Abstract
Objectives
Methods
Results
Conclusions
Introduction
Several studies have suggested that the mini-chromosome maintenance proteins (MCM) may be promising proliferation markers in many different tumor types (1, 2). MCMs are typically induced in the DNA-synthesis phase of the cell cycle, as they are prerequisite for DNA replication and cell-cycle initiation (3). The proteins of the MCM family have similar biochemical functions and consist of 7 major isoforms (MCM2-8).
The identification of new biomarkers that allow the monitoring of molecular events associated with tumor progression in histological or cytological specimens may improve the detection of lesions with higher risk of progression. A number of potential biomarkers for cervical screening, such asp16INK4a and E6-E7 human papillomavirus (HPV) proteins, have been positively related to the identification of women at greatest risk for developing cervical cancer (4, 5). However, to date, there have been only few studies demonstrating the prognostic implications of the new predictive biomarkers in the staging of cervical cancer, as well as identifying patients with high risk of relapse, and optimizing disease management, especially when cervical cancer is initially diagnosed and the clinical stage is either unknown or difficult to assess. To the best of our knowledge, this is the first study to correlate MCM2 expression with the clinical stage of cervical cancer. We aimed to evaluate the immunohistochemical expression of MCM2 in order to examine the hypothesis that a cell cycle-related protein is overexpressed in the uterine cervix of patients with more advanced cervical cancer and evaluate the correlation between MCM2 expression and HPV genotype.
Material and Methods
Tissue Samples and Data Collection
The study material consisted of 130 samples randomly selected from the archives of the Fernandes Figueira Institute, Oswaldo Cruz Foundation, Rio de Janeiro, Brazil. Samples from 87 patients obtained by cervical biopsy or patients' conization specimens were histopathologically reviewed by an expert pathologist and confirmed as invasive cervical cancer. All patients referred to the hospital between 2003 and 2008. Cervice specimens from 43 patients undergoing hysterectomy for benign leiomyomata disease were used as negative controls.
The institutional review board from the Oswaldo Cruz Foundation (Fiocruz), (CAE 0024.0.011.000-09, Rio de Janeiro, Brazil) approved the study, and written consent was required from each patient for inclusion in this study.
Tissue Micro-Array Block Construction
The Tissue micro-array (TMA) blocks were constructed as described by Pires et al in 2006 (6). Briefly, all hematoxylin-eosin (HE) slides were re-examined by a second experienced pathologist and 2 morphologic representative fields of invasive cervical cancer were chosen and encircled with a marker pen. Two cores from each case were punched out from the donor blocks. The corresponding HE slides were overlaid with a custom-built 16 gauge Becton-Dickinson PrecisionGlide® hypodermic needle (area, 1.1 mm2). Then, cores were attached with double-side adhesive tape on a computer-generated paper grid affording alignment on the block mould, which was then filled with liquid paraffin. Three-μm thick sections were obtained from an optical standard rotator microtome (Leica, Bensheim, Germany). Each block provided 40-50 slides containing variable tissue core lengths. Only samples showing the original lesion were used in this study. Normal tissue (placenta and kidney) and position-specific blank cores were adopted for orientation during microscopy analysis. A total of 2 TMA blocks were constructed, 1 with cervical cancer biopsies and the other with controls, in which the absence of cervical intraepithelial neoplasia (CIN) was confirmed by review of the HE-stained slide.
DNA Extraction
From each paraffin-embedded cervical biopsy, 4 slices of 5 µm each were cut. Briefly, after dewaxing with xylene at 48°C for 2 hours, ethanol (100%, 70%, and 50%) was added to remove the residual xylene, followed by rehydration and centrifugation at 13,400 g for 15 minutes. The pellet was resuspended in a solution with 300 µL of proteinase K (100 µg/mL) and 10% SDS for 48 hours at 48°C. Five microliters of additional proteinase K (200 µg/mL) were added after 24 hours. DNA was isolated with a phenol extraction protocol containing 300 µL of phenol/chloroform/isoamyl alcohol (in the proportions: 25/24/1) followed by centrifugation (13,400 g for 10 minutes) to obtain the aqueous phase. After repeating these purification steps twice, the DNA was precipitated at 20°C for 24 hours with 100 µL sodium acetate (final concentration 7.5 M) and twice the aqueous phase volume of cold 100% ethanol. The DNA pellet was collected by centrifugation for 15 minutes at 13,400 g at 4°C, washed with 70% ethanol and resuspended in 50 µL of TE buffer (10 mM Tris base, 1 mM EDTA, pH 7.5). The DNA quantity and quality was analyzed by NanoDrop (Thermo Fisher Scientific, Wilmington, DE, USA).
