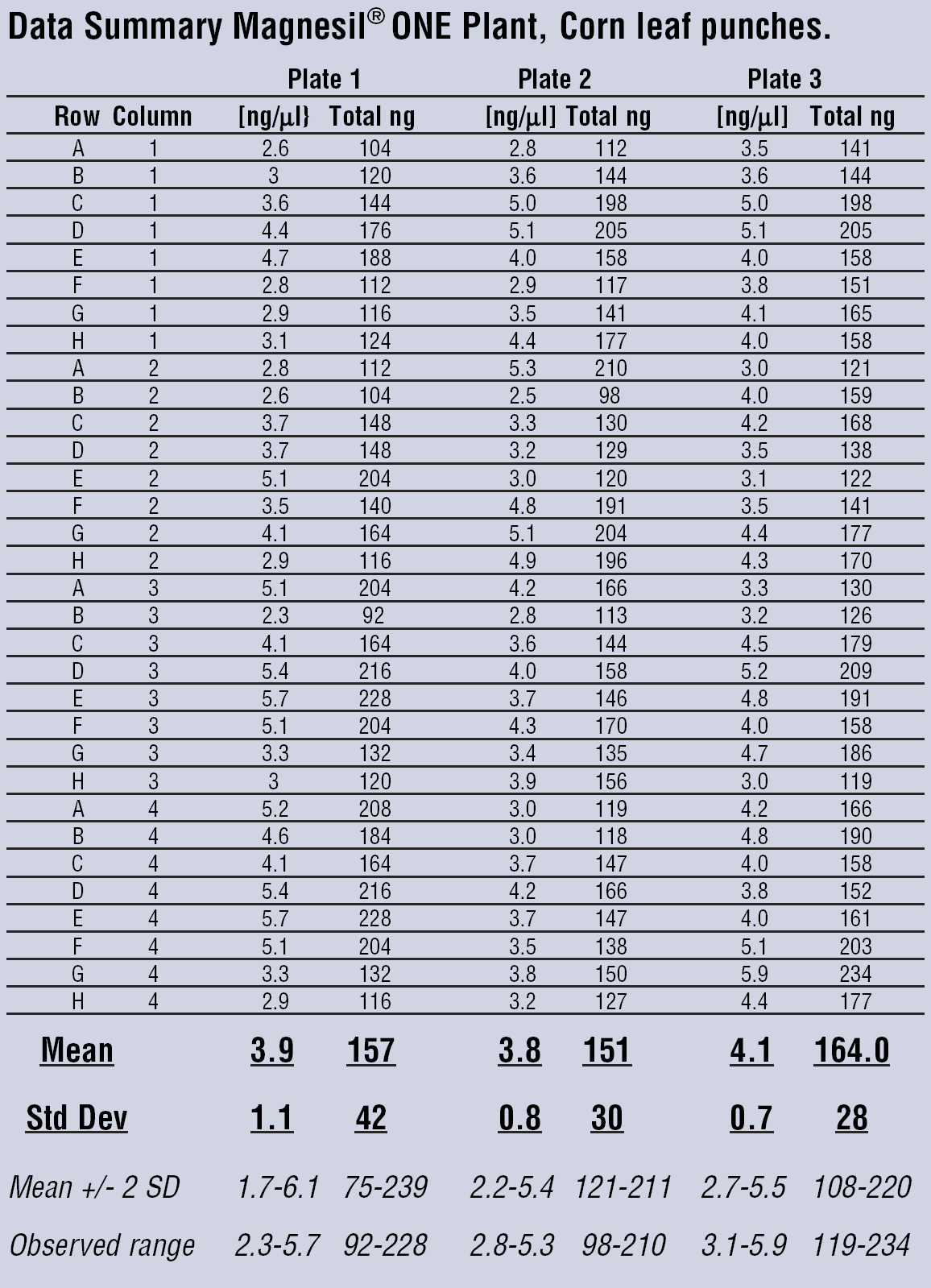
Fixed yield isolation of plant genomic DNA using Magnesil ONE. This data represents three consecutive automated runs of corn punch tissue isolated with the Magnesil ONE technique. Means and standard deviations were calculated, as well as the 95% confidence intervals and the observed ranges. With an average concentration of 4 ng/μl, the volume of DNA solution can be calculated for the particular test and be integrated into a robotic protocol to remove this amount and add it to the reaction 96 wells at a time. For example, if each reaction is to be loaded with 100 ng of DNA, 25 μl of DNA can be removed from the plate and added to reactions. Once this concentration range has been determined experimentally for the particular plant sample (different for corn, canola, wheat, etc.), this amount can be consistently used in subsequent experiments.
DNA purified using the MagneSil ONE System was used in the Third Wave Technology Invader SNP detection system, 5 which is described schematically at www.twt.com. The Cleavase enzyme used in the Invader procedure is known to be sensitive to inhibition from impurities that are different from traditionally known inhibitors of PCR. 19
Figure 14 shows allele determination data from an Invader assay 5 for cauliflower mosaic virus (CaMV) 35S promoter sequences performed on tomato leaf DNA isolated with the MagneSil ONE System. Procedure K is the standard method. Procedures O and L are similar except for the addition of 10 μl of 4 mg/mL RNase A (Promega Catalog # M4261) (Procedure O) or the addition of 10 ml of 4 mg/mL RNase A and 10 μl of 1M dithriothreitol (DTT) (Procedure L). The additives did not improve signal strength over the standard method.
