Abstract
Background
Recent studies have shown that long noncoding RNA (IncRNA) gastric carcinoma highly expressed transcript 1 (GHET1) was involved in the progression of tumors. However, the role of GHET1 in esophageal squamous cell carcinoma (ESCC) remains unclear.
Methods
The expression of IncRNA GHET1 was examined in 55 paired ESCC tissues and adjacent nontumor tissues. Molecular and cellular techniques were used to explore the role of GHET1 on ESCC cells.
Results
Our data showed that GHET1 expression was significantly increased in ESCC tissues and cell lines. High GHET1 expression in ESCC tissues was significantly associated with poor differentiation, advanced tumor nodes metastasis stage, and lymph node metastasis. GHET1 showed high sensitivity and specificity for diagnosing ESCC. Our data from in vitro assays showed that GHET1 inhibition suppressed ESCC cells proliferation, migration, and invasion, and induced cells apoptosis. Furthermore, western blot showed that GHET1 inhibition significantly decreased the expression of vimentin and N-cadherin while it increased the expression of E-cadherin.
Conclusions
Our study indicates that GHET1 acts as an oncogene in ESCC and may represent a novel therapeutic target for the treatment of ESCC patients.
Introduction
Esophageal cancer (EC) is one of the leading aggressive malignancies worldwide, while esophageal squamous cell carcinoma (ESCC) accounts for the most prevalent histopathologic type (1, 2). Although the improvements in the treatment of ESCC have been achieved by radiochemotherapy and surgical resection in recent years, the 5-year survival rate is still less than 50% (3, 4). Therefore, fully understanding the molecular mechanism of ESCC progression is urgent so that new, potential early diagnostic markers and therapeutic targets for the treatment of ESCC can be developed.
Long noncoding RNAs (IncRNAs) represent a diverse type of long RNA molecules lacking protein-coding capacity, with a length greater than 200 nucleotides (5). Recent studies have shown that abnormally expressed IncRNAs in cancer regulate the proliferation, apoptosis, metastasis, and differentiation of cancer cells (6). For example, Sun et al (7) showed that IncRNA RNA NEAT1 acted as an oncogene in non-small cell lung cancer (NSCLC) by regulating the Wnt signaling pathway. Zhang et al (8) showed that IncRNA antisense noncoding RNA in the INK4 locus (ANRIL) indicated a poor prognosis of cervical cancer and promoted carcinogenesis via PI3K/Akt pathways. Cui et al (9) reported that upregulated IncRNA small nucleolar RNA host gene 1 (SNHG1) contributed to the progression of NSCLC through the inhibition of miR-101-3p and the activation of the Wnt/β-catenin signaling pathway.
A novel IncRNA gastric carcinoma highly expressed transcript 1 (GHET1) was found to be aberrant when expressed in gastric carcinoma, and the higher expression of GHET1 was associated with a bigger tumor size and worse overall survival (10). Recent studies showed that GHET1 was aberrantly expressed in various tumors. However, the expression and underlying mechanism of GHET1 on ESCC tumorigenesis are still unclear.
In the present study, we investigated GHET1 expression in ESCC tissues and examined the relationship between GHET1 expression and clinicopathological features of ESCC patients. We also investigated the effect of GHET1 silencing on the proliferation, migration, invasion, and epithelial-mesenchymal transition (EMT) of ESCC cells.
Materials and methods
Patients and tissue samples
A total of 55 patients with ESCC were recruited from The First Hospital of Shijiazhuang between March 2012 and December 2014. No patients had received local or systemic treatment before surgery. The diagnosis of ESCC and clinicopathological features of the patients were confirmed by 2 pathologists. All participants signed informed consent forms, and all experiments were approved by the Ethics Committee of The First Hospital of Shijiazhuang. All specimens were snap-frozen by liquid nitrogen and stored at −80°C until use.
