Abstract
Objective:
We aimed to evaluate the genetic variation of poly(ADP-ribose) polymerase-1 (PARP-1) in the development of gliomas among Chinese individuals.
Materials and methods:
Patients with a confirmed diagnosis of glioma and healthy individuals with no clinical symptoms of glioma were enrolled at Liaocheng People’s Hospital, China. Genetic polymorphisms were studied in plasma samples by polymerase chain reaction-restriction fragment length polymorphism assay. Cytokine levels were measured routinely in serum samples by sandwich ELISA technique.
Results:
A total of 120 Chinese patients with gliomas and 120 healthy Chinese individuals were included. We found that patients with the GG genotype (odds ratio [OR] 2.53, 95% confidence interval [CI] 1.46-4.38, p<0.001) and carriers of the G allele (OR 11.5, 95% CI 6.31-21.3, p<0.0001) were at high risk of developing glioma. A del/ins polymorphism of the NF-κB1 gene (OR 4.27, 95% CI 2.43-7.50, p<0.001) was also found to be associated with glioma. In addition, significantly increased cytokine levels were observed in patients with glioma (p<0.05).
Conclusions:
Our findings showed that PARP-1 polymorphisms are involved in the development of glioma in Chinese individuals. Also serum cytokine levels can be considered among the potential risk factors for developing glioma.
Introduction
Primary brain tumors are a heterogeneous group of malignancies. They have a relatively low prevalence compared with other neoplasms (1). Glioma is a subtype that includes all tumors arising from glial cells. It is the most common and life-threatening malignant primary brain tumor in humans, especially in adults, accounting for approximately 30% of all brain and central nervous system tumors and 80% of all malignant brain tumors (2, 3). Many environmental and lifestyle factors including specific occupations, environmental carcinogens and diet have been found to be associated with an elevated glioma risk (4, 5). In recent years, the oxidative stress response has been of particular interest in gliomas, given the high oxygen metabolism rate in the brain (6). An excess of oxidative stress, which is initiated by reactive oxygen species (ROS) or reactive nitrogen species (RNS), appears to increase the predisposition to glioma, as an increased concentration of ROS/RNS can cause DNA damage, repress the activity of cellular enzymes, influence apoptosis and proliferation, and promote tumorigenesis (7, 8).
It is well known that poly(ADP-ribose) polymerase-1 (PARP-1)-mediated signaling can trigger an outburst of proinflammatory regulators, and the cell death cascade is also known to the scientific community (9). PARP-1, also named adenosine diphosphate ribosyltransferase, catalyzes the process of PARylation by attaching the polymers of ADP-ribose to target protein motifs via ester linkage, and triggers an array of vital cell functions including chromatin structure, DNA repair, transcriptional regulation, apoptosis, necrosis, cell separation and differentiation (10). In humans, 17 members of PARP are expressed, of which PARP-1 is associated with the nucleus and regulates at least 85% of the cellular PARP activities (11). DNA strand nicks/breaks/errors activate PARP-1 and trigger the DNA repair process (12). PARP-1 maintains the genomic integrity by PARylation of histones and other key enzymes, the DNA repair mechanism, interprotein interactions and gene expression to ensure optimum cellular homeostasis (13-16). However, through cellular and/or genotoxic stress, prolonged activation of PARP-1 leads to a decrease in β-nicotinamide adenine dinucleotide (NAD) and adenosine triphosphate (ATP), and causes release of apoptosis-inducing factor (AIF), a mitochondrial proapoptotic protein (17). Such stress-induced dysregulation in cellular homeostasis mediated by PARP-1 triggers the downstream necrosis cascade of cell death (18).
