Abstract
Objective
As the diagnostic significance of microRNAs (miRNAs) in the detection of bladder cancer is controversial, we aimed to perform a meta-analysis to comprehensively assess the diagnostic value of miRNAs in blood and urine for detecting bladder cancer.
Methods
A systematic literature search of public databases was conducted to obtain qualified studies. Sensitivity was utilized to plot the summary receiver operator characteristic (SROC) curve against specificity and the area under the SROC curve (AUC) was generated to evaluate the pooled diagnostic efficiency. Subgroup analyses and meta-regression were applied to investigate the underlying sources of heterogeneity. The STATA 12.0 software was used to perform all statistic analyses.
Results
A total of 58 studies from 22 articles comprising 4,558 bladder cancer patients and 4,456 controls were included in our meta-analysis. MiRNAs in blood and urine manifested relatively good diagnostic efficiency in detecting bladder cancer, with a sensitivity of 0.74, a specificity of 0.78, and an AUC of 0.83. Multiple-miRNA assays were more accurate than single-miRNA ones in bladder cancer diagnosis. Blood-based miRNA assays displayed better diagnostic performance than urine-based ones. In addition, miRNAs showed reduced diagnostic value in bladder cancer among Caucasians compared with Asians.
Conclusions
MiRNAs in blood and urine, especially the combination of multiple miRNAs, may serve as hopeful noninvasive biomarkers for early diagnosis of bladder cancer. Further extensive prospective research is needed to verify their clinical significance in bladder cancer diagnosis.
Introduction
Bladder cancer accounts for the second most common tumor diagnosed and the second leading cause of death among patients with genitourinary tract malignancies worldwide (1). Frequent recurrence and high mortality are characteristic of bladder cancer when it becomes invasive (2). It is well known that cystoscopy and urine cytology are the primary diagnostic and surveillance approaches for bladder cancer (3). However, cystoscopy is uncomfortable, invasive and costly, and it is difficult to detect carcinoma in situ (CIS) by cystoscopy; urine cytology is more specific but less sensitive. These limitations underscore a pressing need for novel noninvasive biomarkers with high sensitivity and specificity for bladder cancer diagnosis.
MicroRNAs (miRNAs) are short non-coding RNA molecules that can modulate protein expression at the posttranscriptional level and serve as oncogenes and/or tumor suppressors (4). Numerous studies have provided evidence that miRNAs can exist in a stable form in various body fluids (including urine, serum, plasma, tears, bronchial lavage fluid, colostrum, breast milk, amniotic fluid, etc.) and be exploited as noninvasive biomarkers for detection of bladder cancer (5-6-7-8-9-10-11-12-13-14-15-16-17-18-19-20-21-22-23-24-25-26-27). However, there have been inconsistencies and discrepancies among these studies regarding the application and reliability of miRNAs for early detection of bladder cancer. We therefore conducted this meta-analysis to comprehensively elucidate the diagnostic value of miRNAs for early noninvasive detection of bladder cancer based on previous publications.
Materials and methods
Literature Search
This meta-analysis was performed in accordance with guidelines for diagnostic meta-analysis (28). A systematic literature search of PubMed, Embase and the Cochrane Library for studies published in English up to November 6, 2015 was conducted to obtain eligible original articles focusing on the diagnostic value of miRNAs for bladder cancer. The retrieval strategy adopted was as follows: (“urinary bladder neoplasms” OR “bladder cancer” OR “bladder tumor” OR “transitional cell carcinoma of bladder” OR “bladder carcinoma” OR “urinary tract transitional cell carcinoma”) AND (“microRNA” OR “miRNA” OR “miR”) AND (“blood” OR “serum” OR “plasma” OR “urine” OR “urinary” OR “cell-free” OR “circulating”) AND (“diagnosis” OR “sensitivity and specificity” OR “ROC curve”). Furthermore, the reference lists of all relevant publications were manually retrieved to obtain additional articles.
