Abstract

Molecular diagnostics is the fastest-growing segment of laboratory medicine principally because of the exquisite sensitivity and specificity inherent in nucleic acid biochemistry; in other words, disease management is enhanced. The wide use of molecular diagnostics beyond the venues of the clinical and reference laboratory will continue to be inhibited by the logistical and economic reasons stated above. A low-cost, automatable solution is essential for more of the customers of the healthcare system, whether they be healthcare providers or patients, to exploit the advantages molecular diagnostics offer. To that end, one must consider the core components of molecular diagnostic investigation, whether that be for infectious disease detection, genotyping for genetic disease or pharmacogenetic applications, oncology or any of the other key applications of nucleic acid technology.
The components of molecular diagnostics are nucleic acid preparation (the complexity of which is a function of the downstream components of a test); target complexity reduction (again a function of the downstream biochemistry; for example, primer selection in PCR or restriction enzyme digestion in Southern blot analysis); amplification (which is becoming standard in molecular investigation with a only a few exceptions for which Southern blotting remains a “gold standard”); and target detection. Target detection often depends on a hybridization event. Array-based, so-called “chip technology” is being offered as a solution that can be applied to automating some components of molecular diagnostics tests. Chip technologies that have been developed for high density applications are so high in the cost of manufacturing and optical detection as to be untenable options for the clinical diagnostics environment; to suggest otherwise is to ignore the economic realities of the clinical laboratory. Other chip technologies that have been developed with clinical diagnostics in mind, are so technology- and material-intensive as to make economical solutions for all except the clinical reference or hospital laboratory out of reach. Even in these venues, potential customers will have to consider the impact such devices could have on test volumes and labor costs to justify the associated large capital and operating costs.
The full potential of diagnostic applications of biochip technology will only be achieved with (i) absolutely inexpensive diagnostics, regardless of venue, and (ii) improved diagnostics that reduce the overall cost of disease management, so-called outcomes-based management. Clinical Micro Sensors, Inc. (CMS; Pasadena, CA) has developed a bioelectronic detection system for nucleic acid targets that is (i) a homogeneous system requiring minimal specimen preparation, (ii) specific and sensitive, and (iii) amenable to manufacturing scale-up with very inexpensive chemistries, electronics and materials. All of these characteristics are important ingredients in the eventual automation and adoption of molecular diagnostics using a universally applicable detection platform. The system is described more fully later in the article. The technology involved, though “molecular” in nature, is being engineered so that it is simple to perform, obviating the need for specialized training and expensive equipment. Thus, bioelectronic detection could be made part of a complete diagnostics system, amenable not only to the clinical laboratory, but also to the point of care, including the POL or STD clinic. With bioelectronic detection, specimen preparation consists of lysis of the cells, bacteria or viruses of interest, followed by denaturation and fragmentation, potentially all in one step. Ultimately, an automated system or in tube test would incorporate lysis of the specimen and presentation of the nucleic acid for analysis. A complete test would consist of specimen introduction, sample preparation (lysis and nucleic acid denaturation/fragmentation), amplification (if necessary, depending on the application), hybridization incubation time, signal generation (initiated by application of requisite voltage to generate electrical current) and reading the result displayed on the (handheld or benchtop) device and interfacing of the instrument with the laboratory information system. The CMS prototype handheld instrument has been fashioned with electrical contacts that interface with a cradle that supplies a serial port connection to a PC (Figure 1 and cover); larger, benchtop devices are also in use. Bioelectronic “chips” are inexpensive to manufacture and the associated cost-savings will help encourage wide use of molecular diagnostics in non-traditional venues.
