Abstract
Prostate cancer (Pca) is one of the noncutaneous cancers occurring worldwide. Its high morbidity and mortality make it a concern. X-ray repair cross-complementing group 1 (XRCC1) Arg399Gln polymorphism (rs25487) has been reported to be related to Pca. However, the conclusions are controversial. In this study, PubMed, HuGENet and Chinese National Knowledge Infrastructure (CNKI) databases were combined with a comprehensive literature search. Four models including dominant (AA + AG vs. GG), recessive (AA vs. AG+GG), codominant (AA vs. AG, AA vs. GG) and per-allele analysis (A vs. G) were applied. Finally, 15 studies with 18 sets of data were included. A positive association was discovered in pooled results for recessive (odds ratio [OR]=1.202, 95% confidence interval [95% CI], 1.060-1.363, I2=46.20%), codominant (AA vs. AG; OR=1.258, 95% CI, 1.099-1.439, I2=38.50%; AA vs. GG; OR=1.283, 95% CI, 1.027-1.602, I2=51.70%) and allele analysis (OR=1.116, 95% CI, 1.001-1.244, I2=58.00%). In ethnicity subgroup analysis, these 4 models were also significant in the Asian subgroup. However, for whites, only 2 models seemed to be significant (AA vs. AG+GG: OR=1.525, 95% CI, 1.111-2.093, I2=52.60%; AA vs. AG: OR=1.678, 95% CI, 1.185-2.375, I2=30.70%). In further analysis, we regrouped the data based on race, in which pooled results and Asian subgroup were again shown to be positive. In the next analysis, expression quantitative trait loci (eQTL), linkage disequilibrium (LD), TagSNP and functional analysis were used. The results showed that the SNP was a tag and functional SNP with LD block in both Asians and whites. In summary, we suggest that XRCC1 Arg399Gln might be significantly associated with development of Pca.
Introduction
Prostate cancer (Pca) is a common noncutaneous cancer among men resulting in near 10% of male cancer-related deaths worldwide (1). Just in 2002, 189,000 men with Pca and 30,200 deaths directly attributable to this disease were identified in the United States (2). As a multifactorial disease, both genetic and environmental elements have been identified to be etiological factors (3), among which DNA repair gene polymorphism might be significantly associated with Pca risk (4, 5). DNA repair genes are a group of genes that may alter protein function and DNA repair capacity. The mutation of common polymorphisms would lead to genetic instability (gene rearrangements, translocations, amplifications and deletions) and carcinogenesis (6–7–8). There are 3 main pathways of DNA repair (base excision repair [BER], nucleotide excision repair [NER], and homologous recombinational repair [HRR]), among which X-ray repair cross-complementing group 1 (XRCC1) is an essential one involved in BER (9).
The XRCC1 gene is located on chromosome subband 19q13.2 from 43543312 bp to 43575578 bp. Many studies have shown a significant association in the XRCC1 gene polymorphisms and Pca risk. In 2012, Zhou et al (10) suggested that XRCC1 gene polymorphism was associated with susceptibility to Pca in Chinese men. Other polymorphisms such as Arg399Gln etc. have also also identified (4, 5). However, conclusions regarding an association between XRCC1 Arg399Gln polymorphism (rs25487) and Pca risk are controversial. To investigate the potential relation between Pca risk and XRCC1 Arg399Gln polymorphism, this study was conducted. Our study not only included the results of meta-analysis, but also elucidated the single nucleotide polymorphism (SNP) concerned more clearly using expression quantitative trait loci (eQTL), linkage disequilibrium (LD) and functional analysis.
Methods
Study filtration
PubMed, HuGENet and Chinese National Knowledge Infrastructure (CNKI) databases were combined with a comprehensive literature search. “XRCC1”, “X-ray repair cross-complementing group 1”, “RCC”, “Pca”, “prostate cancer”, “prostatic carcinoma” and “prostate carcinoma” were used as search terms. The remaining studies were discovered via the reference lists in the identified articles. Our study was updated in May 2014. When selecting the studies, the inclusion criteria were as follows: (i) a case-control study and (ii) focused on the association of XRCC1 Arg399Gln polymorphism and Pca risk. The exclusion criteria were (i) lack of raw data about the number of each genotype for XRCC1 Arg399Gln polymorphism among the cases and controls; (ii) review paper; (iii) duplicate publication; (iv) only case samples in the studies.
