Abstract
Influenza A (H5N1) virus, has spread to several countries in the world and has a high mortality rate. Meanwhile, the virus has evolved into several clades. The human influenza A (H5N1) virus circulating in Indonesia is a member of clade 2.1, which is different in antigenicity from other clades of influenza A (H5N1). An analysis of the antigenic variation in the H5 hemagglutinin gene (HA) of the influenza A (H5N1) virus strains circulating in Indonesia has been undertaken. Several position of amino acid mutations, including mutations at positions 35, 53, 141, 145, 163, 174, 183, 184, 189, and 231, have been identified. The mutation Val-174-Iso appears to play an important role in immunogenicity and cross-reactivity with rabbit antisera. This study shows that the evolution of the H5HA antigenic variation of the influenza A (H5N1) virus circulating in Indonesia from 2005 to 2011 may affect the immunogenicity of the virus.
Introduction
The cases of avian influenza (AI) in humans from 2003 to August 2012, there were 608 cases of AI with 359 dead victims in 15 countries in the world. 1 There were 192 human AI cases with 160 deaths in 15 provinces throughout Indonesia from July 2005 to December 2012.1,2 AI in humans occurs concurrently with AI in poultry in the local vicinity. As reported by the Food and Agriculture Organization of the United Nations (FAO), the influenza A (H5N1) virus continues to circulate in countries where there is AI in humans, including in Indonesia. 3
Because the characteristics of the influenza virus constantly evolve and change unpredictably, it is very important to control the virus. The influenza A virus is an RNA virus with eight gene segments. The determination of subtypes is based on two surface proteins, HA and NA. To date, there are 16 subtypes of HA and 9 subtypes of NA.4–6
As the HA gene mutates, antigenic drift occurs. As a result, the influenza A (H5N1) virus is divided into 9 clades, based on the phylogenetics of the HA gene. In Indonesia, the H5N1 virus is endemic in poultry and in humans, and it is a member of clade 2.1. 7 Clades and subclades of the AI virus have different antigenic properties, presenting a problem for the development of new vaccines.8,9
H5N1 that was isolated between 2005 and 2011 evolved from clade 2.1.2 to clade 2.1.3. 10 In all types of clades, the antigenicity of the circulating H5N1 consists of four groups based on the phylogenetic tree classification: group A, viruses deriving from Indonesia; group B, viruses circulating in Vietnam and Thailand; group C, viruses deriving from Middle Asia and North Africa; and group D, viruses deriving from Southern China. 11
The spread of H5N1, which occurs continuously, is of particular concern for public health as the diagnosis is often delayed, leading to delays in administering the antiviral treatment. 12 From the studies mentioned above, it is very clear that all of the viruses circulating in Indonesia belong to one distinct and unique cluster which is different from the types of viruses in other countries. Therefore, to develop the next vaccine in Indonesian, we need to identify the profile of the H5N1 antigenic viruses that were in circulation and infecting humans between 2005 and 2011.
Comprehensive research is required to determine the characteristics of the H5N1 antigenic virus that infects human beings in Indonesia, as well as to determine the changes in the antigenicity of the H5N1 virus. In this study, the antibodies to various viral strains will be elaborated upon to determine the variation in the antigenicity of H5N1 from different years. Thus, it can be observed that the antigenic variation will affect the immunogenicity and will provide significant information in deciding the appropriate vaccine for use in Indonesia.
Materials and Methods
Viruses
Seven H5N1 strains isolated between 2005 and 2011 (A/Indonesia/FKUI-NIHRD01/2005, A/Indonesia/ FKUI-NIHRD08/2006, A/Indonesia/FKUI-NIHRD03 /2007, A/Indonesia/NIHRD8987/2008, A/Indonesia/ NIHRD9160/2009, A/Indonesia/NIHRD10612/2010, and A/Indonesia/NIHRD11167/2011) were studied. The viruses were grown in the allantoic cavity of 10-day-old embryonated chicken eggs for 30 hours in an incubator at 37°C in the Biosafety Level 3 (BSL3) facility at the National Institute of Health Research and Development (NIHRD) of the Ministry of Health in the Republic of Indonesia. After harvesting, the viruses were aliquoted and stored at −80°C until use. The preparation of the seven viruses was heat inactivated for the immunization of rabbits. 13
Preparation of Antibodies
Twenty-two five-month-old male New Zealand white rabbits were immunized intramuscularly three times at three-week intervals with 500 μL of inactivated H5N1 that had a titer of 32 HAU suspended in PBS with seven strains of the virus (each strain was injected to three rabbits and one rabbit injected with sterile PBS as negative control). To confirm inactivation, the absence of infectivity after the inactivation of the H5N1 virus within the embryonated chicken eggs must be shown. Blood was collected from each rabbit before injecting the antigen to ensure there was no anti-H5N1 antibody. The antibodies were tested by the hemagglutination inhibition (HI) test.14–16 To obtain the titer of the virus required for immunization, the experiment was performed with different titers, from titers 8, 16, 32, 64 to titer 128. It was determined that titer 32 was the lowest titer which produced good immunity.