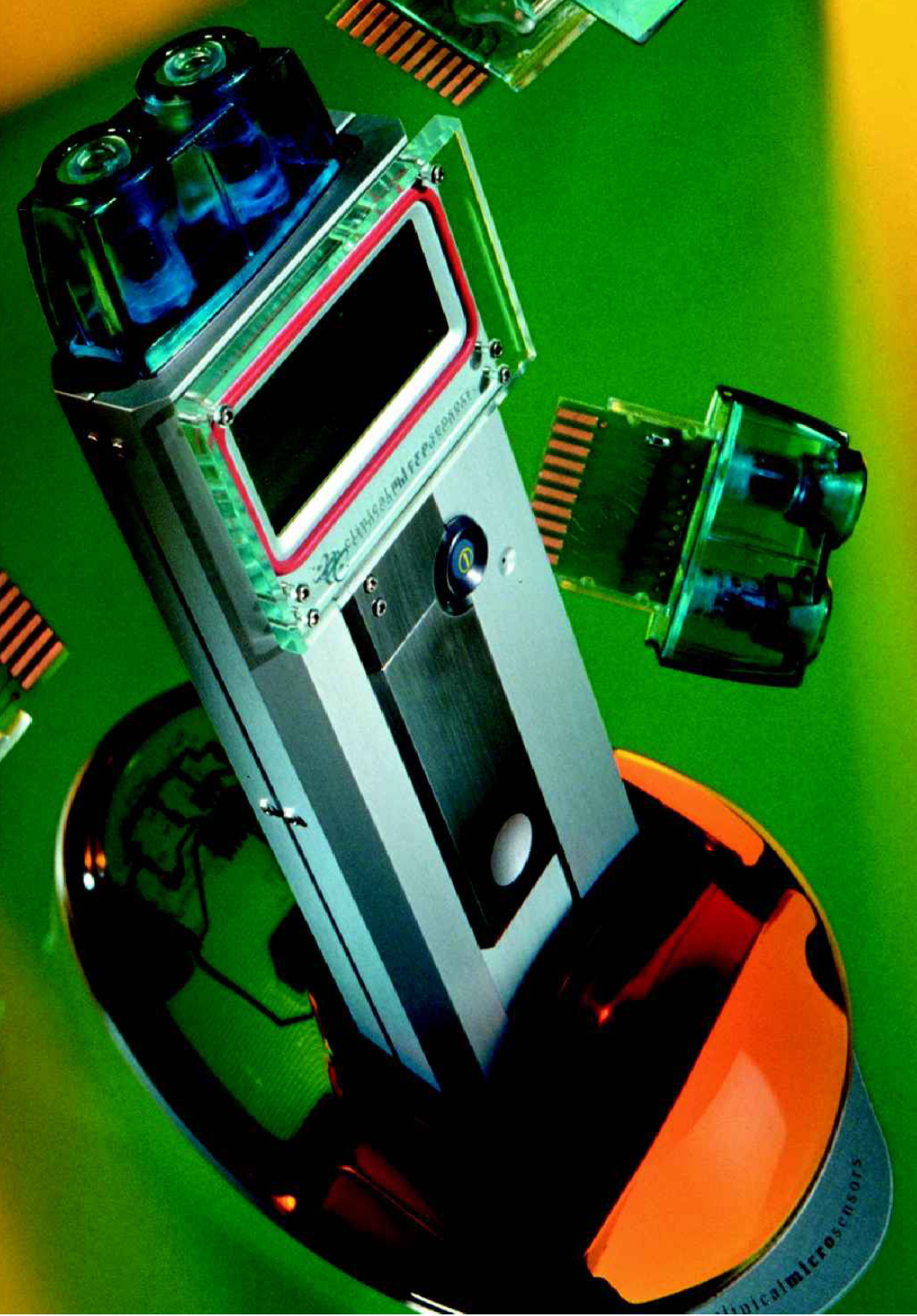

Clinical Micro Sensors (Pasadena, CA) working prototype handheld reader device. This instrument applies voltage to CMS biochips. One chip is inserted in the top of the reader and parts of three others appear in the photo. The result is displayed on the built in screen near the top of the device. The portable reader is shown resting in a CMS cradle which serves as an interface to a PC. Photo by Henry Blackham
BIOELECTRONIC DETECTION OF HYBRIDIZATION
“[I]f an adenine forms one member of a pair, on either chain, then on these assumptions the other member must be thymine; similarly for guanine and cytosine. [I]f only specific pairs of bases can be formed, it follows that the sequence on the other chain is automatically determined” (Watson and Crick, 1953). In addition to so many other profound implications from Watson's and Crick's landmark 1953 paper on the double helical nature of DNA, it gave birth to the notion of nucleic acid hybridization. Simple denaturation of DNA leads to reannealing of sister DNA strands. Competition for that sister strand reannealing can be achieved with introduction of a molar excess of single-stranded nucleic acid probe, complementary to a given region of interest. Simple competition, under favorable experimental conditions, promotes hybrid formation (at the expense of sister strand reannealing) between the probe and target of interest. Much pioneering work has gone into this, one of the most fundamental molecular biology tools: nucleic acid hybridization. Grunstein and Hogness (1975) described colony hybridization for the isolation of cloned genes. Southern (1975) described detection of specific DNA sequences on solid supports by nucleic acid hybridization. Hybridization has followed a miniaturization trend from nitrocellulose and nylon membranes to probes in the wells of microtiter plates to on-board chemistries of the most recently released limited menu, automated or semi-automated molecular infectious disease analyzers. Now the miniaturization trend is moving the community towards “chip” and glass supports, which are variations on the hybridization theme. Work at CMS has shown that DNA hybridization and detection can also be done on a unique support, printed circuit board material, with a novel reporter tag or label