Abstract
Background
The human regenerating gene 1B (
Methods
Paraffin-embedded specimens from 30 pairs of colorectal cancer tissues and adjacent colon tissues were used to investigate the expression of REG1B by immunohistochemistry. We also examined whether REG1B itself may be related to cell proliferation, cell cycle arrest, apoptosis, migration and invasion in colon cancer HCT116 cells.
Results
Our results showed that REG1B was highly expressed in colorectal carcinoma and was significantly associated with cell differentiation status. The results also illustrated that REG1B silencing with shRNA inhibited cell proliferation, migration and invasion but did not induce apoptosis. Furthermore, down-regulation of REG1B induces G1-phase cell cycle arrest in colon cancer cells.
Conclusions
Knockdown of REG1B can inhibit cell proliferation, migration and invasion. It may act by a mechanism regulating cell cycle progression. Thus, REG1B may be a novel candidate therapeutic target for colorectal cancer.
Introduction
Colorectal cancer (CRC) is the third most commonly diagnosed cancer in males and the second in females in the world, and the fifth most common cancer among patients in China (1). Though stage I and stage II CRC have high cure rates after surgery, the rate of recurrence is about 50% for stage III CRC after surgery alone, and most patients with metastatic disease will ultimately succumb to their cancer (2). In recent years, treatment with adjuvant therapy for CRCs have improved survival, especially in stage III, because some gene and protein biomarkers may increase the sensitivity of chemotherapy (3). Therefore, a better understanding of the molecular mechanisms underlying CRC formation and progression should be helpful in developing more effective treatments for this disease.
The regenerating gene
The human regenerating gene 1B
Moreover, REG1B was not only suggested to be involved in inflammatory diseases but also in carcinogenesis in various digestive tissues. For example, It is overexpressed in peritumoral acinar tissue compared with normal acinar tissue and with acini affected by chronic pancreatitis (15). Also, it has been identified as a new biomarker for pancreatic cancers, because it may be directly or indirectly associated with the secretory pathway in normal pancreatic cells and is consistently increased in adenocarcinomas of the pancreatic ductal fluid compared with normal fluid, and that in pancreatitis and intraductal papillary mucinous neoplasm (16). Aside from up-regulation in pancreatic cancer, REG1B is also increased in CRCs (17) and colorectal adenomas, which are the precancerous lesions of CRC (18). As the
The purpose of this study was to determine whether REG1B expression is associated with clinicopathological factors in patients with CRC and whether the biological functions of REG1B are related to its specific expression in tumors.
Materials and Methods
Clinical Samples
For immunohistochemistry, 30 pairs of CRC tissues and adjacent colon tissues were randomly chosen from the archival samples of the Shanxi Cancer Hospital (Taiyuan, China). After surgery, the removed bowel segment was brought to the Department of Clinical Pathology, and a pathologist sampled tissue from the tumor. Morphological criteria were used to determine the degree of resemblance between the neoplasia and its tissue of origin. A high degree of differentiation meant that the neoplasia was morphologically similar to the native organ, forming neoplastic organoid structures, whereas a low degree of differentiation meant that cells were gradually losing the capacity for structural organization and starting to display reduced cohesiveness (19). All patients provided written informed consent, and the study was approved by the Research and Ethics Committee of the Shanxi Cancer Hospital and the Ethics Committee of Shanxi Medical University. The protocol followed was consistent with the principles of the Declaration of Helsinki.
Cell Culture
Human colon cancer cells sw480 and HCT116 were purchased from the American Type Culture Collection (ATCC Inc., Rockville, MD, USA). Sw480 were cultured in 1640 Medium (Hyclone), and HCT116 were cultured in Iscove's Modified Dulbecco's Medium (IMDM; Hyclone) supplemented with 10% fetal bovine serum (FBS) and 100 U/mL penicillin plus 0.1 mg/mL streptomycin at 37°C in 95% humidity and 5% CO2 humidified incubator.