In addition, inflammation is the immune system’s response to external or internal aggressors such as injury or infection in certain tissues. The body’s response to cancer shows many parallels with inflammation and repair; the inflammatory cells and cytokines present in tumors are more likely to contribute to tumor growth, progression and immunosuppression than to build an effective antitumor defence (19). Cytokines have been involved in various stages of glioma progression, participating in tumor onset, growth enhancement, angiogenesis and aggressiveness (20). Among the cytokines, interleukin (IL)-6, IL-1β and tumor necrosis factor alpha (TNF-α) play an important role in tumor growth and glioma angiogenesis (20). Several lines of evidence have shown that increased expression of IL-6 in glioma stimulates the growth of glial tumor cells by an autocrine or paracrine mechanism. IL-6 is mitogenic for astrocytes and together with its soluble receptor (sIL-6Rα) induces massive reactive gliosis in IL-6/IL-6Rα double transgenic mice (21-24). Interleukin-1β is one of the cytokines most commonly present in glioma, and evidence has shown that IL-1β modulates glioma progression by interacting directly with the tumor cells (25). TNF is an endogenous tumor promoter which stimulates cancer cell growth, proliferation, invasion and metastasis as well as tumor angiogenesis. It was found that TNF-α induced proliferation of C6 glioma cells via beta-adrenergic receptors in rodent models (26, 27). Another study strongly suggests that TNF-α induces IL-6 synthesis, which further stimulates the growth of glial tumor cells (28). A report suggested the involvement of nuclear factor kappa-light-chain-enhancer of activated B cells (NF-κB) in the activation of tumor growth and discussed its potential role in the etiology of cancer (29).
Galia et al (30) suggested that PARP-1 gene polymorphisms were involved in the development of glioblastoma multiforme in Caucasian patients. The main objective of our study was to ascertain whether such polymorphisms also play a role in Chinese patients with glioma, given the different involvement of gene polymorphisms based on ethnicity according to the literature (31). This pilot study was designed to investigate whether there is any ethnic difference in gene involvement between Chinese and Caucasian patients with glioma. In addition, we assessed whether cytokines and NF-κB were involved in glioma development. Our study may serve as a basis for conducting large multicenter and multicountry clinical studies to assess the association of PARP-1 with glioma.
Materials and methods
Patients of both genders aged between 18 and 65 years with a confirmed diagnosis of glioma and healthy individuals with no clinical signs or symptoms of glioma (control group) were enrolled at Liaocheng People’s Hospital, China. The study was approved by the institutional ethics committee and written consent was obtained from each study participant. All participants were informed about the study procedures and the potential benefits to society. All participants underwent laboratory tests to confirm their eligibility as healthy individuals and patients with glioma.
Plasma samples were collected from the study population and DNA from leukocytes was extracted by the high salt DNA extraction method. All isolated DNA samples were kept at −80°C for further assessment. Using efficient molecular biology techniques, polymorphisms of the PARP-1, NF-κB1 and NF-κBIA genes were studied in DNA samples by polymerase chain reaction-restriction fragment length polymorphism (PCR-RFLP) analysis. For the PARP-1 C410T and G1672A promoter regions, 297 and 187 base-pair PCR fragments were amplified in a 25-μL reaction volume at pH 8.3 consisting of 50 mM potassium chloride (KCl), 100 ng genomic DNA, 200 pM deoxynucleotide (dNTP), 200 pM of 10 mM Tris-hydrogen chloride (HCl), 1 U Taq polymerase (Sigma) and 2 mM MgCl2 (Fermentas). PCR was performed using an optimized sequence of cycles: a denaturation phase at 95°C (2 min) followed by 30 cycles at 95°C (30 s), 60°C (30 s), 72°C (1 min), and a final incubation phase at 72°C (5 min). The PCR products were treated with 5 U of HpyF3I (DdeI) (Fermentas) and Bsh1236I (Fermentas) at 37°C overnight for digestion. The resulting fragments were run on ethidium bromide-stained 3% agarose gel for 45 minutes at 90 V and quantified through direct detection under ultraviolet (UV) light. Likewise, the NF-κB1 and NF-κBIA genes were amplified by 285 and 424 base-pair PCR fragments using the same reagents. The PCR running sequence included a denaturing step at 95°C (1 min) followed by 35 cycles at 95°C (30 s), 61°C (30 s), 72°C (1 min) and a final incubation at 72°C (5 min). The PCR products were treated with 5 U PflMI (Van91I) and HaeIII (BsuRI) (Fermentas) at 37°C overnight for digestion and the digested products were run on ethidium bromide-stained 3% agarose gel for 45 minutes at 90 V and quantified directly under UV light.
