Abstract
Purpose
To compare circulating tumor cell (CTC) detection rates in patients with early (M0) and metastatic (M+) breast cancer using 2 positive-selection methods or size-based unbiased enrichment.
Methods
Blood collected at baseline and at different times during treatment from M0 patients undergoing neoadjuvant therapy and from M+ women starting a new line of treatment was processed in parallel using AdnaTest EMT-1/ and EMT-2/Stem CellSelect/Detect kits or ScreenCell Cyto devices. CTC positivity was defined according to the suggested cutoffs and cytological parameters, respectively.
Results
Higher CTC detection rates were obtained with the AdnaTest approach when using for CTC-enrichment antibodies against ERBB2 and EGFR in addition to MUC1 and the classical epithelial surface marker EPCAM (13% vs. 48%). In M0 patients mainly, CTC positivity rates further increased when EMT- and stemness-related marker expression (PIK3CA, AKT2 and ALDH1) was evaluated in addition to EPCAM, MUC1 and ERBB2. When the physical properties of tumor cells were exploited, CTCs were detected at higher percentages than with positive-selection-based methods, without any difference between clinical stages (78% in M0 vs. 72% in M+ cases at baseline). Circulating tumor microemboli (CTMs) were detected in addition to single CTCs with significantly higher frequency in M0 than M+ samples (78% vs. 27%, p = 0.0002).
Conclusions
Different approaches for CTC detection probably identify distinct tumor cell subpopulations, but need technical standardization before their clinical validity and biological specificity may be adequately investigated. The distinct role of CTMs compared with CTCs as prognostic and predictive biomarkers represents a further challenge.
Introduction
The number of new methods for circulating tumor cell (CTC) detection is rising vertiginously, but few products or devices designed for CTC evaluation have provided robust and independently validated clinical results (1). The CellSearch™ (Janssen Diagnostics, LCC, Raritan, NY, USA) approach was granted FDA approval for CTC detection and monitoring in metastatic solid tumors, but although it produced ample evidence on the prognostic role of CTCs in metastatic (2) and early breast tumors (3-5), CTC enrichment with this approach is limited to epithelial cells and no CTCs are found in a variable rate (35%-55%) of patients with metastatic disease.
In large trials such as SWOG500 (6), a change of treatment based on CTC status did not benefit the patient, which questions the actual biological role of the CTC subpopulation defined according to the CellSearch criteria. Different explanations have been given for such a failure (7), including adequacy of treatment, but maybe it is time to concentrate our efforts on the approx. 50% CTC-negative subset of patients with metastatic breast cancer, which might contain a CTC population undetectable with CellSearch (8). The present work focuses on the issue of going beyond the simple presence/enumeration of CTCs so that the molecular peculiarities of “missed” CTCs can be identified and further pieces of clinically important information can be unraveled.
Methods
Case Series
Patients with histologically confirmed breast cancer and no evidence of metastatic disease (M0) who were receiving anthracycline/taxane-based neoadjuvant therapy and trastuzumab if HER2 positive, and breast cancer patients with metastatic disease (M+) who were starting a new line of systemic treatment (mostly endocrine treatment) were prospectively recruited at the Department of Medical Oncology at Fondazione IRCCS Istituto Nazionale dei Tumori, Milan (INT).
Ethical Statement
The study was approved by the ethics committee of INT and written consent was obtained from all patients.
Blood Sample Collection
Samples of peripheral venous whole blood were drawn from all patients using a 26 G needle and collected in K3EDTA or K2EDTA BD Vacutainer tubes, if processed with the AdnaTest (AdnaGen, AG, Langenhagen, Germany) or ScreenCell® Cyto (ScreenCell, Paris, France) kits, respectively. The first blood tube was discarded to minimize the risk of contamination with skin epithelial cells. Samples were stored in the dark at 4°C and processed within 1 hour according to the manufacturer's instructions. For each patient, samples were collected at baseline (before starting treatment) and during treatment at established times: 3 and 6 months after initiation of neoadjuvant treatment and around 4 weeks after mastectomy in M0 women, and at the beginning of a new line of treatment and at 3 months from treatment start or at progression in M+ women.