Immunohistochemical Staining and Evaluation
Paraffin-embedded sections of adjacent and tumor tissues were stained for REG1B expression. Immunohistochemistry for REG1B was performed according to standard procedures as described (20). Briefly, after deparaffinization and rehydration of the tissue sections, immunohistochemistry was conducted with an anti-REG1B rabbit polyclonal primary antibody (1:500; Santa Cruz), followed by incubation with biotinylated secondary antibody against rabbit IgG. Negative controls were set up by omitting the primary antibody. The signal visualization was performed using DAB chromogen for 2 to 3 minutes. The REG1B staining was determined independently by 2 senior pathologists. All of the sections were observed and photographed with a microscope (Olympus, Japan). Ten fields were selected, and the expression was evaluated in 1,000 tumor cells (100 cells per field) with ×100 microscopy (21). The scoring system used in grading the REG1B expression was described previously (22). Tumors with strong and moderate immunostaining intensity were classified as having positive (+) expression – that is, the expression of REG1B was defined when the area of REG1B staining was 10% or more. And tumors with absent and weak immunostaining were classified as having negative (-) expression – that is, the area of REG1B staining was less than 10%. The determination of positive or negative was done under the microscope.
RNA Interference
To silence REG1B in cells, a small interfering RNA (siRNA) was transfected into the cells. The short hairpin RNA (shRNA) sequence that we used was as follows: REG1B, sh1: GAGCAGTGCTAATGCTGGCTA; sh2: TAGCACTGAT GACAGCAATGT; sh3: AAGGATGAATCTTGTGAGAAG; the negative control duplex TTCTCCGAACGTGTCACGT targeted a nonspecific sequence. These were purchased from Shanghai Genechem Company. HCT116 cells were plated in 6-well plates, cultured to 50%-60% confluence and transfected with shRNA or control shRNA for 48 hours using Lipofectamine 2000 (Invitrogen, USA). Transfection was performed as previously described (23).
Real-time RT-PCR Analysis
Total RNA was extracted from the HCT116 cells (with or without transfection) with TRIzol reagent (Invitrogen, USA) according to the manufacturer's protocols. The mRNA expression level of REG1B was detected using a quantitative 2-step reverse transcriptase polymerase chain reaction (RT-PCR) assay. Real-time PCR was performed using SYBR Green PCR Master Mix (Takara, Japan) with the ABI Prism 7500 SDS apparatus (Applied Biosystems, Foster City, CA, USA). The primers used for real-time PCR in this study were the following: 5′-AGC TCC TAC TCT GTC TGA CCT G-3′ (upstream primer) and 5′-CTT AGC GGC AGT GAA TCT CTT G-3′ (downstream primer) for REG1B; 5′-CAA GGA GTA AGA CCC CTG GAC-3′ (upstream primer) and 5′-AAC TGT GAG GAG GGG AGA TTC-3′ (downstream primer) for glyceraldehyde-3-phosphate dehydrogenase (GAPDH). The relative copy number of target sequences was calculated according to the comparative threshold CT (Δ ΔCT) method. The starting relative copy number of the target tumor DNA is given by 2-Δ ΔCT, where Δ ΔCT = ΔCTtumor− ΔCTnormal and each ΔCT = CTtarget−CTreference (24). All reactions were done in triplicate, and all experiments were repeated at least 3 times.
Western blot Analysis
Whole cell lysates were prepared in lysis buffer (50 mM Tris-HCl, pH 7.4, 150 mM NaCl, 1 mM MgCl2, 100 μg/mL PMSF and 1% Triton X-100) (Beyotime, China) for 30 minutes on ice. Equal amounts (50 μg) of lysate proteins were separated on 15% sodium dodecyl sulfate–polyacrylamide gel electrophoresis (SDS-PAGE) gels and then transferred onto nitrocellulose membrane (Millipore, USA). After blocking with 5% non-fat dry milk in TBST buffer (10 mM Tris, pH 7.5, 150 mM NaCl and 0.05% Tween 20) for 2 hours at room temperature, the membranes were incubated overnight at 4°C with primary antibodies specific for REG1B (an anti-REG1B rabbit polyclonal; Santa Cruz, USA) and β-actin (an anti-β-actin rabbit polyclonal; Sigma, USA). After incubation in a 1:2,000 dilution of secondary antibodies conjugated to horseradish peroxidase for 1 hour at room temperature, protein bands were visualized using enhanced chemiluminescence system (ECL; Boster, China).
