Abstract
Background:
MicroRNA-21 (miR-21) has previously been demonstrated as a potential biomarker in diagnosis of various human tumors. This meta-analysis was performed to evaluate the possibility of miR-21 as a biomarker for early detection of lung cancer.
Methods:
Relevant lung cancer-related miRNA microarray datasets were collected from the NCBI Gene Expression Omnibus (GEO) database and EBI ArrayExpress database up to February 2014. Quality control of the output data was estimated using Limma package and ExiMiR package in R. Standardized mean difference (SMD) with 95% confidence intervals (CIs) from selected datasets was pooled. Heterogeneity was assessed using Cochran's Q test and the I2 statistic, and a p value <0.0.05 or I2 >50% was defined as significant heterogeneity. Furthermore, sensitivity analysis was conducted to evaluate the stability of the pooled results. Four miRNA datasets (GSE24704, GSE17681, GSE27486 and GSE40738) from blood samples were selected, including 153 lung cancer patients and 109 healthy people.
Results:
The pooled results generated by random-effects model revealed that no significant difference was observed between case and control groups (SMD = 0.58; 95% CI, −0.04 to 1.19; p = 0.07) with significant heterogeneity (p = 0.0032, I2 = 78.2%; p = 0.06). Sensitivity analysis indicated that the results of the meta-analysis were stable.
Conclusions:
MiR-21 expression levels in whole blood and peripheral blood cells did not show significant differences between lung cancer patients and healthy controls, and it might be ineffective to measure miR-21 expression to achieve an early diagnosis of lung cancer.
Introduction
Lung cancer is one of the most common causes of cancer-related death in the world (1). In clinical practice, cancers, including colorectal cancer and lung cancer, are often diagnosed at an advanced stage or diagnosed as metastatic disease (2, 3). Therefore, the best period for operation (accounting for almost 70%) is nearly always lost since cancer cells have already spread (4). Thus, development of a method for early detection of lung cancer is urgently needed to reduce the high morbidity and mortality rates associated with this cancer.
MicroRNA (miRNA) is a small noncoding RNA molecule (containing about 19-25 nucleotides) found in plants, animals and some viruses. The role of miRNA in cancer development has also been proven (5, 6). Given stable and consistent levels of miRNAs in serum among individuals of the same species (7), miRNA is recognized as an useful biomarker for the detection of various cancers or other diseases (8). For example, MicroRNA-21 (miR-21), an oncogenic miRNA, has been shown to be overexpressed in various human tumors (9, 10). The up-regulated miR-21 promotes tumorigenesis through inhibition of apoptosis and negative regulators of the Ras/MEK/ERK pathway (11). Furthermore, the expression levels of miR-21 in other blood samples, including whole blood and peripheral blood cell, have been evaluated (12, 13). However, the small sample size in the previous individual studies have limited the reliability of any conclusion regarding miR-21 expression levels in blood samples.
Therefore, a meta-analysis was performed to evaluate the suitability of miR-21 as a biomarker in early diagnosis of lung cancer. In the present review, several miR-21 gene expression profiling datasets from lung cancer patients' blood samples were pooled to evaluate the expression levels of miR-21 in lung cancer patients through comparison with healthy controls.
Materials and Methods
Data Acquisition
Lung cancer-related miRNA microarray datasets from blood samples were collected from the National Center of Biotechnology Information (NCBI) Gene Expression Omnibus (GEO; http://www.ncbi.nlm.nih.gov/geo/) database and the European Bioinformatics Institute (EBI) Array Express (http://www.ebi.ac.uk/arrayexpress/) database up to February 2014. The following keywords were used in the meta-analysis: lung cancer, serum, blood and non-coding RNA profiling.
Inclusion Criteria
Eligible datasets were included if they met the following criteria: (i) both lung cancer patients and healthy people were included in each dataset, and each group contained more than 2 samples; (ii) the expression profiling data of miRNA from the case and control groups were provided or could be calculated; (iii) the subjects involved in the study were humans.
Quality Control and Data Extraction
Two investigators independently extracted the data from all eligible datasets according to the criteria listed above. Disagreements were resolved through discussion with a third researcher. Quality control was conducted using Limma package (14) and ExiMiR package (15) in R, including background correction and normalization processing. Expression values of miR-21 and sample size in both case groups and control groups were extracted. If multiple probes were mapped to a single miRNA, average value of probes were used as the expression value of miRNA. Further, means and standard deviations of these values were calculated.