Cytokine levels (IL-6, IL-1β and TNF-α) were measured routinely in serum samples by a specific sandwich ELISA technique using commercial kits (Assaypro Human IL-6, IL-1β and TNF-α) coated with antibodies specific to human leptin and adiponectin antibodies according to the manufacturer’s protocols (St. Charles).
Statistical analysis
The study was designed as a preliminary pilot study to assess the association of PARP-1 with glioma in a Chinese population. Hence, no formal sample size calculation was performed. We planned to enroll at least 100 individuals in each group (glioma patients and healthy controls). We designed our study to serve as a basis for large genetic clinical studies to assess PARP-1 in patients with glioma. The data of each study participant were coded and analyzed using the GraphPad Prism statistical software (version 6.0). Quantitative variables were presented as means (± standard deviation) and analyzed by parametric and nonparametric statistical tests depending on the number of groups for comparison and the distribution of data; 2-sided statistical tests were used. Normality testing (Kolmogorov-Smirnov test or Shapiro-Wilks test) was used to check the distribution of quantitative data. Categorical variables were presented as absolute numbers and/or percentages of subjects in each category, and analyzed by the chi-square or Fisher exact test depending on the size of the data; again, 2-sided statistical tests were used. Odds ratios (ORs) were calculated based on univariate analysis using the chi-square or Fisher exact test.
Results
A total of 120 Chinese patients with glioma and 120 healthy individuals were enrolled and completed the study. The data of all individuals of both groups were subjected to statistical analysis. The mean age in the glioma and healthy groups was 44.6 (±8.3) years and 48.7 (±3.6) years, respectively. Gender distribution was similar in the patient and control groups. Body mass index was comparable among individuals of both groups (Tab. I).
Demographic characteristics of healthy individuals and glioma patients
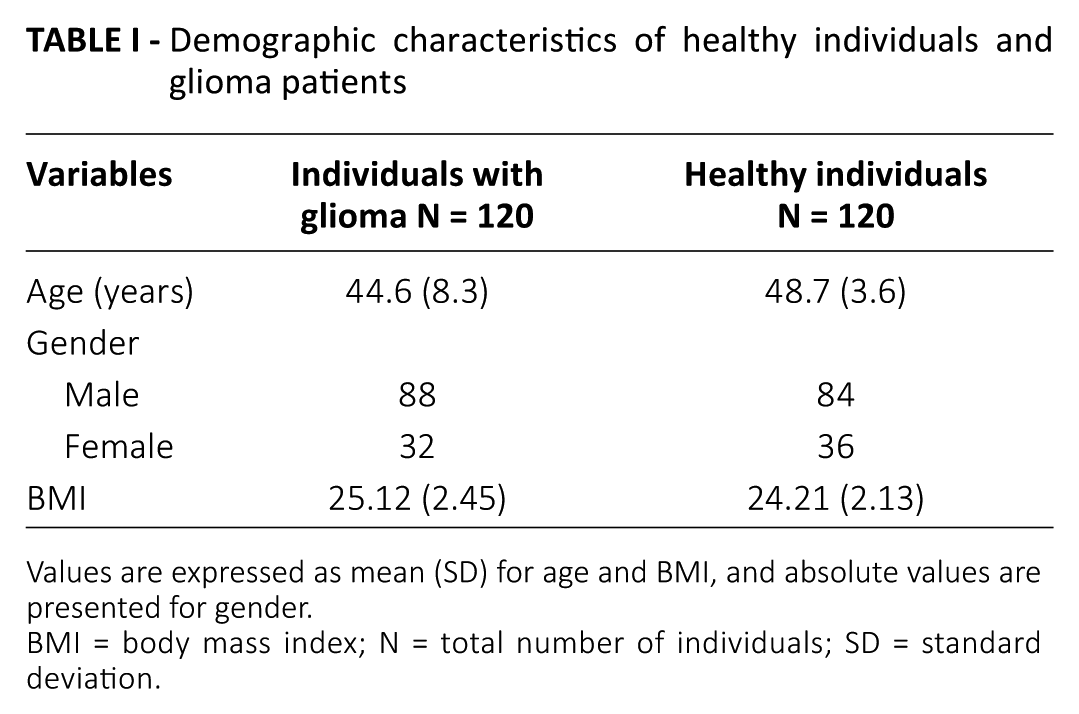
Values are expressed as mean (SD) for age and BMI, and absolute values are presented for gender.
