Abstract
Background:
Leptomeningeal disease (LMD) is a devastating complication of metastatic breast cancer (MBC). Metastatic tumors to the central nervous system (CNS) are not routinely assessed for the presence of actionable mutations, hence the frequency and concordance of actionable mutations in the cerebrospinal fluid (CSF) versus extracranial sites are not well characterized.
Objective:
To evaluate the frequency and concordance of actionable mutations in the CSF versus extracranial sites in patients with MBC and LMD.
Methods:
In this single-center non-therapeutic prospective observational study, we enrolled 15 patients with MBC and known or suspected LMD from 2020 to 2024 and collected CSF, blood, and archival tumor samples. We enumerated CSF circulating tumor cells (CTCs) and analyzed circulating tumor DNA (ctDNA) via next-generation sequencing (NGS) using the CNSide technology (Biocept). When available, we compared CSF NGS results with available NGS data from matched blood and tumor samples.
Results:
Of the 15 patients enrolled, 14 were determined to have LMD based on CSF cytology, radiographic, and clinical assessment (7 hormone receptor-positive (HR+)/human epidermal growth factor receptor 2 negative (HER2−), 6 triple negative breast cancer, 1 HER2+). CSF CTC enumeration was performed in a subset of patients and was positive in 5/7 (71.4%), of which 4/5 had negative CSF cytology. Median CTC enumeration was 0.29 CTCs/mL (range 0–136.4). Of the 14 patients with confirmed LMD, analysis of CSF ctDNA detected pathogenic variants in half of the patients (7/14, 50.0%). Comparison of CSF versus blood and/or tissue-based NGS testing was available for 10 patients: 5 patients (50%) had concordant CSF versus extracranial mutations, 3 patients (30%) had discordant CSF versus extracranial mutations, and 2 patients (20%) had both concordant and discordant CSF versus extracranial mutations.
Conclusion:
CSF CTCs can be detected and enumerated in patients with MBC and LMD, even in patients with negative CSF cytology. We were able to detect pathogenic mutations in the CSF ctDNA in half of the patients. We observed variable concordance and heterogeneity in the detection of actionable mutations between the CSF, blood, and tumor tissue, supporting the value of investigating the CSF to identify novel targets and resistance mechanisms. Larger studies are needed to assess the clinical utility of these observations, particularly with the development of novel targeted CNS-penetrant agents.
Plain language summary
Leptomeningeal disease (LMD) is a devastating complication of metastatic breast cancer (MBC), so novel therapeutic strategies are needed. It is not common to evaluate the tumor cells in the fluid around the brain (cerebrospinal fluid (CSF)), so it is not well known how mutational profiles may differ between this fluid and other tumor sites in the body. Here, we enrolled 15 patients and collected extra CSF in order to perform analysis of CSF circulating tumor cells and their mutational profiles. We found that we could detect tumor cells in the CSF, and we were able to characterize the mutational profiles. There were some instances where the mutational profiles were similar to other areas in the body, but other times the mutational profiles were different. Therefore, it is important to consider specific analysis of tumor cells in the CSF to identify unique mutations that may inform treatment for patients with LMD.
Keywords
Introduction
Leptomeningeal disease (LMD) is a devastating complication of metastatic breast cancer (MBC). LMD is characterized by the infiltration of cancer cells into the leptomeninges, including along the pia mater and arachnoid planes or within the subarachnoid space. 1 The prevalence of LMD in patients with MBC varies from 5% to 20% in different studies, with newer reports showing a higher incidence, likely due to improved magnetic resonance imaging (MRI) detection and higher incidence as patients are living longer with metastatic disease.1–3
LMD is diagnosed via a combination of diagnostic tests, including cerebrospinal fluid (CSF) cytology and/or via radiographic evidence on MRI brain and/or spine studies. 1 However, these tests can have limited sensitivity, leading to difficulty in diagnosis, with some patients undergoing multiple lumbar punctures (LPs) to establish a diagnosis by positive CSF cytology. 1 To improve the limit of detection for accurate diagnosis of LMD, novel assays are needed, and the evaluation of circulating tumor cells (CTCs) in the CSF has been of interest. Multiple retrospective studies across tumor types have demonstrated that the detection of CTCs in the CSF outperformed diagnosis via positive cytology in diagnosing LMD,4–10 but these assays are not yet standard of care, and larger studies are needed to validate these findings and determine optimal clinical utility for LMD diagnosis.