Cell culture and siRNA transfection
ESCC cell lines (EC109, EC9706, KYSE30, and KYSE450) and a normal esophageal epithelial cell (Het-1A) were obtained from the Cell Bank of the Chinese Academy of Sciences (Shanghai, China). All cells were cultured in RPMI 1640 (Gibco) supplemented with 10% FBS (Gibco), 100 U/mL penicillin, and 100 μg/mL streptomycin at 37°C in a humid atmosphere of 5% CO2.
GHET1 siRNA (si-GHET1-1, 5′-CGGCAGGCATTAGAGATGAACAGCA-3′ si-GHET1-2, 5′-GUGAUCCAGACUCUGACCUU-3′) and negative control siRNA (si-NC, 5′-UUCUCCGAACGUGUCACGUTT-3) were purchased from RiboBio (Guangzhou). ESCC cells were transfected with the siRNAs by using Lipofectamine 2000 (Invitrogen), according to the manufacturer's instructions. After 48 hours of transfection, GHET1 expression was determined by quantitative real-time polymerase chain reaction (qRT-PCR).
qRT-PCR
Total RNA was extracted from ESCC tissues and cell lines by using TRIzol reagent (Invitrogen), according to the manufacturer's instructions. Next, mRNA was reverse transcribed to cDNA by using PrimeScript RT Reagent kit with cDNA Eraser (TaKaRa), according to the manufacturer's protocol. GHET1 expression was determined by performing qRT-PCR with SYBR Premix Ex Taq (TaKaRa) and 7500 real-time PCR system (Applied Biosystems). Sequences of primers used for performing qRT-PCR are as follows: GHET1, 5′-CCCCACAAATGAAGACACT-3′ (forward) and 5′-AGTGGCTGGCATATAACCAACA-3′ (reverse); GAPDH, 5′-TTCCCAACACCCTATAAGAT-3′ (forward) and 5′-GTTGTCATGGATGACCTTGGC-3′ (reverse). GAPDH was used as an internal control for normalizing GHET1 mRNA expression. Relative GHET1 expression is expressed as fold induction by using the 2−ΔΔCt method.
Cell proliferation assay
A cell proliferation assay was performed using Cell Counting Kit-8 assay kit reagent (CCK-8; Beyotime), according to the manufacturer's instructions. Cells in the si-NC and si-GHET1 groups were cultured in 96-well plates in triplicate at a density of 1 × 10 4 cells per well. After culturing for 24, 48, 72, and 96 hours, 10 μL CCK-8 reagent was added to each well and the cells were incubated further for 4 hours at 37°C. Absorbance was measured at 450 nm by using a microplate reader.
Flow cytometry assay
The transfected cells were harvested and were washed twice with cold PBS and re-suspended in a binding buffer. Then the cells were stained with FITC Annexin V apoptosis detection kit (BD Bioscience) for 30 minutes at room temperature in the dark. The apoptosis detection was determined using flow cytometry (BD Bioscience) within 1 hour.
Cell migration and invasion assays
A cell migration assay was carried out using a wound-healing assay; 1 × 106 cells were seeded into 6-well plates. A wound was made by scraping the cell monolayer with a 200 μL pipette tip. Cell motility was evaluated by observing at intervals of 0 and 24 hours.
A cell invasion assay was carried out using Transwell invasion assay, cells (1 × 105) in a serum-free medium were placed into the upper chamber of a 24-well Transwell Chamber (8-μm pore size; Corning). RPMI-1640 medium containing 10% FBS as a chemo-attractant was added to the lower chamber. After incubation for 48 hours, nonmigrating cells were removed by a cotton swab, and the migrating cells on the lower side of the insert filter were fixed using methanol, and were stained with 0.1% crystal violet for 20 minutes. The number of migrating cells was counted in 5 random visual fields and imaged using the light microscope (Olympus).