Inclusion and Exclusion Criteria
Two reviewers independently assessed the eligible publications. Any disagreement between the 2 reviewers was resolved by discussion with a third reviewer to arrive at a consensus. Studies qualifying to be included had to fulfill the following criteria: (1) studies using the reference standard to confirm the diagnosis of bladder cancer patients; (2) studies concerning the diagnostic significance of miRNAs in blood and urine for bladder cancer; (3) studies providing sufficient data for reconstruction of 2-by-2 tables, including true positive (TP), false positive (FP), true negative (TN), and false negative (FN). Exclusion criteria were (1) case reports, reviews, letters or seminar articles; (2) studies unrelated to the diagnostic performance of miRNAs in blood and urine for bladder cancer; (3) studies with duplicate data published in other articles.
Data Extraction
The following data were independently extracted from all eligible publications by 2 reviewers using standardized forms: (1) basic study characteristics consisting of first author, publication year, country of publication, ethnicity, sample size, mean age, gender ratio, cancer type, specimen type, miRNA profiling, methods of miRNAs assay; and (2) diagnostic value, including area under the curve (AUC), sensitivity, specificity, TP, FP, FN, and TN.
Quality Assessment
Each included study was evaluated systematically and scored independently by 2 reviewers according to the revised Quality Assessment of Diagnostic Accuracy Studies 2 (QUADAS-2) criteria (29). This scoring system consists of 4 basic domains: patient selection, index test, reference standard, and flow and timing, adopting 7 questions to be answered with “yes”, “no”, or “unclear” to assess the quality of the included publications. Each domain was assessed in terms of the risk of bias, and the first 3 domains were assessed in terms of applicability. An answer “yes” indicated that the risk of bias could be judged as low, while “no” or “unclear” corresponded to a high risk and an unclear risk, respectively. In the event of controversy, a third reviewer was consulted and consensus was reached through multilateral discussion.
Statistical Analysis
The Q test and I2 statistic were conducted to evaluate significant heterogeneity among the included studies (30). A p value less than 0.10 for the Q test or an I2 value ≥50% demonstrated substantial heterogeneity, in which case the random-effects model should be applied. In addition, subgroup analyses and meta-regression were employed to further investigate potential sources of heterogeneity. The pooled sensitivity, specificity, positive likelihood ratio (PLR), negative likelihood ratio (NLR), and diagnostic odds ratio (DOR) of all included publications were calculated with a bivariate meta-analysis model. The sensitivity and specificity of each included study were applied to plot the summary receiver operator characteristic (SROC) curve and to figure out the AUC representing a pooled evaluation index of diagnostic performance. Fagan's nomogram was used to estimate the postdiagnostic effect after pooled analysis. Deek's funnel plot asymmetry test was performed to evaluate the publication bias with p<0.10 implying statistical significance (31). All analyses were performed by means of the STATA 12.0 software (StataCorp LP).
Results
Literature Retrieval
The flowchart of the literature search is presented in Figure 1. A total of 331 potentially relevant articles were obtained by the initial database search, of which 42 duplicates were excluded. After reviewing abstracts and keywords, 66 letters, reviews and meta-analyses, 10 articles that were not about humans, and 160 articles unrelated to our research were further excluded. Fifty-three articles remained for full-text review, of which 31 were excluded: 13 articles irrelevant to diagnosis, 14 articles without sufficient data, and 4 articles about tissue-miRNA assays. Finally, 22 eligible articles were employed in this meta-analysis, including 5 articles involving blood-based miRNAs (16, 17, 20, 24, 27) and 17 articles concerning urine-based miRNAs (6-7-8-9-10-11-12-13-14-15, 18, 19, 21-22-23, 25, 26).
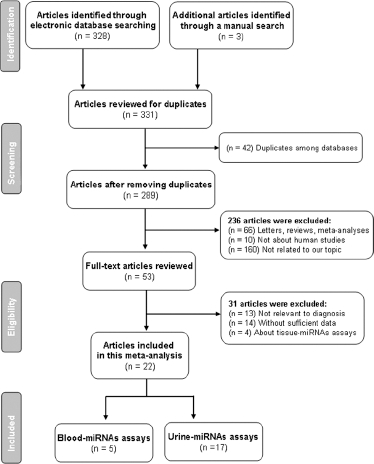
Flowchart of publication selection process.
