Abstract
We have developed a method for the automated purification of DNA sequencing reactions using the RapXtract(tm) 384 Dye Terminator Removal Kit and the Quadra 3(tm) Workstation. The process enables purification of 384 reactions in five minutes, significantly impacting the throughput potential of sequencing laboratories. The RapXtract technology utilizes superparamagnetic particles, (i.e., particles that are not themselves magnetic but that respond to a magnetic field) and eliminates the need for centrifugation, vacuum filtration, or modified primers. The Quadra 3 Workstation is a 384-channel liquid handling system, fitted with a retractable magnetic nest designed to incorporate a 384 magnetic separator. The combined technologies result in reduced variability associated with manual methods for sequencing reaction purification.
INTRODUCTION
Automated DNA sequencing is an important research and discovery tool. Improved throughput and reliable data quality of DNA sequencing will result in accelerated advances in biosciences including medical, agriculture and environmental research. In order to realize the full potential of current DNA sequencing technology, it is necessary to achieve high throughput and reliability through automation of methods that are critical to the entire DNA sequencing process. One example is the method for removing unincorporated dye terminators from cycle sequencing reactions. This process should be rapid, reliable, and compatible with 384 well plates, reduced-volume sequencing reactions, and reduced dye terminator concentrations. Furthermore, the process should integrate smoothly within the high throughput sequencing laboratory environment.
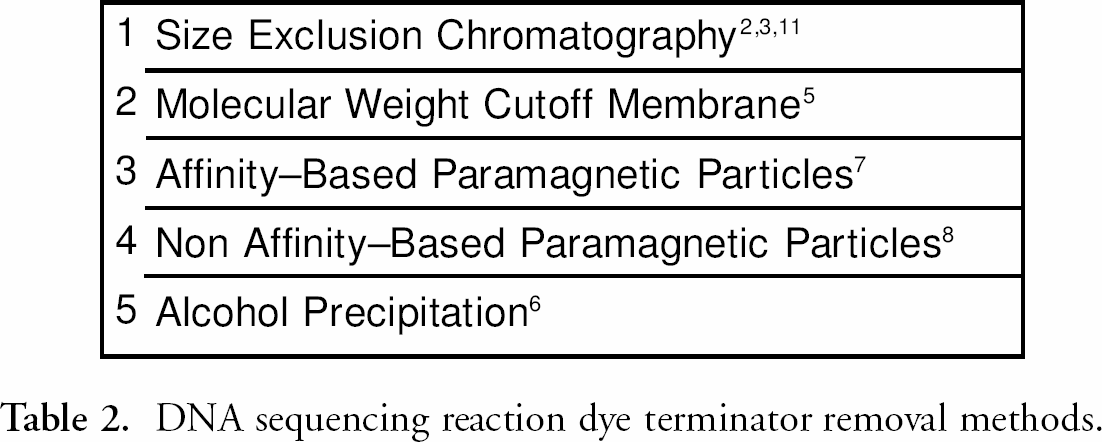
Listed below (table 1) are the steps involved in DNA sequencing. Many of the steps listed in the process can be automated resulting in an increase of throughput and concomitant reduction of human error. One exception to date is the removal of dye terminators from cycle sequencing reactions in a 384 well format. 1 table 2 lists methods for dye terminator removal. Each method listed in table 2 has steps that can be automated, but no method listed can be fully automated in a 384 well format. For example, size exclusion chromatography 2––4 uses either centrifugation, which cannot be easily automated, or vacuum filtration, which typically requires manual intervention for the adjustment of the vacuum manifold setting as outlined in the user guide “Montage Seq96” (Millipore Corporation, Cambridge, MA). In addition, this method does not work well with sample volumes of less than 10 μL and is difficult to configure in a 384 well format. Membrane-based size exclusion methods 5 require a vacuum manifold and a vacuum source that can control the applied negative pressure; this limits its compatibility with some liquid handling platforms. This approach also suffers from minimum volume constraints, requiring the addition of water or buffer to sequencing reactions to ensure adequate recovery of sequencing products from the wells. A membrane-based size exclusion method is available for an automated liquid handling system fitted with a monitored vacuum manifold, 5 however the purification process takes over 25 minutes and requires technician intervention. Alcohol precipitation 6 requires high-speed refrigerated centrifugation, specialized rotors and carriers, and is difficult to automate. Affinity-based 7 and non-affinity-based superparamagnetic particles 8 often use flammable solvents, undesirable in unattended automated systems. These methods also typically require multiple wash steps and an elution step, which can limit throughput and result in poor product recovery.
Steps involved in DNA sequencing.
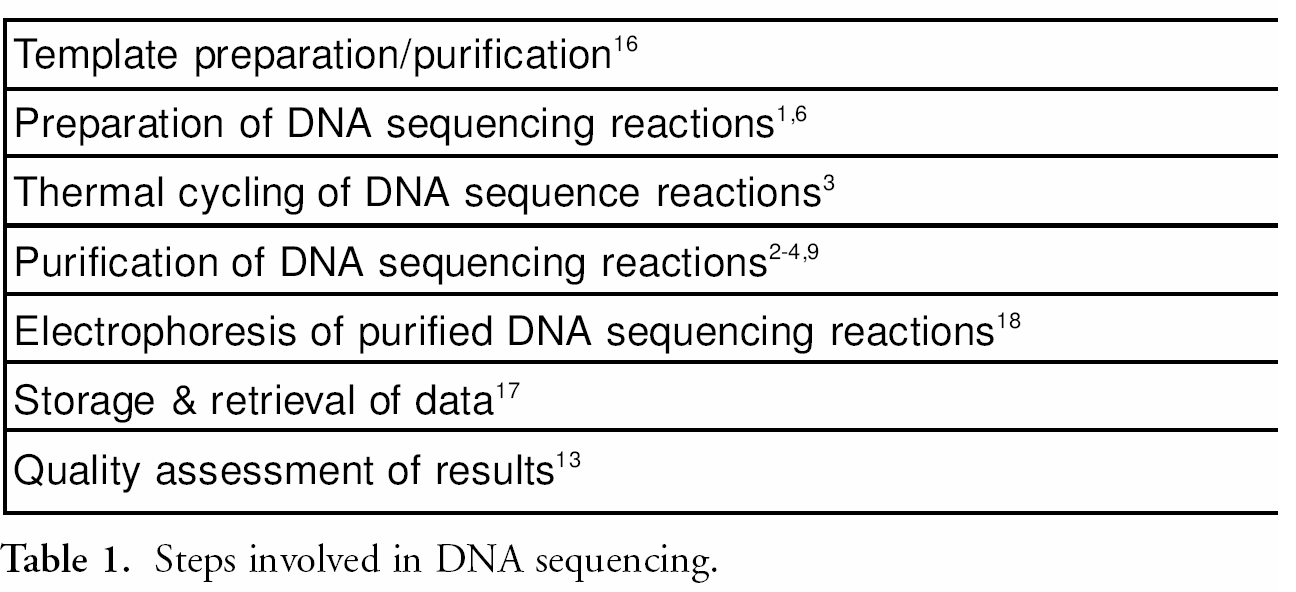

DNA sequencing reaction dye terminator removal methods.

