Abstract
Background
The CD24 gene has been correlated with poor prognosis of various malignancies. The significance of CD24 in esophageal cancer remains unknown. Our aim was to evaluate the association between CD24 genetic polymorphism and esophageal cancer.
Materials and Methods
Between June 2011 and May 2012 patients with esophageal cancer and healthy controls were prospectively enrolled and clinicopathological data were collected. Genomic DNA was extracted and restriction fragment length polymorphism (RFLP) analysis was performed to determine CD24 polymorphism at the coding region of CD24, which results in a substitution of the amino acid Ala by Val. Statistical significance was determined by unpaired t-test, χ2-test, and Fisher's exact test.
Results
A total of 102 patients were included, of whom 51 had esophageal cancer and the rest comprised a healthy control group. The incidence of the polymorphism variant (Val/Val) among the healthy subjects and the esophageal cancer cohort was 6% in both groups. The incidence of N3 (metastasis in 7 or more regional lymph nodes) was markedly higher in those esophageal cancer patients who carried the polymorphism variant compared with those who did not carry it (66% and 2%, respectively, p=0.007). No significant difference was found between the groups with regard to age, gender, histology type, tumor location, tumor stage, and other histological characteristics of the tumor.
Conclusions
This CD24 polymorphism may serve as a novel prognostic marker identifying esophageal cancer patients with poor prognosis. Further studies are warranted to evaluate CD24 function and to validate its predictive potential with regard to esophageal cancer.
Keywords
Introduction
Esophageal cancer is the eighth most common cancer worldwide (1). At presentation, the overwhelming majority of patients have locally advanced, regionally advanced, or disseminated cancer, irrespective of histologic type (2, 3). The lack of a serosal envelope and the rich submucosal lymphatic network of the esophagus lead to extensive local infiltration and lymph node involvement, whereas the lung, liver, and bone are the most common sites of distant disease. As a result of this aggressive behavior, multidisciplinary treatment modalities including surgery, chemotherapy, biologic therapy, and radiotherapy are utilized to counter the disease. Unfortunately, despite treatment, the overall 5-year survival rate ranges between 20% and 25% (4, 5).
Currently, the most commonly used conventional prognostic factors for patient survival are tumor grade and stage at the time of diagnosis, including depth of tumor invasion, involvement of regional lymph nodes, and metastatic spread to distant organs (6). Thus, it is vital to uncover novel prognostic markers than might allow earlier identification of esophageal cancer and facilitate an individualized multidisciplinary approach.
The human CD24 gene translates into a small peptide, of 31 amino acids, homologous to the mouse CD24/heat stable antigen (HSA) (7, 8). CD24 is a short mucin-like heavily glycosylated protein, anchored to the plasma membrane by a glycosylphosphatidylinositol (GPI) linkage. Its peptide backbone harbors multiple sites for both N- and O-glycosylation. The glycosylation pattern vastly diverges within cells and among cells from different tissues (9, 10).
CD24 overexpression has been demonstrated in various human malignancies and often correlates with poor survival (9, 11). High expression of CD24 is reported with a high prevalence in B-cell lymphomas, gliomas, small-cell and non small-cell lung, hepatocellular, uterine, ovarian, breast, prostate, colorectal, and pancreatic carcinomas (11). CD24 is usually associated with a more aggressive course of the disease (12, 13).
Functionally, CD24 has been identified as an alternative ligand of P-selectin, an adhesion receptor on endothelial cells (14–16), and might therefore enhance the metastatic potential of tumor cells (9, 14). CD24 expression has also been shown to regulate tumor cell proliferation and migration (17).