AdnaTest Approach
CTC Positive Enrichment
CTC enrichment with the AdnaTest EMT-1/Stem CellSelect (EMT1) and EMT-2/Stem CellSelect (EMT2) kit was performed in parallel on two 5-mL blood samples from the same patient according to the manufacturer's instructions.
CTC Detection
For CTC detection, the expression levels of EPCAM, MUC1, ERBB2, PIK3CA, AKT2, TWIST1 and ALDH1 transcripts were evaluated by semiquantitative multiplex PCR according to the manufacturer's instructions and using the suggested thresholds for positivity (0.15 ng/µL for EPCAM, MUC1, ERBB2 and ALDH1 and 0.25 ng/µL for PIK3CA, AKT2 and TWIST1). Samples with ACTB (beta-actin) expression <0.70 ng/µL were excluded from the analysis, whereas those passing the quality control criteria where considered as positive for at least 1 of the above-mentioned markers.
ScreenCell Cyto Approach
CTC Enrichment by Size
All blood samples were processed within 1 hour after collection using the ScreenCell® Cyto kit (ScreenCell, Paris, France) according to the manufacturer's instructions (9). Briefly, blood was diluted in 4 mL of red blood cell lysis and fixation buffer and incubated 8 minutes at room temperature. Three filtrations of 3 mL of blood each were separately performed for each patient; microporous membranes were rinsed with PBS, collected from the device, air-dried and immediately stained at room temperature for 1 minute with Hematoxylin Solution S (Merck, Darmstadt, Germany) and then for 30 seconds with Shandon Eosin Y Aqueous Solution (Thermo Fisher Scientific Inc., Waltham, MA, USA). Microporous membranes were stored at −20°C until cytological evaluation by a certified pathologist (JW).
Cytomorphological Analysis and CTC Counts
All membranes were analyzed by the same pathologist (JW) without knowledge of the clinical data. Major criteria for CTC identification were a high nuclear-to-cytoplasmic ratio (≥0.75) and large nuclear size (≥20 µm), whereas minor criteria included irregular nuclear contours and nuclear hyperchromatism. The cytomorphological analysis and CTC count were based on the previously reported criteria of malignancy (10). Circulating tumor microemboli (CTMs) were defined as clusters of at least 2 CTCs, often mixed with platelets and various leukocytes, showing criteria of malignancy like those described for single CTCs. The nuclear-to-cytoplasmic ratios between single CTCs and CTC aggregates are similar (11).
Results were expressed as numbers of CTCs and CTMs for single membranes. For each patient, total CTC or CTM numbers derived from 3 membranes (corresponding to 9 mL of blood) were added together to better meet the criteria for accurate detection of rare events following the Poisson probability distribution (12). Membranes showing poor quality of cytology, estimated on the basis of poor preservation of the leukocytes, were excluded from the analysis. Samples were rated as CTC or CTM positive if at least 1 CTC or CTM was detected in the 3 membranes.
Statistical Analysis
Data were analyzed using conventional descriptive statistics. Contingency tables were analyzed by Fisher's exact test and the quantitative measure of the agreement between categorical variables was evaluated by Cohen's kappa statistics, κ. Differences among ordinal variables were assessed by Mann-Whitney's U test. Two-tailed p≤0.05 was adopted as the significance threshold.
Results
The AdnaTest EMT-1/Stem Cell Select + Detect (hereafter referred to as EMT1) and the AdnaTeast EMT-2/Stem Cell Select + Detect (hereafter referred to as EMT2) kits were used for a head-to-head comparison of CTC identification methods using 2 distinct blood samples from the same patient.
Data on CTC status were separately analyzed for 21 blood samples derived from 13 nonmetastatic breast cancer patients (M0) undergoing neoadjuvant treatment with conventional anthracycline/taxane schemes, and for 9 samples derived from 6 women with metastatic disease (M+) who were starting systemic treatment.