The RapXtract Dye Terminator Removal Kits overcome many of the previously mentioned limitations as they are compatible with most automated liquid handling systems, and do not require centrifugation or vacuum filtration. The RapXtract Kits are available in 96 and 384 well formats (Prolinx, Inc., Bothell, WA), and are compatible with using reduced sample volumes and samples containing diluted dyes. The RapXtract Dye Terminator Removal Kits are based on proprietary superparamagnetic particles that bind unincorporated dye terminators from cycle sequencing reactions, leaving the purified extension products in the supernatant; a “reverse purification” process. 9 Figure 1 illustrates the three step RapXtract purification process. The mixing step (step two) can be performed on a liquid handling system by pipet mixing allowing for sample purification without technician intervention.
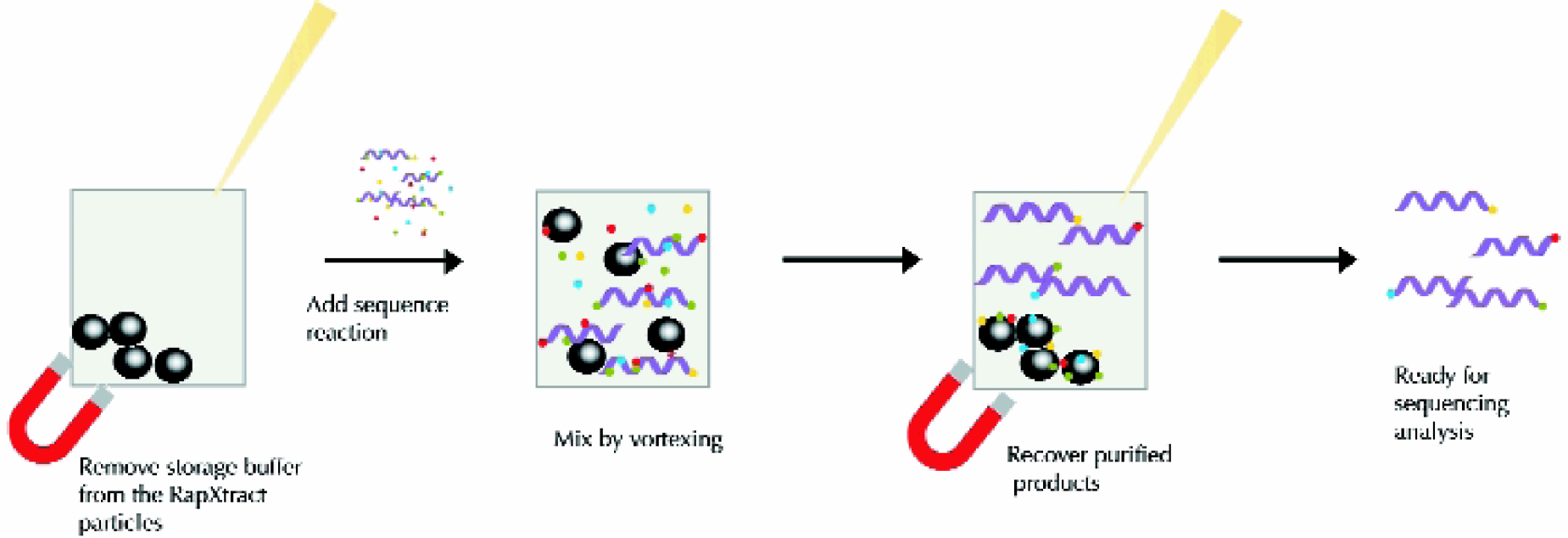
3 step “reverse purification” process: 1. Remove storage buffer. 2. Add sequencing reaction and mix by pipet agitation. 3. Remove purified extension products.