Cell Counting and Cell Proliferation Assay
HCT116 cells were plated into 96-well plates at 103 cells/well in triplicate in 100 µL of culture medium and allowed to adhere overnight. The cells were transfected with control (NC-shRNA) or plasmid DNA (REG1B-shRNA). Then cells were collected 0 hours, 24 hours, 48 hours and 72 hours after transfection by trypsinization, mixed in phosphate-buffered saline (PBS) and trypan blue was added. Cells were counted using a microscope counting chamber. Additionally, in another set of experiments, cell proliferation was assessed 0 hours, 24 hours, 48 hours and 72 hours after transfection using CCK8 (Dojindo, Japan) according to the manufacturer's instructions. The absorbance was measured by a microplate reader at 450-nm wavelength.
Cell Cycle and Apoptosis Analysis
Forty-eight hours after transfection, cells were collected, washed with PBS, fixed with 70% (v/v) ethanol at 4°C overnight, and centrifuged at 1,500 rpm/min for 5 minutes. Then ethanol was removed, and cells were stained with propidium iodide (PI). Cell cycle analysis was performed using a flow cytometer (BD, USA).
Apoptosis analysis was assessed using phycoerythrin (PE) staining Annexin-V by flow cytometry. The assay was performed using an Annexin-V PE Apoptosis Detection Kit (Keygen Biotech, China) following the manufacturer's protocol. In brief, at 48 hours after the treatment, 5 × 105 trypsinized cells of each group were stained and analyzed by a flow cytometer (FACS Calibur, BD).
Migration and Invasion Assay
Cell migration and invasion capacity were measured
Statistical Analysis
Values are expressed as means ± SD. All data represent the results of at least 3 independent experiments. Statistical analysis was performed using SPSS statistical software (version 17.0). The differences in characteristics between the 2 groups were examined by the chi-square test and Fisher's exact test. All p values were determined from 2-sided tests, and statistical significance was based on a p value of 0.05.
Results
Clinical Significance of Positive REG1B Expression in CRC Tissue
To investigate whether REG1B was involved in the tumorigenesis of CRC, we examined the REG1B expression in CRC specimens. Thirty colon cancers from both the left and the right side and the adjacent normal mucosa were analyzed for expression of REG1B. Among the 30 patients, 21 patients (70%) showed positive REG1B expression. Additionally, only 3 (10%) of 30 adjacent tissues showed positive REG1B expression (Fig. 1). The expression of REG1B was localized to the cytoplasm and expressed in the basal crypt epithelium or some of the proliferating goblet cells. REG1B-positive staining was only detected in 3 of 9 (33.33%) well-differentiated tissues, whereas the detection of positive staining in moderately differentiated and poor differentiated tissues was 11 of 13 (84.62%) and 7 of 8 (87.50%), respectively. We then used Fisher's exact test to evaluate the relationship between REG1B expression and cell differentiation. These results suggest that REG1B expression was significantly associated with cell differentiation status in CRC (p = 0.020). However, REG1B expression was not significantly correlated with sex, age or tumor location (Tab. I).
Clinical correlation between REG1B expression and clinicopathological features in colorectal carcinoma
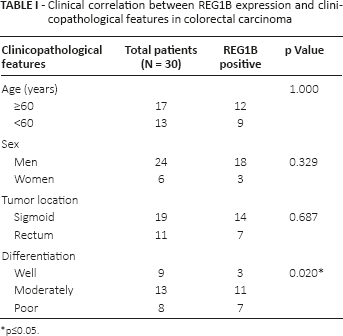
p≤0.05.
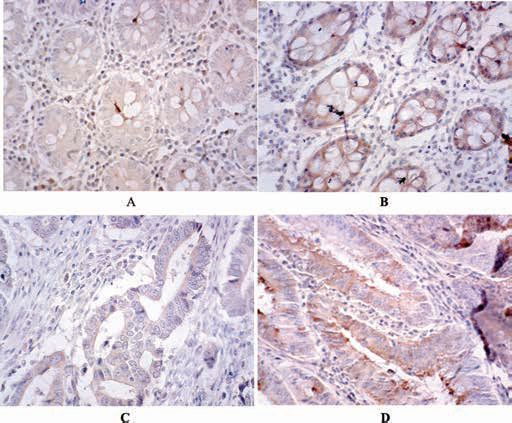
Representative immunohistochemical staining of REG1B positive and negative expression in colon cancers or adjacent tissue (original magnification ×100). A) REG1B protein was negative in adjacent tissues; B) REG1B protein was positive in adjacent tissues; C) REG1B protein was negative in tumor tissues; D) REG1B protein was positive in tumor tissues.
