Abstract
Prolinx,® Inc. of Bothell, WA has developed the RapXtract™ 384 Dye Terminator Removal Kit for full automation of DNA sequencing reaction purification. The RapXtract product line is based upon proprietary superparamagnetic particle technology that eliminates the need for centrifugation, vacuum filtration, or modified primers to achieve purification of sequencing reactions. The kit described here is pre-dispensed in a 384-well microtiter plate and run on the TECAN GENESIS Workstation 150 (Tecan U.S. Inc., Research Triangle Park, NC). This system enables rapid purification of up to 384 sequencing reactions in a single run.
As the completion of the Human Genome Project nears, it is imperative for biotechnology and pharmaceutical companies to increase throughput of DNA sequencing in order to be competitive in the drug discovery and validation process. The “race to market” requires a shift from standard DNA sequencing processes-including DNA sequencing reaction purification-towards complete walk-away automation for all steps.
Existing sequencing reaction purification methods (Table 1) require considerable resources including: plastic and other laboratory consumables; specialized equipment, such as high-speed centrifuges or vacuum filtration apparatus; and labor-intensive protocols requiring large amounts of technician time. As a result, walk-away automation of standard purification methods is difficult and expensive.
AUTOMATING CURRENT PURIFICATION METHODS
Size exclusion chromatography, the process of separation using beads with a defined particle size and chemical stability, requires centrifugation or application of pressure or a vacuum to move the solution through the chromatography bed. 1 -3 This type of chromatography in a single-tube format is an efficient method for removing impurities from sequencing reactions, but is not cost effective for use with samples in a 96-well plate preparation and would be prohibitive in the 384-well format. Formats using 96-well columns exist, but these still require more manual intervention than is optimal for sequencing facilities using a 384-well format. The 96-well format also requires specialized equipment (e.g., centrifuges or vacuum stations) to be automated. Finally, the ability to manufacture 384-well size exclusion plates is limited by the difficulty of preparing a stable column of chromatography media in a 384-well plate suitable for the complete removal of unincorporated dyes from the extension products.
Molecular weight cut-off membrane plates 4 can be automated, but doing so requires several specialized robotic tools, including a rotary platform shaker and a vacuum manifold. Not all automation platforms are compatible with these accessories, so implementation is somewhat limited. In addition, small sequencing fragments are often retained by the membrane, leading to little or no signal early in the sequence.
Affinity-based paramagnetic particle dye terminator removal systems 5 are completely amenable to automated liquid handling, but these require modified primers, which adds to their cost. The protocols also generally contain washes using flammable solvents, such as ethanol, a feature that may be undesirable in an automation system. Furthermore, the wash steps and an additional elution step add to the time required for purification. Non-affinity based paramagnetic particle dye terminator removal systems 6 do not require the modified primers, but share the other limitations of the affinity-based paramagnetic particle systems.
Alcohol precipitation methods 7 cannot be automated fully, nor easily adapted to a 384-well format. The method requires high-speed centrifugation, which is not compatible with automation. In addition, centrifuge carrier incompatibilities for 384-well plates can result in cycle sequencing sample plates breaking while being centrifuged and, therefore, result in a loss of sequencing samples. This procedure is very labor-intensive and requires a specialized rotor capable of high-speed centrifugation.