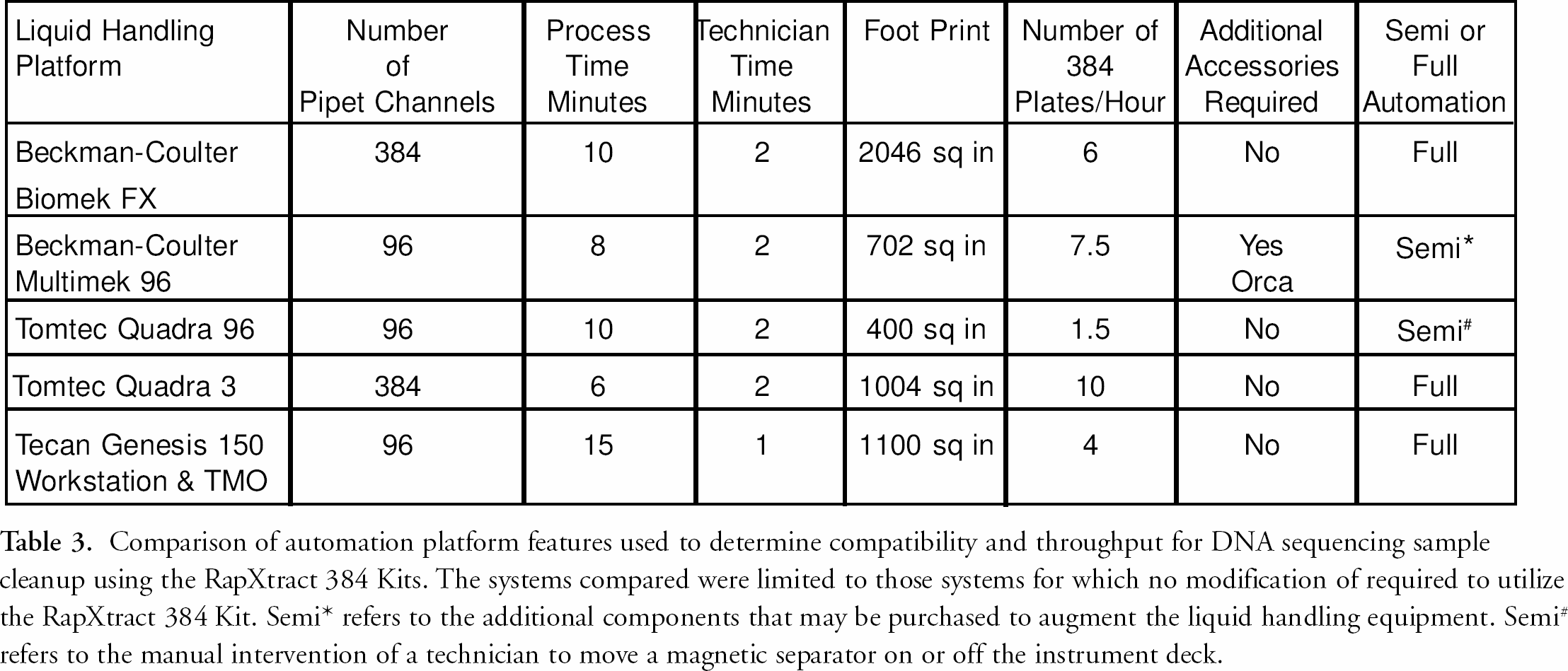
We have previously reported a fully automated method for removing unincorporated dye terminators from DNA sequencing reactions using the RapXtract 384 Dye Terminator Removal Kit. 9 The process was demonstrated on a TECAN Genesis 150 Workstation (TECAN U.S. Inc., Research Triangle Park, NC). The method allowed for the purification of 384 DNA sequencing reactions in an hour. Although a significant achievement, this throughput was unsatisfactory to a subset of scientists, 5 particularly those in DNA sequencing genome centers. In one instance the goal was to process 384 DNA sequencing samples, five microliters in total volume, in less than ten minutes in a fully automated format. In order to achieve this, it was necessary to increase significantly the rate-limiting step for DNA sequencing reaction purification when using the RapXtract 384 Kit, which is the mixing step. Since, in the case of the previous study using the Tecan Genesis system we were limited to eight pipet channels, resulting in a modest throughput of 384 samples to 60 minutes it became necessary to evaluate additional automated liquid handling technologies. We had experience with several liquid handling platforms (table 3), however, in order to achieve the goal of one sample per second, we recognized the need to purify 384 samples simultaneously. The Quadra 3 (Tomtec Inc., Hamden, CT), configured with a 384 probe pipet head, afforded the ability to purify 384 samples at once, but was limited by its design to a semi-automated method. In the standard configuration a technician was needed to move a magnetic separator on and off the instrument deck to facilitate pelleting the RapXtract particles during the process. In a collaborative effort, Tomtec Inc., and Prolinx Inc., designed the “Auto-Mag” nest, a software controlled magnetic separator device utilizing the Prolinx Magnetic Separator 384 that integrated with the Quadra 3 liquid handling platform affording a fully automated purification method.
Comparison of automation platform features used to determine compatibility and throughput for DNA sequencing sample cleanup using the RapXtract 384 Kits. The systems compared were limited to those systems for which no modification of required to utilize the RapXtract 384 Kit. Semi* refers to the additional components that may be purchased to augment the liquid handling equipment. Semi# refers to the manual intervention of a technician to move a magnetic separator on or off the instrument deck.
Additional features of interest of the Quadra 3 include its software that afforded the ability to program methods on a personal computer (PC) using the QWinEdit PC program. This method program could then be uploaded to the Quadra 3 system via its integral disk drive, or controlled directly via a PC using the Quadra 3 Windows Command Software. Optionally, the system can be programmed using its Text User Interface (TUI) on the VGA flat panel display, thereby allowing for independent operation of the unit without a PC interface. Other features of the Quadra 3 that make it ideal for use with the RapXtract 384 kit are described below:
A pipet head consisting of 384 individual positive displacement piston/pipet tip assemblies controlled by a stepper motor. This offers the precision that is essential for the sub-microliter aspirating, dispensing, and mixing of low volume cycle sequencing reactions required in the RapXtract 384 process.
An indexing stage located under the pipet head, which controls the z-axis or height of any of the shuttle positions on the deck relative to the pipet tips, affords maximum control and precision of mixing and transfer of the purified extension products.
A shuttle deck that can access stacker cassettes that have the capacity to hold twenty RapXtract 384 plates, twenty 384 well thermal cycler plates containing reactions and twenty unused 384 well thermal cycler plates. This arrangement affords con tinuous operation for approximately one hour with minimal technician intervention.
An ultrasonic tip washer, which provides efficient washing inside and outside of the pipet tips to prevent cross contami nation between samples, as well as affording a source of clean deionized water.
A program-controlled retractable magnetic nest, the “Auto-Mag,” that is compatible with the Prolinx Magnetic Separator 384, required to interface with the RapXtract 384 plate.
All of these features contributed to the reduction in sample processing time and led to the ability to achieve the goal of purifying 384 DNA sequencing samples in less than ten minutes 100% success rate and with a Phred Q20 score of greater than 550 +/−50 for DS plasmid sequencing samples.