BMI = body mass index; N = total number of individuals; SD = standard deviation.
For the G1672A genotype, we found that individuals with a GG genotype in the PARP-1 gene were at very high risk of developing glioma (OR 2.53, 95% confidence interval [CI] 1.46-4.38, p<0.001). We observed that individuals with a G allele in PARP-1 were also at very high risk of developing glioma (OR 11.5, 95% CI 6.31-21.3, p<0.0001). There was no involvement of other genotypes such as GA or AA among patients with glioma. None of the other genotypes of NF-κB1, NF-κBIA and PARP-1 were found to be involved (Tab. II). No evidence of PARP-1 polymorphisms was observed in healthy subjects.
Genotypes involved in gene polymorphism in patients with glioma
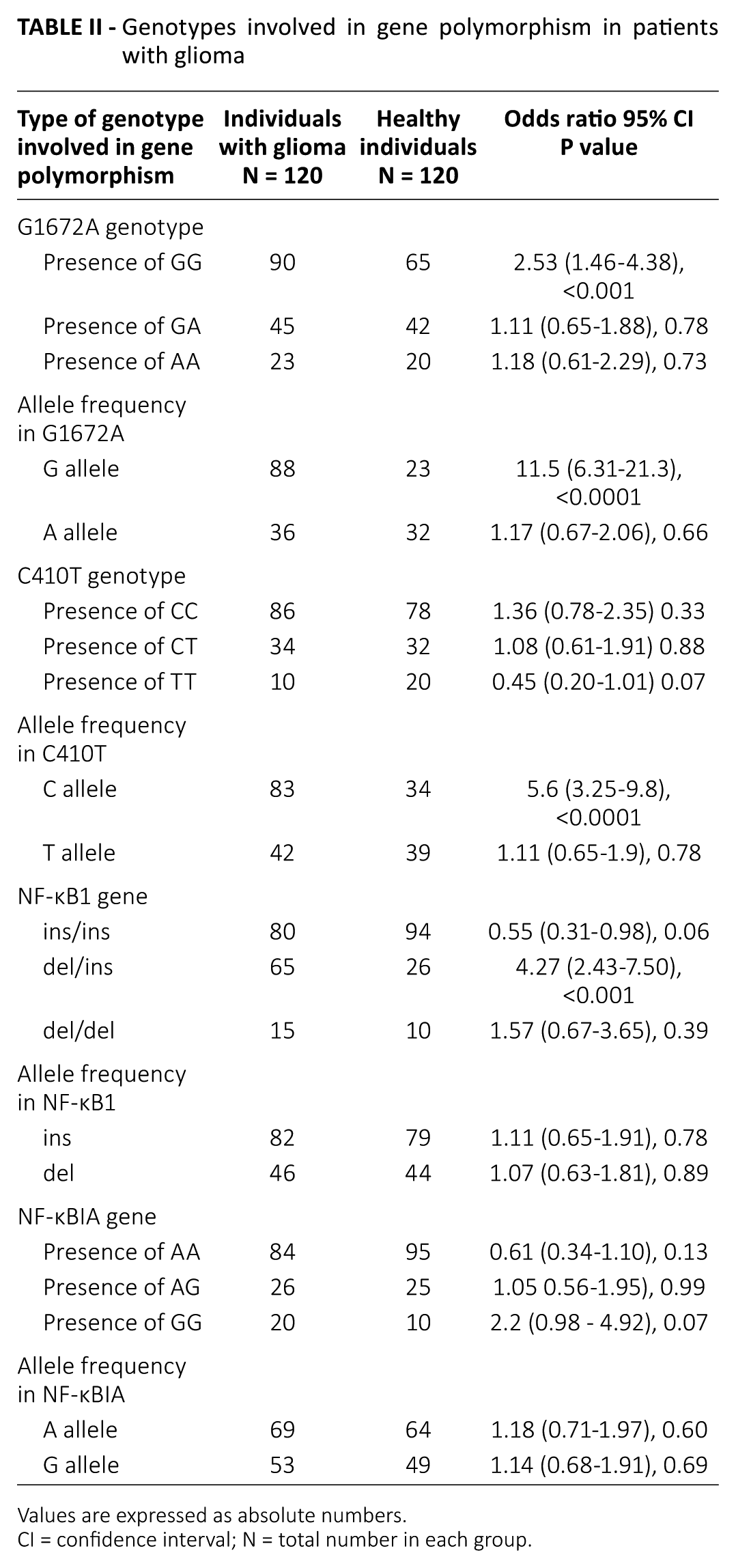
Values are expressed as absolute numbers.