RAPXTRACT KITS WORK VIA “REVERSE PURIFICATION”
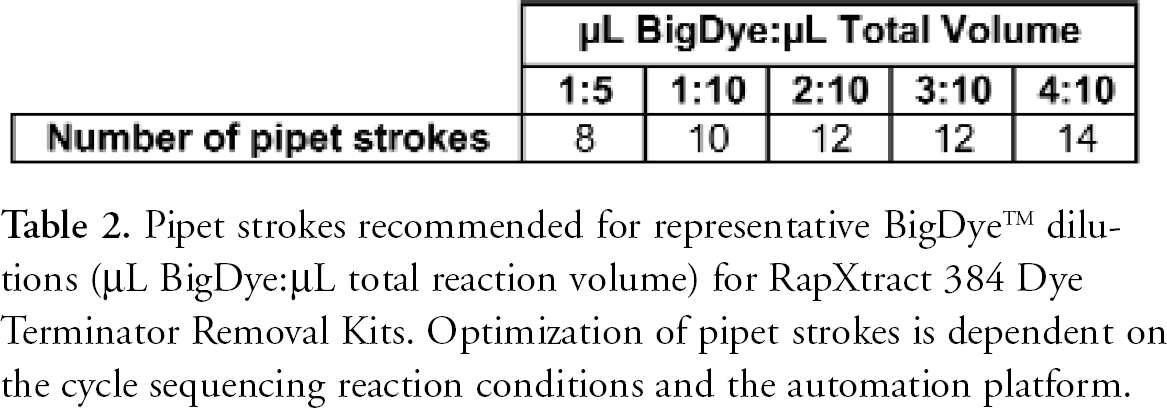
RapXtract Kits are based on proprietary paramagnetic particles that enable a “reverse purification” of sequencing reactions. Unlike other magnetic particle-based purification methods that bind the extension products, RapXtract particles remove the unincorporated dyes from the reaction, leaving behind the purified extension products in the supernatant (Figure 1). The extraction process occurs when the particles are mixed with the sequencing reactions. Mixing can be accomplished by agitation on a vortex mixer or by pipeting. The pipeting process is easily automated and is highly reproducible on most robotic platforms. The amount of mixing required is dependent on the amount of dye in the cycle sequencing reaction (Table 2). As shown in Table 2, the number of pipet strokes varies with the changes in dye terminator dilutions.
Pipet strokes recommended for representative BigDye™ dilutions (μL BigDye:μL total reaction volume) for RapXtract 384 Dye Terminator Removal Kits. Optimization of pipet strokes is dependent on the cycle sequencing reaction conditions and the automation platform.
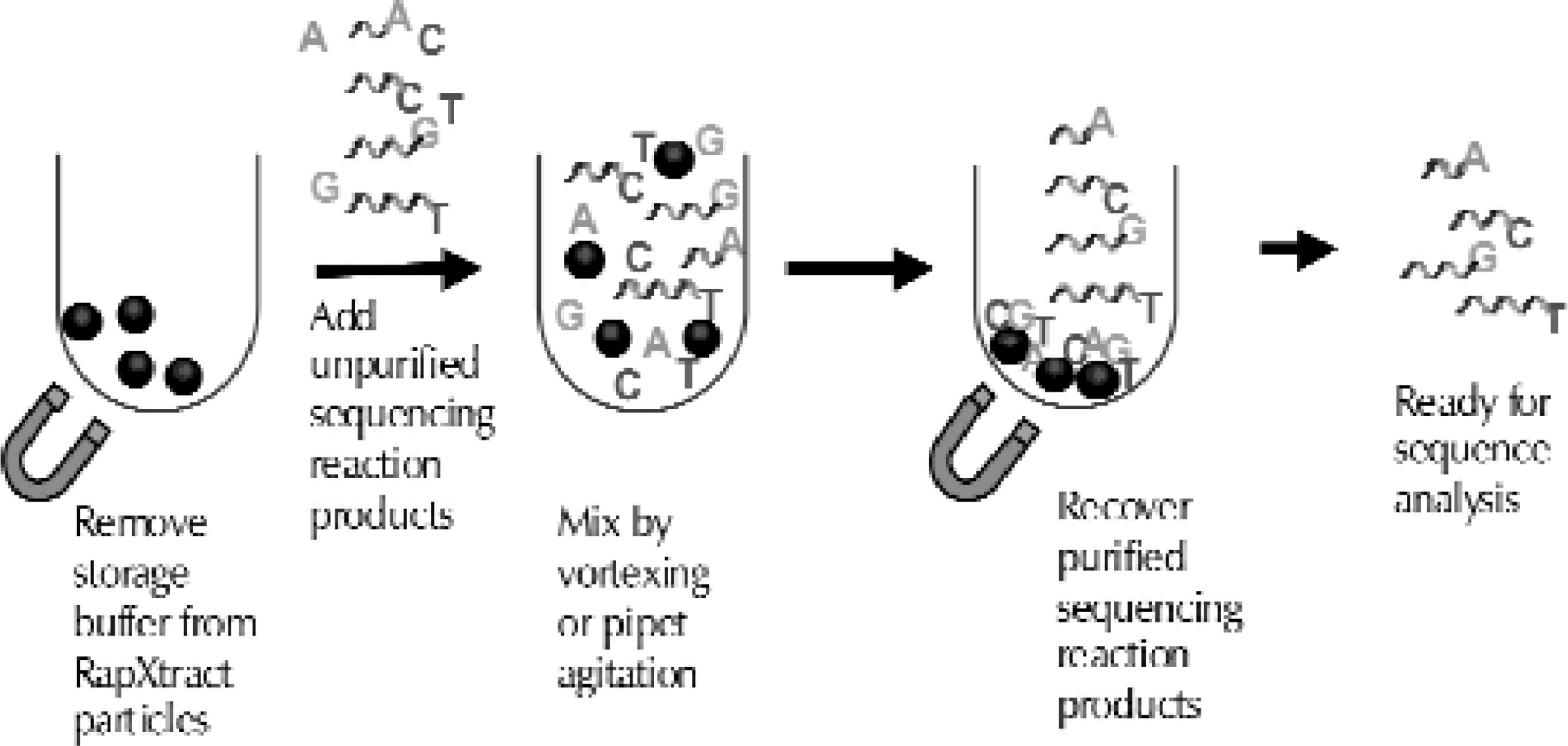
The RapXtract kit works via a “reverse purification” process.