After the diagnosis of LMD, prognosis is poor with a median survival of 4–6 months.1,11 Clearly, novel treatment strategies are needed. LMD remains the most challenging site of MBC to treat, in part due to the inability of many therapeutic agents to accumulate in sufficient concentrations in the CSF for effective treatment. Additionally, leptomeningeal spread is most often a late manifestation of MBC, when patients have been exposed to and progressed on multiple lines of treatment and when the tumor cells have developed resistance. For this reason, it is valuable to understand the CSF tumor receptor characteristics and circulating tumor DNA (ctDNA) mutational status to identify possible resistance markers (such as an ESR1 mutation) or additional targets for therapy (such as human epidermal growth factor receptor 2 (HER2) amplification or mutation, or a somatic BRCA mutation).
Several studies across tumor types have evaluated CSF ctDNA mutational profiles to date,12–14 but the frequency, variation, and concordance of actionable mutations present in the CSF versus extracranial sites have not been well described in patients with MBC and LMD. Identifying targetable mutations and directed therapies is a key strategy toward improving survival in patients with LMD. To address this unmet clinical need, we developed a single-center, non-therapeutic, prospective registry to identify and enroll patients with MBC and LMD to collect CSF for research purposes. We used the CNSide platform (Biocept; San Diego, CA, USA) to perform CTC enumeration and analysis of ctDNA next-generation sequencing (NGS). We compared these results with available NGS data from matched blood and systemic metastatic tumor samples.
Methods
Study design and objectives
This was a single-center, prospective non-therapeutic study that enrolled patients with MBC and known or suspected LMD treated at the University of California San Francisco (UCSF) between 2020 and 2024. Patients were eligible to enroll in this study if they had MBC with known or suspected LMD with a clinical indication for a LP or Ommaya-tap per standard of care and if there was no contraindication for the collection of additional CSF for research purposes. Patients were enrolled prior to their planned LP/Ommaya tap and consented to collect 5–10 cc additional CSF for research purposes. The primary objective of this study was to characterize the CSF molecular profile in patients with LMD. Secondary objectives included comparison of central versus extracranial mutational profiles. This study adheres to the Equator network STROBE guidelines for observational studies 15 ; STROBE checklist is provided as Supplemental Table 1.
CSF CTC enumeration and HER2 status
We collected 5–10 cc additional CSF for research purposes and placed it into a proprietary Biocept tube and then shipped it to Biocept (San Diego, CA, USA). We evaluated CSF CTC enumeration and HER2 status by fluorescent in situ hybridization (FISH; ⩾2.0 HER2 to CEDP17 ratio or ⩾HER2 signals) in a subset of patients using the CNSide assay (Biocept).
Next-generation sequencing
We used the following CLIA-approved NGS assays to evaluate the status of actionable mutations: CNSide (Biocept) for CSF; Guardant360, Foundation One CDx, and CNSide for blood; and Caris and the UCSF CLIA-validated UCSF500 panel for the systemic metastatic tumor samples. Biocept CSF testing was limited to identifying the following mutations: AKT1, ESR1, SF3B1, CCND1, EGFR, FBXW7, ERBB2/HER2, KRAS, TP53, FGFR1, ERBB3, and PIK3CA. Of note, all platforms contained the gene mutations that are a focus of this manuscript (ERBB2, ESR1, PIK3CA, and TP53).
Clinical chart abstraction
Detailed information including patient demographics, tumor characteristics, treatment history, clinical outcomes, and survival dates were obtained via manual chart abstraction. Survival data was updated on January 15, 2025.
Statistics
Data were analyzed using Prism Software (GraphPad, San Diego, CA, USA). Descriptive statistics were used to summarize numeric responses as rate of events (%) and median (range) as appropriate. Survival distributions were estimated by the Kaplan–Meier non-parametric method.
Institutional review board statement
This research was approved by the UCSF Institutional Review Board (CC #21-34354).