CI = confidence interval; N = total number in each group.
For the C410T genotype, we observed that individuals with a C allele in the PARP-1 gene were at high risk of developing glioma (OR 5.6, 95% CI 3.25-9.8, p<0.0001). Among patients with glioma, a DNA sequence variation, namely del/ins of the NF-κB1 gene, was observed (OR 4.27, 95% CI 2.43-7.50, p<0.001). This suggests a role of del/ins of NF-κB1 in the development of glioma. No statistically significant association of other NF-κB1 genotypes with glioma was observed (Tab. II).
We also studied the role of cytokines in glioma in relation to different genes. The results showed a significantly higher level of cytokines in glioma patients compared with healthy individuals (Tab. III). Our study suggested a significant association between cytokines and the development of glioma irrespective of the DNA sequencing results.
Plasma level of cytokines in individuals with glioma and healthy individuals by different genotypes
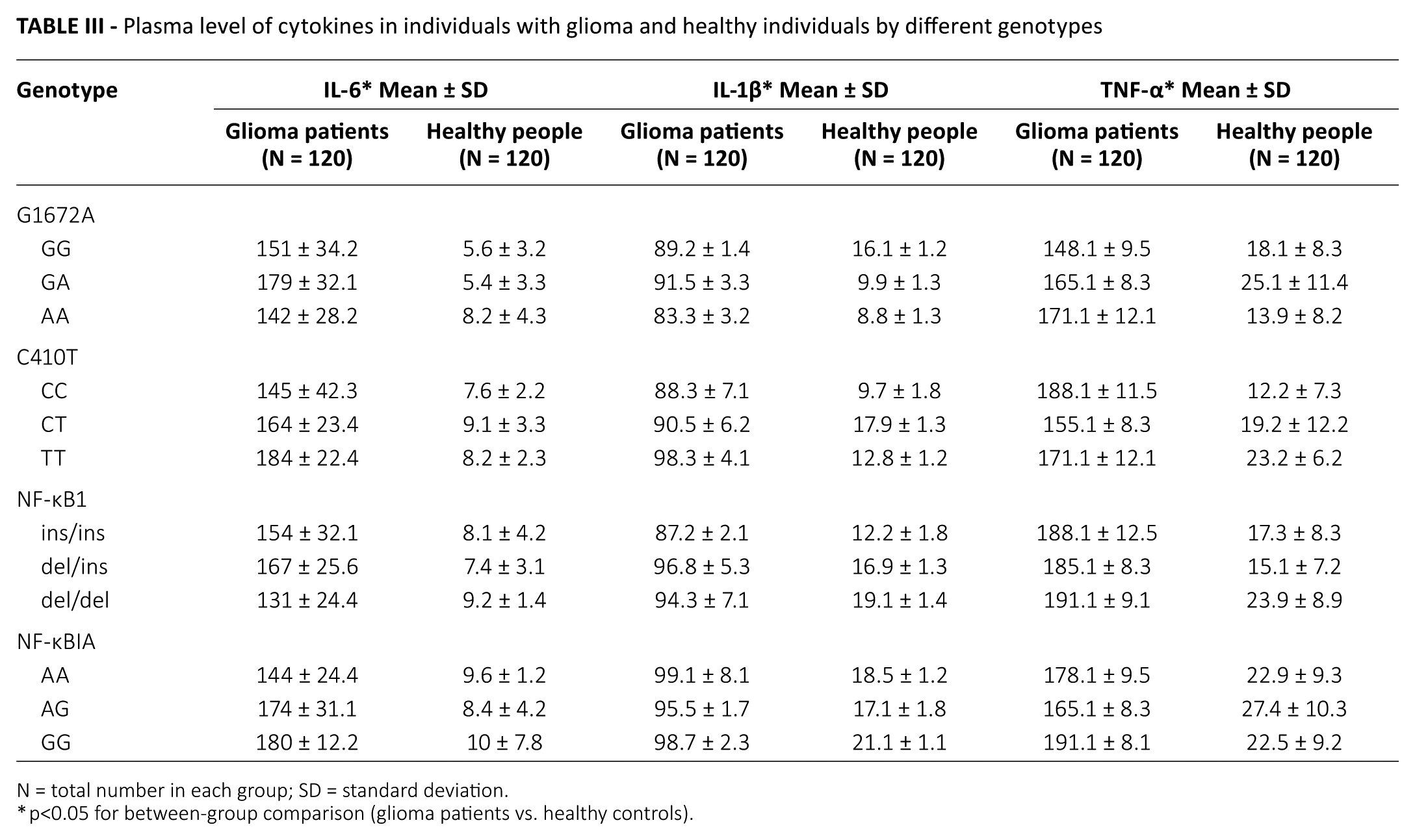
N = total number in each group; SD = standard deviation.