The RapXtract 384 Dye Terminator Removal Kit, combined with the Prolinx Magnetic Separator 384 enables walk-away automation of DNA sequencing reaction purification on the TECAN GENESIS Workstation 150. Results shown in Figure 2 demonstrate the quality of data obtained on this workstation after purification with a RapXtract Kit using the BigDye chemistry from Applied Biosystems (Foster City, CA). Sequences represented in the figure were obtained from a range of dilutions of the BigDye Ready Reaction Premix. 8 , 9 Quality (Q) scores, presented in Figure 2, are a means of quantifying the accuracy of sequence data. The Q scores used in this study were obtained using the algorithm contained in the base-caller software Phred. 10 The Q20 score for a particular sequence represents the number of peaks in that sequence that have an accuracy of 99% or greater.
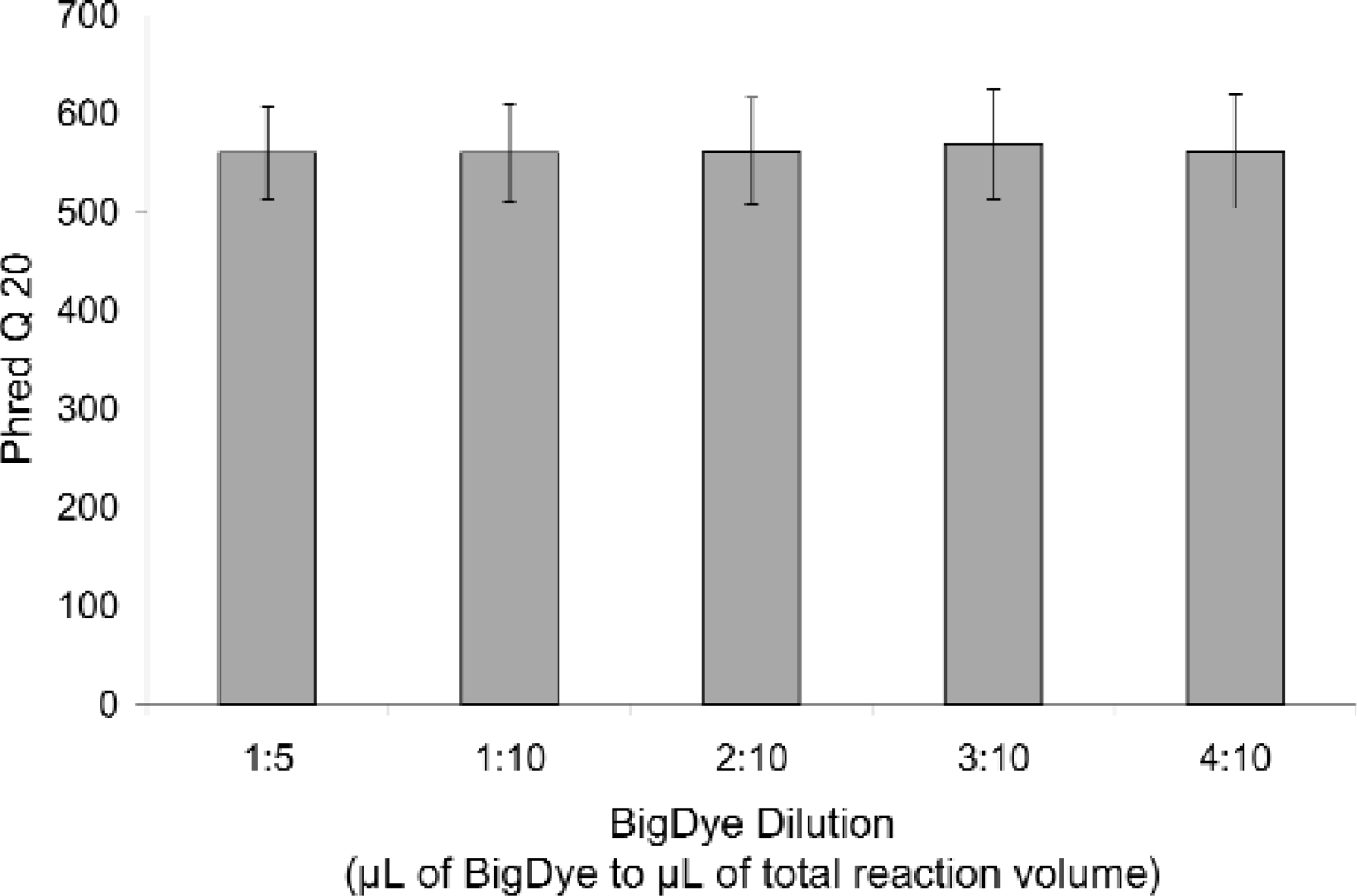
Quality scores (Phred Q 20) from sequencing reactions in a range of BigDye Ready Reaction Mix dilutions (μL BigDye:μL total reaction volume) and run on the ABI PRISM 3700 DNA Analyzer (96 samples/dilution). Samples were purified prior to sequencing on a TECAN GENESIS Workstation 150 using the RapXtract 384 Dye Terminator Removal Kit.
The RapXtract Kits are compatible with other dye terminator chemistries, such as the IRDye™ 800 Terminators infrared chemistry (LI-COR, Lincoln, Nebraska) and with the DYEnamic™ ET terminator chemistries (Amersham Pharmacia Biotech, Inc., Cleveland, Ohio). Data are not shown.
GUIDELINES FOR AUTOMATION
We used the TECAN GENESIS Workstation 150 11 to illustrate the required automation features for full automation of the RapXtract 384 Kit (Figure 3). Ideally, to fully automate the RapXtract 384 Dye Terminator Removal Kit, a liquid handling platform should have the following attributes:

A typical deck layout on the TECAN GENESIS Workstation 150; note the LiHa (on the left), RoMa (on the right), and the RapXtract 384 plate.
A Liquid Handling pipet head (LiHa), consisting of 8 Teflon-coated stainless steel tips (Standard Tips). These Standard Tips are controlled by the XP 3000 precision pumps with 500 μL syringes, affording volume accuracy of ± 0.2 μL. The LiHa also provides the ability to recognize 384 well plate spacing formats in either a 384-well thermal cycling plate or a RapXtract 384 plate.
A Robotic Manipulator (RoMa), capable of moving 384-well thermal cycling plates and RapXtract 384 plates both on and off the automation deck.
The ability to set z-axis control for the LiHa during liquid transfer and mixing.
The ability to wash both the inside and outside of the LiHa pipet tips in a fast and efficient manner using a wash station.
The ability to control aspirating, dispensing, and mixing speeds.
A deionized water reservoir able to hold 200 mL or more.
Outlined below is a brief protocol for the TECAN GENESIS Workstation 150, used to purify 1:10 (μL BigDye: μL total reaction volume) diluted sequencing reactions.
NOTE: Reagents used in a standard reaction mix were: template DNA at 250 ng, pUC19 (New England Biolabs [NEB], Beverly, MA); M13 −47 primer at 4 pmol (NEB), BigDye Ready Reaction Premix 1 μL and 5X buffer mix 1.5 μL (ABI) to a total volume of 10 μL. The sample was thermal cycled as follows: 96 °C for 5 minutes; 30 cycles through a sequence of 96 °C for 20 seconds, 50 °C for 20 seconds, and 60 °C for 4 minutes; then hold at 4 °C.