Results
Patient demographic characteristics and tumor characteristics
Fifteen patients with MBC and known or suspected LMD were enrolled in this study and underwent CSF collection and analysis. Of the 15 patients enrolled, 14 were ultimately determined to have LMD; one patient did not meet criteria for the diagnosis of LMD based on standard CSF cytology, radiographic, and clinical assessment but was included in the analysis due to the presence of unique CSF genetic alterations; this patient did have brain metastases. Patient demographics are summarized in Table 1. All patients were female (n = 15, 100%). Most patients were non-Hispanic (n = 14, 93.3%) and white (n = 10, 66.7%). Median age at MBC diagnosis was 53.2 years (range 38.5–64.6 years). Most patients had ductal histology (n = 8, 53.3%). Patients had the following MBC subtypes: HR+/HER2− (n = 7, 46.7%), triple negative breast cancer (TNBC; n = 7, 46.7%), and HER2+ MBC (n = 1, 6.7%). Of the 15 patients enrolled, 9 patients (60.0%) had positive CSF cytology at time of study enrollment, 5 patients (33.3%) were CSF cytology negative (no serial CSF samples obtained) and diagnosed via suspicious MRI imaging only, and 1 patient (6.7%) was not diagnosed with LMD but did have brain metastases. Most patients had concurrent parenchymal brain metastases (n = 12, 80.0%) at the time of CSF collection.
Patient demographic and disease characteristics.
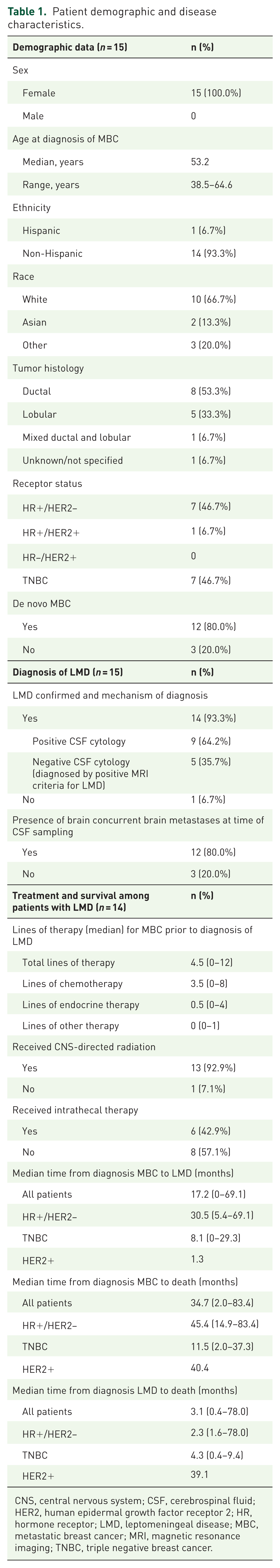
CNS, central nervous system; CSF, cerebrospinal fluid; HER2, human epidermal growth factor receptor 2; HR, hormone receptor; LMD, leptomeningeal disease; MBC, metastatic breast cancer; MRI, magnetic resonance imaging; TNBC, triple negative breast cancer.
Patients received a median of 4.5 lines of prior therapy for MBC prior to the diagnosis of LMD (range 0–12 lines) including a median of 3.5 prior lines of chemotherapy (range 0–8 lines). Of those diagnosed with LMD, most patients received central nervous system (CNS)-directed radiation (n = 13, 92.9%) and many received intrathecal therapy (n = 6, 42.9%). Among the 14 patients with LMD, median time from diagnosis of MBC to LMD was 17.2 months (range 0–69.1 months). For all patients with LMD, median time from diagnosis of MBC to death was 34.7 months (range 2.0–83.4 months) and from diagnosis of LMD to death was 3.1 months (range 0.4–78.0 months).
CSF tumor cell enumeration, HER2 FISH testing, and ctDNA NGS results
Results of the CSF tumor cell enumeration, HER2 FISH testing, and ctDNA NGS testing are summarized in Table 2 and Figure 1.
NGS from the CSF, blood, and tumor among patients with MBC and known or suspected LMD.

CSF, cerebrospinal fluid; CTC, circulating tumor cell; HER2, human epidermal growth factor receptor 2; HR, hormone receptor; LMD, leptomeningeal disease; NGS, next-generation sequencing; TNBC, triple negative breast cancer.

NGS results from the CSF, blood, and/or tumor tissue among patients with MBC and known or suspected LMD.
A subset of patients in this cohort (n = 7) underwent CSF CTC enumeration testing via the Biocept platform, and 5/7 (71.4%) had positive CSF CTC enumeration in at least one sample. Median CTC enumeration was 0.29 CTCs/mL (range 0–136.4). Of note, 4/5 patients with positive CSF CTC enumeration were negative by CSF cytology but were diagnosed with LMD by MRI imaging criteria; the patient with the highest CTC enumeration (136.4 CTCs/mL) was positive by CSF cytology. Of the two patients with negative CSF CTC enumeration, one had equivocal MRI findings that were ultimately deemed to be diagnostic for LMD, and the other was the patient who was LMD negative.