Detection of REG1B in CRC Cell Lines
To test the expression of REG1B in CRC cell lines, Western blot and quantitative reverse transcriptase polymerase chain reaction (qRT-PCR) analysis of REG1B were performed on HCT116 and sw480 cells. The results revealed that the level of mRNA and protein of REG1B in HCT116 cells were both higher than that in sw480 cells (Fig. 2). In the next experiment, we used HCT116 as the target cells of RNA interference.
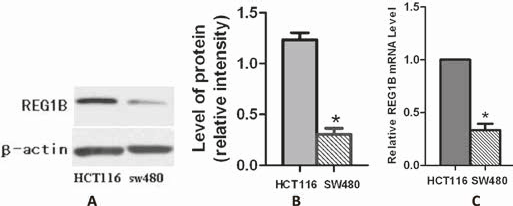
Western blot and quantitative reverse transcriptase polymerase chain reaction (qRT-PCR) analyses of REG1B expression in HCT116 and sw480 cells. A) The cells were analyzed by immunoblotting with specific antibodies to REG1B and β-actin; β-actin was the internal loading control. B) Western blots were scanned and quantified. Data present densitometric analyses of REG1B relative to β-actin for 3 independent experiments. *p<0.05. C) Real-time RT-PCR was performed to detect REG1B mRNA levels in HCT116 and sw480 cells. Glyceraldehyde-3-phosphate dehydrogenase (GAPDH) was the internal control. The bar graph shows the results of 3 independent experiments. *p<0.05.
Suppressed Expression of REG1B by Employment of shRNA in HCT116 Cells
To study the function of REG1B in CRC cell biology, we used shRNA targeting REG1B to inhibit the REG1B expression. Three REG1B-shRNA plasmids and control plasmids were transfected into HCT116 cells. The overall transfection rates were estimated to be >60% following 48 hours of incubation. To investigate whether REG1B was knocked down by shRNA, the transfection results were confirmed by real-time RT-PCR and Western blot. The results showed that the interference effects of sh3 were better than those of sh1 and sh2. Thus, in subsequent studies we used sh3 as our target sequence (each experiment was performed 3 times, and a typical result is present in Fig. 3).
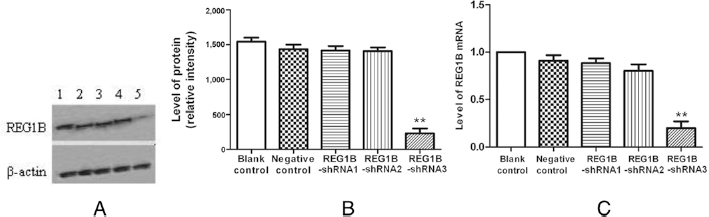
Effective knockdown of REG1B by shRNA in HCT116 cells. A) Western blot of REG1B protein in HCT116 cells at 48 hours after transfection. Lane 1: nontransfected cells; Lane 2: cells transfected with negative control shRNA; Lane 3: cells transfected with REG1B-shRNA1; Lane 4: cells transfected with REG1B-shRNA2; Lane 5: cells transfected with REG1B-shRNA3. β-actin antibody was used as a loading control (bottom panel). B) The Western blots were scanned and quantified. Data are densitometric analyses of REG1B relative to β-actin for 3 independent experiments. **p<0.01, vs. blank control. C) REG1B mRNA levels in nontransfected cells and cells at 48 hours after transfection of negative control shRNA or REG1B shRNA. REG1B mRNA level was significantly down-regulated in REG1B-shRNA3; **p<0.01. Values are from at least 3 independent experiments.
Inhibition of Cell Proliferation by REG1B shRNA
After confirming the knockdown efficiency of the shRNAs targeting REG1B, we examined the effect of a reduced REG1B level on cell proliferation with cell counting and cck8 assay. Growth curves demonstrated that HCT116 cells which were transfected with shRNA targeting REG1B showed a dramatic inhibition of cell proliferation compared with blank control or NC-shRNA-transfected cells (Fig. 4). This result demonstrated that the REG1B had a direct effect on cell proliferation in HCT116 cells.