PCR Amplification and HPV Genotyping
Amplification of the HPV L1 consensus region was performed with generic primers GP5+ and GP6+ (synthesized by Invitrogen) to generate a PCR product of approximately 150 bp as previously described (7, 8); the negative samples were subjected to a nested PCR with primers PGMY09 and PGMY11, as previously described (9), synthesized by Invitrogen. Each PCR assay included a positive control as HeLa cellular DNA (HPV 18 DNA) and a negative template control. The integrity of the specimen's DNA was verified by amplification of a 110-bp fragment of the β-globin gene with primers PC03 and PC04 (synthesized by Invitrogen). Table I depicts the forward and reverse primers used in this study.
OLIGONUCLEOTIDES USED FOR HPV DNA DETECTION
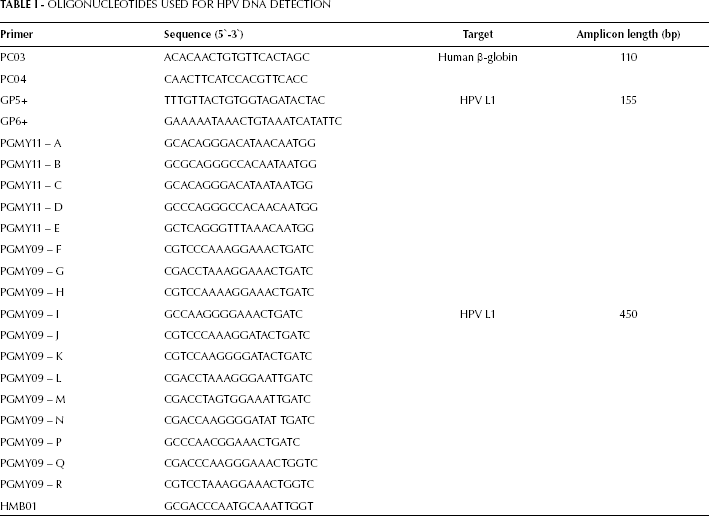
The amplifications were carried out in a GeneAmp PCR System 9700 thermal cycler (Applied Biosystems), and all PCR products were analyzed by ethidium-bromide stained electrophoresis in 1.5% agarose gel, as previously described.
A final reaction volume of 75 µL was purified by the GF-1 DNA Recovery Kit (Vivantis, Oceanside, CA, USA), in order to determine the HPV genotype. For sequencing, we prepared a solution containing 3.2 pmol of each primer (GP5+ or GP6+), 5-10 ng of PCR product, 2.5 µL Big Dye Terminator v 3.1 Cycle Sequencing Kit (part number 4337455; Applied Biosystems, Foster City, CA, USA), and MilliQ water completing for a final volume of 10 µL. The purified mixtures were analyzed on an ABI Prism 3730 Genetic Analyzer (Applied Biosystems) available at the DNA PDTIS/Fiocruz Platform (10). Alignments were obtained directly from online Blastn server or phylogenetically analyzed by the software MEGA5 (11). INNO-LiPA HPV Genotyping v2 (Innogenetics, Gent, Belgium) was used in 9 cases that were not amplified by PCR or were not able to be sequenced. Additionally, 3 other doubtful cases were assayed by the PapilloCheck Kit (Greiner Bio-One, Frickenhausen, Germany) in order to confirm the HPV DNA type. The distribution of HPV types detected by these kits are displayed in Table II.