Hemagglutination Inhibition Test
The seven strains of H5N1 were used to test all of the rabbit sera in the BSL3 containment facility. The sera were treated by using a receptor-destroying enzyme (Denka Saiken Co., Japan) and incubated overnight at 37°C. The sera were then incubated at approximately 56°C for 30 minutes and diluted 1:10 with 0.85% NaCl. Then, the sera were tested by the HI assay using 0.5% packed fresh chicken red blood cells (CRBC) provided by Bogor Agricultural Institute (IPB). 17
PCR Amplification and Sequencing
Viral RNA was extracted from 140 μL of allantoic fluid using the QiAMP Viral Mini Kit extraction system (Qiagen, Germany), according to manufacturer's instructions.
Reverse transcription PCR was performed using primers specific for the HA gene segment from 1–1707 nucleotides (primer sequences are available upon request). The amplification protocol was as follows: 55°C for 30 minutes, hot start at 94°C for 2 minutes, followed by 35 cycles of amplification including denaturation at 94°C for 30 seconds, annealing at 55°C for 30 seconds, and extension at 72°C for 2 minutes. The final step was 72°C for 10 minutes. The RT-PCR reaction was conducted using the SuperScript III Reverse Transcriptase PCR with Platinum Taq kit (Invitrogen, Carlsbad, CA, USA) in a total volume of 25 μL of reaction mix, consisting of 5.5 μL 2 X RT-PCR mix buffer, 0.5 μL of 10 μM forward and reverse primers, 1 μL of 200 U/μL Superscript III enzyme, and 5 μL of template RNA. Then, the PCR products were purified using ExoSAP-IT purification kits (Affymetrix, Inc., Santa Clara, CA) under the following conditions: 37°C for 15 minutes and 80°C for 15 minutes. The DNA templates were sequenced using a DNA ABI Prism 3130 xl sequencer (Applied Biosystems, Foster City, CA) and the Big Dye X Terminator V3.1 Cycle Sequencing kit (Applied Biosystems, Foster City, CA) in a total volume of 10 μL of reaction sequencing mix consisting of 3 μL of 5X dilution buffer, 1 μL of 1.6 pmol primers, and 0.5 μL of Big Dye terminator mix. The reactions were then put in the thermal cycler (Biorad, USA) using the following reaction conditions: 96°C for 1 minute, followed by 25 cycles of 96°C for 10 seconds, 50°C for 5 seconds, and 60°C for 4 minutes. The PCR sequencing product was then purified using the Big Dye X Terminator Purification Kit (Applied Biosystems, Foster City, CA), according to the manufacturer's instructions. The DNA was sequenced and edited using the Bio Edit 7.0.8.0 sequence analysis software (Ibis Biosciences, USA). 18 Phylogenetic analyses were carried out using the neighbor-joining (N-J) method with 1000 bootstrap replicates implemented in the MEGA 4 software. 19
Nucleotide Sequence Accession Numbers
The nucleotide sequences obtained during this re search have been deposited in the GenBank database (accession numbers ABW06367.1 and ABL07030.1) and in the GISAID database (accession numbers EPI423757, EPI423755, EPI340962, EPI420777, and EPI420778).
Results
Analysis of the Antigenicity of the Viruses
The seven viruses used in this research are from clade 2.1, which includes viruses that circulate in Indonesia. 20 To obtain a better understanding of the molecular antigenic variation of the H5N1 virus, it is necessary to analyze the genetic mutations occurring in the HA protein. According to the sequencing results from those seven viruses compared with the sequences of the viral protein vaccine candidate Indo/05 reported by WHO, there were only a few amino acid differences at specific positions (Table 1). The similarities in antigenicity from the viral strains that infect humans from different years are found at the antigenic sites A, B, C, D, and E.11,21,22 At amino acid position Pro141, Val174, Asp183 and Arg189, which may be associated with antigenic changes,11,22 there is a substitution from Pro to Ser at position 141 that only occurs in the A/Indonesia/7/05 strain. This is also true for the Iso174 position in A/Indonesia/ NIHRD8987/2008, A/Indonesia/NIHRD9160/2009, and A/Indonesia/NIHRD10612/2010. An amino acid substitution also occurs at Asn183 and Met189 in the viral strains isolated between 2007 and 2011, including A/Indonesia/FKUI-NIHRD03/2007, A/Indonesia/ NIHRD8987/2008, A/Indonesia/NIHRD9160/2009, A/Indonesia/NIHRD10612/2010, and A/Indonesia/ NIHRD11167/2011.