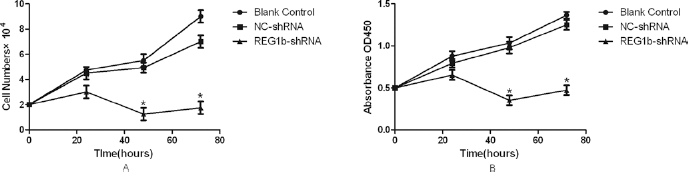
Effect of REG1B knockdown on cell proliferation in HCT116 cells. A) Cell counting to detect cell proliferation. Cell numbers were counted at hours 0 to 72 as indicated after transfection into HCT116 cells with either REG1B-shRNA or a negative control (NC). B) CCK-8 analysis to detect cell proliferation. CCK-8 cell proliferation assay was performed at hours 0 to 72 after transfection into HCT116 cells with either REG1B-shRNA or a negative control (NC); *p<0.05, vs. blank or negative control group.
REG1B Silencing does not Trigger Obvious Apoptosis in HCT116 Cells
To determine whether apoptosis is involved in the growth inhibition, we performed cell apoptosis analysis. Forty-eight hours after transfection, cell apoptosis was examined with an Annexin-V PE/7-AAD apoptosis detection kit and flow cytometric analysis. The apoptotic rate of REG1B-shRNA was 10.91% ± 1.85%, and the rate of control group was 6.28% ± 1.32%. There was no difference between the 2 groups, and the results indicated that down-regulation of REG1B had no effect on cell apoptosis (Fig. 5).
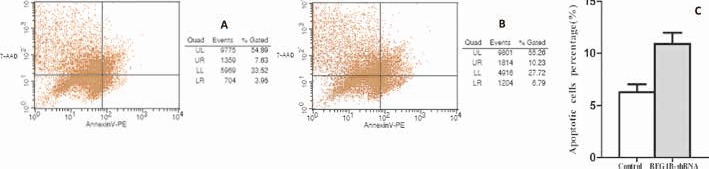
Effect of REG1B silencing on cell apoptosis. Apoptosis was estimated with Annexin V-PE staining by flow cytometry. A) Apoptotic analysis of control group. B) Apoptotic analysis of REG1B-shRNA group. C) Data represent the mean apoptotic cell percentage (n = 3).
Down-Regulation of REG1B Induces G1-phase Cell Cycle Arrest in Colon Cancer Cells
We further tested whether the down-regulation effect of REG1B expression cell proliferation was via cell cycle arrest. The knockdown of REG1B in HCT116 cells using a sequence-specific shRNA against REG1B caused a striking decrease of cells in the S phase, from 28.2% to 13.15%, accompanied by an increase in the G1 phase from 39.18% to 50.17%. However, no obvious alteration was found in the percentage of G1-phase cells in NC-shRNA-transfected HCT116 cells compared with blank controls. Therefore, REG1B silencing may inhibit HCT116 cells proliferation by inducing cell cycle arrest at the G1/S phase (Fig. 6).
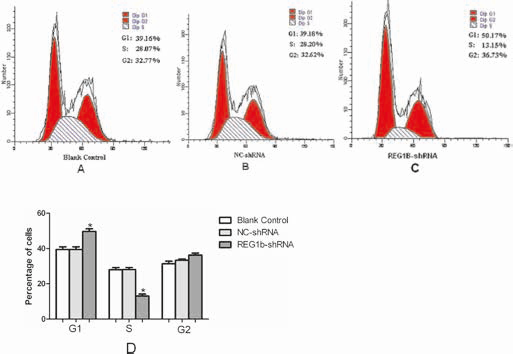
Effect of REG1B shRNA on cell cycle of HCT116 cells. Flow cytometric analysis showed that knockdown of REG1B with shRNA induced G1 phase arrest in the HCT116 cell line (A-C). D) Bar graph showing means of 3 independent experiments; *p<0.05.
REG1B Silencing Inhibited in vitro Migration and Invasion of HCT116 Cells
To determine whether repression of REG1B expression inhibits cell migration or invasion, the transwell assay was performed on HCT116 cells either untransfected or transfected with REG1B shRNA or negative control shRNA. As shown in Figure 7, after incubation for 24 or 48 hours, the number of HCT116 cells passing through the inserts in the REG1B shRNA group was significantly lower than that in the untransfected group and negative control group. The results suggest that shRNA-mediated inhibition of REG1B expression decreased the migration of HCT116 cells.