After filtration, the following data from eligible studies were recorded: author's name, year of publication, country, races (categorized as white, Asian or mixed), type of case and control, methods for experiments and numbers of samples for cases and controls. This information is presented in Table I.
Characteristics Of Eligible Studies In This Study
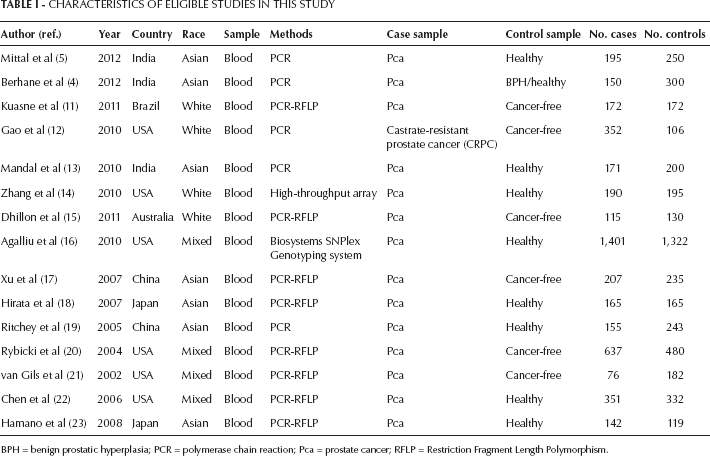
BPH = benign prostatic hyperplasia; PCR = polymerase chain reaction; Pca = prostate cancer; RFLP = Restriction Fragment Length Polymorphism.
Statistical analysis
To get comprehensive results, 4 genotype models were applied in this analysis, including dominant model (AA+AG vs. GG), recessive model (AA vs. AG+GG), codominant model (AA vs. AG, AA vs. GG) and per-allele analysis (A vs. G). Association was evaluated by the odds ratio (OR) and corresponding 95% confidence interval (95% CI). Based on the individual ORs, the pooled OR was estimated. When selecting the fixed and random effects, we used the value of I2 as criterion. The fixed effect was selected when I2<50%. Otherwise random effects were reserved. The effects incorporated an estimate of the interstudy variance and provided wider 95% CIs if the results of the constituent studies differed among themselves. In addition, when estimating heterogeneity, we applied the chi-square-based Q statistic (with p<0.10 as the standard) (24), which represented the weighted sum of the squared difference in the overall effect sizes from each study. On the basis of similar characteristics (i.e., ethnicity and control sample, etc.), subgroup analysis was conducted. In our eligible studies, 6 countries were classified: India, Brazil, United States, Australia, China and Japan. Regarding ethnicity, 3 groups were differentiated: whites, Asians and mixed.
Publication bias was assessed both visually by Funnel plots and statistically by Begg's unweighted regression tests. All of the allele frequencies were calculated according to genotype data. Stata 9.0 (Stata Corp., College Station, TX, USA) was used for analysis, and all p values were 2-tailed.
Other analysis for XRCC1 Arg399Gln polymorphism
After conducting the meta-analysis for XRCC1 Arg399Gln polymorphism with the available data, we tried to describe the SNP more clearly. In addition, eQTL, LD, TagSNP and functional analysis were used. For the eQTL analysis, GENe Expression VARiation (Genevar) was applied with the HapMap database (25). The relationship in the distribution of normalized expression levels for genotypes was calculated using Spearman's rank correlation coefficient (rho) in 4 different populations: Han Chinese from Beijing, China (CHB), Japanese from Tokyo, Japan (JPT), Utah residents with Northern and Western European ancestry (CEU) and Yoruba trios from Ibadan (YRI). To avoid false-positive associations, significance was assessed with 10,000-fold permutations.
Then LD and TagSNP were analyzed with the software HaploView and Tagger (26, 27). Using HapMap Release 21 data, the TagSNPs were identified with an allele frequency ≥5% and high r2 ≥0.8 in the CEU and CHB+JPT populations. Meanwhile, the functional SNPs were also predicted (http://manticore.niehs.nih.gov/snpinfo/snpfunc.htm). Combining all these analyses, we could present more comprehensive descriptions of the SNP, which can help to understand the association between the XRCC1 Arg399Gln polymorphism and Pca risk more clearly.
