Abstract
Kaposi's sarcoma associated herpesvirus (KSHV or human herpesvirus 8 [HHV-8]) is a gammaherpesvirus highly associated with KS, primary effusion lymphoma (PEL), and multicentric Castleman's disease, an aggressive lymphoproliferative disorder. KSHV, like other gammaherpesvirus latently infects predominantly B-cells and endothelial cells. Infected cells retain the virus from one generation to the next existing as a multicopy circular episomal DNA in the nucleus, expressing a limited subset of viral genes. Of these latently expressed genes, LANA1, the latency associated nuclear antigen is highly expressed in all forms of KS-associated malignancies. Various studies so far show that LANA1 tethers the viral episomes to host chromosomes and binds to specific sites within and close to the TR elements contributing to the stable maintenance of the viral episomes in successive daughter cells. Anti-LANA1 intrabody strategies might represent a new therapeutic approach to treatment of KSHV infections, since LANA1 is regained for KSHV latency. In addition, the use of intrabodies can help drug development by mapping LANA1 inhibiting regions. We report development of several LANA1 specific single chain antibodies from immunized rabbits that can be expressed intracellularly, bind to LANA1 epitopes and can be used for functional KSHV studies on viral latency.
Introduction
Kaposi's sarcoma associated herpesvirus (KSHV or human herpesvirus 8 [HHV-8]) is a gammaherpesvirus highly associated with KS, primary effusion lymphoma (PEL), and multicentric Castleman's disease, an aggressive lymphoproliferative disorder.1,2 KSHV DNA is found in the endothelial and spindle cells of KS lesions, as well as lymphocytes and lymphoid tissues, and establishes a predominantly latent state of infection. Infected cells retain the virus from one generation to the next existing as a multicopy circular episomal DNA in the nucleus, expressing a limited subset of viral genes. 3 Of these latently expressed genes, LANA1, the latency associated nuclear antigen is consistently shown by immunohistochemical analysis and in situ hybridization, to be highly expressed in all forms of KS-associated malignancies. 4
LANA1 is a nuclear protein composed of 1162 aminoacids, runs on gels as a doublet, is expressed from ORF73 and encodes an N-terminal proline rich domain, an internal glutamine rich and an acidic repeat domain followed by a leucine zipper motif.5,6 Both the N- and C-terminal LANA1 domains have been described as capable of localizing to the nucleus. 7 Recent studies indicate that LANA1 displays preferential binding to three different regions of the KSHV genome; The N-terminal region is suggested to contain the cis-acting DNA element.8,9 The LANA1 C-terminus binds two adjacent sites within the KSHV TR. 10 The LANA1 C-terminal DNA binding domain is essential for DNA replication, although the N-terminal also has a role in DNA replication.11–13 Amino acids 5–22 have been described as sufficient to mediate a specific interaction of LANA1 with chromosomes during mitosis, 9 and a region in the C-terminus (aa 1129–1143) has also been described as essential for heterochromatin association but is not on it's own sufficient to mediate this tight interaction. 14
In addition to its role in viral episome maintenance, LANA1 modulates host cell behaviour to create a suitable environment for latent KSHV persistence. Biochemical studies show that LANA1 interacts with many cellular factors such as RING3, 15 pRb, 16 p53,17,18 HP1, 12 CREB-binding protein, 19 MeCp2, 18 DEK, 20 histone H1, 21 ORCs, 13 and GSK-3β. 22 Most of these factors are involved in transcription, chromatin structure, and replication, and some or all of them may be required for the viral maintenance as episomes and govern the transcription program in latency.
Intrabodies in a format of single-chain variable region antibody fragments (scFv) can be expressed intracellularly and bind to viral proteins, and other targets interfering with biological processes that take place inside the cell, representing a valuable tool in functional genomics at the protein level (ex. proteomics). Compared to other gene manipulation methods, such as gene knock-out or RNA-based technologies (ribozyme, antisense RNA or RNA interference), intrabodies have the unique advantage of targeting proteins in different cellular compartments as well as specific structural or functional motifs of a protein; 23 mapping protein domains (linear or conformational) intracellularly without mutations; and a potential use as inhibitors.24,25
Anti-LANA1 intrabody strategies might represent a new therapeutic approach to treatment of KSHV infections with the aim of LANA1 knockout and intracellular structure/function analysis. We have developed several LANA1 specific single chain antibodies from immunized rabbits that can be expressed intracellularly. The antibody fragments were able to recognize epitopes both at the N-terminus, repeat regions and C-terminus. Intrabodies recognizing important recognition epitopes were constructed and use to functionally assess its effect in latency neutralization. In addition, to therapeutic applications, our study may provide insights on the structure-function analysis of LANA without the disruptive use of direct mutagenesis.
Results
Bacterial expression and purification of LANA1 proteins
The strategy used for the generation of specific chimeric rabbit-human Fab antibodies against human LANA1 protein from KSHV started out by the amplification and purification of the LANA1 protein. Two separate fragments LANA1000 and LANA2000 were obtained at a considerable yield (data not shown). Therefore, the last step before the immunization of the rabbits consisted in purifying the native form of both these proteins.