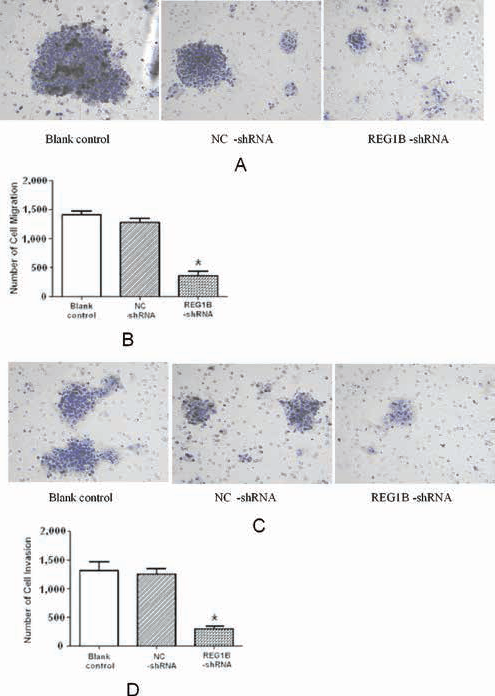
Effect of REG1B inhibition on HCT116 migration and invasion
Discussion
Previous studies have revealed that the expression of REG1B was increased in pancreatic cell lines (25), and in benign diseases such as acute amebic colitis (13), ulcerative colitis and Crohn's disease (26) and adenomatous colorectal mucosa (18). It was also overexpressed in several digestive cancers including pancreas tumors (15) and the CRCs (17). So it was believed that REG1B played a crucial role in the development and progression of carcinogenesis.
In the present study, immunohistochemical analysis was used to investigate REG1B expression in CRCs. We confirmed that the expression of REG1B could be detected in both CRC and adjacent tissues but at a different frequency. REG1B expression was up-regulated in 70% of CRC specimens, whereas only 10% of adjacent tissues showed positive REG1B expression. Zheng et al (27) reported that REG1B was down-regulated in CRC compared with the adjacent tissues, which was in contrast to our results. We consider that this is potentially due to the regulation at the translational level or to the differences in detection approaches, and believe that either increased or decreased expression of these proteins may be involved in the malignant transformation.
It is well known that the traits of all cancers are characterized by increased cell proliferation and decreased apoptosis (28). Thus, it has been an important goal of cancer studies to search for molecular regulators of tumor cell growth. We first examined the expression of REG1B in sw480 and HCT116 cells. Sw480 was isolated from the primary adenocarcinoma arising in the colon (29), and HCT116 cells were used as a tool to dissect the molecular mechanism involved in the metastatic cascade (30). The results showed that the levels of mRNA and protein of REG1B in HCT116 were both higher than in sw480. This means that REG1B may play an important role in the regulation of cell growth in metastatic cells such as HCT116. We then used HCT116 as the target cells of RNA interference in the experiments that followed.
We first found that when REG1B was silenced, the growth of the HCT116 cells was significantly inhibited. This demonstrated that REG1B had a direct effect on cell proliferation, but the specific molecular mechanisms by which REG1B affects tumor growth are not yet fully understood. Defective apoptotic mechanisms are considered to play a central role in the development of malignancy of cancer cells (31, 32). We then examined apoptosis and found that knockdown of REG1B did not influence the rate of apoptosis in cancer cells, suggesting that apoptosis might not be the mechanism underlying the inhibition of tumor growth.
The role of REG1B in regulating CRC cell growth was further assessed by cell cycle analysis. We found that knock down of REG1B inhibits cell growth, arrests cells in G1 phase, but to date, cell cycle analyses and modulation of cell cycle proteins have not been investigated in the context of REG1B.
Cell migration and invasion are one of the common functions required by tumor cells for metastatic progression in target microenvironments (33). Considering the up-regulation of REG1B in many types of human tumors, we speculated that it may play a crucial role in the invasive potential of cancer cells. We investigated the effect of REG1B knockdown on cancer cell migration and invasion with 8.0-μm pore inserts on a 24-well transwell by migration assay. The results showed that REG1B interference in HCT116 cells led to a significant reduction in their
In conclusion, the present study provides evidence that the down-regulation of REG1B expression in colon cancer HCT116 cells can inhibit different aspects of tumor initiation and progression, including cell cycle progression, proliferation and cell migration. Since the expression of REG1B has been detected in many digestive cancers, which is consistent with our data, we can infer that REG1B may be a valuable proto-oncogene and therapeutic target for CRC management.
Footnotes
Financial support: No financial support was received for this submission.
Conflict of interest: The authors have no conflict of interest.