Recently, CD24 has been implicated as a central factor in autoimmune diseases. Two polymorphisms within the CD24 gene were shown to modify disease risk and progression in multiple sclerosis (MS), systemic lupus erythematosus (SLE), giant cell arteritis, and, most recently, chronic hepatitis B (18–24). The first one, a C to T single-nucleotide polymorphism (SNP; rs52812045) at position 170 from the CD24 translation start site, is located in the putative GPI-anchor cleavage site, whereby a valine (V) substitutes an alanine (A). The P170C/T (CD24 V/V) genotype is associated with an increased risk and more rapid progression of MS, SLE, giant cell arteritis, and chronic hepatitis B (18, 20, 21, 23, 24). The second polymorphism, a dinucleotide deletion (TG/Del) (rs3838646) within the 3′ untranslated region (UTR), highly reduces CD24 mRNA stability and is seemingly protective (18, 20, 21, 23, 24). This data suggest that CD24 plays an important role in modulating the immune response and autoimmunity. Moreover, we have previously reported that the risk of developing colorectal cancer increases significantly in individuals with a polymorphism both in the CD24 and APC genes compared to individuals with an APC polymorphism alone (25).
The clinical significance of the CD24 gene in esophageal cancer remains unknown. A single study, by Sano et al (26), has evaluated the correlation between esophageal cancer and the CD24 gene. The authors demonstrated an association between CD24 expression and pathologic stage, as well as disease-free survival, of patients with esophageal squamous cell carcinoma. To our knowledge, no other study has evaluated the correlation between the CD24 gene and esophageal cancer. Thus, in this study our aim was to examine the association between the P170C/T CD24 genetic polymorphism and esophageal cancer.
Materials and Methods
Patients
Between June 2011 and May 2012 a total of 102 subjects, of whom 51 with esophageal cancer and 51 healthy controls, were prospectively enrolled. The 51 patients with esophageal cancer were recruited from the Department of General Surgery at the Rabin Medical Center. Age and gender-matched controls were enlisted from healthy subjects attending the Integrated Cancer Prevention Center at Tel Aviv Sourasky Medical Center; these subjects had no evidence of inflammation or neoplasia.
All participants signed a written informed consent form, underwent physical examination, completed a lifestyle-habits questionnaire, and provided a blood sample. Preoperative clinical staging of esophageal cancer included CT and endoscopic ultrasound (EUS). Neoadjuvant treatment was offered to patients with stage IB or higher. Pathological examination of esophageal specimens was performed by a pathologist who was blinded to both the results of the CD24 polymorphism and the health status of the participants. Final staging was judged by the highest combination of the preoperative clinical staging and the postoperative pathological staging. Approval for this study was provided by the local institutional review board (IRB) and the Israeli Ministry of Health. Blood samples from the participants were preserved and stored at 4°C until processing on the same day of sample collection.
Assay methods: isolation of peripheral blood leukocytes
Peripheral blood leukocytes (PBLs) were isolated from whole blood samples by collecting white buffy coats obtained after blood centrifugation for 3 minutes at 3,000 rpm and discarding the plasma supernatant.
DNA extraction and genotyping
Genomic DNA was extracted from peripheral blood leukocytes using standard procedures. DNA samples were examined for their CD24 genotype using polymerase chain reaction restriction fragment length polymorphism (PCR-RFLP) analysis as described below.
Determination of the CD24 polymorphism
The reported SNP at the coding region of CD24 replaces a C nucleotide with a T (C→T) at position 170 (GenBank accession no. NM_013230), which results in a substitution of the amino acid Ala by Val at position 57 near the GPI anchorage site of the mature protein. DNA samples were analyzed for their CD24 genotype by PCR amplification, using primers embracing CD24 exon-2 followed by a restriction reaction using BstX1, the cleavage site of which is created only in alleles that harbor the polymorphic T.
Briefly, DNA fragments bearing the SNP were amplified by a PCR reaction using a forward primer (TTG TTG CCA CTT GGC ATT TTT GAG GC) and a reverse primer (GGA TTG GGT TTA GAA GAT GGG GAA A). The predicted CD24 PCR fragment is 522-bp long. The C→T change yields a BstXI restriction enzyme site at nucleotide 327, allowing the differentiation between the 2 different CD24 alleles by RFLP analysis.
Statistical analysis
Categorical variables were compared using the χ2 or Fisher's exact test depending on the number of observations, while the Student's t-test with a two-tail distribution was used for continuous variables. A p value of less than 0.05 was considered significant. The statistical software for the social sciences (SPSS) version 21 was used to calculate significance.