DISTRIBUTION OF HPV GENOTYPING BY INNO-LiPA AND PAPILLO CHECK KIT
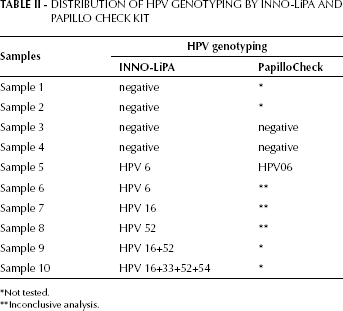

Not tested.
Inconclusive analysis.
In Situ Hybridization
HPV DNA detection was also determined by in situ hybridization (ISH), following a previously published protocol (12). In brief, the biotin-labeled probe cocktails can detect the high-risk HPV types (HPVs 16, 18, 31, 33, 35, 45, 51, 52, 56, 58, 68) (Enzo Life Sciences, Farmingdale, NY, USA). The probe-target complex is detected due to the action of alkaline phosphatase on the chromogen nitroblue tetrazolium and bromochloroindolyl phosphate (NBT/BCIP) yielding a dark blue color with a pink counterstain for the HPV negative cells due to nuclear fast red.
Immunohistochemistry (IHC)
IHC reactions were performed on TMA silane-coated slides (Sigma, St. Louis, MO, USA). Briefly, slides were dewaxed in xylene, rehydrated through a graded ethanol series, washed with distilled water and then treated in solution with methanol containing 0.3% hydrogen peroxide for 10 minutes to eliminate endogenous peroxidase activity. Antigen retrieval for IHC was performed by boiling the tissues with the Target Retrieval Solution (Dako, Carpinteria, CA, USA). Non-specific antibody binding was inhibited by incubating sections with serum-free protein block (1% BSA). Tissue sections were sequentially incubated overnight in a humidified chamber at 4°C with the specific primary monoclonal antibody anti-MCM2 (dilution, 1/100; Abcam, Cambridge, MA, USA). A secondary biotinylated multilink antibody was applied for 30 minutes followed by streptavidin-peroxidase incubation for 30 minutes. The enzymatic reaction was developed in a freshly prepared solution of 3.3'-diaminobenzidine using DAKO Liquid DAB Substrate-Cromogen solution as a chromogen (DAKO, Glostrup, Denmark) and exposed for 5 minutes. Finally, slides were counterstained with hematoxylin for 1 minute and permanently mounted in Entellan (Merck, Darmstadt, Germany).
Assessment of IHC Slides and Cell Counting
Two independent observers reviewed and quantified the expression of the IHC stains. Only the nuclear-based signal for MCM2 was scored. Two representative areas of each core were manually counted by the observers and classified as: negative, less than 25% of positive cancer cell nuclei, 25%-75% of positive nuclei, or more than 75% of positive nuclei.
Statistical Analysis
Data were analyzed by means of the STATA/SE 10.1 software, with the non-parametrical tests of Kruskall-Wallis, Student's t, Fisher's exact, and chi-square, as appropriate. A test for trend across the ordered groups (nptrend) was used to determine if MCM2 expression among FIGO stages followed a linear trend. P values <0.05 were considered statistically significant.
Results
A total of 130 women (average age 51.1±13.1 years; range 24-88 years) were included in this study. The relative degree of MCM2 expression, as determined by IHC, was compared between invasive cervical cancer cases and controls, among the different FIGO stages in cervical cancer, and among HPV genotypes in the HPV-positive invasive cancer specimens.
HPV Prevalence and Distribution
The overall prevalence of HPV DNA in the control samples and invasive cancer specimens was 29.3% (12/41) and 95.4% (84/87), respectively. Two of the 43 control samples yielded unsatisfactory results for DNA extraction and were excluded from HPV DNA prevalence analyses. Three negative samples by PCR with GP5+/6+ were positive for HPV DNA by a nested PCR using the PGMY09/11 primers. The HPV genotypes' distribution in the control group and the invasive cervical cancer group is shown in the Table III. Infections with more than 2 types of HPV in a single sample were rare, with only 2.2% (2/92) of HPV-positive samples showing multiple infections. In the control group, HPV type 16 was the most prevalent (66.7%) followed by types 35, 67, and 6 (8.3% of prevalence for each type). In women with cervical cancer, HPV type 16 was also the most common (66.6%), followed by type 33 (11.8%), and type 35 (3.6%). HPV type 58 and 6 were found in 2 cases (2.4%), while HPV 18, 31, 52, 67, 70, 69, and 30 were found in only 1 case (1.2%). Women infected with HPV 16 exhibited a statistically significant association (p=0.012) with a lower average age (mean, 50.9 years; SE, 1.9 years) compared to women with other types of HPV (mean, 59.9 years; SE, 2.8 years).