Selection of LANA1 specific antibody fragments from a phage display chimeric rabbit/human fab library
The first step in the construction of an antibody-reactive library is a successful course of animal immunization with an antigen of choice. Antibodies from immune animals are subjected to in vivo selection and, thus, are more likely to recognize a given antigen selectively, that is, with less cross-reactivity to other antigens. Rabbits are widely used for the generation of polyclonal sera to antigens, as they elicit better immune response, and generation of antibody diversity does not rely on the use of many V-segments. Most rabbit B-lymphocytes recombine the same VH gene in the V(D)J gene rearrangements. These genes are then diversified through gene conversion and hypermutation.
Chimeric rabbit/human Fab libraries were generated from cDNA from the spleen and bone marrow of two New Zealand white rabbits after a series of immunizations with both purified LANA1 fragments. 26 Phage display libraries were constructed based on the knowledge of the different immunoglobulin families of variable region sequences of rabbits, known at the time, as described in Material and Methods. Three independent libraries were constructed with each of the immune repertoires represented by a complexity of approximately 2.7 x 10 8 independent transformants (supplementary material; Table 1). Single clones, in a total of 88 phage-Fabs, obtained from the two libraries were then produced as soluble Fabs by transformation in E.coli non-supressor strain TOP10. Protein expression was analyzed for each of these Fab clones by ELISA using goat anti-human IgG Fab for capture and a rat anti-HA mAb conjugated to HRP for detection. Of these 88 Fabs, a total of 34 clones (Fig. 1) with signals higher than background were isolated and further analyzed by DNA fingerprinting, which consisted in the analysis of different sizes of fragments after amplification by PCR. AluI digestion (4-base cutter) of these antibody fragments encoding PCR products was performed to evaluate the diversity among the clones.
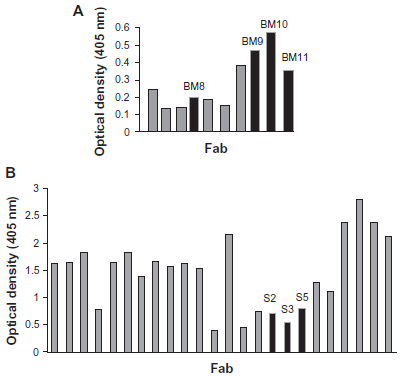
Relative binding affinities of A) anti-LANA1000 Fab antibody fragments and B) anti-LANA2000 Fab antibody fragments. 34 clones with signals higher than background were isolated and analyzed by PCR screening, Alul digest and sequencing; the clones marked in black were chosen for further studies (scFv constructs).
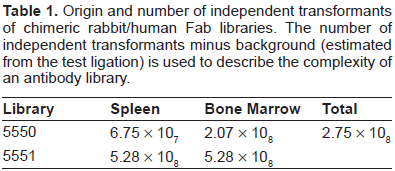
Origin and number of independent transformants of chimeric rabbit/human Fab libraries. The number of independent transformants minus background (estimated from the test ligation) is used to describe the complexity of an antibody library.

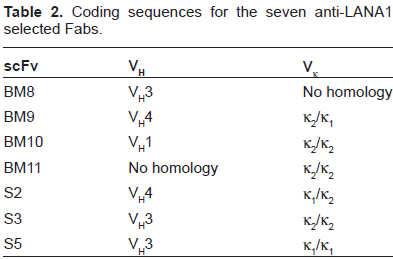
Among 34 positive clones from the 2 different libraries 12 distinct Fabs were identified by the fingerprinting. All 12 positive clones were subsequently analyzed by DNA sequencing of both VH and VL coding regions. Four Fab clones that bound to LANA1000 (Fig. 1A) and three Fab clones that bound to LANA2000 (Fig. 1B) revealed unique sequences that were clonally unrelated. The clones with distinct sequences that bound to LANA1000 were designated BM8, BM9, BM10 and BM11; and the clones that bound to LANA2000 were designated S2, S3 and S5.
The identification of the different coding sequences was done by analysis of the sequence homology existent between the rabbit variable family regions known at the time. Figure 2B shows a schematic N-J tree of the 5 VH and VL scFv families. BM10, BM11 and S3 shared Vκ coding sequences and BM8, S3 and S5 shared VH family coding sequences (supplementary material; Table 2). BM9 and S2 both have an identical VH4 coding sequence.