MATERIALS AND METHODS
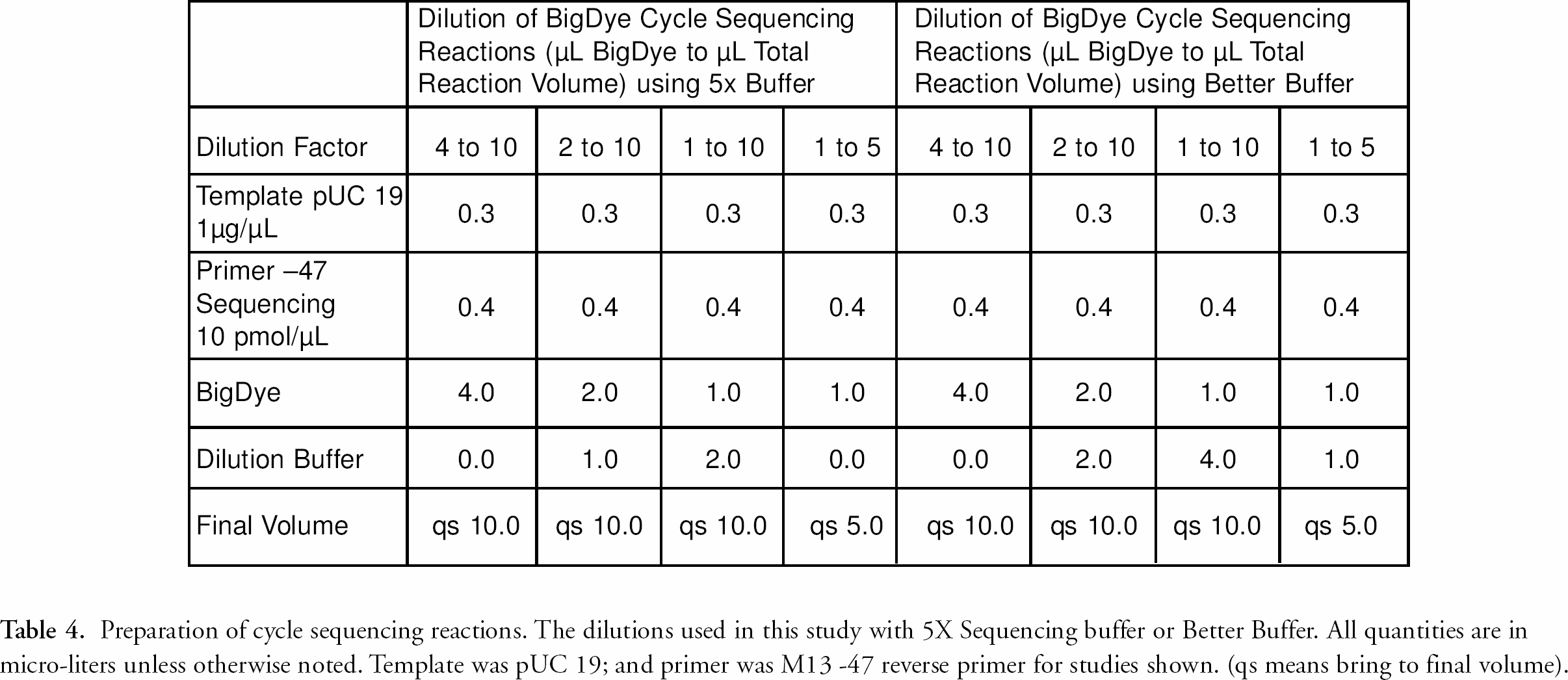
Cycle sequencing reactions were prepared as summarized in table 4. 10 BigDye Ready Reaction Mix versions 111 and three (d-rhodamine fluorescent energy transfer dye terminator dNTPs) were used for these studies (Applied Biosystems Incorporated, Foster City, CA). Diluted reactions using less than the 8 μL of the BigDye Ready Reaction Mix were prepared using 5X Sequencing Buffer (ABI, Foster City, CA,) or Better Buffer (The Gel Company, San Francisco, CA). 12 Reactions contained 250 ng pUC19 double-stranded plasmid and four pmoles of (-47) M13/pUC forward sequencing primer (New England Biolabs, Beverly MA). 10 A master mix containing all reagents for 200 reactions was prepared. The reactions were mixed then dispensed by a repeat pipettor into MicroAmp 384-Well Reaction Plates and sealed with MicroAmp Clear Adhesive 384 well sealing tape (ABI, Foster City, CA). Plates were centrifuged for 20 seconds to pellet the reactions, then thermally cycled using a PCR Express Thermal Cycler (Hybaid, Middlesex, UK) as follows: denature at 96° C for five minutes; followed by 30 cycles of [96° C for 20 seconds, 50° C for 20 seconds, and 60° C for four minutes; ending with a 4° C hold]. Reactions were purified using the RapXtract 384 Dye Terminator Removal Kit and a Quadra 3 automated pipetting system. The specific protocol varied based on the dye dilution in the cycle sequencing reactions. Figure 2 shows the deck layout of the Quadra 3. The basic process for the purification of a 1 to 5 cycle sequencing reaction dilution, (1 μL BigDye to 5 μL total reaction volume) on the Quadra 3 is as follows (see Figure 3 for complete protocol):
Preparation of cycle sequencing reactions. The dilutions used in this study with 5X Sequencing buffer or Better Buffer. All quantities are in micro-liters unless otherwise noted. Template was pUC 19; and primer was M13 −47 reverse primer for studies shown. (qs means bring to final volume).
