Abstract
Objective
Owing to inconsistency between reports, a meta-analysis was designed to appraise the clinical implications of long non-coding RNAs (lncRNAs) in urine and blood for the diagnosis of bladder cancer.
Methods
Studies that met the criteria were acquired by bibliographic retrieval through PubMed, Embase, and the Cochrane Library. The pooled diagnostic performance was evaluated by calculating the area under the summary receiver operator characteristic (SROC) curve. The potential sources of heterogeneity were approached through meta-regression and subgroup analyses. All statistical analyses and plots were performed by RevMan 5.3, Meta-DiSc 1.4, and STATA 12.0.
Results
A total of 43 studies from 15 articles consisting of 3370 bladder cancer patients and 3212 controls were incorporated in our meta-analysis. lncRNAs in urine and blood performed relatively well in diagnosing bladder cancer, with a pooled sensitivity of 0.78, a specificity of 0.79, and an area under the SROC curve (AUC) of 0.86. H19 displayed the best diagnostic accuracy with a pooled AUC of 0.90, followed by UCA1 and MALAT1. The heterogeneity among studies was partly conducted by sample size, lncRNA existence form (cell-free or intracellular lncRNA), lncRNA origin (exosome- or non-exosome-based lncRNA), lncRNA profiling (single- or multiple-lncRNA), specimen types, and ethnicity.
Conclusions
lncRNAs in urine and blood may serve as noninvasive diagnostic biomarkers with great promise for bladder cancer, while their clinical values need to be examined through further synthetic forward-looking studies.
Introduction
Bladder cancer is the 13th deadliest and 9th most common cancer worldwide. 1 The features are frequent recurrence and high mortality when the tumor progresses to the invasive stage. 2 The limitations of cystoscopy (invasive and hard-to-detect carcinoma in situ) and urine cytology (less sensitive for low-grade tumor) greatly affect their clinical application as diagnostic and surveillance tools for bladder cancer. 3 Thus, a probe into noninvasive biomarkers with high diagnostic value for bladder cancer was required.
lncRNAs (>200 nt long) play key roles in regulating chromatin dynamics, gene expression, growth, differentiation, and development, and are abundantly expressed in a variety of cancers. 4 Many studies have reported that lncRNAs could be stable in body fluids and serve as diagnostic noninvasive biomarkers for bladder cancer;5–19 however, difference existed among these studies in relation to the dependability of lncRNAs for detecting bladder cancer. Given this, we performed this meta-analysis to synthetically illuminate the diagnostic performance of circulating lncRNAs for bladder cancer based on published articles.
Materials and methods
Bibliographic retrieval
This meta-analysis was carried out in compliance with guidelines for diagnostic meta-analysis. 20 Qualified articles concerning the diagnostic significance of circulating lncRNAs for bladder cancer were acquired by systematic bibliographic retrieval of PubMed, Embase, and the Cochrane Library for research published in English until January 6, 2020. The following retrieval strategy was utilized: (“urinary bladder neoplasms” OR “bladder cancer” OR “bladder tumor” OR “transitional cell carcinoma of bladder” OR “bladder carcinoma” OR “urinary tract transitional cell carcinoma”) AND (“rna, long noncoding” OR “long noncoding rna” OR “lncrna”) AND (“blood” OR “serum” OR “plasma” OR “urine” OR “urinary” OR “cell-free” OR “circulating”) AND (“diagnosis” OR “sensitivity and specificity” OR “ROC curve”). Additional articles were acquired by manually searching reference lists of all relevant publications.
Inclusion and exclusion criteria
Two reviewers independently evaluated the eligible articles, while a third reviewer was consulted to arrive at consensus by multilateral discussion when any dispute arose. The inclusion criteria were: (a) histopathology was exploited to definitely diagnose patients with bladder cancer; (b) diagnostic value of lncRNAs in urine and blood for bladder cancer was investigated; and (c) adequate data could be extracted to reconstruct two-by-two tables comprising true positive (TP), false positive (FP), true negative (TN) and false negative (FN). The exclusion criteria were: (a) studies with insufficient data, duplicate data, or low quality; (b) review, meta-analysis articles, seminar articles, case reports, or meeting reports; and (c) studies irrelevant to the diagnostic performance of circulating lncRNAs for bladder cancer.