Basic Characteristics of Included Studies
The main characteristics of the selected articles are listed in Table I. A total of 58 studies from 22 articles were included in our meta-analysis. Only 12 of these 58 studies investigated the diagnostic value of multiple-miRNA assay in bladder cancer detection, while 46 studies focused on single-miRNA assay. The specimens used by the included studies comprised voided urine, urine sediment, urine supernatant, serum, plasma and blood, which can be classified into 2 general types, “urine” and “blood”. Fifty studies employed urine-based specimens and the other 8 studies used blood-based specimens. Twenty-four studies selected Asian participants and 34 studies had Caucasian participants. The total numbers of bladder cancer patients and controls were 4,558 and 4,456, respectively. The publication years of the included articles ranged from 2010 to 2015. All studies applied quantitative real-time reverse transcription polymerase chain reaction (qRT-PCR) to detect the expression levels of miRNAs. Quality evaluation of the included studies with the QUADAS-2 tool is presented in Figure 2. The majority of studies selected in this meta-analysis fulfilled at least 4 items in QUADAS-2, implying good overall quality of the included studies.
Main characteristics of the 58 studies included in the meta-analysis
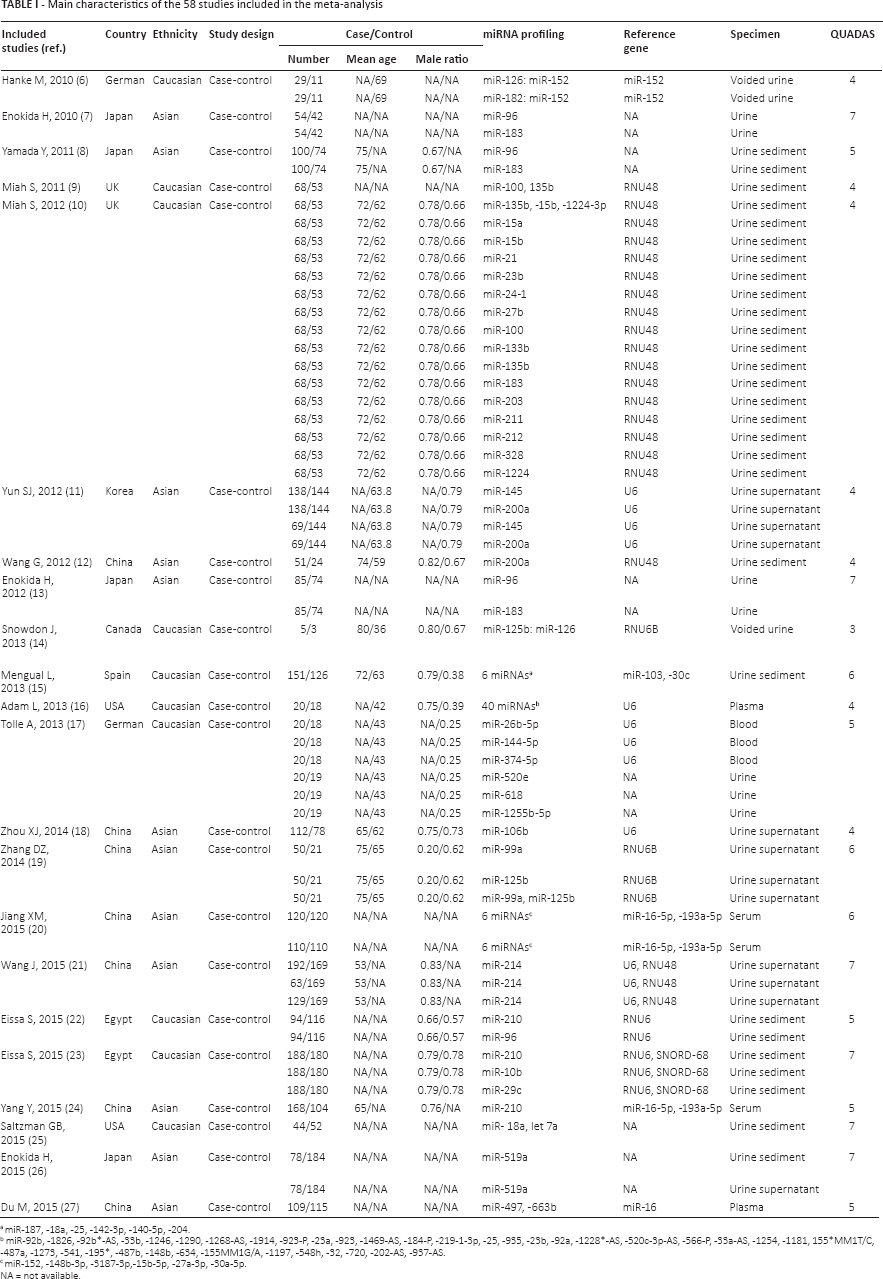
miR-187, -18a, -25, -142-3p, -140-5p, -204.