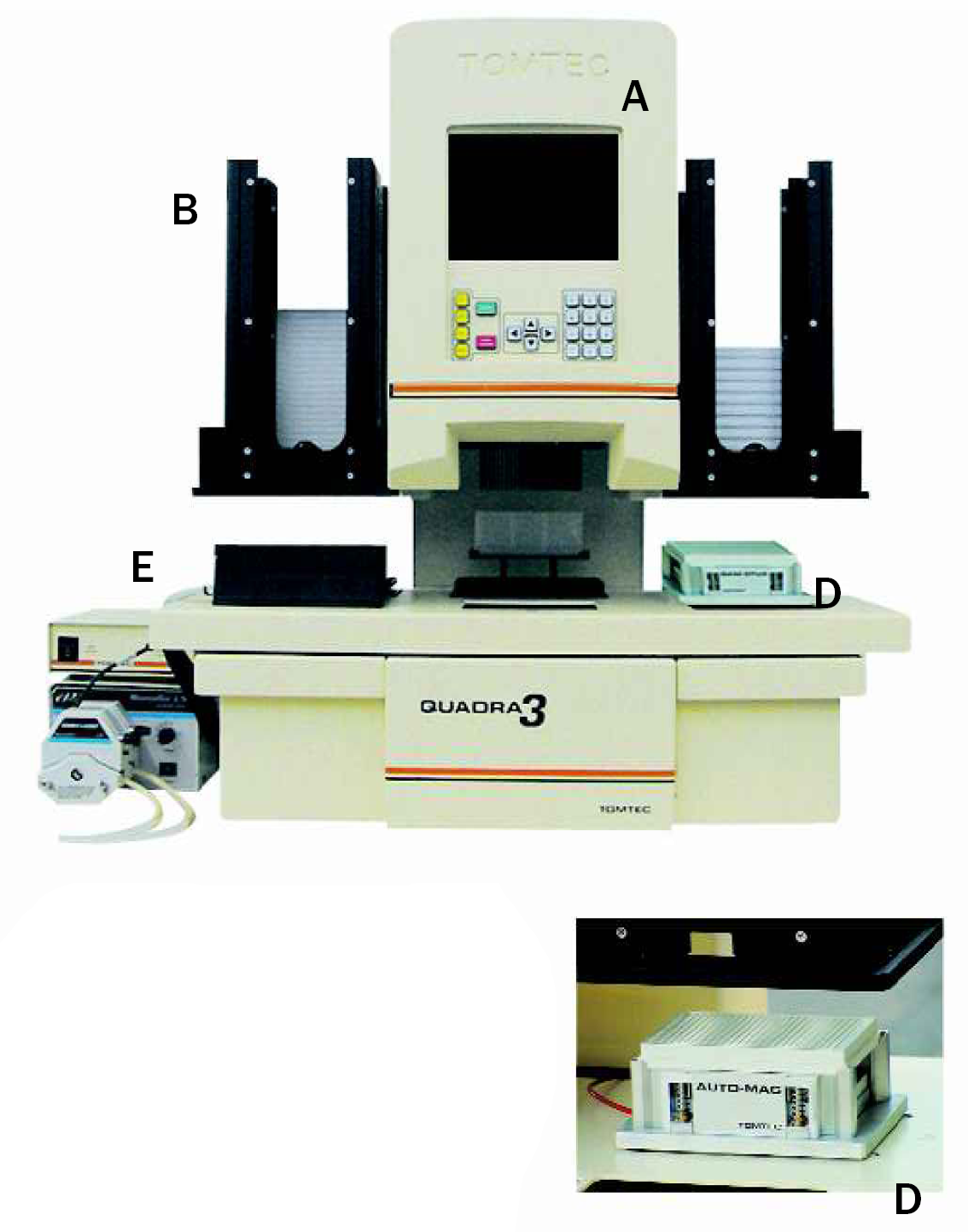

The Tomtec Quadra 3; the VGA flat panel display-A, stacker cassettes-B, 384 channel pipet head-C. The right rear deck position shows the “Auto Mag” unit-D. The small inset shows the RapXtract 384 plate on top of the retractable magnetic nest the “Auto Mag” containing the Prolinx Magnetic Separator 384. The left front deck position, shuttle position 6, contains the ultrasonic tip washer-E.

Robotic protocol for purifying a 384 well plate containing 1 to 5 (1 μL BigDye to 5 μL total reaction volume) reactions using a new plate, providing maximum throughput for this system.
Retrieve an unsealed RapXtract 384 plate from the right rear stacker (station 3) containing the retractable magnetic nest; then engage the Prolinx Magnetic Separator 384 to bring it into contact with the RapXtract plate.
Retrieve the cycle sequencing reaction plate from the left rear stacker (station 1).
Remove all but five microliters per well of storage buffer from RapXtract plate and discard it into the ultrasonic tip washing station.
Retract the Prolinx Magnetic Separator 384 from the RapXtract 384 plate.
Transfer cycle sequencing reactions (station 1) to the 384-well RapXtract plate (station 3).
Mix by pipet agitation, displacing the full reaction volume eight times
Note: For other reaction dilutions this step will vary.
Engage the Prolinx Magnetic Separator 384 to pellet the particles.
Remove the cycle sequencing plate and place into the left front stacker.
Note: The user can either wash the existing cycle sequencing plate or retrieve a new plate. In this example, a new plate is used. This option provides the most rapid processing time.
Retrieve a new 384-well plate from the left rear stacker (station 1).
Wash the pipet head in the tip washer (station 6).
Aspirate a 5 μL air gap followed by 10 μL of water at the tip washing station.
Aspirate an additional 5 μL air gap and then aspirate and transfer the purified cycle sequencing reactions from the RapXtract 384 plate on the magnetic separator to a clean 384-well plate along with the water already in the pipet head.
Off load the injection plate with the purified sequencing samples into the stacker.
Remove the plate containing purified reactions from the stacker, cover, and centrifuge plate for two seconds in preparation for sequencing.
The plate containing the purified reactions was covered with a Thermowell sealer (Corning Inc., Corning, NY), mixed for five seconds, and centrifuged for 15 seconds to remove any air voids in the wells. Purified samples were then sequenced on an ABI PRISM 3700 DNA Analyzer with POP 5 separation medium (ABI, Foster City, CA) using electrokinetic injection parameters of two kV for 45 seconds. The sequence quality was accessed using the Phred Quality Scoring Program, (available at: http://www.phrap.org; University of Washington, Seattle, WA 13 ).