Data extraction
Standardized forms were adopted by two independent reviewers to collect the following information from all qualified studies: (a) basic features of studies, including first author, publication year, publication journal, country of studies, sample size, ethnicity, mean age, gender ratio, specimen type, cancer type, tumor stage, tumor grade, lncRNA profiling, assay methods, threshold and reference gene; and (b) diagnostic significance comprising TP, FP, FN, TN, sensitivity, specificity, and area under the curve (AUC).
Quality appraisal
Two reviewers systematically appraised and independently graded each included study in line with revised Quality Assessment of Diagnostic Accuracy Studies 2 (QUADAS-2) criteria 21 as we reported in a previous article. 22 Any dissent was resolved by discussing with a third reviewer to reach an agreement.
Statistical analysis
The I2 statistic and Q test were applied to estimate significant heterogeneity (I2 value ≥ 50% or P value < 0.10 for the Q test, then adopting the random-effects model) among the included studies. 23 Furthermore, the potential sources of heterogeneity were elucidated by subgroup analyses and meta-regression. Pooled sensitivity, specificity, positive likelihood ratio (PLR), negative likelihood ratio (NLR) and diagnostic odds ratio (DOR) of all included studies were calculated by employing bivariate meta-analysis model. The sensitivity and specificity of each included study were utilized to draw the summary receiver operator characteristic (SROC) curve with AUC indicating pooled diagnostic efficacy. The post-diagnostic effect after pooled analysis was assessed by Fagan’s nomogram. The publication bias was evaluated by Deeks’ funnel plot asymmetry test with P < 0.10 demonstrating significant difference. 24 All analyses and drawings were carried out by the RevMan 5.3 (Nordic Cochrane Centre, Copenhagen, Denmark), Meta-DiSc 1.4 (Ramony Cajal Hospital, Madrid, Spain), and STATA 12.0 (StataCorp LP, College Station, TX, USA).
Results
Bibliographic retrieval
Supplementary Figure 1 indicates the process of document retrieval. Through an initial database search, 63 potential articles were acquired, of which duplicates (6) were ruled out. There were 12 reviews and meta-analysis; 3 articles were not about bladder cancer; and 20 articles were unconnected to our topic (regarding mRNA, molecular mechanism or prognosis, etc.) were further excluded through evaluation of their abstracts. Then, 22 articles were kept for the full-text review, of which 7 articles were eliminated (2 articles unrelated to diagnosis, 4 articles without enough data, and 1 article about tissue-lncRNA assay). Eventually, 15 articles that qualified were included in this meta-analysis, comprising 5 articles concerning blood-based lncRNAs10,11,15,16,18 and 10 involving urine-based ones.5–9,12–14,17,19
Essential features of included studies
Table 1 demonstrates the general characteristics of the included articles. A total of 43 studies from 15 articles published from 2006 to 2019 were included in our meta-analysis, including 3212 controls and 3370 bladder cancer patients; 21 studies involved Caucasian participants and 22 studies involved Asian participants. One study assessed the diagnostic value of circulating lncRNAs in muscle invasive bladder cancer (tumor stage T2–T4); and 26 studies involved both non-muscle invasive bladder cancer (tumor stage Tis, Ta–T1) and muscle invasive bladder cancer. One study referred to high-grade bladder cancer, and 36 involved both low- and high-grade ones. Only 11 studies exploited multiple-lncRNA assays in bladder cancer detection; 32 applied a single-lncRNA assay, of which 12 studies concerned the diagnostic significance of circulating single UCA1 in bladder cancer; 5 studies involved single MALAT1; and 5 studies referred to single H19. Thirty studies explored cell-free lncRNAs and 13 studies probed into intracellular lncRNAs. Quantitative real-time reverse transcription polymerase chain reaction (qRT-PCR) was utilized to detect the expression level of lncRNAs in all included studies, 42 studies of which normalized the lncRNA concentration to a reference gene, and 1 study normalized to a standard curve. The specimens used by included articles consisted of urine sediment, serum, serum exosomes, urine supernatant, urine exosomes, and plasma exosomes, which could be classified into two basic types: “blood” and “urine.” Eleven studies investigated blood-based specimens and the other 32 studies focused on urine-based specimens. lncRNAs derived from exosomes were examined in 22 studies, while lncRNAs not originating from exosomes were examined in 21 studies. As indicated in Supplementary Figure 2, the majority of studies adopted in this meta-analysis met at least four items in the QUADAS-2 tool, demonstrating the good overall quality of the included studies.
Essential features of 43 studies included in the meta-analysis.