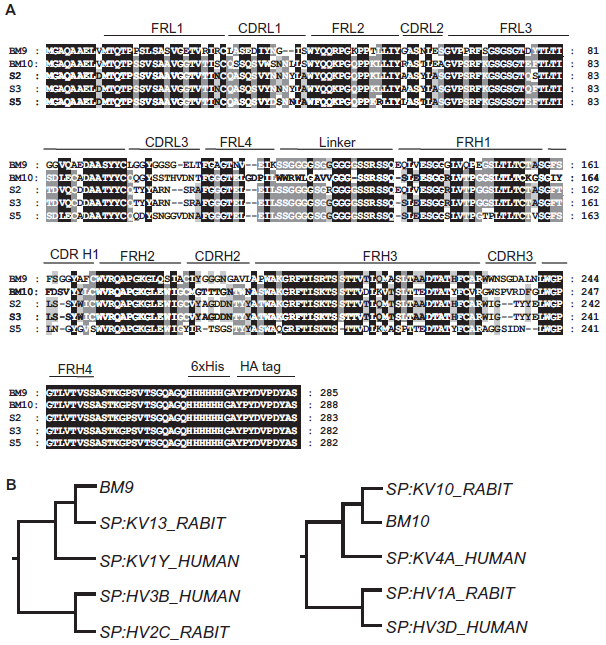
A) Amino acid sequence alignment of the VL and VH sequences of LANA1 specific antibodies. Shown are the CDRs and FRs of VL and VH sequences of 5 of the antibodies; BM8 and BM11 had unspecific readings. Dashes indicate shorter sequences. Diversified positions are shown in grey boxes and conserved positions in black boxes. Alignment of the sequences emphasizing the different FRs and CDRs was based on the numbering according to the Kabat numbering. B) N-J Tree of anti-LANA1 BM9 and BM10 scFvs. Rabbit VH sequence of BM9 and BM10 were found to be most homologous to the human VH3 family. The BM9 VL sequence was found to be most homologous with the human Vκ1 family, and BM10 with the human Vκ4 family.
Coding sequences for the seven anti-LANA1 selected Fabs.
As under the reducing conditions of the cytoplasm the conserved intra and inter disulfide bond of Fabs cannot form, these were transformed into scFvs which are linked through a peptidic bridge allowing the correct folding in sufficient amounts to be active as cytoplasmic intrabodies. The heavy chain complementarity determining region 3 (CDR H3) and light chain complementarity region 3 (CDR L3) length diversity ranged from 10 to 12 and 7 to 9 amino acid residues, respectively, and revealed a similar distribution in both immune repertoires (BM-bone marrow and spleen-S) (Fig. 2A).
LANA1 mapping
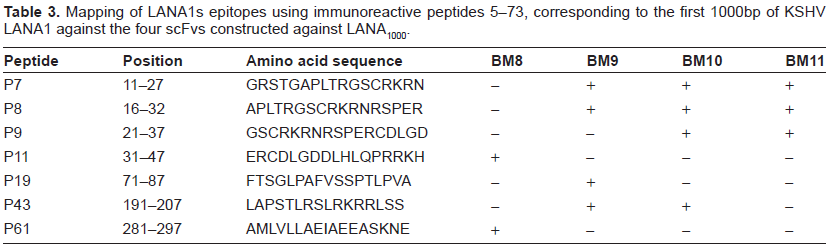
Peptide screening of LANA1 using 171 peptides covering the whole LANA1 protein except part of the repeat region was performed by ELISA, to map anti-LANA1 immunoreactive epitopes of all seven scFv using purified proteins. Similar peptides were previously used as solid-phase antigens in an ELISA assay to test sera from KSHV-infected patients by Olsen et al. 29 Beginning at the amino terminus, these 171 cleavable biotinylated (N-terminus SGSK) 17-mer peptides offset by 5 were sequentially numbered P5 to P96 and P101 to P179. Identical peptides from the repeat region were eliminated. Two biotinylated (N-terminus SGSK and SGSG) 10-mer peptides from the HIV gp41 protein were synthesized and used as internal controls. In order to identify immunogenic epitopes, peptides 5–73 were screened by ELISA against BM8, BM9, BM10 and BM11 (scFvs that bound to LANA1000 fragment which corresponds to the first 1000bp of LANA1). Immunogenic linear epitopes of LANA1 that interacted with these four scFvs are shown in Table 3. Peptides 74–179 were screened against S2, S3 and S5 which correspond to scFvs that bound to LANA2000 fragment, corresponding to 2000bp of the repeat region and carboxyl terminus of LANA1. Immunogenic linear epitopes of LANA1 that interact with these three scFvs are shown in Table 4.
Mapping of LANA1s epitopes using immunoreactive peptides 5–73, corresponding to the first 1000bp of KSHV LANA1 against the four scFvs constructed against LANA1000.
Mapping of LANA1s epitopes using immunoreactive peptides 74–179, corresponding to the repeat region and carboxyl terminus of KSHV LANA1 against the 3 scFvs obtained against LANA2000.