for detection of hybridization events based on the generation of electrical current. The fact that the hybridization event and detection generates a signal so ubiquitous to the electronics industry as electrical current goes hand in hand with the notion of molecular electronic detection of hybridization as a routine, broad-based platform for nucleic acid investigation.
Bioelectronic detection technology rests on the specific generation of electrical current through the reversible oxidation and reduction of metal complex labels on nucleic acid targets. In the case of one of the earliest tools of modern molecular biology, the Southern blot, the solid support for nucleic acid hybridization was cellulose nitrate; eventually evolving to nylon membranes. In bio-electronic detection, inexpensive printed circuit board can be used as the support material, and may be colloquially thought of as a “chip.” Deposition of DNA reagents on electrodes embedded in the printed circuit board allows one to think of the entity then as a “DNA chip.” Due to inexpensive materials and reagents and routine electronic detection mechanisms and instrumentation, it is inappropriate, however, to associate “DNA chip”-based bioelectronic detection with the relatively high costs of other DNA array systems, often employing high-cost optical detection technology.
Working prototype versions of CMS bioelectronic “DNA chips” have fourteen electronically active gold electrodes as well as reference and auxiliary electrodes (Figure 2). Each electrode is coated with specific DNA capture probes. The capture probes (approximately 25 bases in length) are deposited on the electrode surface through a process of self-assembly that forms a monolayer with relevant chemistry (Bain, et al. 1989).
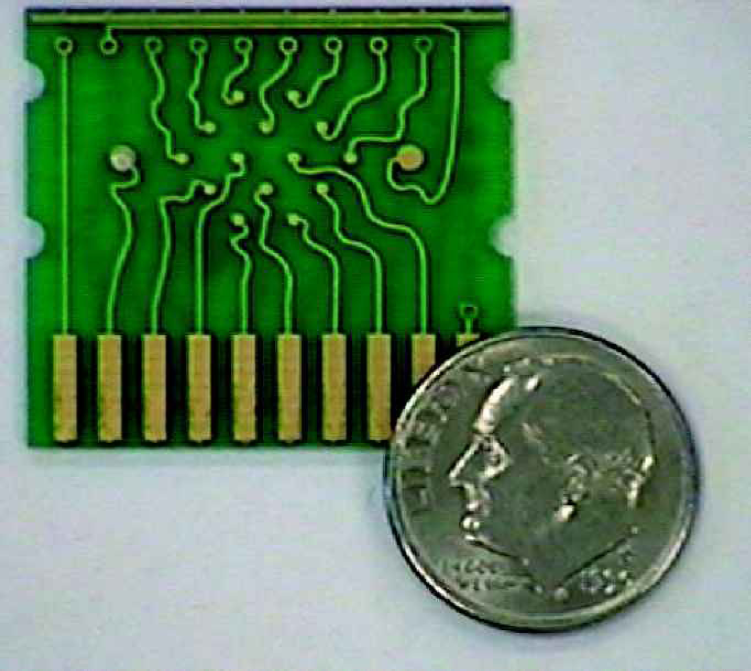

Photograph of a working prototype, printed circuit board-based, Clinical Micro Sensors “DNA Chip” used as the platform for bioelectronic detection of nucleic acid hybridization. Note 14 microelectrodes (solid gold dots in the center and top center of the chip) with one larger reference and one larger auxiliary electrode on each end. The rectangular bars at the bottom of the chip serve as the interface between the microelectrodes and the Reader device shown in Figure 1 and on the cover of this issue of JALA.