lncRNA: long non-coding RNA.
aInc-MEG3, SNHG16, MALAT1
blnc-uc004cox.4, GAS5
clnc-MALAT1, PCAT-1, SPRY4-IT1
dlnc-PCAT-1, UBC1, SNHG16
elnc-UCA1-201, UCA1-203, MALAT1, LINC00355
NA: not available; SCC: squamous cell carcinoma; TCC: transitional cell carcinoma; UCB: urothelial carcinoma of the bladder.
Diagnostic efficacy of circulating lncRNAs for bladder cancer
Since there was significant heterogeneity among studies in sensitivity (I2 = 77.47%) and specificity (I2 = 78.02%) (P < 0.01), the random-effects model was exploited. As presented in Table 2, the pooled parameters worked out from all 43 studies were: sensitivity, 0.78 (95% CI 0.75, 0.81); specificity, 0.79 (95% CI 0.76, 0.83); PLR, 3.8 (95% CI 3.2, 4.5); NLR, 0.27 (95% CI 0.23, 0.32); DOR, 14 (95% CI 11, 19); and AUC, 0.86 (95% CI 0.82, 0.89) (Figure 1(a)), signifying that lncRNAs in urine and blood may serve as a good diagnostic index for bladder cancer with moderate accuracy. As demonstrated in the Fagan’s plot (Figure 1(b)), the pre-test probability was 51%, the post-test probability of bladder cancer for a positive test result was 80%, while the negative test result was 22%, implying that both the post-test probabilities and likelihood ratios were moderate. The PLR of 3.8 displayed that a person with bladder cancer is 3.8-times more likely to have a positive test result than a healthy person. Furthermore, the DOR value was 14 (95% CI 11, 19), which revealed that lncRNAs in urine and blood can be qualified to distinguish bladder cancer patients from controls. Besides, circulating single H19 exhibited the best diagnostic accuracy with pooled AUC of 0.90, followed by UCA1 with AUC of 0.86, and MALAT1 with AUC of 0.76 (Table 2 and Figure 2). Supplementary Figure 3 displays the weight and sensitivity of each study with the pooled sensitivity of 0.78 (95% CI 0.75, 0.81) (P < 0.001).
Summary estimates of diagnostic performance for lncRNAs profiling in bladder cancer detection.
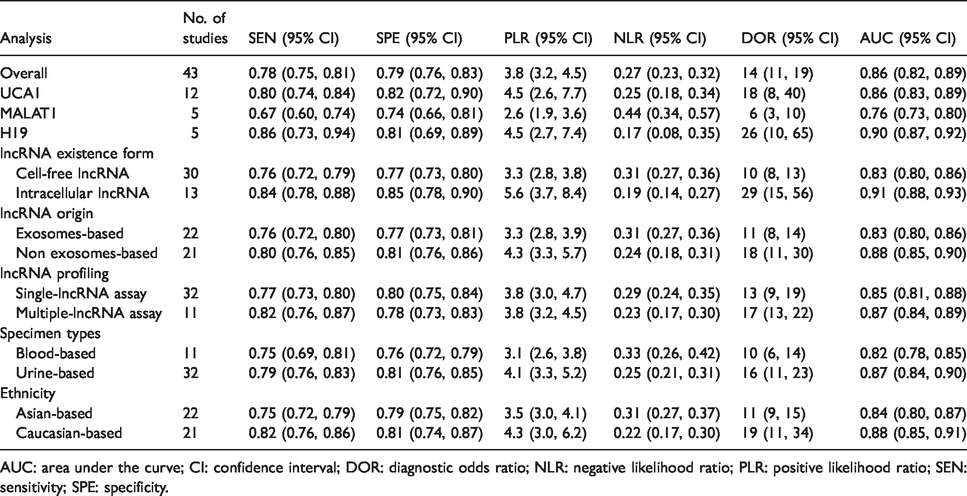
AUC: area under the curve; CI: confidence interval; DOR: diagnostic odds ratio; NLR: negative likelihood ratio; PLR: positive likelihood ratio; SEN: sensitivity; SPE: specificity.