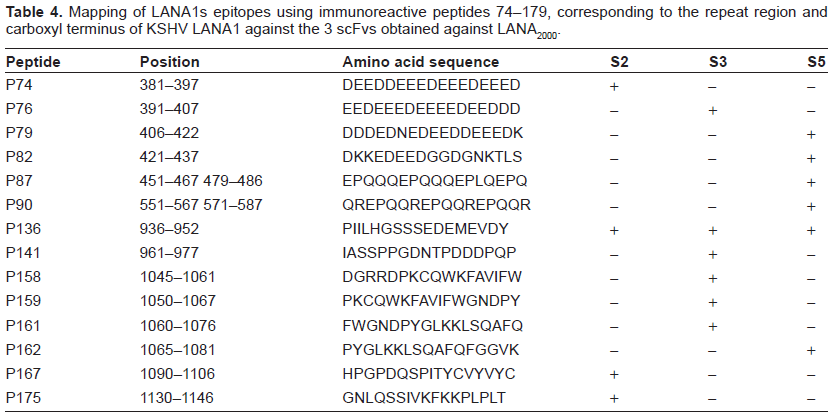
The primary analysis of all 171 biotinylated peptides mapped against the different purified scFvs, identified 20 potential immunoreactive peptides which correspond to 15 potential epitope sequences. These 20 peptides are distributed equally over the entire LANA1 protein, including 7 peptides from the amino terminus, 6 from the repeat region and 8 from the carboxyl terminus. Figure 3 depicts the candidate peptide sequences that interact with the different scFvs.
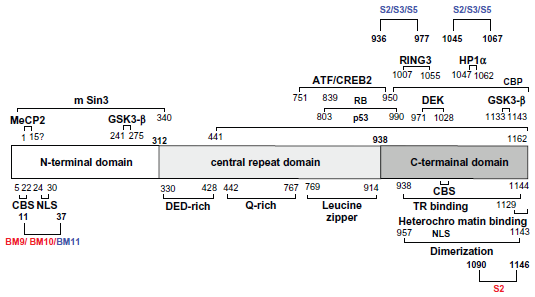
LANA1 is a 222–232 kDa nuclear protein expressed from ORF73 and encodes an N-terminal proline rich domain, an internal glutamine rich and an acidic repeat domain followed by a leucine zipper motif. 30 ScFv binding-epitope regions are indicated under antibody names.
Anti-LANA1 scFvs BM9, BM10 and BM11 interact with the N-terminal sequence motif (aa 5–22) described as functioning to tether LANA1 to chromosomes during mitosis (chromosome binding site (CBS)); 9 and also interact with the nuclear localization signal (NLS) of the protein (aa 24–30). 30 S2 interacts with the TR binding domain (aa 938–1144) described by Garber et al 31 and the heterochromatin binding region (aa 1129–1144) later identified by Viejo-Borbolla et al. 14 One of the epitope regions (aa 936–977) of S2, S3 and S5 is described as the binding site of DEK and CBP close to the C-terminal domain of LANA1. 20 These results indicate a broad interaction of isolated scFv against the LANA protein, raising the possibility that scFv-derived intrabodies can neutralize different functions of the latency antigen.
Expression and functional analysis of scFv
To analyze the functionality of different epitope recognizing intrabodies, studies were first performed in 293 cell line to evaluate in vivo expression.
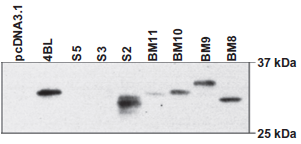
Expression of anti-LANA1 scFvs in pcDNA3.1NLS in 293 cells. BM8, BM9, BM10, BM11 and S2 express in 293 cells. For immunodetection HRP-conjugated anti-HA monoclonal antibody was used. 4BL is a HIV specific scFv used as control.
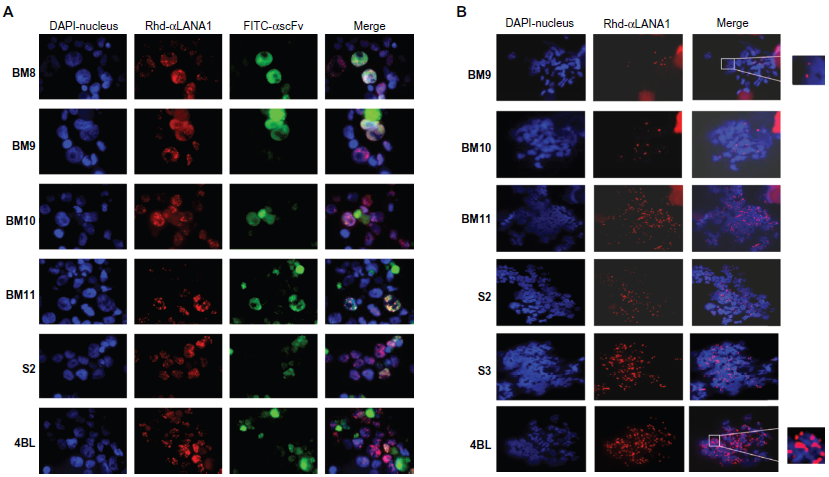
A) Immunofluorescence of BC-1 cells transfected with scFvs. Distribution of LANA1 and anti-LANA1 scFv in BC-1 cells with immunofluorescence microscopy after simultaneous detection with mLANA1 and monoclonal HA. LANA1's speckled pattern becomes dispersed when BM9 and BM10 are expressed. scFv is shown in green (FITC) and LANA1 in red (Rhodamine). B) Metaphase spreads of BC-1 cells transfected with scFvs. LANA1 in red in BC-1 chromosome spreads after detection with mLANA1. The amount of LANA1 protein binding to the chromosomes diminishes and loses its doublet pattern when BM9 and BM10 are expressed. Magnification 1000x.
evaluating dispersion of LANA1 due to scFv inhibition in BC-1 interphase cells. LANA1 spots were counted and student t-test were performed to statistically compare between scFvs and controls a .