Genetic variation in the HA gene of the H5N1 virus.
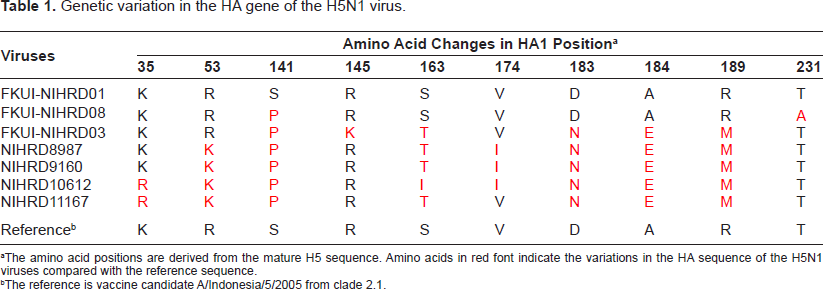
The amino acid positions are derived from the mature H5 sequence. Amino acids in red font indicate the variations in the HA sequence of the H5N1 viruses compared with the reference sequence.
The reference is vaccine candidate A/Indonesia/5/2005 from clade 2.1.
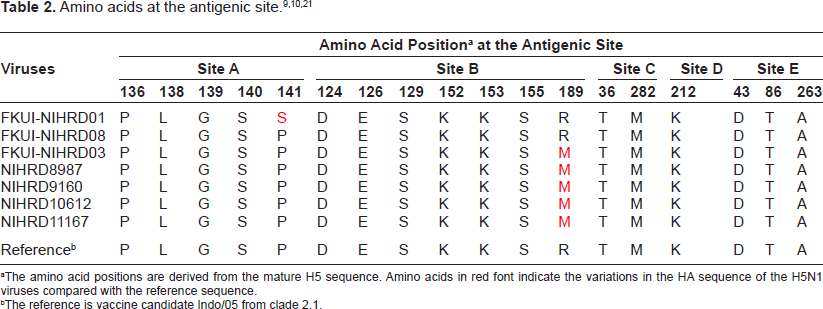
Based on the nature of the influenza virus, which continuously changes antigenic form, in these seven strains, there is antigenic drift. The antigen that changes only occurs at site B at position 189-Met in strains isolated from 2007 to 2011 (Table 2). Some amino acid positions that are outside of the antigenic site also have substitutions, including amino acids 35, 53, 141, 163, 174, 183, and 184. Phylogenetic analysis results as shown in Figure 1 put all of the seven viruses in clade 2.1 subclade 2.1.3.2.
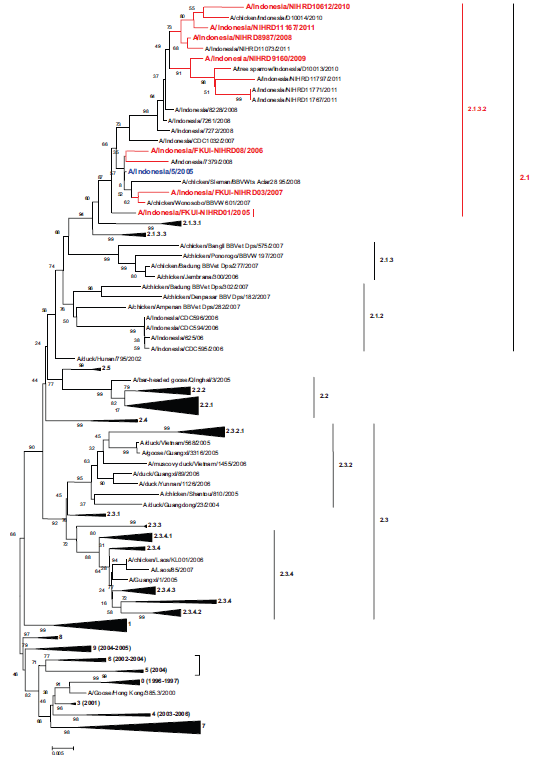

The phylogenetic tree of the clade of Indonesian strain viruses. The phylogeny was constructed using the Mega 4 version AA Neighbor-Joining bootstrap analysis (1000 replicates), and the Kimura 2 parameter model is shown.
