Abstract
Background
Accumulating evidence indicates that circulating pericyte progenitor cells (CPPCs) may be angiogenic biomarkers in cancer and diabetes. Their validity as biomarkers depends on the accuracy of techniques used for enumeration. In this report, absolute CPPC counts were performed by 2 single-platform technologies. The reliability of the 2 methods, including retest reliability and intraobserver and interobserver variability, was assessed according to the intraclass correlation coefficient (ICC). The linear correlation and agreement among both methods were assessed, and the stability of CPPC numbers in blood samples was analyzed.
Methods
The blood samples were obtained from ICR mice. The samples were processed through a no-lyse, 1-wash procedure, and Syto16+CD45-CD31-CD140b+ CPPCs were analyzed by exclusion of dead cells and by fluorescence-minus-one control. CPPCs were enumerated by 2 methods: bead-based 123count eBeads count (eBioscience) and direct volume–based Accuri C6 Flow Cytometer count (BD). The cells were measured immediately and after storage of blood samples for 24 and 48 hours.
Results
There were excellent retest correlations and intraobserver and interobserver agreement in both methods. The 2 methods showed a high linear correlation (R2 = 0.923) and with a high level of agreement (0.986). It was demonstrated that CPPCs are unstable in blood samples.
Conclusions
In this study, 2 reproducible protocols for CPPC quantification were established. These protocols should facilitate future studies to further define the role of CPPCs as cellular biomarkers.
Keywords
Introduction
Pericytes were described in the late 1800s by the French scientist Charles-Marie Benjamin Rouget and are important cellular constituents of capillaries and postcapillary venules. Pericytes wrap around vessels and control their stabilization and hemodynamic processes (1). Most pericytes, expressing the platelet-derived growth factor receptor beta (PDGFR-β), are recruited by PDGF-B–expressing endothelial cells (ECs) (2). As a mesenchymal stem cell (MSC)–like cell, pericytes can differentiate into fibroblasts, smooth muscle cells, osteoblasts, chondroblasts, adipocytes or macrophage as well if required (3).
Interestingly, there is evidence that pericyte progenitor cells (PPCs) circulate in the peripheral blood (4). These cells have been identified and isolated from human or murine peripheral blood, reside in the nonhematopoietic (CD45-neg) compartment, and are distinct from ECs as they lack endothelial antigens (CD31-neg), but express the typical pericyte marker PDGFR-β (CD140b). Circulating PPCs (CPPCs) were found to be increased in patients and mice with malignant tumors and also increased after chemotherapy (4). In type 2 diabetic patients with microangiopathy, glucose control is associated with a transient increase in CPPCs. So in clinical and preclinical studies of cancer and diabetes, CPPC enumeration might be used as a tool for the definition of the optimal biologic dose of drugs and as a possible prognostic tool.
We used to isolate circulating endothelial cells from peripheral blood using immunomagnetic beads (5). But peripheral circulating stem/progenitor cell analysis is gradually being undertaken by flow cytometry, employing either a dual- or a single-platform approach. The dual-platform technique utilizes immunophenotypic data derived from the flow cytometer analysis together with the total white blood cell (WBC) count obtained from a hematology analyzer. The total WBC count generated by hematology analyzers has been recognized as a significant factor in the high coefficient of variation (CV) for absolute enumeration (6). In contrast, single-platform technology derives the absolute cell count directly from the flow cytometry, using either precision fluidics or beads technology (7). Theoretically, single-platform techniques should overcome the limitations of the dual-platform approach and therefore be the preferred method of analysis (8).
In this report, absolute CPPC counts in mice peripheral blood were performed by 2 types of single-platform technologies: (i) 123count eBeads count (eBioscience, San Diego, CA, USA), in which a defined number of beads are added to a known volume of peripheral blood, and (ii) Accuri C6 Flow Cytometer count (BD, Franklin, NJ, USA), a volumetric flow cytometer that determines absolute counts by accurate measurement of the direct volume aspirated from samples. Then, the agreement and the linear correlation between the 2 methods were assessed. Finally, the stability of CPPC numbers in peripheral blood samples was analyzed.