Results
A total of 102 subjects were included, of whom 51 had esophageal cancer, and the remaining 51 represented a healthy control group. The demographic data of the esophageal cancer patients are summarized in Table I. The greater part of cancer patients were males (31 patients, 61%), and 32 (61%) had an adenocarcinoma histology. The most common tumor location was at the gastroesophageal junction (GEJ). Neoadjuvant treatment was given to 30 patients (59%), and the greater part of the patients was at stage III (44%).
Demographic Data Of The Esophageal Cancer Cohort (51 Patients)
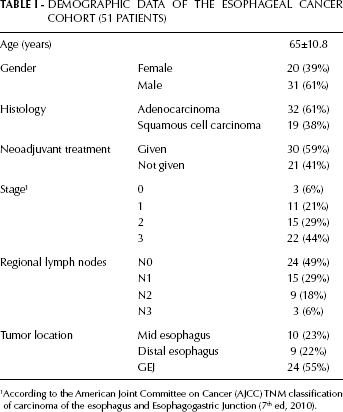

According to the American Joint Committee on Cancer (AJCC) TNM classification of carcinoma of the esophagus and Esophagogastric Junction (7th ed, 2010).
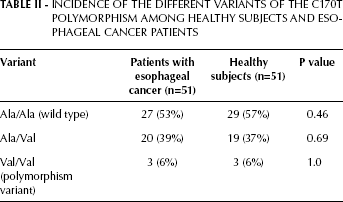
The incidence of the different variants of the C170T polymorphism among healthy subjects and esophageal cancer patients is summarized in Table II. The polymorphism variant rate resulted identical in the 2 groups (6%). The other variants, i.e. the wild type (Ala/Ala) and the heterozygote (Ala/Val), did not show any significant difference between the 2 groups.
Incidence Of The Different Variants Of The C170T Polymorphism Among Healthy Subjects And Esophageal Cancer Patients
The clinicopathological differences between the esophageal cancer patients who carry the polymorphism variant (Val/Val variant of the C170T polymorphism) and those who do not carry it are summarized in Table III. Despite the higher incidence of stage III patients among the polymorphism variant carriers (66%) compared to the non-carriers (50%), no statistical significance was demonstrated (p=0.57). Similarly, no significant difference was found between these groups with regard to age, gender, histology type, tumor location, and other histological characteristics of the tumor. Interestingly, we found that the incidence of N3 (metastasis in 7 or more regional lymph nodes) was markedly higher in those esophageal cancer patients who carry the polymorphism variant compared with those who do not carry it (66% and 2%, respectively, p=0.007). Of note, the heterozygous variant (Ala/Val) did not show any clinicopathological differences.
Clinicopathological Differences Between Esophageal Cancer Patients Carrying Or Non-Carrying The C170T Val/Val Polymorphism Variant
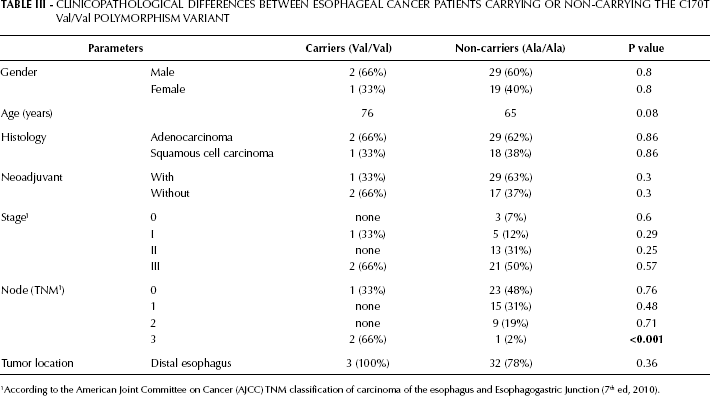
According to the American Joint Committee on Cancer (AJCC) TNM classification of carcinoma of the esophagus and Esophagogastric Junction (7th ed, 2010).