Gold coated electrodes work best for monolayer formation. One example of the specific chemistry associated with the gold electrodes that enables bioelectronic detection is shown in Figure 3 Through a process of self-assembly (Creager, et al., 1999) mono-layers are formed as shown in Figure 3. Thus, a layer of alkane thiol insulator molecules, covalently attached to the gold micro-electrode, is interspersed throughout the electrode surface with phenylacetylene molecular wires (Yu, et. al, 1999), similarly sulfur bonded to the microelectrode. Covalently bound to the end of the phenylacetylene molecular wires are single-stranded DNA capture probes designed to be complementary to a particular target of interest. Typically, the capture probes present on a given electrode are identical to each other and may or may not be identical to the capture probes present on other electrodes on the same chip. Depending on the arrangement of capture probe deposition on the array of electrodes, varying levels of redundancy may be built-in to a particular chip.
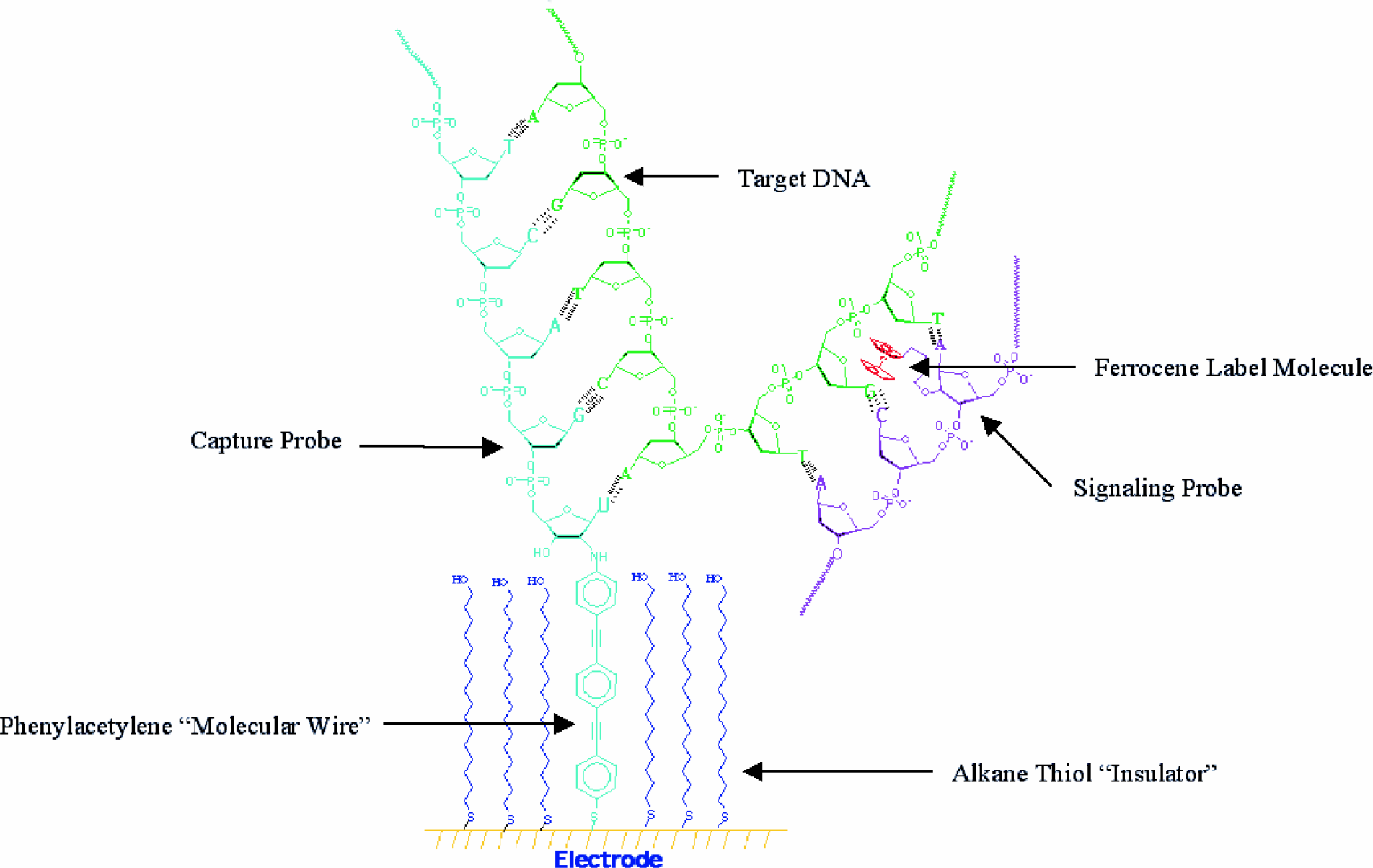
Chemical Surface of CMS Electrode: Ferrocene label (shown in red) covalently attached to a signaling probe (purple). The capture probe (aqua) is attached to the gold microelectrode through phenylacetylene “molecular wires” (aqua) that maintain the desired electrical contact between the probe and the electrode (gold) surface. The electrode surface is electrically insulated with a monolayer coating of alkane thiols (blue) to prevent unwanted redox species that may be present in the specimen from interfering with the measurements of the test system.
Bioelectronic detection proceeds via a traditional sandwich assay where a nucleic acid target of interest is simultaneously bound by capture probes on the electrode surface and a second probe in the system referred to as a signaling probe, also shown in Figure 3 Signaling probes are single-stranded