Materials and Method
Animal Handling
All animal handling and procedures were approved by the Experimental Animal Care and Ethics Committee of the Institute of Microcirculation, Chinese Academy of Medical Sciences (CAMS) & Peking Union Medical College (PUMC). Eight-week-old male ICR (18-22 g) mice were purchased from the Institute of Laboratory Animal Science, CAMS-PUMC. The mice were caged in groups of 3 with free access to chow and water. They were acclimated for 1 week before any experiment. The animal containers were kept constantly at 26°C, 38.5% humidity, with a 12-hour light, 12-hour dark cycle (7:30 AM to 7:30 PM). The mice received no hormonal treatment.
Collection of Blood Samples
After induction of anesthesia using pentobarbital sodium (40 mg/kg, intraperitoneally [i.p.]), 400 µL to 600 µL of blood was collected by retro-orbital puncture and anticoagulated with use of ethylenediaminetetraacetic acid (EDTA). Samples were processed immediately after collection. To assess retest reliability, every subject was tested twice by the same person. For the evaluation of variability over time, fresh samples were evaluated immediately after collection and after 24 and 48 hours stored at room temperature (22°C).
Monoclonal Antibodies and Reagents
Monoclonal antibodies directly labeled with fluorochromes were as follows: CD140b-PE, CD45-APC and CD31-PE/Cy7 (Biolegend, San Diego, CA, USA) and nuclear staining Syto16-FITC (Invitrogen, Carlsbad, CA, USA). To minimize nonspecific staining by antibody binding via Fc-receptors, all samples were treated with FcR-blocking reagent (Miltenyi Biotec, Bergisch Gladbach, Germany). Red blood cells were lysed using a commercially available reagent RBC Lysis Buffer (eBioscience). For all staining and washing steps, cells were kept in phosphate-buffered saline (PBS) supplemented with bovine serum albumin (0.5%) and EDTA (1.5 mM). For absolute counts, the reagent 123count eBeads containing approximately 1,000 beads/μL was used.
Immunostaining of Blood Samples
Immunostaining was performed by a modification of previously described methods (9, 10). Briefly, the volume of original blood sample was 200 μL, which was defined as the initial volume. The blood was added to a tube and centrifuged at 500 g for 10 minutes. The upper plasma phase was removed. The lower phase was resuspended with 100 μL of cold PBS and incubated on ice for 30 minutes. Then, the FcR-blocking agent was added at a concentration of 1 μg/mL, and incubated on ice for 10 minutes. Samples and the fluorescence-minus-one (FMO) control were labeled with antibodies and fluorescence dye. After incubation, red blood cells were lysed using 2 mL RBC Lysis Buffer in the dark, according to the manufacturer's directions. The reaction was stopped by diluting the lysis buffer with 2 mL of PBS. The cells were centrifuged at 300 g and resuspended in 200 μL of PBS. Then 50 μL of beads was added to each sample. The final volume in the tube was 250 μL. Each tube was briefly vortexed 6 times (for ~5 seconds) before collection to ensure equal dispersion of cells and beads.
Flow Cytometry Analysis
Accuri C6 Flow Cytometer and the Accuri CFlow analysis software (version 1.0.264.15) were used. All flow cytometric data were reanalyzed in a blinded fashion and by the same person to assess the intraobserver variability. To assess the interobserver variability, all data were reanalyzed in a blinded fashion by another observer. Before analysis, the flow cytometer was thoroughly cleaned to remove residual cells. Samples were analyzed at a medium flow rate (<3,000 events per second) using an FSC-H threshold of 550,000 to exclude debris and electronic noise. In each analysis, 1,000,000 total events were collected. For each blood sample, FMO-matched controls were analyzed to set the appropriate regions. CPPCs were defined as Syto16 (DNA) positive, negative for the hematopoietic marker CD45 and the EC marker CD31 and positive for CD140b. For analysis of CPPCs (Fig. 1), all leukocytes were first gated on a FSC/SSC dot plot (P1). This P1 population was shown on a SSC/CD45 dot plot, and the CD45- events were gated as P2. Then, the P2 cell set was shown on a CD31/Syto16 dot plot following gating of the CD31-Syto16+ events as P3. Finally, the CD31-Syto16+ events were shown on a CD31/CD140b dot plot and the cutoff for CD140b+ events (P4) was assessed by the FMO control. For bead counting, the beads region (P5) was gated on a CD31/CD140b dot plot showing all events.