An increase in the percentage of samples defined as CTC positive was observed in samples processed using the EMT2 kit, where immunomagnetic enrichment of CTCs takes advantage of the addition of antibodies against EGFR and ERBB2 besides the classical EPCAM/MUC1 antibodies employed in the EMT1 kit (Tab. I). In the overall series, CTC positivity rates roughly passed from 13% to 47%. Such an increase in the number of samples defined as CTC positive was also observed by stratifying patients according to the clinical setting.
CTC positivity in blood samples processed in parallel with EMT1 and EMT2 kits
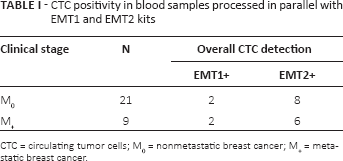
CTC = circulating tumor cells; M0 = nonmetastatic breast cancer; M+ = metastatic breast cancer.
Consistent with the above reported observations, there was indeed poor overall agreement between EMT1 and EMT2 on CTC status (κ = 0.159; 95% confidence limits −0.159-0.417). Only 60% of samples were concordantly rated for CTC status. The discordant samples were enriched in EMT1-/EMT2+ cases (11 EMT1-/EMT2+ versus 1 EMT1+/EMT2-).
These results clearly show that, by using the EMT1 kit, 10 samples (6 M0 and 4 M+) would have been classified as CTC negative. Such missed CTCs, whose clinical relevance needs to be defined, deserve careful molecular characterization.
To comply with the reported heterogeneity of the CTC population (8, 13), different CTC subpopulations identified by distinct gene expression patterns were separately evaluated. To this end, besides the classical epithelial-like/tumor-associated phenotype for CTC definition (EPCAM, MUC1, ERBB2), we also considered cell subpopulations expressing EMT-related genes (PIK3CA, AKT2, TWIST1) and stemness-related genes (ALDH1).
Overall, considering the optimal CTC selection with EMT2, an increase in positivity rates was observed when adding the expression of EMT-related genes (from 20% to 40%, NS), which reached 47% when expression of ALDH1 was also included (Tab. II). The identification of CTCs including genes associated with EMT and with stemness, compared with epithelial-like and tumor-associated markers only, increased the positivity rates in the overall population, although the increase was more evident in M0 than M+ patients.
EMT2 CTC positivity rates according to different CTC identification criteria
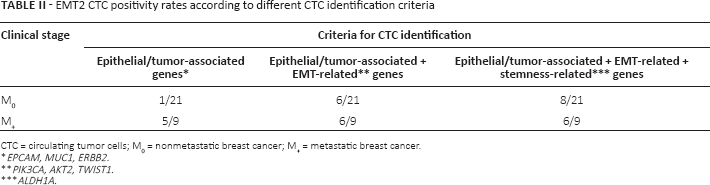

CTC = circulating tumor cells; M0 = nonmetastatic breast cancer; M+ = metastatic breast cancer.
EPCAM, MUC1, ERBB2.
PIK3CA, AKT2, TWIST1.
ALDH1A.
In a distinct case series of 14 M0 and 18 M+ breast cancers, CTCs and CTMs were identified based on cytological criteria only, after unbiased CTC enrichment by size-selection. The results, expressed as total number of CTCs and CTMs in 9 mL of blood, are reported in Table III using 1 CTC and 1 CTM in 9 mL of blood as positivity cutoff.
CTC and CTM positivity rates in samples processed by size-based selection
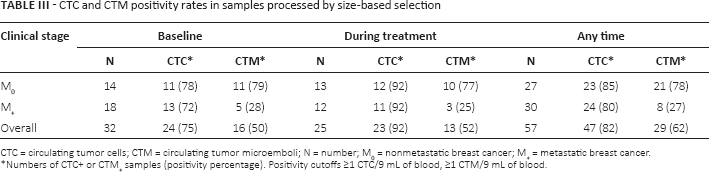
CTC = circulating tumor cells; CTM = circulating tumor microemboli; N = number; M0 = nonmetastatic breast cancer; M+ = metastatic breast cancer.
Numbers of CTC+ or CTM+ samples (positivity percentage). Positivity cutoffs ≥1 CTC/9 mL of blood, ≥1 CTM/9 mL of blood.