HPV GENOTYPE DISTRIBUTION
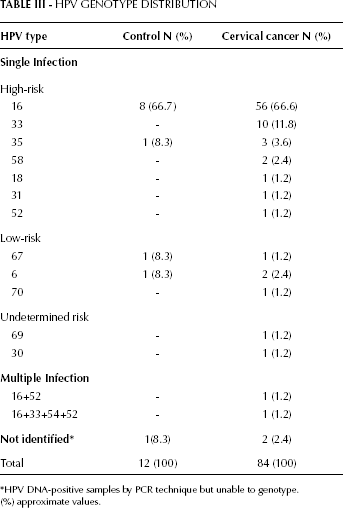
HPV DNA-positive samples by PCR technique but unable to genotype.
(%) approximate values.
MCM2 Expression
MCM2 was quantified by 2 different techniques: manual quantification and Image Pro-Plus Software. The MCM2 was expressed exclusively in the nuclear compartment of all positive cases, in 32 (78.1%) of the controls, and 87 (100.0%) cervical cancer samples. There was a marked difference in the histologic distribution of MCM2 in the normal cervical epithelia when compared to the cervical carcinoma epithelia. In control samples, more than 50% of positive cells were strictly restricted to the basal layer of the epithelium. In the cervical cancer tissue, MCM2 expression was most often found throughout the entire thickness of the epithelia (Tab. IV and Fig. 1).
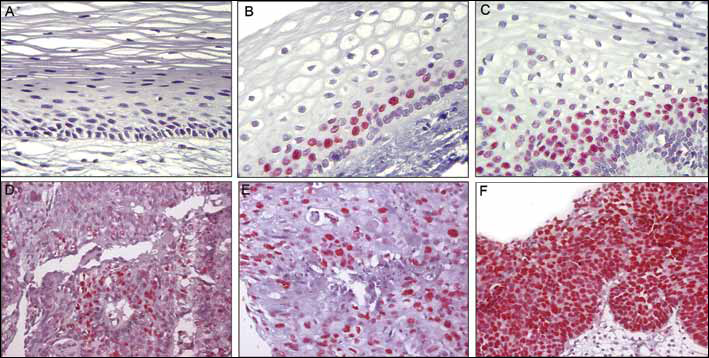
MCM-2 expression.
MCM2 FREQUENCY
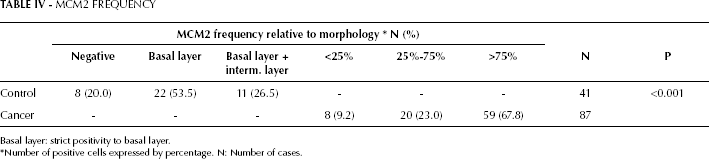
Basal layer: strict positivity to basal layer.
Number of positive cells expressed by percentage. N: Number of cases.
We observed a statistically significant association between the ISH signal for HPV, as defined by the percentage of cancer cells positive for viral DNA, and MCM2 expression (p<0.001) (Fig. 2 and Tab. V). However, no statistically significant association was observed between MCM2 expression, FIGO stages (p=0.462), or the specific genotype of HPV when comparing HPV 16 to the cancer cases positive for HPV types other than HPV 16 (Tab. VI).
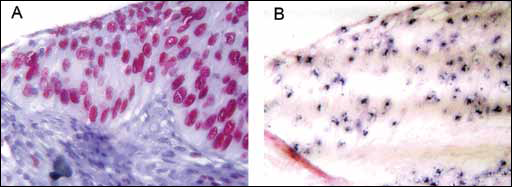
MCM-2 and In situ hybridization stain in a unique spot. (
CORRELATION BETWEEN MCM2 EXPRESSION AND HPV DNA PREVALENCE BY IN SITU HYBRIDIZATION
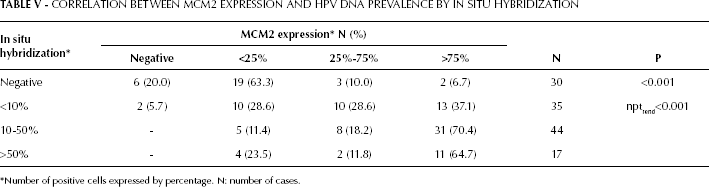
Number of positive cells expressed by percentage. N: number of cases.