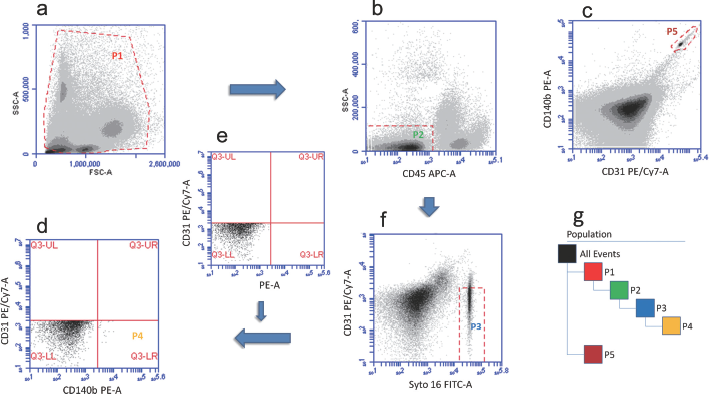
Multicolor flow cytometric analysis of circulating pericyte progenitor cells (CPPCs). All leukocytes were first gated on a FSC/SSC dot plot as P1
The absolute enumeration of CPPCs was defined as the concentration of cells in the original blood sample added to the tube. Based on reference to the manual for 123count eBeads and other references, counting was determined in 2 ways (11, 12):
For bead counting, calculations were carried out using Equation 1 in Figure 2;
For direct volume counting, calculations were carried out using Equation 2 in Figure 2.
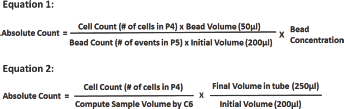

Equation for calculation of absolute counts. The absolute count is defined as the concentration of cells in the original blood sample added to the tube. For bead counting, the calculations were carried out with Equation 1. For direct volume counting, the calculations were carried out with Equation 2.
Statistical Analysis
Statistical analyses were performed with IBM SPSS Statistics for Windows (version 19.0; SPSS Inc, Chicago, IL, USA). The normality of continuous data sets was assessed using the 1-sample Kolmogorov-Smirnov text. Logarithmic transformation was performed where necessary. Continuous data are presented as the means ± SD. Retest correlation, intraobserver agreement and interobserver agreement were assessed by the intraclass correlation coefficient (ICC). Linearity of data as obtained by both methods was assessed, and the agreement among both methods was determined by ICC. A 2-way mixed, type consistency ICC was selected. ICC interpretation was similar to Cohen's Kappa for interrater reliability: An ICC of <0.20 means poor agreement, 0.20 to 0.39 fair, 0.40 to 0.59 moderate, 0.60 to 0.79 substantial, and above 0.80 outstanding interrater reliability (13).
Results
There were excellent retest correlations and intraobserver and interobserver agreement for both methods, with 95% confidence intervals ranging from substantial to almost perfect. Table I shows the mean values of CPPCs levels measured by different methods. The 2 methods showed a high linear correlation (R2 = 0.923) (Fig. 3) and with a high level of agreement (0.986 by ICC).
Mean values of CPPCs measured by different methods, and retest correlations and interobserver and intraobserver agreement for each method
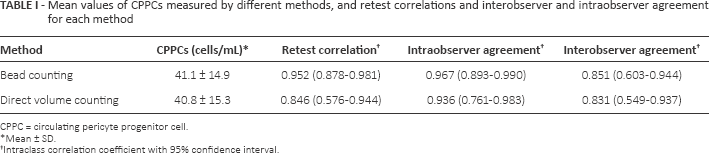
CPPC = circulating pericyte progenitor cell.
Mean ± SD.
Intraclass correlation coefficient with 95% confidence interval.
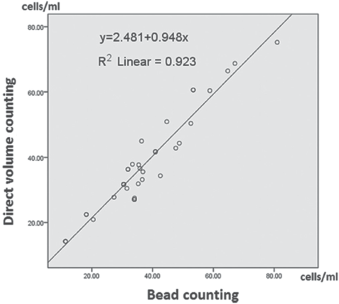
Comparison of bead counting to direct volume counting. The 2 methods showed a high linear correlation (R2 = 0.923).
CPPC numbers were measured in fresh blood samples and samples stored for 24 and 48 hours at room temperature (22°C). It was found that the CPPC count had already decreased significantly after short-term storage for 24 hours (Fig. 4).