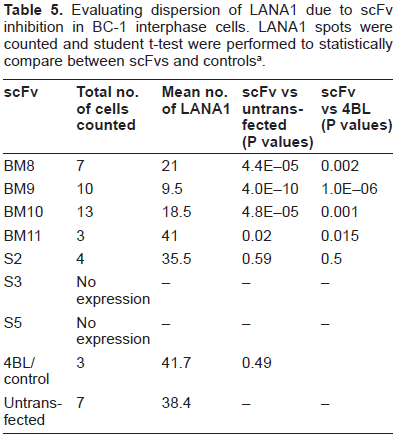
student t-tests p-values for average differences, assuming equal variances and unpaired. Two-tailed for among controls or among treatment comparisons and one-tailed for treatment versus controls.
Evaluating dispersion of LANA1 due to scFv inhibition in BC-1 metaphase cells. LANA1 spots were counted and student t-test statistically compare between scFvs and controlsb.
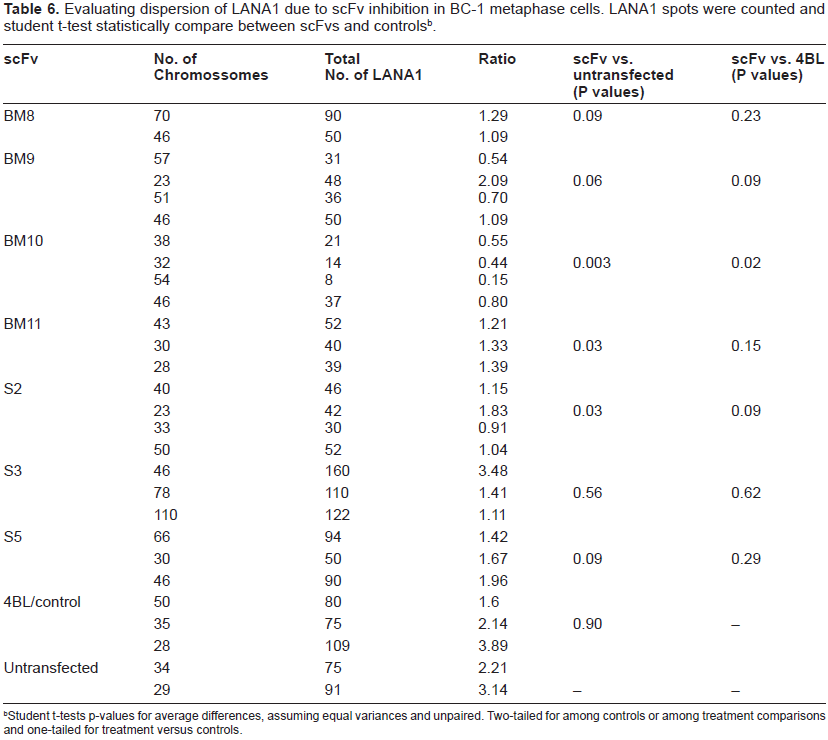
Student t-tests p-values for average differences, assuming equal variances and unpaired. Two-tailed for among controls or among treatment comparisons and one-tailed for treatment versus controls.
The efficacy of intrabody expression for binding and inhibition of LANA1 was examined by a cell viability assay for the quantification of cell proliferation and cell viability (WST-1 Assay). As shown in Figure 6A, LANA1 and Z6 expressing BJAB cells do not survive when anti-LANA1 scFvs are expressed. In contrast, BJAB cells expressing control (pcDNA3.1 vector) do not show a reduction in cell viability. This result indicated that the expression of anti-LANA1 scFvs BM9, BM10 and S2 targeted the latency antigen, with probable episomal elimination from the cell, since LANA1 tethers the episome to the chromosome during cell division.