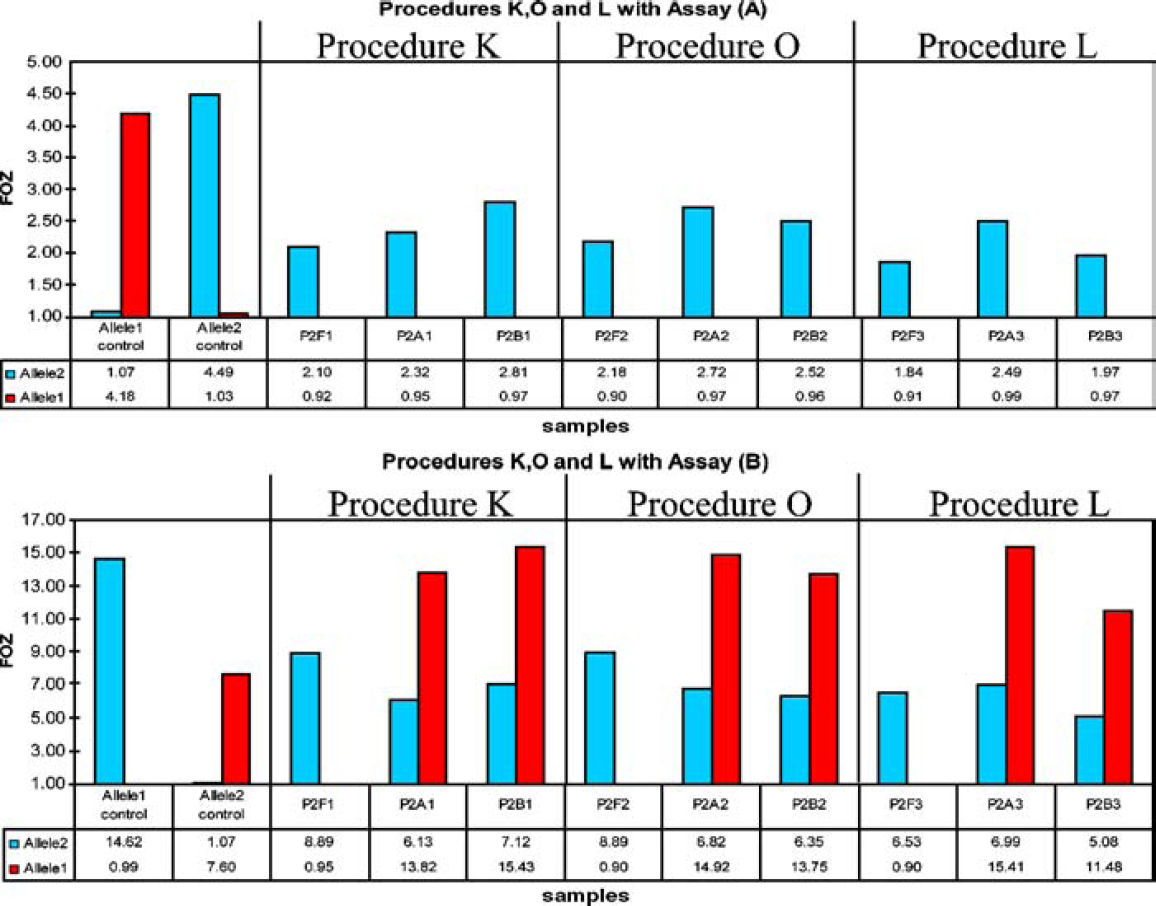
Comparison of DNA purified by different methods in TWT Invader Assays.
Discussion
MagneSil and MagneSil ONE paramagnetic particles are suited to high-throughput automated DNA purification from plant material and provide an attractive alternative to commonly used time-and labor-intensive methods based on organic extraction. The Wizard Plant System uses MagneSil paramagnetic particles (PMPs) in a 96-well format to purify sufficient DNA for PCR-based plant genotyping. The system can be scaled-up for high yield to provide genomic DNA for large numbers of PCR-based tests or archiving. In contrast, the MagneSil ONE System uses MagneSil ONE PMPs in a 96-well format to purify a fixed amount of ultraclean DNA for high sensitivity in Third Wave Technology Invader SNP genotyping assays. We have described automated methods that allow MagneSil-based purification from small to large scale and in low- to high-throughput automated formats. Purified DNA can be used in PCR, RAPD, and SSR analysis. We also show the usefulness of adding PVPP to MagneSil PMP chemistry for purification of high-quality DNA from polyphenolic-rich cotton leaf. MagneSil ONE Fixed Yield PMPs can be used to purify DNA for more challenging applications such as Third Wave Invader genotyping assays.
The flexibility and scalability inherent in MagneSil PMP-based methods result in lower costs compared to filter-plate methods and provide the level of throughput critical for efficient high-throughput plant genotyping applications.