The amino acid positions are derived from the mature H5 sequence. Amino acids in red font indicate the variations in the HA sequence of the H5N1 viruses compared with the reference sequence.
The reference is vaccine candidate Indo/05 from clade 2.1.
Cross-Reactive Antibodies
The 21 rabbits that were immunized with the seven strains of the virus all demonstrated a strong immune response. It can be observed from the anti-H5 antibody titers that there were approximately 10 – 320 with the HI test. Anti-H5 also showed cross reactivity with the seven strains of the H5N1 virus. Antisera from each rabbit were reacted with the seven strains of the virus, and all of them showed high antibody titers (Table 3).
Cross-reactive rabbit antisera by the Hemagglutination Inhibition test.
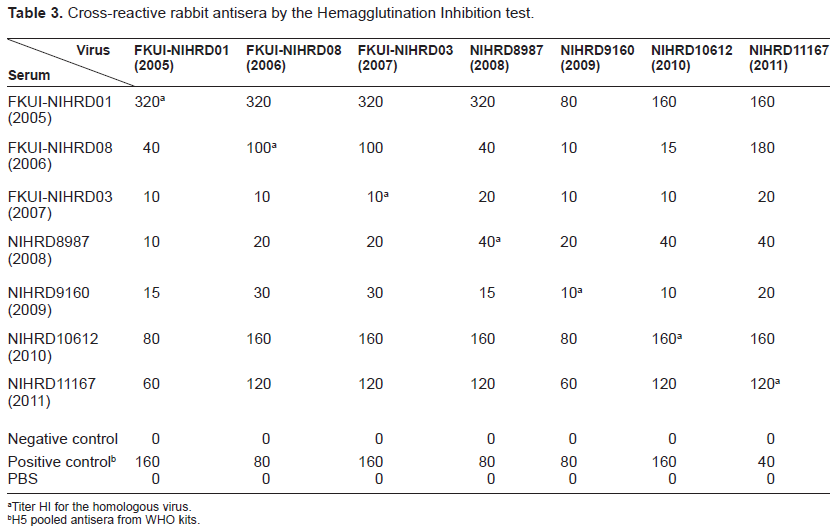
Titer HI for the homologous virus.
H5 pooled antisera from WHO kits.
In this research, we also conducted an analysis of the immune response produced by each strain by using the rabbit antibody panel from those seven strains, even though unusual rabbit antisera cannot be used to evaluate the antigenicity of the influenza virus. By observing the changes of the amino acids in the antigenic site, which also represents the location of antigenic epitope binding, the immune response could be observed to be different. Therefore, the HI test was conducted to determine the reaction of the antibodies with the amino acid changes (Table 3).
The nucleotide mutations cause genetic differences from year to year, especially in the antigenic area, to avoid the immune response of the host. The evolution of the antigenic region needed to be analyzed and compared with the HI test results to determine the activity of antigen-antibody binding. Table 3 shows that the HI test for the anti-H5N1 antibodies demonstrated cross-reactions with the seven strains that were used in this study. Most of the antisera reacted well and gave the results of the high titer antigen used for the immunization (homologous virus). From the seven strains of the virus, it can be observed that there are three viral strains, FKUI-NIHRD01, NIHRD10612, and NIHRD11167, which cause a high immune response and that they cross-react with the complete antisera (Table 3).
For the seven HA protein sequences compared with the reference virus, there are ten different amino acids at the HA1 position (Table 1). The strains NIHRD10612 and NIHRD11167 are different from the other four strains and the reference K35R. It is necessary to conduct a further study of the possibility of amino acid substitutions at position 35 which may contribute to increasing the immunogenicity of the virus since K35R is the only position that distinguish between two strains (NIHRD10612 and NIHRD11167) and other strains.
Whole rabbit antisera obtained in this study cross-reacted with a high HI titer of the viruses FKUINIHRD01, NIHRD10612 and NIHRD11167.
Discussions
Infection of avian influenza A (H5N1) in Indonesia and some other countries in the world has a high rate of human mortality. H5N1 is transmitted from birds to humans and the absence of a suitable vaccine which prevents infections from this virus is alarming.