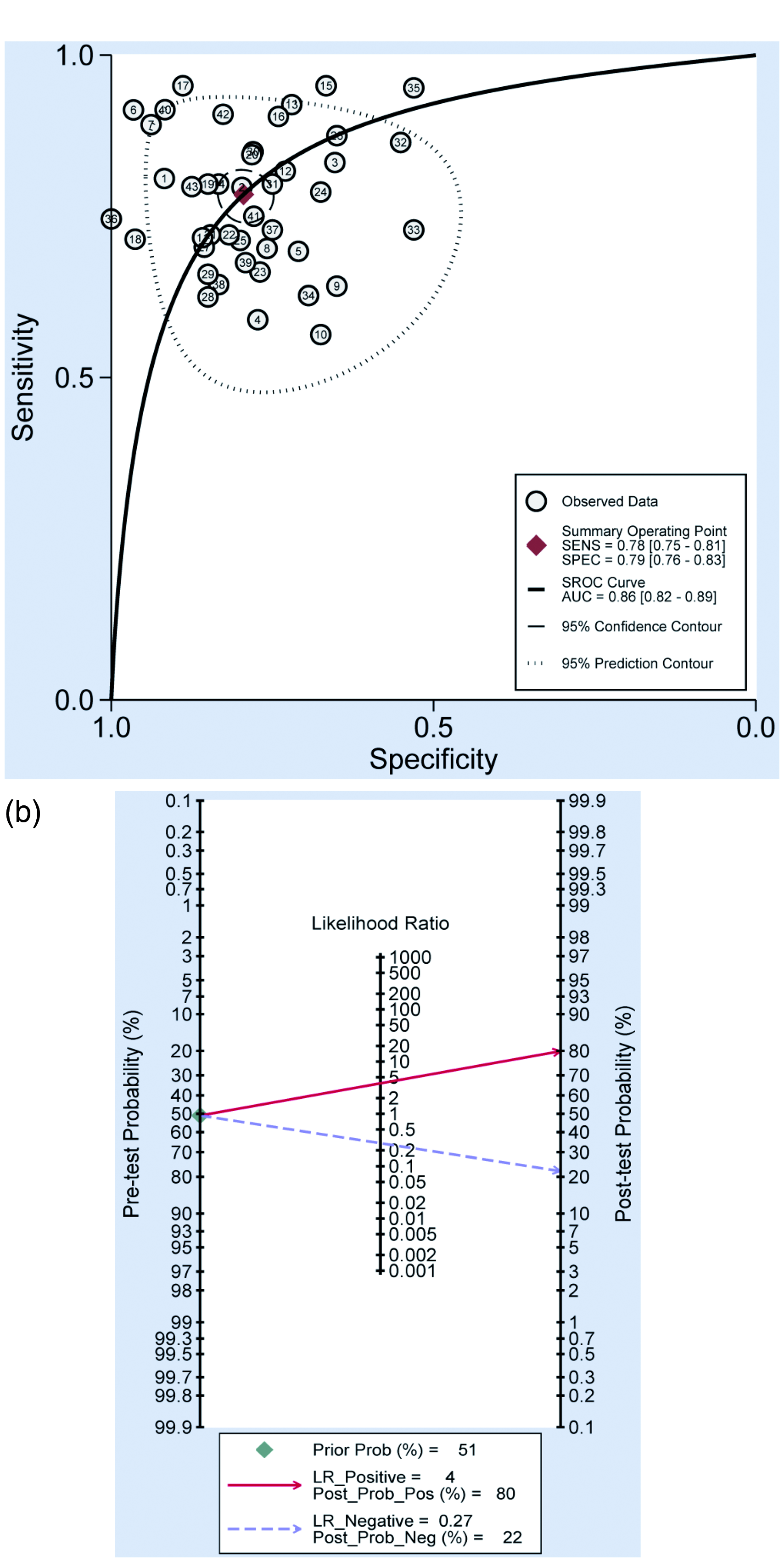
Diagnostic performance of circulating lncRNAs for bladder cancer. (a) SROC curve with pooled estimates of sensitivity, specificity and AUC of overall studies. (b) Fagan’s nomogram for evaluation of post-test probabilities based on pooled estimates of PLR and NLR of overall studies.