Results
Characteristics
Three databases (PubMed, HuGENet and CNKI) were retrieved with key words searches. From these, 209 studies (PubMed: 128; HuGENet: 27; CNKI: 54) were found. After reading the abstracts, 152 articles were removed with no evidence of any study of the association between Pca risk and XRCC1 gene polymorphism. Then the duplicate studies in the 3 databases were identified, resulting in 28 papers. While scanning the full texts, 13 studies were removed for the following reasons: (i) 5 articles had no raw data about the genotypes of XRCC1 Arg399Gln polymorphism (28–29–30–31–32); (ii) 3 meta-analyses regarding the XRCC1 polymorphism and Pca risk (33–34–35); (iii) 1 study was focused on the association between c.910A>G polymorphism of the XRCC1 gene and Pca risk (10); (iv) a case-only study (36, 37); (v) 1 study was a duplication with the same data presented (38); (vi) in another report, although investigating the association of Pca and XRCC1 Arg399Gln polymorphism, the data for each genotype could not be differentiated (39). As a result, 15 studies with 18 sets of data were treated as those eligible for consideration (4, 5, 11–12–13–14–15–16–17–18–19–20–21–22–23).
Among these collected studies, DNA came from the blood of subjects. Genotyping was mainly conducted with polymerase chain reaction (PCR) except for in 2 studies (17, 22). The cases were identified as Pca based on pathological examinations at local hospitals. Controls were defined as those with benign prostatic hyperplasia (BPH) or cancer-free and healthy. In addition, 7 studies were mainly focused on Asian patients (4, 5, 13, 17–18–19, 23). Four were on whites (11, 12, 14, 15). And the others were of mixed races with whites, African Americans or other races involved (16, 20–21–22). The participants came from India, Brazil, United States, Australia, China and Japan.
XRCC1 Arg399Gln polymorphism and prostate cancer
Based on the 4 different genotype models, the meta-analysis was conducted. A positive association was discovered in the pooled results for the recessive model (AA vs. AG+GG; OR=1.202, 95% CI, 1.060-1.363, I2=46.20%), codominant model (AA vs. AG; OR=1.258, 95% CI, 1.099-1.439, I2=38.50%), codominant model (AA vs. GG; OR=1.283, 95% CI, 1.027-1.602, I2=51.70%) and allele analysis (A vs. G; OR=1.116, 95% CI, 1.001-1.244, I2=58.00%). However, the significance in the dominant model (AA+AG vs. GG) was not presented (OR=1.000, 95% CI, 0.918-1.090, I2=49.60%). To analyze the data more comprehensively, subgroups were considered. Based on ethnicity, 3 groups (Asians, whites and mixed races) were identified. Among Asians, the recessive model (AA vs. AG+GG; OR=1.546, 95% CI, 1.248-1.915, I2=0.00%), codominant model (AA vs. AG; OR=1.614, 95% CI, 1.279-2.037, I2=0.00%), codominant model (AA vs. GG; OR=1.545, 95% CI, 1.209-1.974, I2=0.00%), allele analysis (A vs. G; OR=1.208, 95% CI, 1.070-1.363, I2=6.20%) were identified to be significant. As for whites, only 2 models seemed to be significant (AA vs. AG+GG: OR=1.525, 95% CI, 1.111-2.093, I2=52.60%; AA vs. AG: OR=1.678, 95% CI, 1.185-2.375, I2=30.70%). However, in the mixed races group, there seemed to be no evident relationship between XRCC1 Arg399Gln polymorphism and Pca risk. All of the results are shown in Table II (Fig. 1).