oligonucleotide probes of approximately 25 bases in length that are complementary to a different, adjacent portion of the target than the region bound by the capture probe. Signaling probes may be added simultaneously with the specimen being interrogated for target or may be part of the “cocktail” present prior to specimen introduction. In either case, signaling probes serve to label the target upon hybridization.
Covalently bound to signaling probes (see Figure 3) is the organometallic, electronically-active moiety, ferrocene. More than one ferrocene molecule can be present on each signaling probe and the optimum number that balances synthesis, reagent solubility, high signal and low non-specific background is a function of the application. Hybridization between target and signaling probe electronically couples the ferrocene-labeled target to the underlying electrode. Upon hybridization of the regions of the target complementary to the capture probe and the signaling probe, the ferrocene labels are brought into sufficient proximity to the microelectrode surface for detection. Application of a voltage to the electrode (in the millivolt range) results in a reversible reduction and oxidation of ferrocenes. The result is that electrons are transferred through the insulating monolayer and to the electrode surface only when the target is present and hybridized by both signaling probe and capture probe. The current (in picoamps to microamps) generated by this system is detectable with standard or customized electronic detection systems, analog and digital signal processing equipment and computer software. A proprietary handheld reader device (Figure 1) supplies the voltage that initiates these events and has on-board software that interprets the electrical signal to both identify and quantitate, if appropriate, the target nucleic acid (the amount of current generated is proportional to the quantity of target bound).