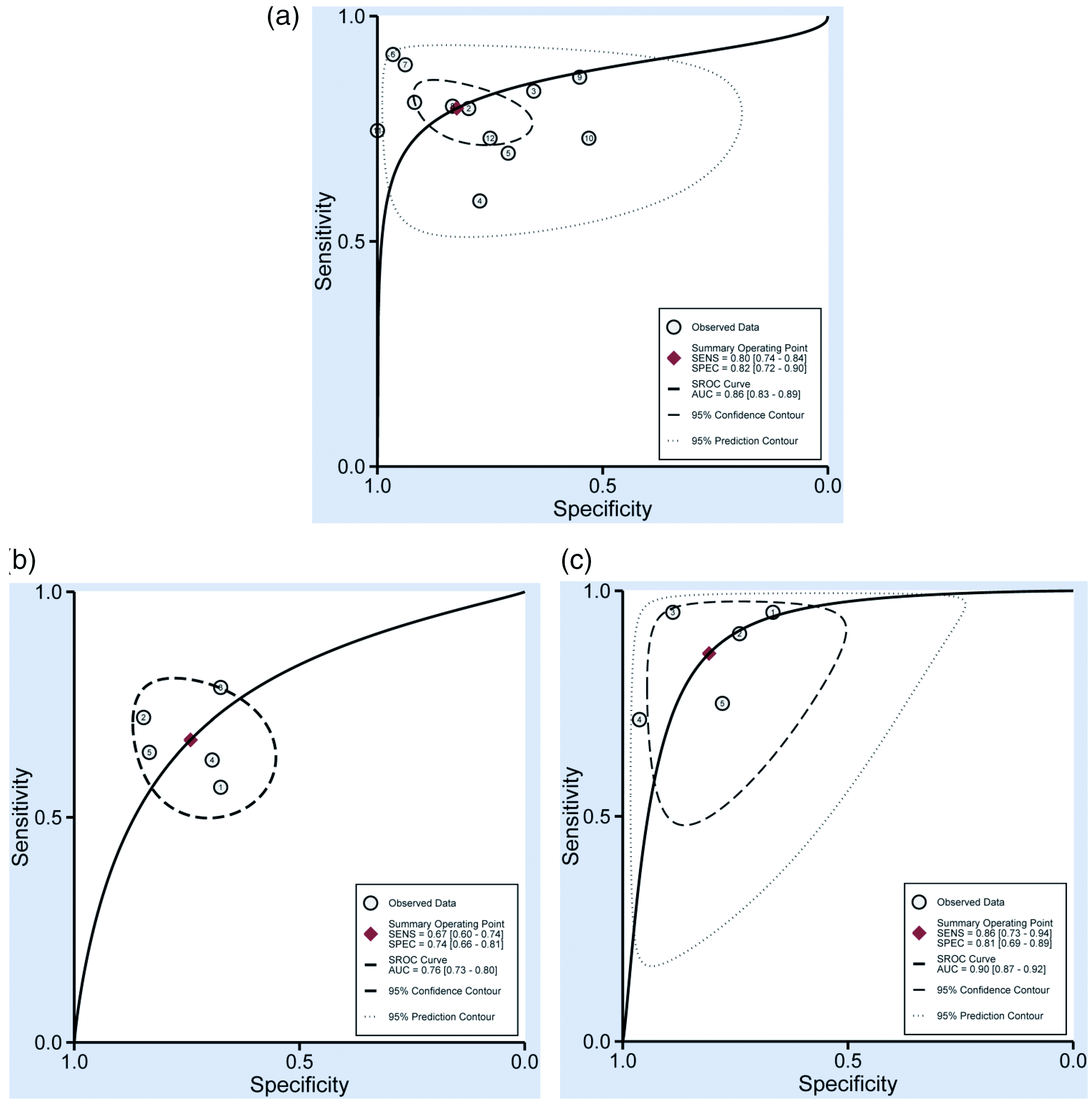
SROC curves based on diagnostic studies of (a) UCA1, (b) MALAT1, and (c) H19.
Subgroup analysis
Subgroup analyses on the basis of lncRNA existence form (cell-free or intracellular lncRNA), lncRNA origin (exosome- or non-exosome-based lncRNA), lncRNA profiling (single- or multiple-lncRNA), specimen types, and ethnicity were performed separately. The pooled results for diagnostic value in different subgroups are shown in Table 2. Cell-free lncRNAs generated a pooled AUC of 0.83, while intracellular lncRNA gave rise to a pooled AUC of 0.91. lncRNAs originating from exosomes exhibited good diagnostic accuracy with a pooled AUC of 0.83, while that not from exosomes had a pooled AUC of 0.88. Diagnostic performance of multiple-lncRNAs assays versus single ones was 0.87 versus 0.85 for AUC. Diagnostic property of urine-based assays versus blood-based ones was 0.87 versus 0.82 for a pooled AUC. In addition, Asian- and Caucasian-based lncRNA assays manifested a pooled AUC of 0.84 and 0.88, respectively.