CTCs were identified in a high percentage of samples, ranging from 72% to 92% without any differences between clinical stages and between samples collected at baseline or during treatment. CTMs, too, were present at similar rates in samples derived from untreated or treated women; however, overall the CTM presence rates were different (p = 0.0002) between M+ (27%) and M0 (78%) patients, and the difference was perceptible at baseline (p = 0.0113) and during treatment (p = 0.017).
Median CTC counts did not differ between M+ and M0 patients (9 vs. 8.5 at baseline, 10 vs. 15 during treatment, and 9 vs. 11 overall), whereas median CTM counts were higher in patients without clinical evidence of metastases at all tested times (3 vs. 0, p = 0.0056 at baseline; 2 vs. 0, p = 0.0278 during treatment; and 2 vs. 0, p = 0.001 overall). These results suggest early dissemination of the disease and highlight the importance of better knowledge of the molecular heterogeneity of CTCs for predicting progression.
Discussion
CTCs, enriched and identified by their epithelial features, have reached a high level of evidence as prognostic tools in different clinical stages. However, we would like to give further importance to previous observations that, besides the purely epithelial CTCs, there is an additional, missed CTC population (8) whose clinical relevance is still unexplored and deserves further attention. Using immunomagnetic CTC enrichment in 2 small case series in the present study, we were able to demonstrate that by i) optimizing the antibody composition of cocktails used for CTC enrichment by immunobeads and ii) shifting the CTC identification criteria from purely epithelial features (CellSearch criteria) to include also mesenchymal and stem cell features, a significantly higher number of blood samples was defined as CTC positive. Optimized enrichment and less strict CTC definition reduce the differences between clinical stages, suggesting that in patients without clinical evidence of metastases CTCs may show mesenchymal and stemness traits and lose epithelial features.
The extreme approach of unbiased CTC enrichment by a size-based filtration method yields a further increase in CTC positivity rates, paradoxically abolishing the differences in CTC status and numbers between M0 and M+ patients. This raises interesting questions about distinct molecular features of CTCs in the 2 settings, which might not only depend on the stage of the disease but also help in predicting progression. Whereas this is a possible explanation, the absence of standardization in the CTC assay (with the exception of the CellSearch approach) delays the achievement of clinical validation and represents a limit of the present study.
CTMs, representing heterogeneous clusters of CTCs with blood cells and platelets, are an additional, promising feature worth being explored. It was recently suggested in animal models that CTC clusters are more proficient in generating metastases than single CTCs (14): from such a perspective, the significantly lower number of CTMs in M+ compared to M0 patients represents a new finding that (although apparently not intuitive, similarly to the increases in CTC positivity reported above for improved detection methods) supports the necessity to uncover the message of these missed CTCs. We also speculate that CTMs are particularly worth characterizing from the molecular point of view in M0 patients, whereas in M+ patients single CTCs that could derive from metastatic sites may harbor the molecular makeup of the metastasis, recapitulating the clonal evolution of the disease.
It is important to mention that the currently employed technical approaches for CTC detection, albeit based on distinct properties, share a lack of biological specificity. Whereas it can be affirmed that the identified CTCs are by all criteria cancer cells, data on their ability to invade, proliferate or cause metastases are as yet scanty. A different way of investigating the clinical role of such cells is an urgent need if we want to fulfill the promises of the liquid biopsy approach.
Footnotes
Acknowledgment
The authors would like to thank all patients who participated in the study and their families.
Financial support: Financial support was received from Associazione Italiana per la Ricerca sul Cancro, AIRC (IG 10611 - M.G. Daidone); European Commission under the 7th Framework Programme, grant agreement No. 260791 Eurocan Platform; Italian Ministry of Health. E.F. is the recipient of an AIRC fellowship.
Conflict of interest: Janine Wechsler is a consultant pathologist at ScreenCell. Maria Grazia Daidone is a member of the International Journal of Biomarkers Editorial Board. The authors confirm that her involvement as editor of the journal in no way alters the adherence to the journal policies.