RESULTS AND DISCUSSION
Dye terminator removal from cycle sequencing reactions has been automated using the Prolinx RapXtract 384 kit and the Quadra 3 liquid handling system. These combined technologies allow for reliable processing of 384 cycle sequencing samples in approximately five minutes. The RapXtract 384 Dye Terminator Removal kit is compatible with automated liquid handling and robotic platforms. Depending on the system, the process can be fully automated. The Quadra 3 is a liquid handling platform that can be used to fully automate the RapXtract 384 Dye Terminator Removal Kit. The system can function without technician intervention independently for over one hour and process twenty plates when the stacker cassettes are used. One cassette can be used to feed the twenty cycle sequencing plates, alternately stacked with twenty new thermal cycle plates for the electrokinetic injection, while the other stacker cassette can hold twenty of the RapXtract 384 well plates. Using the method described here an entire 384 well cycle sequencing plate of samples is purified and ready for analysis in five minutes or twenty 384 well plates to be purified in approximately one hundred minutes. This output can supply ten ABI 3700 DNA Analyzers using POP 5 polymer for 24 hours.
The RapXtract 384 Dye Terminator Removal Kit and the Quadra 3 method afford high quality reproducible data for BigDye versions one or three over a wide range of reaction dilutions and volumes (Figure 4). BigDye version three was formulated to extend run length on the ABI 3700 DNA Sequencer. In addition, the RapXtract 384 kits have been shown to be compatible with the Amersham DYEnamic™ ET terminator chemistry (Amersham Pharmacia Biotechnology, Cleveland, OH) as well as the LI-COR IR™ Dye Terminator infrared chemistry (LI-COR, Inc., Lincoln, NE). 14
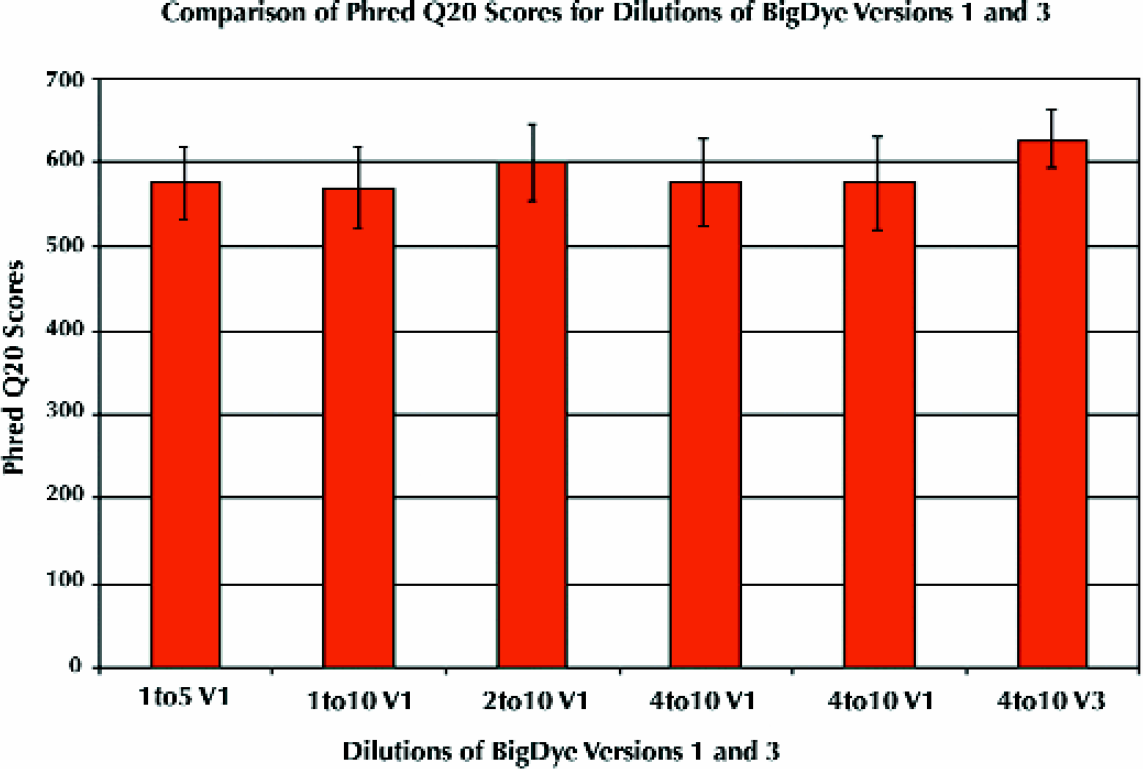
Reproducible high Phred Q 20 scores for dilutions of the BigDye Terminator (versions 1 and 3) produced by the RapXtract 384 kit on the Quadra 3; n = 384 per Dilution.