Meta-regression and publication bias
The potential sources of heterogeneity were further explored through meta-regression analysis. As displayed in Figure 3(a), sample size, lncRNA existence form, lncRNA origin, lncRNA profiling, specimen types, and ethnicity seemed to be the primary sources of heterogeneity for lncRNAs assays in bladder cancer. The publication bias was investigated by the Deeks’ funnel plot asymmetry test and there was no significant publication bias (P = 0.38) in our meta-analysis (Figure 3(b)).
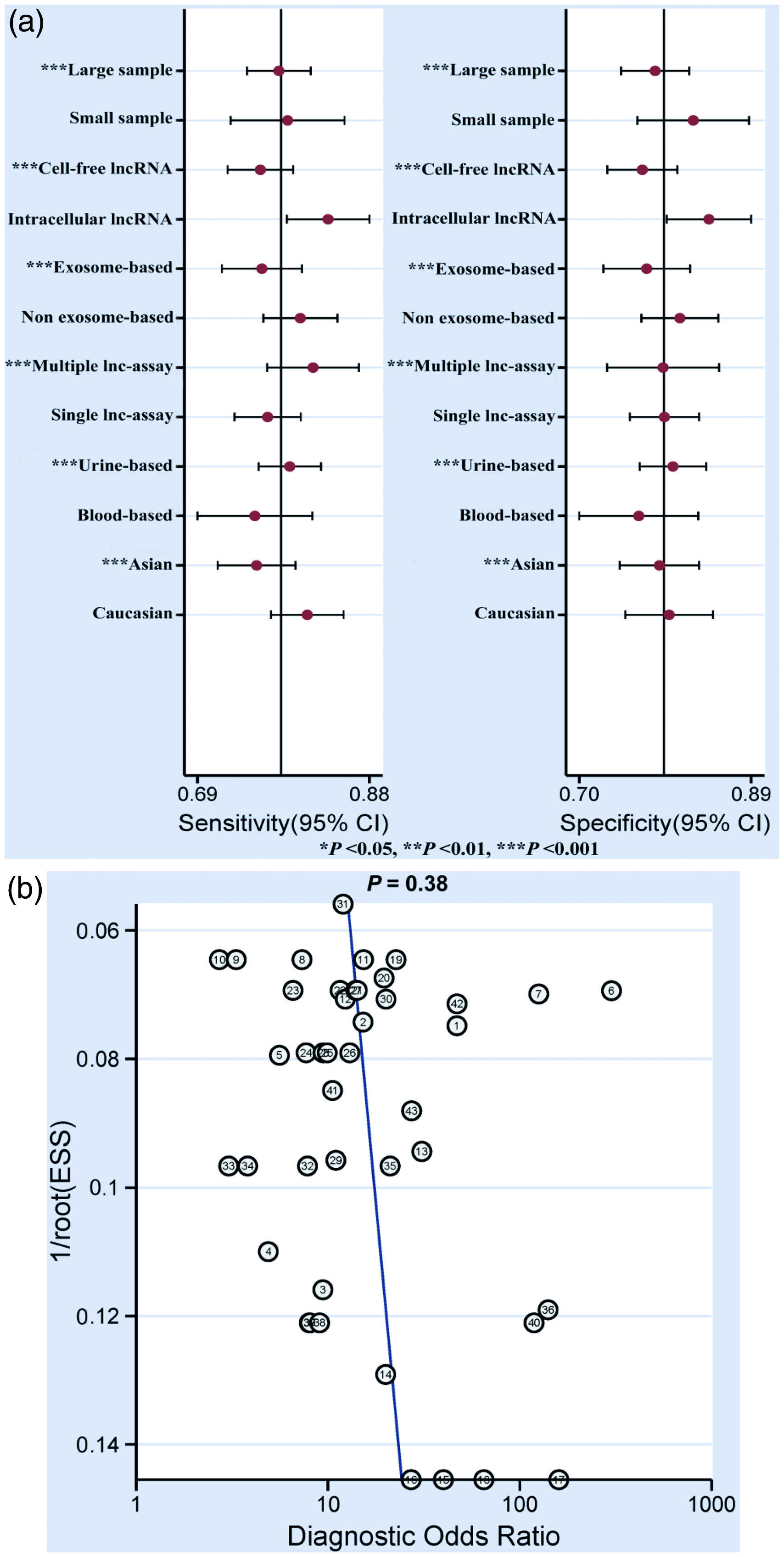
Meta-regression and publication bias based on overall studies. (a) Forest plots of multivariable meta-regression analyses for sensitivity and specificity (vertical lines signify pooled estimates of sensitivity and specificity respectively). (b) Deeks’ funnel plot asymmetry test.