Results Of The Statistical Analysis For Association Between Prostate Cancer Risk And Xrcc1 Arg399Gln Polymorphism
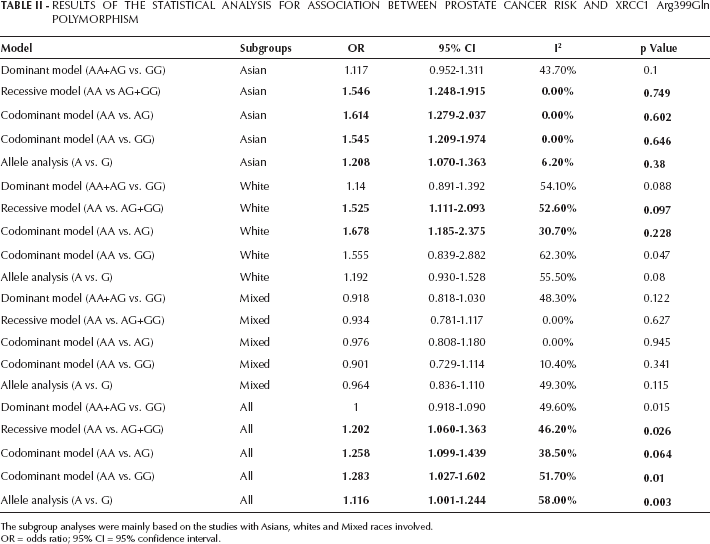
The subgroup analyses were mainly based on the studies with Asians, whites and Mixed races involved.
OR = odds ratio; 95% CI = 95% confidence interval.
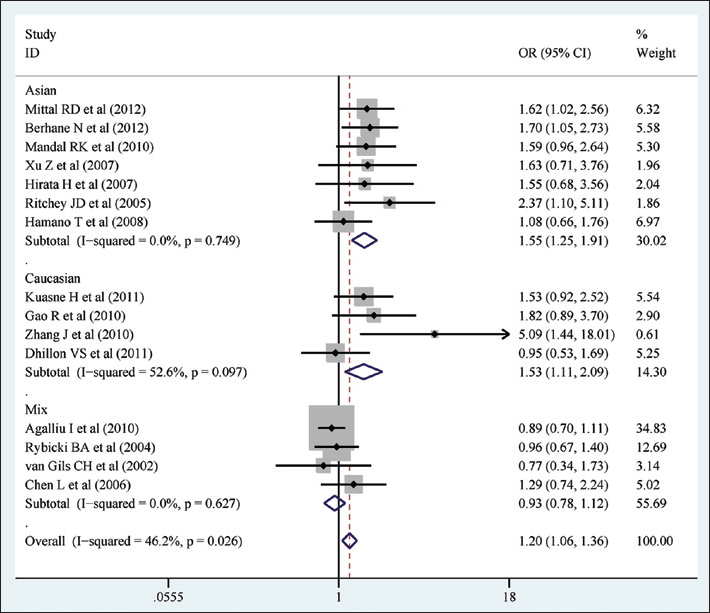
Meta-analysis with fixed effects and recessive model (AA vs. AG+GG) for the association between the XRCC1 Arg399Gln polymorphism and prostate cancer. The first author and year of publication for each study are shown. In this analysis, 3 subgroups are shown: Asian, white and mixed. Odds ratio (OR) and accompanying 95% confidence interval are also shown for the association of the XRCC1 Arg399Gln and prostate cancer.
There are 4 studies investigating mixed races (16, 20–21–22). Three of them provided data for whites independently (16, 20, 22). As for the remaining 1 study (21), the individual data for whites or other races could not be differentiated. Considering this, our data were rearranged. Finally, 7 sets of data for Asian and Whites, 2 for African Americans and mixed races were included. In the new results, the positive associations were again found in the pooled analysis and Asian subgroups. However, the significant associations for whites found in the first stage analysis disappeared (Tab. III).