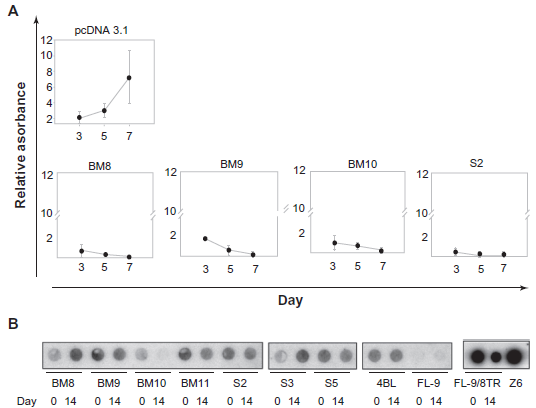
A) Outgrowth Assays to detect episome loss in FL-9/8TR cells. FL-9/8TR cells were transfected by nucleofection with each scFv. Empty vector pcDNA3.1 NLS was transfected as control. Cell counts were done every 2 days in 24 replicas of each using cell proliferation reagent WST-1 and measured by spectrophotometer at 450 nm. ScFvs that are effective in neutralizing LANA1 inhibit cell growth. S.D. is indicated by error bars. B) Southern Blot to detect episome loss in FL-9/8TR cells. FL-9/8TR cells were transfected by nucleofection with each scFv 4BL was transfected as control. Day 0–-zeocyn selection at 48 h pos-transfection. Day 14 is 14 days after selection with zeocyn was initiated.
To assess whether episomal loss was a consequence of scFv expression, FL-9/8TR cells were independently transfected with the several scFvs and control vectors and selected with zeocyn, hygromycin and neomycin for maintenance of single-chain antibodies, pSG5-LANA1 plasmid and Z6 cosmid respectively. As depicted in Figure 6B, Southern Blot experiments using a TR probe that detected Z6 cosmid, showed a reduction in episomal presence 14 days after selection. In contrast, cells transfected with an irrelevant scFv (4BL) that recognizes a HIV-1 antigen did not show an elimination of Z6 episome after 14 days, and BJAB cells which did not contain Z6 (FL-9) but expressed BM9 or BM10, did not show a reduction in cell viability. Episome loss is detected in BM9, BM10, BM11 and S5, fourteen days after having been transfected into FL-9/8TR cells and selected.
As fibroblasts and B cells stably expressing the several KSHV/LANA1 constructs indicated loss of episome after interaction of LANA1 with the different scFvs, outgrowth assays were performed to confirm these results in PEL cell lines which contain endogenous KSHV and LANA1. Two cell lines were used in these assays–-BC-1, a B-cell tumor line co-infected with KSHV and EBV and, BCBL-1, a B-cell tumor line infected only with KSHV. LANA1 knockdown by expression of anti-LANA1 scFvs BM8, BM9, BM10 and BM11 in BC-1 cells, showed marked short-term growth inhibition as depicted in Figure 7A. In BCBL-1 cells BM9, BM10 and S2 expression promote LANA1 knockdown and consequently growth inhibition, as shown in Figure 7B.
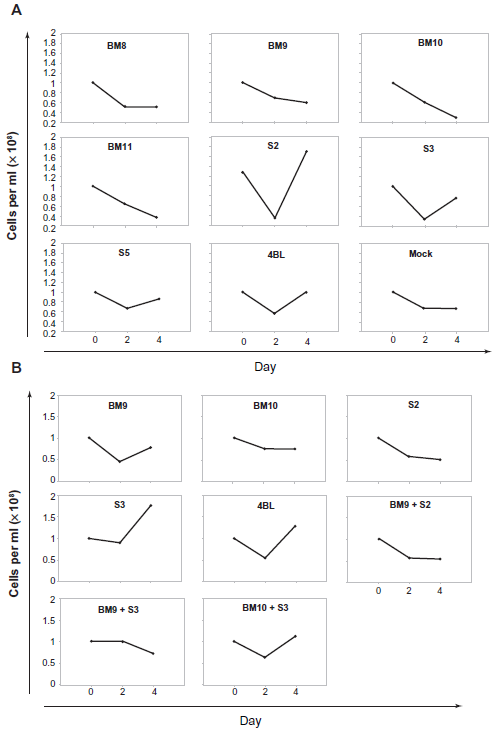
Outgrowth assays in A) BC-1 cells and B) BCBL-1 cells transfected with ScFvs. ScFvs that are effective in neutralizing LANA1 inhibit cell growth.
Discussion
The phage display approach with its natural linkage of phenotype and genotype provides ready access to antibody sequences and facilitates further in vitro optimizations such as humanization or affinity maturation. Using phage display, we selected Fab antibodies from rabbits immunized with LANA1, a human viral antigen from KSHV, and transformed these Fabs into scFv antibodies. Using Fab display, we were able to select antibodies that have a higher affinity to the LANA1 antigen then when using scFv display. This is based on the fact that Fab are displayed monovalently at the phage surface and are selected based on their affinity and expression, whereas the selection of scFv is also influenced by avidity due to their tendency to dimerize or form higher order aggregates. The conversion from Fab to scFv format yields scFv with high affinity and does not affect the antigen-binding site.