Statistical Analysis
Meta-analysis was performed by using the meta package in R (16). Continuous outcomes were estimated as standard mean difference (SMD) with 95% confidence interval (CI), and effect sizes were pooled using random- or fixed-effects model. Heterogeneity across studies was assessed using the chi-square test of Q (17) and the I2 statistic (18). A p value <0.05 or I2 >50% was considered to be heterogeneous, and then the random-effects model (DerSimonian-Laird method) was used for pooling the data (19). Otherwise, the fixed-effects model (Mantel-Haenszel method) would be chosen (20).
To further explore the source of heterogeneity, subgroup analyses according to sample sources were performed. Additionally, SMD and its 95% CI were pooled using fixed-effects model or random-effects models to evaluate the robustness of the meta-analysis. Finally, sensitivity analysis was performed by omitting 1 study at a time to evaluate the possible bias caused by the individual study.
Results
Characteristics of the Selected Datasets
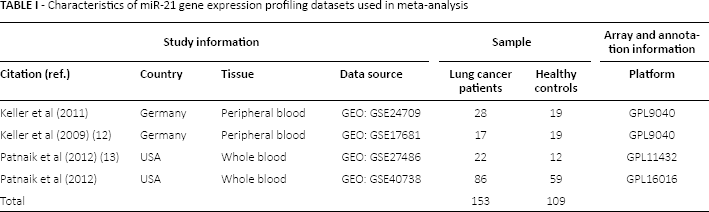
The characteristics of the enrolled datasets are shown in Table I. In total, 4 eligible datasets, including GSE24709 (Germany), GSE17681 (Germany) (12), GSE27486 (USA) (13) and GSE40738 (USA), were involved in the meta-analysis. The data for GSE24709 (Germany) and GSE17681 (Germany) were both derived from peripheral blood cells, and the data for GSE27486 (USA) and GSE40738 (USA) were from whole blood. In total, 153 lung cancer patients and 109 healthy people were enrolled in the 4 datasets.
-Characteristics of miR-21 gene expression profiling datasets used in meta-analysis
Diagnostic value of microRNA-21 as a Biomarker for Lung Cancer
According to the heterogeneity test, significant heterogeneity occurred among individual datasets (p = 0.0032, I2 = 78.2%; Fig. 1). Thus, the random-effects model was selected to calculate the pooled SMD and 95% CI. Results showed that no significant difference was observed between case group and control group (SMD = 0.58; 95% CI, −0.04 to 1.19; p = 0.07), suggesting that miR-21 expression levels in lung cancer patients were similar to those in healthy controls.
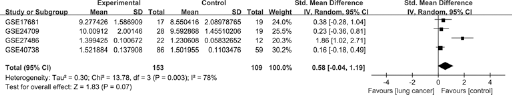
Forest plot of studies evaluating standard mean difference (SMD) of miR-21 expression between lung cancer group and control group (a random-effects model).
The results of subgroup analyses stratified according to sample source are shown in Figure 2. No significant difference in miR-21 expression level was observed between lung cancer patients and healthy controls when we pooled datasets from peripheral blood cells (SMD = 0.29; 95% CI, −0.14 to 0.73; p = 0.19) using the fixed-effects model. Significant heterogeneity was found between the 2 datasets derived from whole blood (p = 0.0002, I2 = 93%). Thus, the random-effects model was chosen to pool effect sizes. Compared with healthy controls, there were no significant differences in miR-21 expression level in lung cancer patients (SMD = 0.96; 95% CI, −0.71 to 2.64; p = 0.26).
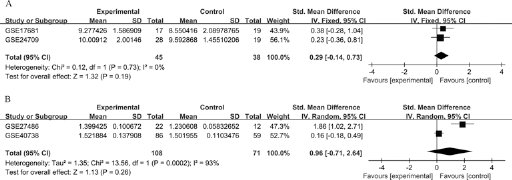
Forest plots of studies evaluating standard mean difference (SMD) of miR-21 expression both in peripheral blood cells (A) and whole blood (B) between lung cancer group and control group.