Discussion
Cystoscopy and urine cytology serve as the most commonly utilized diagnostic and surveillance strategies for bladder cancer at present; however, they have been proven to be far from clinically satisfactory. The minimally invasive “liquid biopsy markers” (including cell-free DNA, circulating tumor cells, circulating RNAs, cell-free proteins, peptides, and exosomes) hold great promise for personalized medicine in bladder cancer owing to their ability to provide multiple non-invasive global snapshots of the primary and metastatic tumors. 25 Recently, there has been a growing trend towards investigating the potential of circulating lncRNAs as biomarkers for bladder cancer.5–19 In light of discrepancies existing among these studies, we carried out this meta-analysis to probe into the possibility of circulating lncRNAs as noninvasive diagnostic biomarkers in bladder cancer.
As one of the hitherto few evidence-based meta-analysis expounding the diagnostic implication of circulating lncRNAs in identifying bladder cancer, our report verified the potential diagnostic performance of lncRNAs in blood and urine as noninvasive biomarkers for differentiating between bladder cancer and controls with a pooled AUC value of 0.86 (pooled sensitivity = 78%; pooled specificity = 79%). Moreover, the DOR value of 14 (95% CI 11, 19) signified that circulating lncRNA test-positive patients have a 14-times higher chance of bladder cancer compared to controls.
Twelve studies concerned the diagnostic value of circulating UCA1 in bladder cancer, 5 studies involved single MALAT1, and 5 studies referred to single H19; therefore, independent meta-analyses of these three lncRNAs were conducted separately. Circulating single H19 did best with a pooled AUC of 0.90, a sensitivity of 0.86, a specificity of 0.81, a PLR of 4.5, an NLR of 0.17, and a DOR of 26, exhibiting relatively high diagnostic accuracy. Also, H19 even displayed relatively higher value of sensitivity, specificity, PLR, DOR, and AUC than overall lncRNAs and a subgroup of single-lncRNA assay. Likewise, single UCA1 and single MALAT1 also manifested satisfying diagnostic efficacy with an AUC of 0.86 and 0.76, respectively. The above-mentioned results indicated that single H19, UCA1, and MALAT1 showed their potential to serve as powerful noninvasive diagnostic biomarkers for bladder cancer.
Then, subgroup analyses and meta-regression analysis were carried out to explore the potential sources of the heterogeneity. Our results demonstrated that both pooled sensitivity and specificity were influenced by sample size, lncRNA existence form, lncRNA origin, lncRNA profiling, specimen types, and ethnicity, manifesting that factors discussed above may be the principal sources of heterogeneity for lncRNAs assay in bladder cancer. The existence form of lncRNAs seemed to contribute to the heterogeneity for circulating lncRNA assays, and the diagnostic property of intracellular lncRNAs (all from urine sediment) was 0.91 for pooled AUC. Since lncRNAs in urine sediment mainly mirrored intracellular expression from exfoliated tumor cells, renal tubular cells, normal urothelial cells, red blood cells, and white blood cells, heterogeneous cellular components may influence their repeatability as dependable biomarkers. Meanwhile, as only 13 included studies involved intracellular lncRNAs, the results should be treated with discretion. Furthermore, lncRNAs originating from exosomes exhibited a pooled AUC of 0.83. As lipid membrane-bound satchels (40–150 nm) with a multivesicular endosomal origin, exosomes are secreted by both normal and neoplastic cells to mediate cell-to-cell communication, contain soluble materials (such as proteins, nucleic acids, and lipids), and hence protect the latter from degradation in body fluid.26,27 lncRNA profiling also had an impact on both the pooled sensitivity and specificity of circulating lncRNAs assays. Diagnostic accuracy of multiple-lncRNA assays was 0.87 for pooled AUC, elucidating the strength of utilizing panels of lncRNAs to get the whole picture. As presented in Table 1, MEG3, SNHG16, and MALAT1 from serum; 10 uc004cox.4 and GAS5 from urine supernatant; 13 MALAT1, PCAT-1, and SPRY4-IT1 from urine exosomes; 14 PCAT-1, UBC1, and SNHG16 from serum exosomes; 16 as well as 4-lncRNAs panel from urine exosomes 17 performed well in diagnosing bladder cancer. Nevertheless, because there were just 11 included studies regarding multiple-lncRNA assay, it is best to treat the results cautiously. The molecular mechanism underlying the weakness of a single-lncRNA biomarker may be that abundantly expressed single lncRNA might be widely linked to a variety of cancers. 4 Incidentally, now that the development of cancer can be deemed as the result of an intricate multi-stage process of genomic and epigenetic abnormalities, it also should be targeted by multiple lncRNAs. Moreover, diagnostic performance of urine-based assays was 0.87 for pooled AUC, which may be because urine is a particularly satisfying source of biomarkers for bladder cancer for the following reasons: urine is far more convenient and less invasive to obtain, and contains lower levels of proteins than blood, thus reducing protein interference during RNA isolation. 3 Since urine is likely to pass through the malignant lesions in the urologic system before being discharged, its lncRNA content may be specifically linked to the disease status (tumorigenesis, metastasis, and tumor stage) as displayed by tumor tissues. 4 Last but not least, ethnicity seemed to be another possible source of heterogeneity, generating pooled AUC of 0.88 for Caucasian-based lncRNA assays and 0.84 for Asian-based ones, which might be attributed to different hereditary factors and living backgrounds between the two races. Since the origin and mechanism of ethnicity-related discrepancy are still unclear, it is necessary to perform extensive investigations to further certify whether this discrepancy really exists.