p<0.05 for between-group comparison (glioma patients vs. healthy controls).
Discussion
This was the first study to determine the association of PARP-1 polymorphisms with the development of glioma in Chinese patients. We found that individuals with the GG genotype were at very high risk of developing glioma. Our finding in Chinese patients was consistent with the findings of Galia et al (30) in Caucasians and indicated no ethnic differences in this respect. In this Chinese study, we also observed that individuals carrying a C allele were at high risk of developing glioma. Additionally, we observed the involvement of a del/ins polymorphism in NF-κB1 in glioma development among Chinese individuals. Significantly higher cytokine levels were observed in glioma patients compared to healthy controls, which indicates a positive correlation between cytokine levels and development of glioma.
Several lines of evidence point to a role of cytokines; it was reported that cytokine mediators and related genes were positively involved in glioma development, and high levels of any cytokine are considered a risk factor for glioma (24). Our study supports these previous findings. The etiology of glioma was not only related to polymorphisms of single genes: in many instances researchers have found that the interaction of polymorphisms of multiple genes may result in the development of glioma (32, 33).
Individuals with a GG genotype in DNA sequencing and carriers of a G allele were more susceptible to and at very high risk of developing glioma. This indicates that a GA genotype might have a defensive influence on the disease. The possible involvement of the GG genotype as a risk factor for glioma could be explained by molecular heterosis, which is observed in approximately 50% of cases of gene associations (34).
We found that PARP-1 activates NF-κB1, which leads to the release of chemicals such as IL-1β, IL-1γ, IL-6 and TNF-α (35). Significantly higher levels of cytokines have been reported in glioma and in individuals with PARP-1 polymorphisms; our finding was in line with previous studies since the levels of all types of cytokines were significantly higher in individuals with glioma.
This was the first study to suggest the association of 2 single nucleotide polymorphisms (SNPs) of PAPR-1 (C410T and G1672A) with glioma in Chinese patients. The involvement of these SNPs in other cancer subtypes has not yet been established. Our findings encourage assessing the association of these 2 SNPs in other cancer subtypes. Based on the results of this pilot study, we also recommend conducting large multicenter, multicountry, genetic clinical studies to generalize our findings.
Conclusions
Our results suggest that polymorphisms of the PARP-1 gene are responsible for the development of glioma in Chinese individuals. We also observed the involvement of a NF-κB1 polymorphism in the increased risk of glioma. High cytokine levels were observed in patients with glioma, and an increased serum cytokine level can therefore be considered a potential risk factor for glioma development. Our study results may serve as a basis for conducting large multicenter, multicountry, genetic clinical studies to assess the involvement of PARP-1 and NF-κB1 in glioma.
Footnotes
Acknowledgements
The authors thank all patients for their participation in the study. We would also like to thank Rakesh Ojha (senior consultant medical writer) for writing and editorial support in the preparation of this manuscript.
Disclosures
Financial support: None.
Conflict of interest: The authors declare they have no conflict of interest related to this work.