ASSOCIATION AMONG MCM2 EXPRESSION, FIGO STAGES AND HPV TYPES
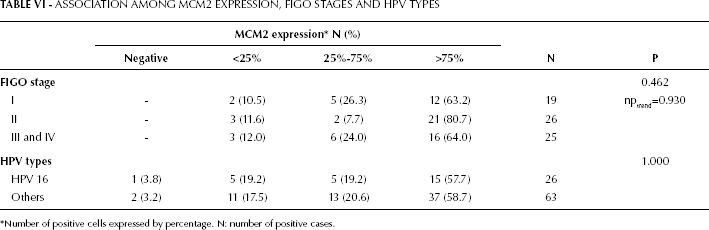
Number of positive cells expressed by percentage. N: number of positive cases.
No difference was observed in terms of MCM2 expression, measured as percentage of positive cells per field, between the 4 HPV-negative cases (case 1: >90%, case 2: <10%, case 3: 50%-75% and case 4: 75%-90%) and the others samples.
Discussion
It is well known that progression to cervical cancer is associated with increased expression of the HPV high-risk proteins E6 and E7, which are able to interact with cellular proteins and disrupt the process of controlled cellular proliferation (13). Thus, many studies have been trying to describe proteins that are overexpressed in cervical cancer in order to better elucidate the molecular pathways that are activated during disease progression (14).
MCM2 has been recognized as a classical marker of proliferation and a component of the pre-DNA replication complex that is overexpressed in cancers (15). Many studies have been directed mainly to the analysis of biomarkers involving precancerous lesions but only few studies have been performed to demonstrate the prognostic implications of MCM2 expression in the progression of invasive cervical cancer. Some well-known prognostic factors including tumor size, depth of invasion, lymph node status, grade of differentiation, and histopathological findings have been correlated with the tumor stage in cervical cancer. These factors have important implications in clinical management. For example, when a cervical cancer is staged between I and IIA, radical surgery or radiation therapy are adopted, and when it is diagnosed as IIB-IV, then chemoradiotherapy is used.
The determination of the clinical stage of cervical cancer is often based on tools such as computed tomography and ultrasonography (16). However, these diagnostic modalities have a relatively low sensitivity and high rates of false negative results. Thus, studies aiming to find new predictive markers for cervical cancer development are important in order to identify patients at high risk of recurrence or progression (17).
In this study, we found that MCM2 was strongly expressed in all layers of the squamous epithelium with invasive cervical cancer. This protein was also found in normal tissue, but only in small proportions and restricted to the parabasal layer region, which is the layer that is responsible for the generation of the overlying, more mature, squamous cells in the cervical epithelia. These data are in agreement with two others studies (4, 18).
In our previous publication, we found that the distribution of MCM2 closely followed the histologic features of the cervical squamous epithelium. Little to no expression of MCM2 was evident in normal and atrophic squamous epithelia of the cervix. In CIN 1 lesions, the basal epithelia and dysplastic squamous cells immediately above this layer showed a strong expression of MCM2. However, the koilocytes at the surface of the CIN 1 lesion showed very little expression of MCM2. As one progresses from CIN 2 to CIN 3, a greater percentage of dysplastic cells were positive for MCM2 to the point that in a lesion showing in situ carcinoma basically all the dysplastic cells were MCM2-positive. These data strongly suggest that MCM2 expression is characteristic of immature dysplastic cells in the cervix as compared to Ki-67, which is more typical of mitotically-active squamous cells both benign and dysplastic. Moreover, analyzing different degrees of cervical lesions (CIN 1, CIN 2/3 and cervical cancer) we found a low specificity (66.0%) and negative predictive value (57.0%), suggesting that MCM2 is not a powerful biomarker for the diagnosis of CIN (19). By contrast, another study showed a considerable advantage using MCM2 for cervical screening to detect abnormal specimens, as cancerous and pre-cancerous lesions, which were not detected by Pap smears (20).