Sensitivity Analysis
To evaluate the stability of the meta-analysis, SMD with its 95% CI was pooled using the fixed-effects model. Notably, as shown in Figure 3, the effect size pooled using the fixed-effects model showed significant differences between the 2 groups (SMD = 0.35; 95% CI, 0.10-0.61; p = 0.006), suggesting miR-21 was significantly overexpressed in lung cancer patients compared with healthy controls. It was shown that the conclusion for the present meta-analysis was unstable according to inconsistent values pooled by random- and fixed-effects models separately.
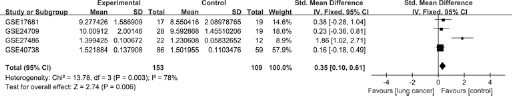
Forest plot of studies evaluating standard mean difference (SMD) of miR-21 expression between lung cancer group and control group (a fixed-effects model).
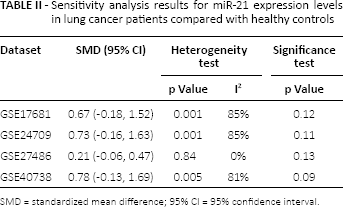
Furthermore, the results for sensitivity analysis are shown in Table II. No significant difference in miR-21 expression was found after omitting each individual dataset. Notably, heterogeneity disappeared when the dataset for GSE27486 was removed (p = 0.84, I2 = 0%), suggesting heterogeneity might be confounded by the GSE27486 dataset.
Sensitivity analysis results for miR-21 expression levels in lung cancer patients compared with healthy controls
Discussion
In the present meta-analysis, a total of 4 gene expression profiling datasets including those of 153 lung cancer patients and 109 healthy people were enrolled. The pooled SMD revealed that miR-21 expression in patients' whole blood and peripheral blood cells showed no significant difference between lung cancer patients and healthy people. In addition, none of the results were reversed when the enrolled individual studies were removed 1 at a time. However, significant heterogeneity was observed across individual studies researching 2 datasets derived from whole blood.
To the best of our knowledge, this is the first meta-analysis to evaluate the suitability of miR-21 as a blood-based bio-marker for early detection of lung cancer by using the micro-array datasets. In 2012, Guan et al performed a meta-analysis evaluating the miR-21 expression levels in lung cancer tissues and demonstrated significantly up-regulated miR-21 expression in lung cancer patients (21). In addition, Selaru et al demonstrated that cholangiocarcinomas and normal tissues could be distinguished on the basis of miR-21 expression level with a sensitivity of 95% and a specificity of 100% (22). However, in our meta-analysis, no significant difference was found in miR-21 expression levels either in whole blood or peripheral blood cells between lung cancer patients and healthy controls. Thus, we concluded that the diagnosis method based on blood-related miR-21 expression levels should be further evaluated.
The strength of the present meta-analysis was limited by significant heterogeneity among individual datasets. Cancer detection methods based on miRNA expression levels have been widely researched. Plasma miRNA profiling shows 64% sensitivity and 89% specificity for miRNA levels, including miR-21, miR-210, miR-155 and miR-196a, from plasma for pancreatic cancer (23). Similarly, miR-21 was significantly overexpressed in serum of hormone-refractory prostate cancer patients (24). Shen et al concluded that miR-21 expression in plasma could be a potential blood-based biomarker for lung cancer (25). Nevertheless, 1 study showed further that blood cells were major contributors to circulating miRNA and that perturbations in blood cell counts and hemolysis could alter plasma miRNA biomarker levels by up to 50-fold (26). Therefore, the possibility of confounding introduced by the sample source or the sample treatment before total RNA extraction in different studies might have contributed to the significant heterogeneity. Further evidence is needed to clarify the confounding technological factors.
Some limitations should be noted in the present metaanalysis: firstly, significant heterogeneity was observed in this study. Although subgroup analyses according to sample source were designed, other confounding introduced by RNA extraction or individual background might disturb the strength of the meta-analysis. Secondly, the small sample size limited the robustness of the meta-analysis. Further studies enrolling large sample size should be designed to confirm miR-21 expression levels in lung cancers.
In summary, miR-21 expression levels both in whole blood and peripheral blood cells did not show any significant difference between lung cancer patients and healthy controls. However, further studies with larger sample size are needed to confirm the conclusion after adjusting confounding technological factors.
Footnotes
Financial support: None.
Conflict of interest: None.