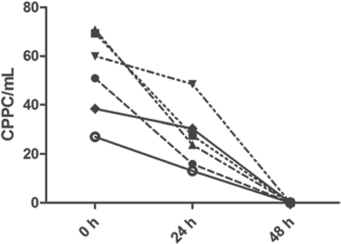
Stability analysis of circulating pericyte progenitor cells (CPPCs) in stored peripheral blood. CPPCs were quantified in peripheral blood of individuals by means of bead counting. For each individual, 2 additional blood samples were analyzed after storage for 24 and 48 hours.
Discussion
Electron microscopy confirmed that CPPCs have progenitor-like morphology (4). But until today, no single antigen has been utilized to discriminate them from other circulating stem/progenitor cells. So, in clinical and laboratory studies, CPPCs have been measured by multiparametric flow cytometry, an approach that requires accurate sequential gating and is prone to operator-induced variability (14).
To optimize the feasibility and reproducibility of our protocol, CPPCs were assessed directly from whole blood and not mononuclear cell preparations which need density gradient centrifugation. Manipulation of samples such as centrifugation and decanting or transferring from 1 tube to another may result in potential cell loss. These manipulation steps should be minimized as much as possible (11). So we employed a no-lyse, 1-wash procedure to process the blood samples in the same tubes. The potential contamination of platelet and red blood cell debris can be gated out during flow cytometric analysis through setting up an FSC-H threshold of 550,000 and using the nuclear stain Syto16 (15) as shown in Figure 1a, f. Flow cytometric assessment of CPPCs belongs to “rare events analysis,” as these cells display weak antigen expression, and their frequency is rather low. Such an analysis requires certain specific steps in order to obtain reliable data (16). For instance, we match high-intensity fluorochromes (PE) to low-density markers (CD140b) and use the FMO control to properly assess the cutoff for true positive cells (17, 18).
123count eBeads, also known as absolute cell count beads, are intended for use in absolute counting of cells by flow cytometry. The beads contain encapsulated dyes that are excited by both blue (488 nm) and violet (405 nm) lasers. They emit bright fluorescence between 500 and 750 nm and can be easily visualized as a compact, bright population (Fig. 1c). In this study, excellent reliability was found, including retest correlation and intraobserver agreement and interobserver agreement, for this bead counting method. But when using these beads, special attention to a few points is needed for accuracy of counts: (i) when adding the blood sample and the beads, use calibrated pipets appropriate for that purposed volume (200 μL or 50 μL); (ii) mix the suspension of beads by vortexing for 3 ~ 5 minutes prior to use; and (iii) be sure to mix cells and beads well before collection by flow cytometry.
The Accuri C6 is a compact flow cytometer that uses a peristaltic pump with a laminar flow fluidic system, and which can compute sample volume and cell concentration directly so as to measure absolute cell counts (12). This direct volume method requires no additional reference bead counts and offers the benefits of increased speed and reduced cost and complexity. In this direct volume counting by C6, reliability was found to be excellent. The mean number of CPPCs counted by direct volume counting was 40.8 ± 15.3/mL, and by bead counting, it was 41.1 ± 14.9/mL, consistent with the reports in the literature (4). The 2 types of single-platform methods show a high linear correlation and a high level of agreement. Both of the single-platform technologies offer simple, straightforward, new methods for the accurate determination of CPPCs in research laboratories. These alternatives are particularly attractive for laboratories that do not have access to a hematology analyzer.
An important requirement for enumeration of CPPCs in clinical and laboratory situations is stability of this cell population in stored blood samples. A significant decrease of viable CPPC numbers was detected in blood samples stored for more than 24 hours. These results indicated that CPPCs represent a vulnerable cell population necessitating CPPC enumeration at defined time points shortly after peripheral blood collection. In future studies, the potency of previously described buffer systems (19) in stabilizing CPPCs viability/numbers in stored peripheral blood has to be further analyzed.
In this study, 2 reproducible protocols for CPPC quantification were established. These protocols should facilitate future studies with the goal to further define the role of CPPCs as angiogenic biomarkers.
Footnotes
Financial support: This study was supported by the Union Youth Science & Research Fund, Fundamental Research Funds for the Central Universities of China (No. 3332014006).
Conflict of interest: The authors declare no competing interests.