We performed serial CSF CTC analysis on four patients in this cohort (three patients with two CSF samples analyzed, one patient with three CSF samples analyzed; see Table 2). Supplemental Figure 1 shows timelines for two patients (including treatments, clinical response, and CSF sampling results) to provide representative examples.
In the five patients with positive CSF CTCs detected, we also performed CSF HER2 testing via HER2 FISH using the Biocept platform. CSF CTC HER2 was negative by FISH in three patients (two patients with HR+/HER2− and one patient with TNBC). One patient with mTNBC (Pt 35) had 6% of tumor cells that were determined to be HER2 positive by HER2 FISH; the correlation of this testing with standard HER2 IHC testing is unclear, but this may represent HER2-low disease. One patient with low CTCs/mL (Pt 33), had insufficient HER2 probe staining to make a determination, so this was deemed technically insufficient.
All 15 patients in this cohort underwent CSF ctDNA NGS testing via the CNSide (Biocept) assay. Of the 14 patients with LMD, pathogenic variants were detected in the CSF ctDNA of half of the patients (7/14, 50.0%) including at the following loci: ERBB2 amplification (n = 1, 6.7%), ESR1 (n = 1, 6.7%), PIK3CA (n = 4, 26.7%), and TP53 (n = 4, 33.3%). Of the seven patients with HR+/HER2− MBC and LMD, NGS detected actionable mutations in the CSF ctDNA in 57% (4/7), including two patients with PIK3CA E542K, one patient with both PIK3CA E542K and ERBB2 amplification, and one patient with ESR1 Y537S. Of the six patients with mTNBC and LMD, CSF NGS detected actionable mutations in the CSF ctDNA in 17% (1/6), including one patient with PIK3CA E542K. There were no actionable mutations in one patient with HR+/HER2+ MBC and LMD. One patient with mTNBC without LMD per clinical, radiographic, and cytologic criteria, but with brain metastases present (Pt 36), had a mutation in TP53 detected in the CSF.
Concordance/discordance of CSF versus extracranial mutational profiles
Of the 15 patients in this cohort, 10 patients had additional NGS results available from blood and/or tissue sample testing performed per standard of care clinical testing using CLIA-approved NGS testing platforms. Of note, extracranial NGS testing was often performed at other timepoints per standard of care, and NGS assay utilization differed across disease sites (see Table 2 for additional details). Figure 1 and Table 2 describe the concordance among CSF, blood, and tumor tissue NGS testing results for genomic alterations.
Of the 10 patients with available CSF and blood and/or tumor tissue NGS testing, 5/10 patients (50%) had mutations that were concordant between the CSF and extracranial sites, 3/10 (30%) had mutations that were discordant between the CSF and extracranial sites, and 2/10 (20%) had some mutations that were concordant and others that were discordant between the CSF and extracranial sites. Of note, some of these samples were collected at different timepoints (see Table 2).
Concordance between mutational profiles in the CSF and extracranial sites was seen in the following patients: one patient with HR+/HER2− MBC had concordant ESR1 mutations in the CSF and an extracranial site, two patients with HR+/HER2− MBC had concordant PIK3CA mutations in the CSF and an extracranial site, and two patients with mTNBC had concordant TP53 mutations in the CSF and an extracranial site. The patient who was clinically negative for LMD (Patient ID 36) had a concordant TP53 mutation in the CSF, blood, and tumor tissue (Table 2).
Discordance between mutational profiles in the CSF and extracranial sites was seen in the following patients: One patient with HR+/HER2− MBC had an ESR1 mutation detected in an extracranial site but not in the CSF (collected at a later timepoint), and two patients (one HR+/HER2−, one mTNBC) had PIK3CA mutations detected in an extracranial site but not in the CSF, including one patient in which the testing was done concurrently using the Biocept CNSide blood and CSF assays at the same timepoint. Two patients with mTNBC also had discordant TP53 testing, with TP53 mutations detected in an extracranial site but not in the CSF (Table 2).