miR-92b, -1826, -92b*-AS, -33b, -1246, -1290, -1268-AS, -1914, -923-P, -23a, -923, -1469-AS, -184-P, -219-1-3p, -25, -935, -23b, -92a, -1228*-AS, -520c-3p-AS, -566-P, -33a-AS, -1254, -1181, 155*MM1T/C, -487a, -1273, -541, -195*, -487b, -148b, -634, -155MM1G/A, -1197, -548h, -32, -720, -202-AS, -937-AS.
miR-152, -148b-3p, -3187-3p,-15b-5p, -27a-3p, -30a-5p.
NA = not available.
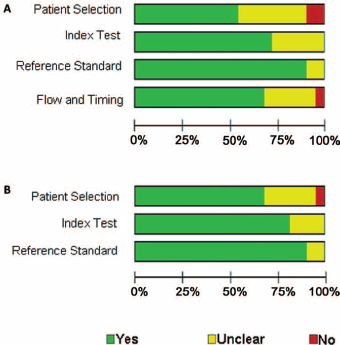
Quality assessment of included publications using QUADAS-2 criteria. (
Diagnostic Accuracy of miRNAs for Bladder Cancer
Because significant heterogeneity between studies was observed in sensitivity and specificity data (I2 = 85.91% and I2 = 84.36%, respectively) (p<0.01), the random-effects model was applied. As shown in Table II, the pooled parameters calculated from all 58 studies were as follows: sensitivity, 0.74 (95% confidence interval [CI]: 0.70-0.78); specificity, 0.78 (95% CI: 0.74-0.81); PLR, 3.3 (95% CI: 2.8-3.9); NLR, 0.34 (95% CI: 0.29-0.39); DOR, 10 (95% CI: 7-13) and AUC, 0.83 (95% CI: 0.79-0.86) (Fig. 3A), indicating that miRNAs in blood and urine may be qualified to distinguish bladder cancer patients from controls with moderate accuracy. Likewise, both the likelihood ratios and post-test probabilities were moderate, as shown in Fagan's plot (Fig. 3B). The PLR of 3 indicated that a person with bladder cancer was 3 times more likely to have a positive test result than a healthy person. Given a pre-test probability of 20%, the post-test probability of bladder cancer for a positive test result was 45%, while for a negative test result this was 8%. Moreover, the DOR of 10 indicated that miRNAs in blood and urine can be utilized as a good indicator of bladder cancer diagnosis.
Summary estimates of diagnostic criteria for miRNA profiling in bladder cancer detection
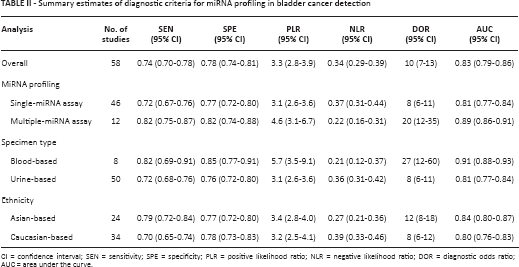
CI = confidence interval; SEN = sensitivity; SPE = specificity; PLR = positive likelihood ratio; NLR = negative likelihood ratio; DOR = diagnostic odds ratio; AUC = area under the curve.