Capture probes are attached to the gold-coated electrode through phenylacetylene “molecular wires” as described above. In this example, except for the protruding molecular wires, however, the microelectrode surface is electrically insulated with the monolayer coating of alkane thiols. This insulation serves to prevent redox species that might ordinarily be present in the specimen from interfering with the measurements of the system. Unpublished observations have demonstrated that concentrations of redox active drugs much higher than would ordinarily be observed in patients taking over-the-counter pain relievers and dietary supplements, for example, do not give rise to false positive electrical signals in the absence of specific target. Inclusion of these potentially interfering substances does not inhibit the ability of the system to detect target specifically. Electronic signals, therefore, depend on specific probe/target interactions.
Another useful feature of bioelectronic detection is its ability to provide specific, sensitive detection in a homogeneous system. Detection is not inhibited significantly in the presence of typical clinical specimens (whole blood, serum, urine) treated to liberate nucleic acids as well as more unusual specimens that would be more likely encountered in industrial microbiology applications (manuscript in preparation). Thus, detection is not dependent on separate labeling and washing steps, and does not suffer inhibition as observed in some nucleic acid amplification technologies.
Other applications of bioelectronic detection will be presented elsewhere (manuscript in preparation). The observations include human genotyping (including point mutation detection useful as a model for single nucleotide polymorphism detection and discrimination); pharmacogenetic applications; post in vitro nucleic acid amplification detection; and detection of viral and bacterial targets.
THE POWER OF AN ARRAY
One of the difficulties associated with some commercially available, kit-based clinical molecular diagnostics assays is the lack of an internal control. In other words, in the case of a negative result, how can one be sure that the result is truly negative or artifactually negative due to the degradation of target nucleic acid in the specimens caused by improper storage or transport, or caused by a high concentration of nucleases present in some tissues? Bioelectronic, array-based DNA detection offers an advantage in this regard. One or more of the electrodes present on the chip may be devoted to capture probes specific for alternate targets that are ordinarily found in all specimens. For example, urine specimens can reasonably be expected to contain sloughed bladder epithelial cells. Appropriate cellular nucleic acid targets could be used as a positive control for specimen integrity and appropriate specimen transport in compliance with College of American Pathologists accreditation requirements. Similarly, blood specimens are rich in a variety of control targets for this purpose. Plasma or serum specimens could be more problematic in this regard as they should not contain significant numbers of cells or nucleic acids, although there are platelet-specific, nucleic acid targets that could be appropriate. Inclusion of control target signaling probes in such a system is necessary for identification of control targets.
The power of array-based diagnostics is not limited to onboard specimen integrity controls. Arrays allow one to develop “panel chips” that could be used for patients exhibiting the appropriate symptoms, e.g., STD panel; delivery room panel; fungal panel; pneumonia panel; respiratory virus panel; Mycobacteria panel; meningitis/encephalitis panel; sepsis (bacteremia and/or fungemia) panel. A hepatitis C virus (HCV) chip for example could easily be engineered to inexpensively contain multiple electrodes with capture probes directed against the highly conserved 5′ untranslated region of HCV; the more such electrodes, the more such redundancy is built into the system. At the same time and on nearby microelectrodes could be capture probes specific for different genotypes of the virus. HCV genotype has been correlated with disease severity, disease progression, and response to therapy (Simmonds, 1995). Such an array could be exploited to ask multiple useful clinical diagnostic questions simultaneously. Similarly, the array may ask the same sorts of speciation or typing questions of other viruses that could reasonably be expected to be present in a plasma specimen, for example, HIV. Based on the distribution of such “blood screening chips,” the geographically dominant clades of HIV in a particular distribution area could be represented by specific capture probes on the chip surface.