Discussion
This prospectively enrolled study represents one of the largest series to date analyzing CSF mutational profiles and CSF versus extracranial mutational concordance among patients with MBC and known or suspected LMD. Of the 14 patients with LMD, pathogenic variants were detected in the CSF of half of the patients (7/14, 50.0%), including actionable mutations in 4/7 (57%) with HR+/HER2− MBC and 1/6 (17%) with mTNBC. We evaluated mutational profiles in the ctDNA from the CSF and extracranial sites (blood and/or tissue) and observed instances of both mutational concordance and discordance.
In our study, the most common mutations detected in the CSF were in TP53 and PIK3CA, consistent with prior studies in MBC.5,6 Similar to previous studies, we found concordance as well as discordance among mutations in CSF, blood, and tissue NGS,6,14,16 supporting the value of investigating the CSF to identify resistance mechanisms and new targets. We detected one case of ERBB2 amplification in the CSF, which clearly represents the most immediately actionable finding since there are multiple drugs (trastuzumab deruxtecan, tucatinib, and neratinib) that have demonstrated efficacy either in clinical trials or case series in HER2 overexpressing or HER2 mutated LMD.17,18 Detection of a PIK3CA mutation in the CSF could be a valuable finding in the near future given the number of CNS-penetrant PIK3CA inhibitors in development. 19
Prior studies have also evaluated the ability of CSF NGS to evaluate the mutational profiles of tumor cells in the CSF. For example, one meta-analysis analyzed CSF sequencing results from 541 patients in 17 studies in patients with metastatic non-small cell lung cancer, a population in which liquid biopsy is critical to treatment decisions throughout the course of metastatic disease. 12 In addition to the expected EGFR mutations, the investigators found other driver mutations and mutations associated with drug-resistance including MET, ALK, ERBB2, KRAS, ROS1, BRAF, RET, and NTRK1. Overall, 155 mutations were detected in 201 patients and the most frequent mutation detected in CSF was MET amplification. In breast cancer, however, fewer studies have reported on CSF ctDNA analysis to date. Pentsova et al. 5 analyzed CSF from 11 patients with MBC and brain metastases (without comment on presence or absence of current LMD) and performed sequencing of cell-free DNA. They detected several mutations in the CSF, including PIK3CA H1047R, PIK3CA E545K, TP53 V272M, HER2 amplification, EGFR amplification, and PTEN deletion. In another study of 35 patients with LMD, CSF sequencing in two patients with MBC revealed PTEN K13Q mutations in both patients and a NOTCH1 mutation in one patient. 13 Sequencing was not reported on matched primary tumor or metastatic samples in this study. Several studies utilized the same CNSide platform as in our study to detect CSF ctDNA and perform mutation analysis using a 12-gene Tumor Suppressor Breast NGS Panel.6,14 In a study by Wooster et al., 6 among 10 patients with MBC and LMD, mutations in CSF ctDNA were detected using the CNSide platform in three patients and included TP53 p.K132R, PIK3CAp.H1047L, and a CCND1 deletion. The PIK3CA mutations were detected in both CSF and plasma, but mutations were discordant between CSF and plasma in the other two cases. A more recent study utilized the CNSide platform to enumerate CSF tumor cells and analyze receptor status and oncogene expression via immunocytochemistry, FISH, and NGS in 105 patients with MBC. 14 This analysis reported that 30% (20/66) of patients had at least one biomarker change in the CSF, with a HER2 “flip” (as detected by FISH) as being the most common among breast cancer patients. This study also detected at least one variant in the CSF samples from the patients with breast cancer 32% of the time, but no details were provided regarding the types of mutations. It is important to note that the sensitivity and specificity of NGS platforms differ in their abilities to detect ctDNA and capture mutational differences. The CNSide assay was only designed to evaluate mutations in 12 specific genes (including those most currently actionable in breast cancer treatment). Other assays may offer a broader diversity of profiling and are currently being used and/or developed.