Western blot analysis
The transfected cells were lysed with an ice-cold lysis buffer; proteins were extracted, and the concentration was detected. Then an equal amount of proteins from each group was separated by SDS-polyacrylamide gel electrophoresis (SDS-PAGE) and transferred to a polyvinylidene fluoride (PVDF) membrane. After incubating with 5% nonfat dry milk in PBST for 1 hour to block nonspecific binding, the membrane was incubated with appropriate primary antibodies overnight at 4°C, followed by HRP-conjugated secondary antibodies for 1 hour. The antibody-reactive bands were visualized using ECL reagent (Millipore). The antibodies against vimentin, N-Cadherin, and E-Cadherin and GAPDH were all obtained from Abcam.
Statistical analysis
Statistical analysis was performed using SPSS 18.0 software (IBM). Continuous variables were presented as mean ± standard deviation. The differences between independent groups were analyzed using an independent t-test; p<0.05 was considered statistically significant.
Results
GHET1 expression was increased in ESCC
GHET1 expression in ESCC tissues and adjacent nontumor tissues was analyzed by qRT-PCR. Our results showed that GHET1 expression was significantly upregulated in ESCC tissues compared with adjacent nontumor tissues (Fig. 1A; p<0.05). To explore the association between the clinicopathological features and GHET1 expression, ESCC patients were divided into 2 groups according to the median expression of GHET1 in ESCC tissues: low GHET1 expression group (<median value) and high GHET1 expression group (≥median value). Noticeably, high GHET1 expression in ESCC was significantly associated with poor differentiation, advanced tumor nodes metastasis (TNM) stage and lymph node metastasis (Tab. I; p<0.05), but was not associated with other clinicopathological features (Tab. I; p>0.05). GHET1 expression was clearly increased (i) in patients with poor tumor differentiation compared with that in patients with moderate + high tumor differentiation (Fig. 1B; p<0.05); (ii) in patients with TNM stage III + IV tumors compared with that in patients with TNM grade I + II tumors (Fig. 1C; p<0.05); and (iii) in patients with lymph node metastasis compared with that in patients without lymph node metastasis (Fig. 1D; p<0.05). Receiver operating characteristic (ROC) curve analysis was performed to determine the sensitivity and specificity of GHET1 for diagnosing ESCC based on its expression. At an optimal expression cutoff value (3.19), the sensitivity and specificity of GHET1 for diagnosing ESCC were 69.7% and 91.3%, respectively, with an area under the curve (AUC) of 0.858 (95% confidence interval, 0.824-0.948; Fig. 1E).
Association between IncRNA GHET1 expression and clinicopathologic features in ESCC patients
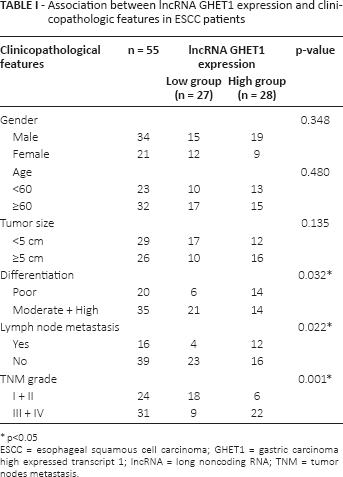
p<0.05
ESCC = esophageal squamous cell carcinoma; GHET1 = gastric carcinoma high expressed transcript 1; IncRNA = long noncoding RNA; TNM = tumor nodes metastasis.