A chimeric rabbit/human Fab library was generated from cDNA derived from bone marrow and spleen RNA of two immunized rabbits that had a strong immune response to LANA1 protein fragments. Animals were immunized with a mixture of LANA1 fragments from aminoacids 1 to 330 (LANA1000) and 330 to 1162 (LANA2000). Antibody libraries of 1 x 10 8 independent clones were constructed and selected. Seven Fab clones that bound LANA1 protein fragments were isolated and converted in single-chain antibody fragments (scFv). The scFv format of the antibody allows the expression in the cytoplasm in a lower molecular weight form. Anti-LANA1 scFvs were constructed by joining VL and VH antibody fragments with a long peptide linker of 18 aminoacids (SSGGGGSGGGGGGSSRSS).
Periplasmic expression of scFv proteins is limited by the overall in vivo folding efficiency. As periplasmically expressed proteins do form intradomain disulfide bonds, they tend to be more stable than when expressed in the reducing cytoplasm. 28 In single-chain antibodies the variable domain of the heavy chain (VH) is connected to the light chain (VL) through a peptide linker, preserving the specificity and affinity of the parent antibody. The presence of the linker is a stabilization factor compared to the Fv fragment. In principle, both the length and sequence of the linker can have an effect on stability, as in antibodies the existence of a ‘domain swapping’ phenomenon, where dimeric, trimeric and even tetrameric assemblies can form with short linkers. Therefore, including linkers of 15–25 residues increases the stability of scFv fragments by favouring monomer formation, which is the thermodynamically stable form of scFv fragments. 34 Stability of scFv fragments is often believed to be directly correlated with expression yield. 28 In the cytoplasm, proteins are made in the reduced form and indeed a good correlation between stability and functional yield of soluble scFv protein was observed in a number of cases.35–37 Most likely, stability is usually the limiting factor for folding of a scFv in the cytoplasm, since the intradomain disulfide bonds cannot form and only intrinsically stable, but not unstable fragments, can fold efficiently. 28 This is confirmed by results from our lab, that show that the intracellular solubility and stability of an intrabody is a critical factor that determines its efficiency in neutralizing intracellular antigens. 24
The antibody diversity generated by VHDJH rearrangements in rabbits is more limited than in mice and humans, since VH1 is predominantly used out of more than 50 functional VH gene segments. 38 In this sense the different families described in the results may have already integrated this balance as BM10 family (VH1; VL– κ2/κ2) is different from BM9 (VH4; VL– κ2/κ1). The scFv protein sequences were blasted and aligned with rabbit and human V genes. The selected rabbit VH sequence of BM9 and BM10 were found to be most homologous to the human VH3 family. The BM9 VL sequence was found to be most homologous with the human Vκ1 family, and BM10 with the human Vκ4 family. The human VH3 and Vκ1 not only gave the best alignment with the selected rabbit sequences, but are frequently found in the human antibody repertoire in vivo, and are highly related to human V genes whose frameworks have been used for a number of mouse antibody humanizations.39,40 In fact, pairs of VH3 family heavy chains and Vκ1 family light chains are the most frequent combinations found in native human antibodies, and so, this framework combination is expected to be non-immunogenic in humans, which would make BM9 a good candidate for humanization.
LANA1 has a central role in KSHV biology. It mediates episome persistence and is known to have transcriptional regulatory properties. LANA1 also interacts with multiple cellular proteins through specific domains. LANA1 associates with mitotic chromosomes,41–43 and immunofluorescence microscopy demonstrated that LANA1 concentrates at sites of KSHV DNA along mitotic chromosomes. It has been described by Barbera et al, that the full length LANA1 N-terminus contains the dominant chromosome binding region because although LANA1 contains a C-terminal chromosome association domain, it plays a different role when in the presence of KSHV episomes. 30 An epitope is the small determinant on the surface of a ligand with which the receptor makes close, geometrically and chemically specific contact. 44 Many epitopes including discontinuous ones, are conformation-dependent because they require the context of the overall protein structure to constrain them in a binding conformation.
To map anti-LANA1 scFv epitopes, LANA1 peptide screening using 171 peptides was performed by ELISA using the different purified single-chain antibodies, as previously described to determine dominant epitopes recognized by sera from KSHV-infected patients. 45 This screening identified 15 different potential linear epitopes, although, LANA1 immunoreactivity may be due to conformational or posttranslational epitopes. Compared to the screening effectuated by Olsen et al 45 using human sera, three of these epitope regions are approximately the same, two of them corresponding to the N-terminal sequence motif of LANA1 and one to the C-terminal.