Relations Of Prostate Cancer Risk And Xrcc1 Arg399Gln Polymorphism In The New Subgroups Analysis
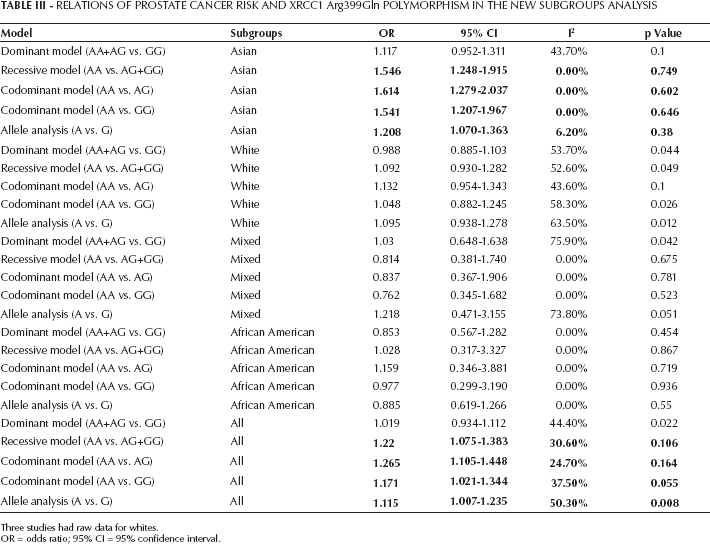
Three studies had raw data for whites.
OR = odds ratio; 95% CI = 95% confidence interval.
Characteristics of XRCC1 Arg399Gln polymorphism
After the traditional meta-analysis, we tried to describe the general characteristics of this SNP. Then eQTL, LD, TagSNP and functional analysis were conducted. The eQTL was analyzed using Genevar with the Hapmap database in 4 different populations (CHB, JPT, CEU and YRI). However, it seemed that rs25487 was not a cis-regulated element of the XRCC1 gene (CEU: rho=-0.161, empirical p value (pemp)=0.0968; CHB: rho=-0.205, pemp=0.0714; JPT: rho=0.127, pemp=0.2576; YRI: rho-=0.05, pemp=0.622) (Fig. 2), even though further analysis suggested that rs25487 might be the tag and functional SNP. In addition, the evident LD among the CHB+JPT and CEU populations hinted that the SNP might be a useful marker in both Asian and whites (Fig. 3).
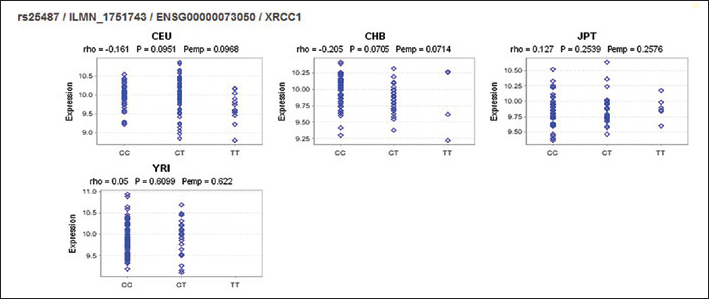
The results of expression quantitative trait loci (eQTL) analysis with Genevar in 4 different populations: Han Chinese from Beijing, China (CHB), Japanese from Tokyo, Japan (JPT), Utah residents with Northern and Western European ancestry (CEU) and Yoruba trios from Ibadan (YRI).
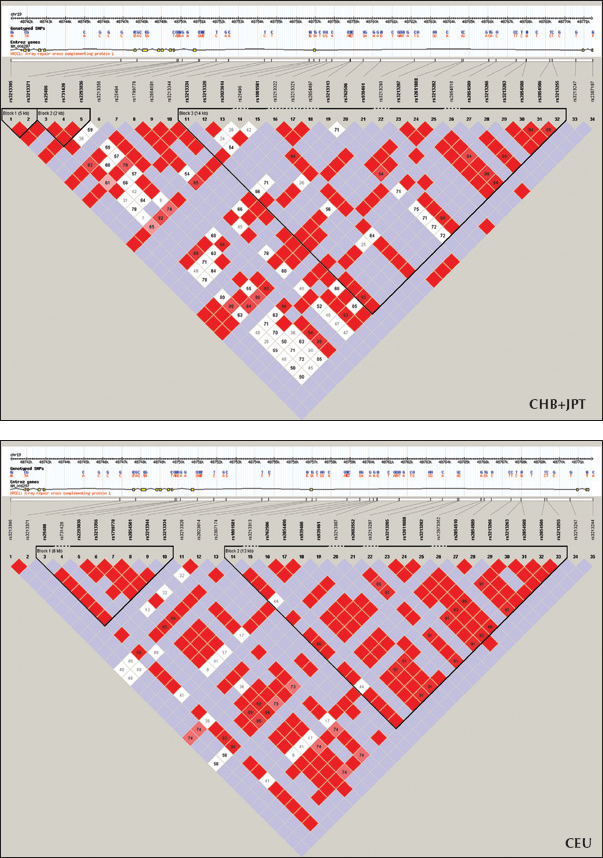
Linkage disequilibrium (LD) maps for the XRCC1 gene for CHB+JPT (top) and CEU (bottom) with Hapmap data. XRCC1 Arg399Gln (rs25487) is in one of the LD block. The stronger LD relationships are shown with a darker color. CHB = Han Chinese from Beijing, China; JPT = Japanese from Tokyo, Japan; CEU = Utah residents with Northern and Western European ancestry.