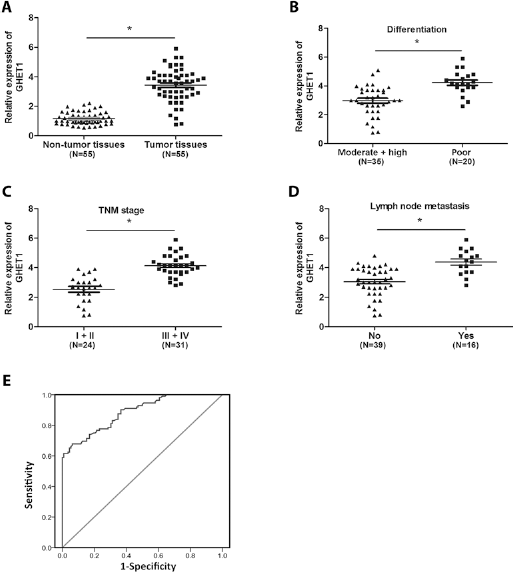
GHET1 was overexpressed in ESCC. (
GHET1 silencing suppressed ESCC cells proliferation
GHET1 expression was determined in ESCC cell lines (EC109, EC9706, KYSE30, and KYSE450) and normal esophageal epithelial cells (Het-1A). Our results showed that GHET1 expression was significantly higher in ESCC cells than that in Het-1A cells (Fig. 2A; p<0.05). To explored the functions of GHET1 in ESCC, GHET1 specific siRNAs were transfected into KYSE30 cells and the knockdown efficiency was confirmed by qRT-PCR (Fig. 2B; p<0.05). A CCK-8 assay showed that cell proliferation was significantly decreased in si-GHET1 transfected cells compared with that in the si-NC group (Fig. 2C; p<0.05). To investigate whether cell apoptosis contributed to the proliferation of ESCC cells in vitro, flow cytometry assay was used to explore the effects of GHET1 on cell apoptosis. We revealed that the apoptosis of KYSE30 cells was greatly enhanced after GHET1 inhibition (Fig. 2D; p<0.05). These data suggested that GHET1 could regulate ESCC cell proliferation by mediated cell apoptosis.
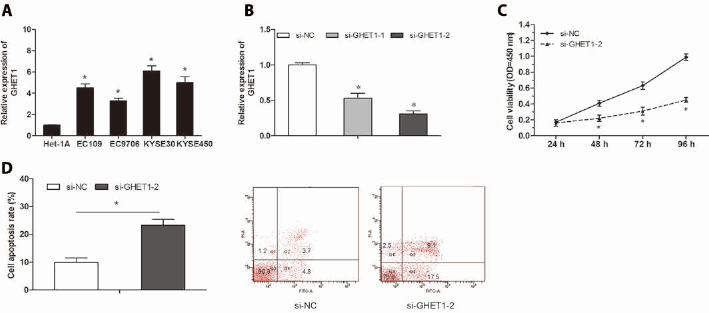
GHET1 inhibition suppressed ESCC cell proliferation in vitro. (
GHET1 silencing inhibited ESCC cells migration, invasion and EMT
To further explore the effect of GHET1 on ESCC cell migration and invasion, a wound healing assay and a transwell invasion assay was performed. As shown in Figure 3A, ESCC cell migration ability was significantly suppressed in KYSE30 cells transfected with si-GHET1 (p<0.05). Furthermore, the transwell invasion assay showed that the invading cells were significantly lower among si-GHET1 transfected KYSE30 cells than the si-NC transfected KYSE30 cells (Fig. 3B; p<0.05). These findings suggested that GHET1 depletion might decrease ESCC cell mobility.
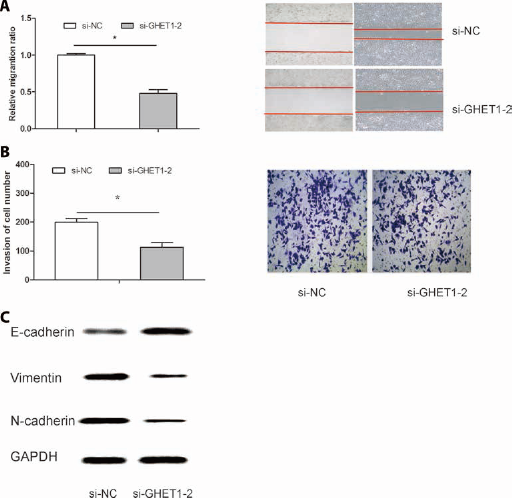
GHET1 inhibition suppressed ESCC cell migration and invasion in vitro. (
Because epithelial-mesenchymal transition (EMT) is the remarkable presentation for cell invasion, whether silencing GHET1 expression inhibited mesenchymal features is still unclear. As shown in Figure 3C, vimentin and N-cadherin were downregulated in KYSE30 cells transfected with si-GHET1, while E-cadherin was overexpressed (p<0.05). Therefore, these data suggest that GHET1 might promote ESCC cell progression by the induction of the EMT process.