The characteristics of the H5N1 are similar to the characteristics of some common influenza viruses that continuously change their antigenicity to avoid the host immune response and are related to antigenic drift and antigenic shift. Antigenic drift is primarily observed to occur where there are mutations that have accumulated in some or all of the influenza virus gene segments, especially in the HA and NA genes, which are the main target of the host's defense mechanisms. The immunity against influenza infection occurs via antibodies that interact with viral surface proteins, that is, HA and NA. 23
For antigenic drift, there is concern that the antibodies against the virus subtype which was previously circulating will not cross-react with a new virus subtype. The immune response will protect the host against reinfection by the same virus or an antigenically similar viral strain. Therefore, in Indonesia, we need to determine whether the anti-H5 antibodies from the H5N1 virus circulating in 2005 can cross-react with the virus circulating in subsequent years. According to these studies, the sera have a high H5 antibody titer, greater than 160. These data indicate that all anti-H5 antibodies can cross-react with the H5N1 virus strains used in this study.
However, the results also show that the H5N1 strain in 2005, can react with the antibodies from the whole rabbits used in this study with low titer, except for the antibodies of the rabbits injected with the virus strain from 2005. Thus, it appears that the antigenicity of the H5N1 virus has changed.
Influenza A viruses are generally highly susceptible to mutation, especially in the HA gene, and these mutations occur frequently and unpredictably. Therefore, the immune response, which plays a major role during influenza virus infection, is the humoral immune response. While cellular immune responses maybe contribute effectively to eradicate cells infected by the virus is not analyze in this study.23,24 Thus, the antibody towards HA and NA is important for protection against reinfection.
The influenza viruses used in this study were derived yearly samples, from 2005 to 2011. After sequencing the HA gene and analyzing the phylogenetic tree, it can be observed that all seven viruses are from clade 2.1, subclade 2.1.3.2. Clade 2.1 is a typical viral clade of the H5N1 virus from Indonesia, isolated from both poultry and human beings. The poultry virus is mainly in the subclades 2.1.3.3 and 2.1.3.1, while the human virus is from the subclade 2.1.3.2.
Based on the results of the phylogenetic analysis and the cross reactivity analysis, it is known that viruses in the same subclade can stimulate different immune responses. Therefore, in addition to discovering the genetic characteristics of H5N1, it is also important to discover the antigenicity of each virus. Compared to the HA gene A/Goose/Guangdong/1/96, which was previously part of H5N1, the change in amino acid 138 from H to L for all Indonesian viruses used in this study, may play a vital role in the antigenicity of H5N1 strains in Indonesia. 7
Conclusion
This study shows that the H5N1 strains isolated from Indonesians from 2005 to 2011 are members of the same clade, 2.1.3, but the antigenicity differ. This can be observed in the test results of the HI virus influenza A from 2005–2011 against rabbit antisera. The H5N1virus from 2005 to 2007 may still positively react to rabbit antisera, injected by the antigen virus from 2008 to 2011. The results are lower, however, than for influenza A from 2010 to 2011. It is important to determine this information to have the best vaccine strain selection.
Author Contributions
Conceived and designed the experiments: VS, FI, AS. Analyzed the data: VS, EP, HAP. Wrote the first draft of the manuscript: VS. Contributed to the writing of the manuscript: VS, EP, HAP. Agree with manuscript results and conclusions: VS, EP, HAP, FI, AS. Jointly developed the structure and arguments for the paper: VS, FI, AS. Made critical revisions and approved final version: VS, EP, HAP, FI, AS. All authors reviewed and approved of the final manuscript.
Funding
This work was supported by grants INO AAE 008 XW 08 EC of the WHO for Avian Influenza Research and by the Incentive Research Strategic Programme No 06/M/Kp/I/2012 RT-2012-1197, through the Ministry of Research and Technology.
Competing Interests
Authors disclose no potential conflicts of interest.
Disclosures and Ethics
As a requirement of publication the authors have provided signed confirmation of their compliance with ethical and legal obligations including but not limited to compliance with ICMJE authorship and competing interests guidelines, that the article is neither under consideration for publication nor published elsewhere, of their compliance with legal and ethical guidelines concerning human and animal research participants (if applicable), and that permission has been obtained for reproduction of any copyrighted material. This article was subject to blind, independent, expert peer review. The reviewers reported no competing interests.
Footnotes
Acknowledgments
The authors are grateful to Dr. Trihono, Dr. Ondri Dwi Sampurno, Prof. Sudomo, PhD and Dr. Graham Tallis for their full support for this study. This paper is dedicated to the late Dr. Endang R. Sedyaningsih, MPH, DrPH.