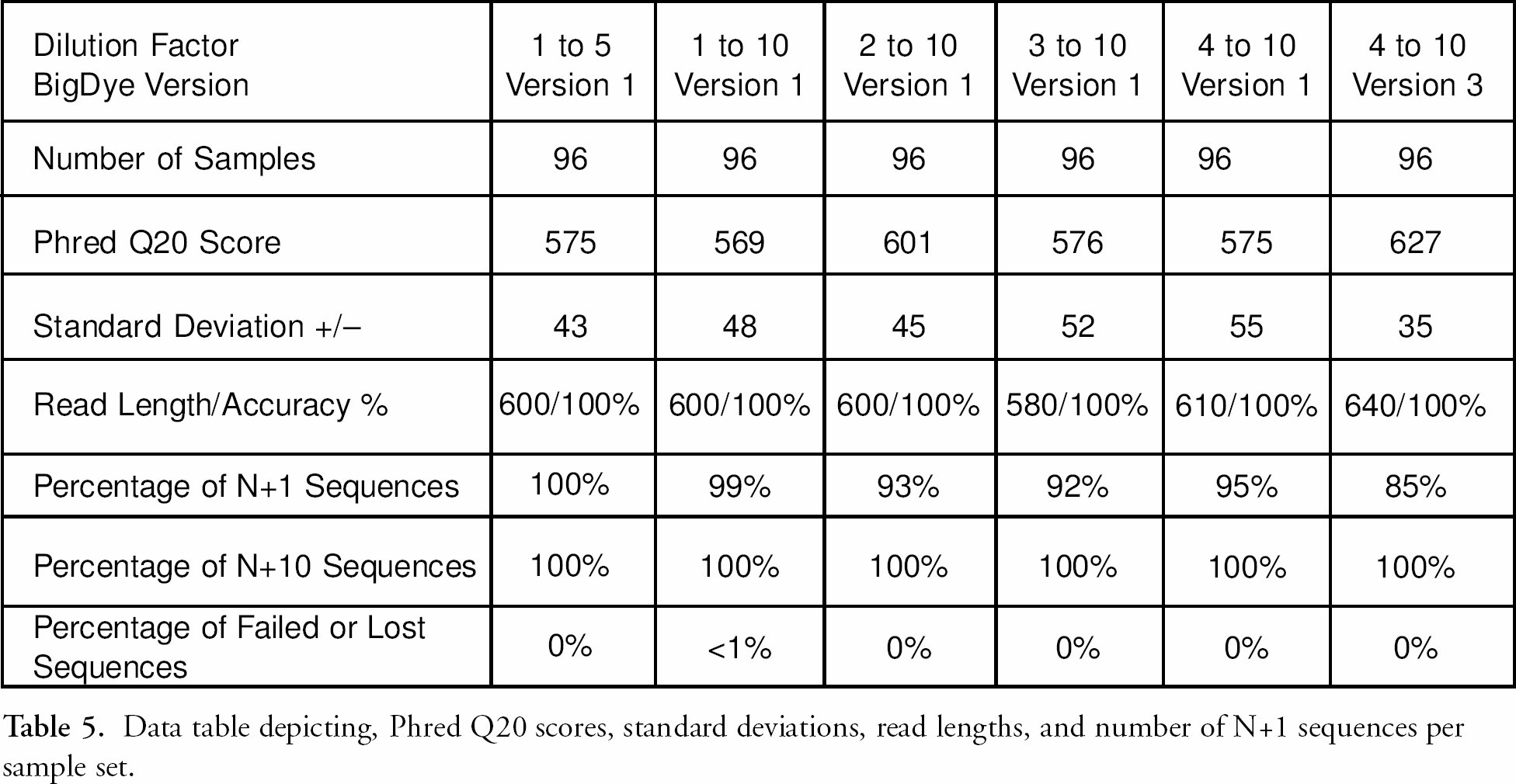
Data was assessed for quality (Q) scores. Q scores are a means of quantifying the accuracy of sequence data; the Q20 score, for a particular sequence represents the number of bases in that sequence called at 99% or greater accuracy. Various algorithms exist for calculating Q scores. In this study the Phred base caller software 13 was used to obtain these data. The Phred Q20 scores for the data set in Figure 4 ranged from 569 to 627 as shown in table 5. table 5 also indicates the range of standard deviations, read lengths, percentages of the N+1 sequences, and percentages of sequencing sample failures from loss during the purification process for this sample set. Reaction volumes were also successfully reduced to five μL total volume. Reduction in both the amounts of BigDye used and the reaction volume are cost saving measures that benefit from the reproducibility offered by automation. Also of note are the savings in time, that is, the reduction of processing time from the one hour to ten minutes for 384 DNA sequencing samples. The minimal changes in quality of Phred scores are easily offset by the overall throughput, (i.e., the ability to perform six times the number of 384 well plates in the same amount of time) thus the overall quality of sequences is increased by virtue of quantity processed.
Data table depicting, Phred Q20 scores, standard deviations, read lengths, and number of N+1 sequences per sample set.
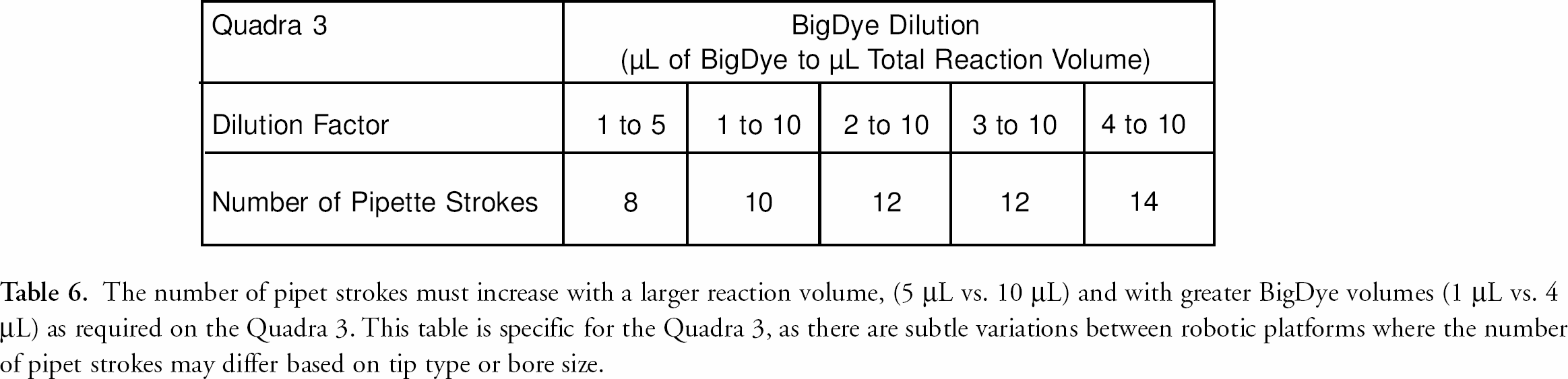
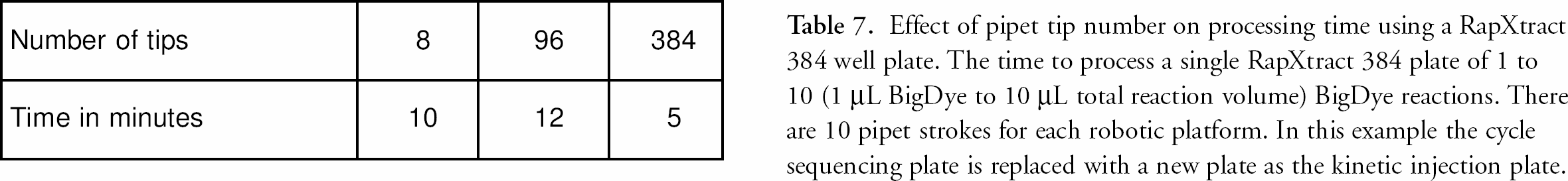
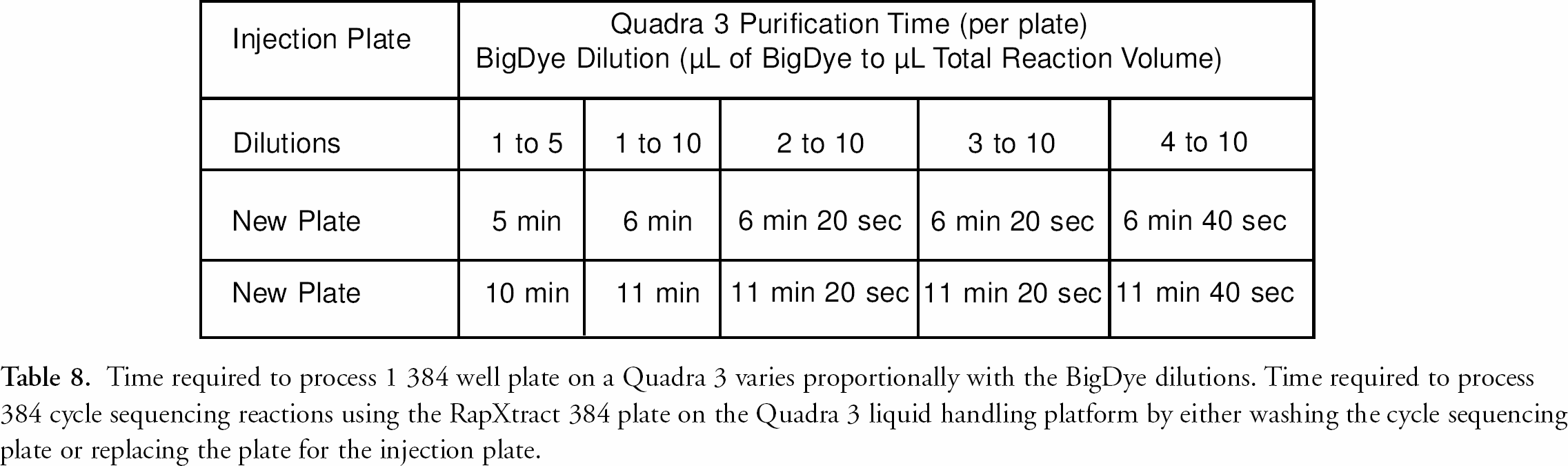
Sample throughput in sequencing laboratories is also critical. With the RapXtract kit and Quadra 3 method, processing time is directly dependent upon the dye dilution. This is because the number of pipet strokes used in the mixing step is dependent on the amount of residual dye terminators left from the initial dilution (see table 6). The number of strokes is optimized to remove unincorporated dNTPs and their degradation products and to minimize the dye front. 9 The number of pipet channels determines the time to process sequencing samples on an automated liquid handling platform (table 7). Liquid handling platforms with fewer tips take longer to process samples than those having more tips as the result of the requirement of washing the tips between each transfer or mixing step (table 7). The Quadra 3 has 384 tips affording rapid processing time which allows for the possible re-use of the thermal cycler plate