Not only can CSF ctDNA be used to improve the understanding of tumor mutations in this compartment, but CTC detection may be a useful tool to improve the sensitivity and specificity of LMD diagnosis and ability to track disease response. In our study, CSF tumor cell enumeration was only performed in a subset of patients and optimization of CTC detection was not the focus of this analysis. However, it is interesting to note that of the 5/7 patients in our study with detectable CSF CTCs by the Biocept platform, almost all were CSF cytology negative and diagnosed with LMD based on radiographic criteria alone. Thus, assays like these may be more sensitive to detect and track CSF tumor cells. This has also been demonstrated by multiple prior studies,4–10 but it is not yet standard of care to use such assays for LMD diagnosis. It is also interesting to note that in one patient in our cohort who was deemed to be LMD negative by standard clinical, radiographic, and cytologic criteria, there were 0 CTCs detected in the CSF but a TP53 mutation was detected in the CSF. It is unclear if this ctDNA detection was due to shedding from other sites (e.g., this patient had extensive brain metastases) and/or subclinical LMD. Future research is needed to better define optimal criteria for LMD diagnosis based on CTC detection and to determine how/if these assays can be used to track leptomeningeal response and resistance.
Our study has several key limitations. First, blood and tissue NGS were performed per standard of care using different NGS platforms. The Biocept CNSside was the only commercially available CLIA-approved NGS platform for CSF available at the time, and they do not perform tissue-based NGS analysis, so it was not possible to use the same commercial assay across disease sites. Therefore, it is possible that differences in NGS platform sensitivities could have contributed to differences in mutational profiles observed among CSF, blood, and tissue. Of note, Biocept filed for bankruptcy in 2023. Another company purchased their CSF CTC and NGS technology, but it is not yet commercially available. In the meantime, other CSF CTC and NGS analysis platforms are currently commercially available, so the findings generated in this study can be hypothesis generating for future work using this or other similar platforms.
Second, the timing of CSF, blood, and tissue NGS analyses differed in many cases as some of these were archival specimens, so some differences in mutational testing may be due to tumor evolution over time; the timing of each tissue collection and NGS platform utilized are indicated in Table 2 for clarity. Third, given the small sample size and heterogeneity in treatment among patients, we could not evaluate the clinical implications of CSF NGS testing on clinical outcomes. Future clinical trials should prospectively evaluate CSF and extracranial NGS testing in larger cohorts and correlate these findings with clinical outcomes.
Conclusion
In conclusion, this study represents one of the largest series to date of CSF ctDNA mutational analysis among patients with MBC and known or suspected LMD. We detected pathogenic mutations in the CSF in half of the patients tested, including many with actionable mutations. We observed concordance and heterogeneity in the status of actionable mutations among CSF, blood, and systemic metastatic tumor tissue, supporting the value of investigating the CSF to identify resistance mechanisms and new targets. Larger studies are needed to assess the clinical utility of these observations, particularly with the development of novel targeted CNS-penetrant agents.
Supplemental Material
sj-docx-1-tam-10.1177_17588359261424678 – Supplemental material for Detection of actionable mutations in the cerebrospinal fluid and concordance/discordance with extracranial mutations in patients with metastatic breast cancer and leptomeningeal disease
Supplemental material, sj-docx-1-tam-10.1177_17588359261424678 for Detection of actionable mutations in the cerebrospinal fluid and concordance/discordance with extracranial mutations in patients with metastatic breast cancer and leptomeningeal disease by Laura A. Huppert, Samantha Fisch, Lindy Her, Christine Hodgdon, Susie Brain, Carol Simmons, Andrew R. Lai, Lev Malevanchik, Ronald Balassanian, Harish N. Vasudevan, Ramin Morshed, Steve Braunstein, Lauren Boreta, Melanie Majure, A. Jo Chien, Hope S. Rugo, Mark Jesus M. Magbanua and Michelle E. Melisko in Therapeutic Advances in Medical Oncology
Supplemental Material
sj-docx-2-tam-10.1177_17588359261424678 – Supplemental material for Detection of actionable mutations in the cerebrospinal fluid and concordance/discordance with extracranial mutations in patients with metastatic breast cancer and leptomeningeal disease
Supplemental material, sj-docx-2-tam-10.1177_17588359261424678 for Detection of actionable mutations in the cerebrospinal fluid and concordance/discordance with extracranial mutations in patients with metastatic breast cancer and leptomeningeal disease by Laura A. Huppert, Samantha Fisch, Lindy Her, Christine Hodgdon, Susie Brain, Carol Simmons, Andrew R. Lai, Lev Malevanchik, Ronald Balassanian, Harish N. Vasudevan, Ramin Morshed, Steve Braunstein, Lauren Boreta, Melanie Majure, A. Jo Chien, Hope S. Rugo, Mark Jesus M. Magbanua and Michelle E. Melisko in Therapeutic Advances in Medical Oncology
Footnotes
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