Discussion
ESCC is the leading prevalent histopathologic subclass of esophageal cancer in central regions of China (11). Despite of the recent rapid promotion in diagnosis and therapy, the prognosis of ESCC remains poor (3). The improvement of the ESCC survival rate requires a clear understanding of pivotal molecular mechanisms from the initiation and progression of ESCC.
Recently, the roles for IncRNA as tumor suppressive and oncogenic functions have been found in prevalent cancer types (12). GHET1 (AK123072) is a recently identified long noncoding RNA (10). Recent studies have shown that GHET1 plays an important role in tumor progression. For example, Li et al (13) showed GHET1 is a potential predictive marker of poor prognosis in bladder cancer patients and promotes cell progression. Zhou et al (14) showed that knockdown of GHET1 inhibited cell proliferation and the invasion of colorectal cancer. Huang et al (15) showed that the knockdown of GHET1 inhibited the cell activation of gastric cancer. However, the role of GHET1 on ESCC remains unclear.
In the present study, we investigated the role of GHET1 in the progression of ESCC. We showed that GHET1 expression was significantly upregulated in ESCC tissues compared to adjacent nontumor tissues. High GHET1 expression was significantly correlated with poor differentiation, advanced TNM stage, and lymph node metastasis, but was not correlated with other clinical features, indicating that GHET1 plays an important role in ESCC progression. In addition, ROC analysis revealed that GHET1 could act as an independent diagnostic marker of ESCC.
In addition, we investigated the effect of GHET1 on ESCC tumorigenesis and progression to determine the potentially therapeutic target for ESCC treatment. We found that GHET1 inhibition suppressed the proliferation, migration, and invasion, and induced the apoptosis of ESCC cells. Proliferation and apoptosis dysfunction play important roles in ESCC development, and cell migration and invasion play central roles in ESCC metastasis. Thus, our data indicated that GHET1 silencing inhibited ESCC cell proliferation and invasion, implying that GHET1 could function as an oncogene in ESCC progression.
EMT, a biological process in which cancer cells lose their epithelial polarity and undergo transition into a mesenchymal phenotype, plays a key role in cancer cell malignant transformation (16, 17). Recent studies revealed that IncRNAs could regulate tumor progression by affecting the EMT process. For example, Xu et al (18) showed that the knockdown of IncRNA HOTAIR suppressed tumor invasion and reversed EMT in gastric cancer. Li et al (19) showed that IncRNA BCAR4 promoted proliferation, invasion, and metastasis of NSCLC cells by affecting EMT. Zhang et al (20) suggested that IncRNA MEG3 inhibited cell EMT by sponging miR-421 targeting E-cadherin in breast cancer. However, the role of IncRNA GHET1 in EMT progress remains unclear. In the present study, western blotting showed that vimentin and N-cadherin (mesenchymal markers) were down regulated in KYSE30 cells transfected with si-GHET1, while E-cadherin (the epithelial marker) was increased, suggesting that the effects of GHET1 on ESCC progression might be partly associated with the EMT process.
Conclusion
Our study has shown that GHET1 is a potential diagnostic marker and therapeutic target for ESCC treatment. However, further studies are required to determine the mechanism underlying GHET1-induced regulation of ESCC tumorigenesis.
Footnotes
Financial support: No grants or funding have been received for this study.
Conflict of interest: None of the authors has financial interests related to this study to disclose.