by washing it and re-using it as the electrokinetic injection plate resulting in a cost savings with a minimal trade-off in total process time. table 8 illustrates the differences in time required for the Quadra 3 to process 384 cycle sequencing reactions based on reaction dilution and the replacement of the reaction plate with a new plate for electrokinetic injection. table 8 also compares the additional time the Quadra 3 requires to wash the cycle sequencing plate so it can be reused for the electrokinetic injection of the samples, if this is desired.
The number of pipet strokes must increase with a larger reaction volume, (5 μL vs. 10 μL) and with greater BigDye volumes (1 μL vs. 4 μL) as required on the Quadra 3. This table is specific for the Quadra 3, as there are subtle variations between robotic platforms where the number of pipet strokes may differ based on tip type or bore size.
Effect of pipet tip number on processing time using a RapXtract 384 well plate. The time to process a single RapXtract 384 plate of 1 to 10 (1 μL BigDye to 10 μL total reaction volume) BigDye reactions. There are 10 pipet strokes for each robotic platform. In this example the cycle sequencing plate is replaced with a new plate as the kinetic injection plate.
Time required to process 1 384 well plate on a Quadra 3 varies proportionally with the BigDye dilutions. Time required to process 384 cycle sequencing reactions using the RapXtract 384 plate on the Quadra 3 liquid handling platform by either washing the cycle sequencing plate or replacing the plate for the injection plate.

CONCLUSION
The goals of automating DNA sequencing are to not only improve throughput of the cycle sequencing samples but to maintain or increase the quality of data as well. The automation of RapXtract 384 Dye Terminator Removal Kits on the Quadra 3 has been able to meet the stated goal of processing 384 DNA sequences in less than 10 minutes and maintaining the high quality data. These combined technologies also offer versatility and afford high quality data regardless of the dilution of BigDye, sample volume, or version of dye chemistry.
The result is that a single research technician can meet the DNA sequencing potential of forty-eight ABI PRISM 3700 DNA Sequence Analyzers in 24 hours using one Quadra 3 and approximately 40 minutes of operator time during a single eight hour shift.
ACKNOWLEDGEMENTS
The authors would like to thank Leslie Linkkila for crit1ical review and assistance in preparation of this manuscript.