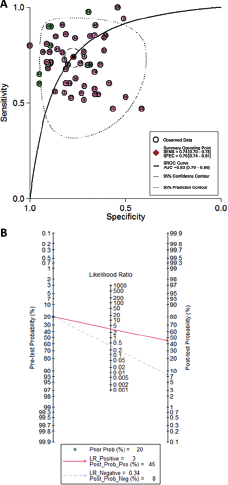
Diagnostic accuracy of miRNAs for bladder cancer. (
Subgroup Analysis
Subgroup analyses based on miRNA profiling (single- or multiple-miRNA), specimen types and ethnicity were conducted and the pooled results for diagnostic performance in different subgroups are displayed in Table II. Subgroup analysis based on miRNA profiling suggested that multiple-miRNA assays (Fig. 4A) had relatively better diagnostic performance than single-miRNA assays (Fig. 4B), with a sensitivity of 0.82 versus 0.72, a specificity of 0.82 versus 0.77, and an AUC of 0.89 versus 0.81 (Tab. II). Notably, the diagnostic properties of blood-based assays (Fig. 4C) were superior to those of urine-based assays (Fig. 4D), suggesting that blood may be a better matrix for miRNA detection. In addition, the diagnostic accuracy of Asian-based miRNA assays (Fig. 4E) might be higher than Caucasian-based ones (Fig. 4F).
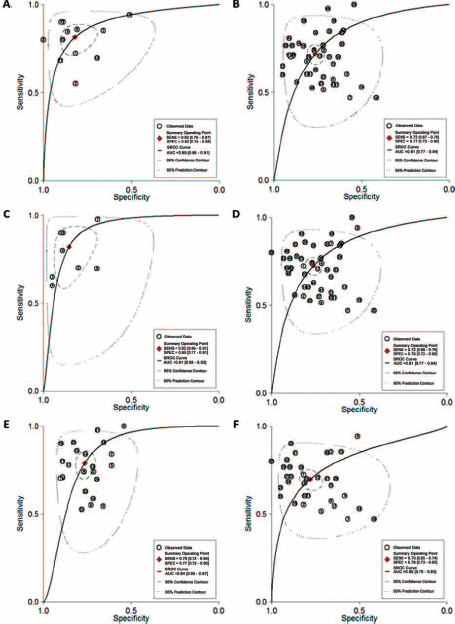
SROC curve with pooled estimates of sensitivity, specificity and AUC. (
Meta-Regression and Publication Bias
Meta-regression analysis was carried out to further investigate potential sources of the heterogeneity and to verify the results of subgroup analyses. As shown in Figure 5A, miRNA profiling, types of specimen and ethnicity may be the major sources of heterogeneity for miRNA assays in bladder cancer.
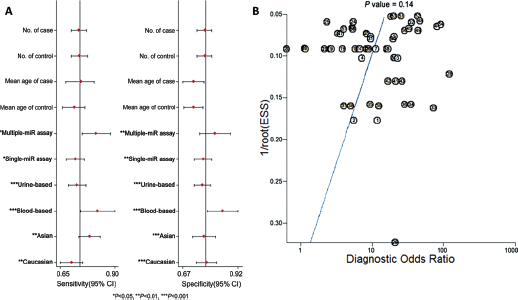
Meta-regression and publication bias. (
Deeks’ funnel plot asymmetry test was conducted to evaluate publication bias. As displayed in Figure 5B, the p value of 0.14 suggested there was no significant publication bias in our meta-analysis.
Discussion
Although random biopsy guided by cystoscopy is currently the most reliable screening means for bladder cancer, its invasive, unpleasant and inconvenient features as well as potential sampling error have severely hindered its wide clinical use. Urine cytology is relatively simple and specific but its diagnostic sensitivity is rather poor. Recently, there has been a vast growth of interest due to the feasibility of blood-based and urine-based miRNAs as biomarkers of bladder cancer (6-7-8-9-10-11-12-13-14-15-16-17-18-19-20-21-22-23-24-25-26-27). Because of the inconsistencies and discrepancies among these studies, we performed this meta-analysis to explore the potential of miRNAs as noninvasive diagnostic biomarkers in bladder cancer.