VARIATIONS ON THE “CHIP” THEME
The prototype for bioelectronic detection is chip-based. While this is certainly an appropriate platform, it is not the only platform that could be used. Based on the capabilities of a homogeneous system described above, many traditional and non-traditional test formats present themselves. For example, one could envision bioelectronic detection occurring on microelectrodes engineered into the bottom of the stopper of a blood collection tube. Dried reagents in the collection tube would include lysis reagents, appropriate buffer components to achieve optimal final hybridization reaction conditions, and appropriate signaling probes and would be reconstituted upon specimen introduction to the collection tube. This test in a tube would include an electrical interface to an outside instrument that could apply the correct voltage to the embedded, immobilized tube and monitor, detect and report the associated electrical current output. This quintessential example of automation for clinical laboratory investigation of multiple targets using molecular bioelectronic detection would take advantage of a test that is begun essentially as soon as a phlebotomist draws a patient's blood sample into the appropriate tube. Other similar scenarios would be a patient providing a specimen into a urine collection cup in the outpatient, STD or family planning clinic, a pharmacy, or potentially in their home.
Incorporating bioelectronic detection as part of an array-based platform for blood cultures would obviate the need for the time and labor involved in streaking out isolates on selective media. Automation of the detection and speciation process could also include incorporation of electrodes with probes specific for antibiotic resistance genes. Such a rapid, automated device would be of economic and scientific value to the clinical microbiology laboratory. It would also be significant to a hospital's medical staff and inpatients for the benefits it could bring, by virtue of more rapid turnaround times and antibiotic resistance and sensitivity profiles, to treatment of septicemia.
THE VALUE OF AUTOMATED POINT-OF-CARE MOLECULAR DIAGNOSTICS
Chlamydia trachomatis and Neisseria gonnorhoeae testing are currently the most popular molecular diagnostic tests based on test volume. Patients presenting with symptoms to physicians or healthcare providers in venues ranging from the OB/GYN office to the STD and family planning clinic may have specimens taken for detection of these organisms. Often, rather than waiting for the results of the molecular test, which may not be available for 24–72 hours, the patient is given a prescription for azithromycin or receives an intramuscular dose of ceftriaxone to treat the suspected infection(s). If the result of the test comes back positive, then appropriate therapy has been provided. If the result returned is negative, then further exacerbation of the problem of inappropriate and unnecessary use of antibiotics has occurred. On the other hand, if the healthcare provider waits for the results of the traditional test before treating, there are associated costs, risks and logistical problems with re-contacting the patient in the case of a positive test result. The benefits provided by a rapid, point-of-care, appropriately sensitive test for these organisms to public health, the worldwide antibiotic resistance problem, the patient, and the economics of the provision of healthcare are immediately obvious and potentially available through the use of a bioelectronic detection platform. Automated, rapid, uncomplicated, point-of-care testing for these organisms would be a boon for rational use of antibiotics and rational, data-based disease management by the physician or healthcare provider.
Point-of-care, bioelectronic detection in hospital delivery rooms for Group B streptococcus and bacterial/viral meningitis panels in emergency centers are natural venues for addition of automated molecular diagnostics and would offer life-saving potential. There are many more examples.
CONCLUSION
Molecular diagnostics is the fastest-growing segment of laboratory medicine for a variety of clinical and business reasons. Molecular diagnostics offers advantages for clinical laboratory medicine; its incorporation into most sections of the pathology department is inevitable. This decentralization of molecular diagnostics from specialized clinical laboratories to the wider pathology laboratory must incorporate the economic realities of today's healthcare environment. Wide use must go hand in hand with simplification and automation to realize the goals of high quality, more efficient operations, and reduced costs. To those ends, nucleic acid preparation, amplification and target detection must become simple and automated. Bioelectronic detection offers a widely applicable platform for these crucial endeavors which must, and will, happen to move clinical laboratory diagnostics forward into the next century.
ACKNOWLEDGMENTS
The author thanks R.H. Levine, Ph.D., J.F. Kayyem, Ph.D., G.F. Blackburn, Ph.D., and H Duong, Ph.D., MBA, for their critical evaluation and helpful comments. The author also thanks S. Dameron for her help in preparation of this manuscript.