Discussion
Prostate cancer is a terrible disease affecting a large number of people. Although enormous damage is induced, an accurate pathogenesis has not been clearly described, in which genetic and environmental elements would play a key role (3). Recently, many studies have shown that the XRCC1 Arg399Gln polymorphism is one of genetic factors. This study was conducted to discover the potential association between the SNP and Pca risk. Combining the results of a meta-analysis and the features of this SNP, we suggest that XRCC1 Arg399Gln polymorphism might be significantly related to Pca risk in both Asians and whites.
After the first-stage meta-analysis, a positive association between Pca risk and XRCC1 Arg399Gln polymorphism was shown. In addition, a relationship was found in Asian and Whites, when the subgroup analysis was conducted. To investigate that relationship more comprehensively, we regrouped the studies with mixed races in which the explicit populations were extracted. However, in the new analysis, the significant association in whites disappeared. Combined with the stage 2 meta-analysis, the potential association between the XRCC1 Arg399Gln polymorphism and Pca was identified especially in Asians. The results have been repeated in recent studies. In 2007, Xu et al (17) suggested that XRCC1 appears to influence the risk of Pca. At the same time, Hirata et al also showed a significant association (18). As key DNA repair genes, polymorphisms are said to be important in the development of cancers. In 2014, Wang and Ai reported on the association of XRCC1 polymorphism and thyroid cancer, which suggested that XRCC1 played a vital role in the development of that cancer (40). In addition, its importance in the development other cancers has also been shown, such as in glioma (41), colorectal cancer (42) and so on. So based on our own results, we have reasons to believe that XRCC1 Arg399Gln polymorphism might be a factor in the development and progression of Pca.
Although the association between the XRCC1 Arg399Gln polymorphism and Pca risk was found among Asians, the results in whites seemed to be unclear. To view the features of this particular SNP, the HapMap database was used. Our further analysis showed that rs25487 was a tag and functional SNP with a distinct LD block in both Asians and whites. Considering these results and our meta-analysis, we still believe that this SNP might also be important in the Pca development in whites. In 2004, Rybicki et al (20) studied the association in XRCC1, XPD and prostate cancer risk. The results showed that XRCC1 could assist the XPD in influencing a risk for Pca. In summary, the XRCC1 Arg399Gln polymorphism is one of the important SNPs associated with the development of Pca, which could provide a clue for further targeted therapy.
Limitations
Although using comprehensive retrieval and analysis, there were still some limitations for this study (1). Three databases were used in this meta-analysis. The limitations of those databases could lead to a selection bias (2). The confusing description and incomplete genotypes data in the studies would also introduce a bias into the results. There are 2 examples: One study investigated the association between XRCC1 Arg399Gln polymorphism and Pca risk, but we could not extract the raw data for every genotype (39). In the other study, the descriptions of SNPs were confusing (Arg309Glu and Arg399Glu). To get accurate data for the Arg399Glu polymorphism, we used the data in Table VI of that study report (4). (iii) Last but not least, the studies had some differences in the definitions of cases and controls, although, the studies did have similar inclusion criteria. So a potential bias could also be introduced there.
Conclusion
Prostate cancer is a worldwide cancer. Genetic factors are said to be associated with its development and progression. As a common polymorphism of DNA repair gene, the XRCC1 Arg399Gln polymorphism has been identified to be a key one. Our study suggested that the XRCC1 Arg399Gln polymorphism could be significantly associated with Pca risk in both Asians and whites. Further studies are needed to confirm this.