So far, our report is among the few evidence-based meta-analyses to testify the diagnostic importance of miRNAs in the early identification of bladder cancer, with a pooled AUC of 0.83 in distinguishing bladder cancer patients from controls (pooled sensitivity = 74%; pooled specificity = 78%), indicating the potential diagnostic value of miRNAs as noninvasive biomarkers. What is more, the DOR was 10 (95% CI, 7-13), implying that miRNA-test-positive individuals have a 10 times higher chance of bladder cancer than those with a negative test result.
There were 3 interesting findings in our meta-analysis. Firstly, multiple-miRNA assays displayed relatively better diagnostic performance than single-miRNA assays, with a sensitivity of 0.82 versus 0.72, a specificity of 0.82 versus 0.77, and an AUC of 0.89 versus 0.81. This demonstrates the advantage of applying panels of miRNAs to obtain a complete picture. It is universally acknowledged that, although detecting an individual cancer-related miRNA as a disease fingerprint is more straightforward and much simpler than employing a combination of miRNAs, the specificity of single-miRNA biomarkers is relatively poor. The molecular mechanism behind the limitation of single-miRNA biomarkers is that aberrant levels of single miRNAs might be associated with several different types of cancer (32). What is more, the development of cancer can be perceived as the result of a complex multistage process of epigenetic and genomic abnormalities, and should therefore be targeted by multiple miRNAs (33). Accordingly, it is rational to exploit panels of miRNAs instead of single miRNAs, thereby circumventing the limitations inherent in utilizing miRNAs as noninvasive biomarkers for cancer detection, especially in localized pathological conditions, when it is difficult to obtain regular biopsies. Secondly, the diagnostic properties of blood-based assays seem to be superior to those of urine-based ones, suggesting that blood may be a better matrix for miRNA detection. However, since only 8 of the included studies concerned blood-based miRNAs, the results need to be taken with caution. In addition, the diagnostic accuracy of Asian-based miRNA assays might be higher than that of Caucasian-based ones, implying that the value of miRNA in the diagnosis of bladder cancer may be different between ethnicities. This might be explained by different living circumstances and backgrounds and different genetic factors between the 2 ethnicities. The origin of the source-related and ethnicity-related differences is still unclear and may be explained by unknown mechanisms. Large-scale investigations are therefore needed in the future to ascertain whether these source-related and ethnicity-related differences truly exist.
Despite our efforts to conduct an accurate analysis, there are several limitations that need to be taken into account. First, no data have been published on African populations. In addition, it is crucial to recognize and exploit panels of miRNAs being able to differentiate cancer from other diseases with similar symptoms. Nevertheless, almost all of the studies included in this meta-analysis sought to distinguish bladder cancer patients from healthy controls. Third, because of their low amount and lack of known endogenous reference genes, accurate quantification of miRNAs in body fluids provides a real challenge for clinical application. Since the lack of conventions for accurate quantification of miRNAs in body fluids limits the cross-comparison of studies conducted by different laboratories, a standardized procedure should be established and preferably followed across all studies to minimize procedure-based bias. Moreover, there was statistical heterogeneity in our analysis ascribed to miRNA profiling, specimen types and ethnicity, which may have influenced the results. Finally, although some studies investigated correlations between miRNA levels and tumor stage and grade, and some even provided cutoff values and reference genes of qRT-PCR, subgroup analyses on the basis of these parameters were restricted due to the deficient data published.
Based on a comprehensive assessment of recent publications in bladder cancer, our meta-analysis demonstrated that miRNAs in blood and urine, especially panels of multiple miRNAs, may serve as promising minimally invasive tools for the diagnosis and monitoring of bladder cancer. Extensive prospective trials are required to verify their diagnostic feasibility in bladder cancer.
Footnotes
Financial support: None.
Conflict of interest: None.