Discussion
CD24 is a small, highly glycosylated, mucin-like protein that is linked to the cell membrane via a GPI anchor. Recent literature has emphasized its involvement in B-cell development and neoplasia, in the developing pancreas and brain, and in regenerating muscle, keratinocytes, and renal tubules (12, 27–33). Moreover, CD24 seems to be functionally involved in the control of cell proliferation, apoptosis, and cell adhesion. In lymphocytes, CD24 can modulate the capacities of early T and B lymphoid progenitor cells to proliferate and survive. Antibody-mediated crosslinking of CD24 induces apoptosis in a process involving the B-cell receptor and mitogen-activated protein (MAP) kinases (33, 34). It has also been demonstrated that CD24 expression in a rat carcinoma system stimulates cell adhesion, migration, and proliferation (14). CD24 has been identified as an alternative ligand for P-selectin and the cell adhesion molecule L1-CAM (16, 35, 36). CD24 is able to support the adhesion of neutrophils or monocytes to activated endothelial cells or thrombin-activated platelets, both of which express P-selectin. CD24-expressing tumor cells can disseminate more readily through their capacities to form thrombi with activated platelets or adhere to endothelial cells in the bloodstream, properties that have been shown for CD24-expressing breast cancer cells (15). The interaction of tumor cells with P-selectin via CD24 may be an important pathway in cancer cell dissemination and metastasis formation. Furthermore, CD24 expression indirectly stimulates cell adhesion to fibronectin, collagen I, collagen IV, and laminin by triggering a3b1 and a4b1 integrin activities (14). CD24 is regulated by Ral GTPases, suggesting that CD24 and other potential transcriptional targets of Ral signaling may mediate important functional aspects of the members of this GTPase subfamily that are emerging as central players in cancer (37).
Recently, CD24 gene polymorphism has been associated with higher risk and progression of multiple sclerosis (MS), systemic lupus erythematosus (SLE), giant cell arteritis, and chronic hepatitis B (18–24). In addition, we previously demonstrated that the risk of developing colorectal cancer increases in the presence of combined CD24 and APC polymorphisms (25). Also, it has been shown that the CD24 Ala/Val SNP is a strong and independent predictor of pathological complete response after neoadjuvant chemotherapy for breast cancer (38). Evaluating polymorphisms as predictive markers is highly requested, as this simple and inexpensive approach could be routinely used in a clinical setting.
High expression of CD24 is reported with a high prevalence in various cancers including: B-cell lymphomas, gliomas, small-cell and non small-cell lung, hepatocellular, uterine, ovarian, breast, prostate, colorectal, and pancreatic carcinomas (11). CD24 is usually associated with a more aggressive course of the disease (12, 13, 39, 40). Only one study by Sano et al has evaluated the relationship between CD24 and esophageal cancer (26). The authors studied 151 patients with squamous cell carcinoma of the esophagus and demonstrated an association between high immunohistochemical expression of the gene and lymph node involvement, pathological stage, lymphovascular invasion, and disease-free survival. Our study has demonstrated an association between esophageal cancer patients who carry the Val/Val variant of the C170T polymorphism and N3 (metastasis in 7 or more regional lymph nodes).
To our knowledge, the relationship between the CD24 gene and esophageal cancer has been evaluated only by the studies of Sano et al (26) and ours. A limitation of these 2 studies is the small sample size. Despite that, the results of both studies are in agreement. Specifically, the involvement of the CD24 gene at different levels (at the level of genetic polymorphism in our study, and at the level of immunohistochemical expression in the study of Sano et al [26]) serves as a negative prognostic factor, as highlighted by the degree of lymph node involvement. Another limitation of our study is the fact that we did not recruit stage IV esophageal cancer patients.
In conclusion, the Val/Val variant of the C170T polymorphism may serve as a negative prognostic marker in esophageal cancer patients. Thus, it might aid physicians with providing an individualized treatment and monitoring high-risk patients. Future research with larger cohorts is needed to validate these preliminary results.