It is clear that MCM proteins are excellent markers to distinguish normal tissues from cancerous tissues. However, the biomarker's capacity to differentiate lesions in progression from invasive cervical cancer is still unclear. Beyond that, only few studies have reported the downregulated expression of MCM proteins in different stages of the invasive cervical cancer. One recent study (21) described the expression of MCM2 in combination with clinical staging according to FIGO classification. Their results are in agreement with those of our study, also confirming our analysis of MCM2 expression by IHC. However, despite the fact that we found a high overall expression of MCM2, we did not find any significant difference among the distinct tumor stages, probably due to the small number of specimens included.
Four out of 87 samples were HPV DNA-negative by ISH and PCR. In these samples, no difference was observed in terms of MCM2 expression. Thus, either there is no correlation between HPV and MCM2 expression levels, or these techniques were not able and sensitive enough to identify HPV DNA, the latter hypothesis suggesting poor DNA quality or low viral load.
Furthermore, we observed that the group with the highest incidence of cervical cancer included women over 55 years of age (37/77, 45.5%). This data is consistent with the percentages reported in Brazil (22, 23). Women with HPV 16 had an average age of 50.8 years, while those infected by other HPV types had an average age of 58.2 years, suggesting that HPV 16-infected women may develop cervical cancer earlier than those infected by other HPV types. Several other studies also noted this association. Liu et al in 2004 showed that tumors positive for HPV 16 and HPV 18 were more often associated with younger age of the women infected (aged less than 45 years), while other HPV types (HPV 31, 33, 35, 39, 45, 51, 35, 39, 45, 51, 67, 68, and 73) were associated with older age of women (24). Sanjose et al in 2010 also observed this association in women with invasive cervical cancer in 34 countries. In this study, women diagnosed with HPV 16, 18, or 45 showed, respectively, a mean age of 50.0, 46.8, and 55.5 years (25).
The role of HPV as an etiological agent of cervical cancer and its earlier lesions has been well documented. The variations in the prevalence of HPV in samples of invasive cervical cancer range from 80.0% to values close to 100.0%, and are probably associated with different detection techniques (23-24-25-26). Among them, the polymerase chain reaction (PCR) has proven to be effective in studies on the prevalence of HPV DNA (27, 28).
In our study, we aimed to determine the prevalence of HPV DNA, and to identify the HPV genotypes. For this, we chose to use the primer pair GP5+/6+. We could detect HPV DNA in many invasive cervical cancer specimens. Among these, 95.4% (83/87) were successfully genotyped. We found a highly satisfactory prevalence compared to other studies that also used paraffin-embedded specimens (29). The 6 more prevalent types of HPV in Central and South America are HPV 16, 18, 31, 45, 33, and 58 (28, 30-31-32).
MCM2 was quantified by 2 different methods in this study: manual quantification and Image Pro-Plus software. The literature has described the utilization of the software for the analysis of various parameters such as total area, density, and integrated optical density. Some authors suggested that these programs may be more advantageous than hand counts (33, 34). The software was programmed to perform the quantification of the marked area (per area) by immunohistochemistry. We found equivalent data when using either quantification method, thus suggesting that either method may be adequate in studies correlating protein, RNA, or DNA expression with clinical variables. Interestingly, we found a strong correlation between the MCM2-positive cells and the presence of HPV DNA detected by ISH, suggesting that the HPV-infected cells are also able to express MCM2.
The present study had some limitations, such as the relatively small number of specimens for each different invasive cancer stages, limiting subgroup analyses. However, our data brought important developments regarding MCM2 expression and cancer stages association, since the literature is still limited on this.
In conclusions, we found a linear correlation between the percentage of HPV DNA-positive cells, determined by ISH, and the increased expression of MCM2. Our data suggest that MCM2 may not be a good biomarker to differentiate the progression of cervical cancer from stage I to stage IV; however, further studies should be performed with different invasive cancer stage specimens in order to evaluate the potential role of this biomarker.