Dispense 10 μL of water per well from the 200-mL water reservoir into a clean 384-well plate on a microtiter plate (MTP) carrier with a 384-well plate locator.
Remove 40 μL of storage buffer from the RapXtract-384 plate on a MTP carrier holding a Prolinx Magnetic Separator 384.
Move the RapXtract 384 plate from the Prolinx Magnetic Separator 384 to a MTP carrier in a non-magnetic position.
Transfer eight cycle sequencing reactions from the 384-well cycle sequencing sample plate to the RapXtract 384 plate.
Mix the first set of eight sequencing samples with the RapXtract particles 10 times (full volume exchanges).
Wash the pipet tips in the wash station (both interior and exterior).
Repeat until all reactions have been processed.
Transfer the RapXtract 384 plate from the non-magnetic MTP carrier position to the Prolinx Magnetic Separator 384.
Transfer the purified cycle sequencing reactions from the RapXtract plate on the Prolinx Magnetic Separator 384 to the 384-well injection plate on a 384-well locator on an MTP carrier.
Remove the 384-well injection plate with the purified cycle sequencing samples and cover with a foil plate seal.
Briefly vortex the 384-well injection plate for two seconds.
Briefly centrifuge the samples (for less than 1 minute at under 200g) to remove any voids within the wells.
Place the sample on the deck of the ABI 3700 DNA Analyzer and inject with the following changes to the default module for the sequencing polymer: Change the injection voltage to 2.5 kV and change the injection time to 45 seconds.
Depicted in Figure 4 is a typical chromatogram resulting from a 1:10 (μL BigDye:μL total reaction volume) diluted sequencing reaction purified on the TECAN GENESIS Workstation 150 using the RapXtract-384 Kit.

A typical sequence (from an ABI PRISM 3700 DNA Analyzer) of a 1:10 BigDye dilution purified with a RapXtract 384 Kit on the TECAN GENESIS Workstation 150. The combined signal strength of the bases is >500. The Phred Q 20 score is 621.
DISCUSSION AND CONCLUSIONS
The paradigm shift from manual or semi-automated DNA sequencing reaction purification to total automation has been driven by the biotechnology and pharmaceutical industries' desire for lowering the cost of DNA sequencing. Reducing the cost of DNA sequencing can be accomplished by altering any one of several steps (reducing the amount of sequencing reagents; automating specific processes for greater accuracy when handling smaller reaction volumes; or increasing throughput by moving from a 96-well to a 384-well format). Given that the traditional methods of DNA sequencing reaction purification are difficult or impossible to automate, a new approach is required. The method using RapXtract Dye Terminator Removal Kits-based on proprietary paramagnetic particles and “reverse purification”-addresses these requirements. In addition, the RapXtract Dye Terminator Removal Kits are comparably priced with other dye terminator removal kits; the cost of the Prolinx Magnetic Separator 384 is similar to that of other lab accessories (such as a vortex mixer).
For complete automation, full-featured liquid handling systems, such as those used in the TECAN GENESIS Workstation 150, are ideal. This particular automation platform has eight pipet channels in the LiHa and as a result takes approximately 60 minutes to purify 384 cycle sequencing samples. The rate limiting step for DNA sequencing reaction purification when using the RapXtract 384 Kit is the number of pipet channels. Automated platforms that have a greater number of pipet channels can purify the 384 samples in less time. For example, the Tomtec Quadra 384, model 230 (Tomtec, Hamden, CT) has 384 pipet channels and can process a RapXtract 384 plate in less than 5 minutes. This 5-minute processing time requires a new injection plate. However, the sequencing sample plate can be re-used by simply washing it. The additional time for doing this is approximately 2 minutes. The trade-off is time versus money and this is left to the end user to decide.
The ability of automated systems to consistently and accurately aspirate, mix and dispense complements the quality-assured performance of RapXtract Kits. This results in reproducible, high-quality sequencing data over a wide range of BigDye dilutions and sequencing reaction volumes.
ACKNOWLEDGEMENTS
The author would like to gratefully acknowledge Donna Larson-Irwin and Leslie Linkkila for their insight, critical review, and help with preparation of this manuscript.