In a meta-analysis on lncRNA and bladder cancer from 2018, Quan et al. 28 analyzed 13 articles on clinicopathological characteristics, 6 articles on diagnosis, and 16 articles on prognosis. They evaluated the diagnostic performance of lncRNAs for bladder cancer based on 8 studies from 6 articles (4 articles on urine, 2 articles on serum) containing 725 bladder cancer patients and 510 healthy controls, and demonstrated that pooled sensitivity, specificity, and AUC of all included lncRNAs were 0.73, 0.78, and 0.82, respectively. However, our report aimed to assess the diagnostic value of circulating lncRNAs from urine sediment, urine supernatant, urine exosomes, serum, serum exosomes, and plasma exosomes, which was different from the Quan et al. 28 meta-analysis. Also, they did not perform subgroup analyses of lncRNA existence form (cell-free or intracellular lncRNA), lncRNA origin (exosome- or non-exosome-based lncRNA), and lncRNA profiling (single- or multiple-lncRNA), which renders their report less comprehensive than ours.
Our best efforts were employed to carry out an accurate analysis; however, our report is far from perfect. Above all, since significantly statistical heterogeneity existed in our analysis on account of sample size, lncRNA existence form, lncRNA origin, lncRNA profiling, specimen types, and ethnicity, it was inevitable that the results would be influenced. Furthermore, it is essential to discern and utilize panels of lncRNAs capable of differentiating between cancer and other diseases with similar symptoms. However, most of the studies included in this meta-analysis just sought to distinguish bladder cancer patients from healthy controls, and did not involve patients with diseases of similar symptoms to bladder cancer. In addition, there is no relevant data published regarding African populations. Moreover, clinical application of circulating lncRNAs is severely limited because it is difficult to acquire accurate quantification due to their low abundance in body fluids and a lack of acknowledged endogenous reference genes. Now that cross-comparison among studies conducted by different laboratories cannot be achieved without convention for accurate quantification of circulating lncRNAs, it is sensible to set up a standardized procedure, which, preferably, should be complied with across all studies to minimize procedure-based bias. Lastly, although some studies provided information of tumor stage and grade, some even supplied reference genes and cut-off values of qRT-PCR; however, subgroup analyses based on these parameters were restricted owing to insufficient published data.
Supplemental Material
sj-pdf-1-jbm-10.1177_1724600820926685 - Supplemental material for Circulating lncRNAs as noninvasive biomarkers in bladder cancer: A diagnostic meta-analysis based on 15 published articles
Supplemental material, sj-pdf-1-jbm-10.1177_1724600820926685 for Circulating lncRNAs as noninvasive biomarkers in bladder cancer: A diagnostic meta-analysis based on 15 published articles by Jinfeng Wang, Yongli Gao, Xiaohua Wang, Yisheng Gao, Luning Li, Jinling Zhang, Lining Zhang and Fengyuan Che in The International Journal of Biological Markers
Footnotes
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by China Postdoctoral Science Foundation (grant number 2018M642671), Shandong Provincial Key Research and Development Project (grant number 2019GSF108275) and Medicine and health science technology development plan of Shandong Province (grant number 2017WS323).
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