Three of the epitopes corresponding to the C-terminal of LANA1 where the scFv interact: aa 1045–1081 by S3; aa 1090–1106 and aa 1030–1046 both by S2 have been described as essential for replication of TR containing episomes. 10 The aminoacid sequence responsible for localizing LANA1 to chromosomes has been mapped to the N-terminus (aa 5–22) 43 and residues 5 to 13 have been described as essential for chromosome association, DNA replication and episome maintenance. 46
Previously, studies have shown that intracellular antibody expression has no obvious negative effects on cell viability or proliferation. 47 Nevertheless, we quantified cell proliferation and cell viability of BJAB human B-cell line stably transfected with LANA1 and Z6 cosmid (FL-9/8TR) compared with BJAB human B-cell line stably transfected with pSG5 empty vector. The assay consists of a colorimetric assay based on the cleavage of the tetrazolium salt WST-1 by mitochondrial dehydrogenases in viable cells. The kinetics of WST-1 metabolism showed that both these cell lines had similar levels of proliferation, however, when either BM8, BM9, BM10 or S2 are transfected into FL-9/8TR, cell killing occurs within 5 days of scFv expression whereas cells transfected with empty vector (pCDNA 3.1) continue to replicate. Cells expressing BM9, BM10, BM11 and S5 have marked loss of TR plasmid by day 14 of culture compared to cells expressing an irrelevant anti-HIV scFv, 4BL. Therefore, it is conceivable that BM9, BM10 and BM11bind conserved epitopes for LANA neutralization and consequent cell killing.
In primary effusion lymphoma cell lines derived from patients LANA1 is predominantly localized in the nucleus. Our results show that anti-LANA1 specific scFvs expressed in the nucleus interact with the N-terminus of LANA1, dispersing and knocking off LANA1 from the chromosomal DNA. LANA1 expression was down regulated in the vast majority of cells expressing scFvs, as determined by IFAs (Fig. 5). The results obtained from the co-transfection of two scFv (one from the N-terminal and another from the C-Terminal-Fig. 6) also support this finding as no additive effect is demonstrated. These results are statistically relevant as shown in Tables 5 and 6 by student t-tests (P values < 0.05) indicating that the knock-off of LANA1 from chromosomal DNA, in both interphase and metaphase arrested cells, is statistically different from both untransfected cells or cells transfected with 4BL (anti-HIV specific scFv).
As a conclusion, the results presented here extend previous studies demonstrating that high affinity single-chain antibodies can be stably expressed and can localize to the nucleus when a nuclear localization signal (NLS) is added. Furthermore, the toxicity of the scFvs was low as assessed by cell viability in intrabody-expressing human PEL cells (4BL/control). LANA1, analogous to the SV40 T antigen, also targets tumor suppressor proteins and other regulatory proteins including pRB 16 and p53. 17 LANA1 has both auto-activating activity and repressor activity for KSHV genes.6, 48, 49 In the latter case, LANA1 expression may repress lytic viral gene expression preventing inadvertent entry into the lytic viral life cycle. BM9 and BM10 scFv expression not only disperses LANA1 protein but also inhibits growth of PEL cell lines as shown by previous results, 26 confirming the requirement for LANA1 expression in KSHV positive cells. Although we have not formally examined whether BM9 and BM10 neutralization may inhibit pRb, p53 or GSK-3β signalling pathways our previous data shows that at least a disruption of the mitotic bridge between cellular chromatin and the viral epissome occurs. Ongoing studies are being conducted to determine precisely how neutralization causes this effect.
The effectiveness of these relatively low-binding scFv in inhibiting latent KSHV infection demonstrates the feasibility of using engineered intracellular antibodies as a novel therapy. Intrabody fragments could potentially be delivered through gene therapy methods allowing direct treatment of latently-infected cells. Manipulation and screening of scFv for high affinity anti-LANA1 antibodies may result in even more efficacious agents.
Materials and Methods
Expression and purification of LANA1 proteins
LANA1 was amplified by PCR from pSG5 LANA1-FLAG, 32 (a kind gift from Dr. Kenneth Kaye, Harvard University) into two separate fragments LANA1000 (1–337aa) and LANA2000 (337–1162aa) due to the existence of a repeat region (region rich in proline and glycine) in between 1000 bp and 2000 bp that complicated both amplification and purification of such a large protein. A polyhistidine tag (His6) was added at the N-terminal end of each fragment. Primers used were as follows: LANA11000: (S) 5‘-ATGGCGCCCCCGGGA ATGC-3’; (AS) 5‘-CTAATGGTGATGGTGATGCT GCTCCTCATCTGTCTCCTGC-3’; LANA12000: (S) 5‘-ATGCAGGAGACAGATGAGGAGGACGAG GAGG-3’; (AS) 5‘-CTAATGGTGATGGTGTGT CATTTCCTGTGGAGAGTCCCCAGGTGG-3’. These products were cloned into pBad-TOPO (Invitrogen, Carlsbad, CA) and expressed in bacteria by induction with arabinose (Invitrogen, Carlsbad, CA). LANA1000 was purifed by FPLC-affinity cromatography using Ni-NTA resin (Invitrogen, Carlsbad, CA); and LANA2000 was purified by ionic exchange cromatography followed by molecular exclusion chromatography.
Phage display library
Two New Zealand White Rabbits were immunized with purified LANA11000 and LANA12000 proteins. Spleens and bone marrows were harvested separately 50 for total RNA using TRI Reagent (Molecular Research Centre Inc., Cincinnati, Ohio). First strand cDNA was synthesized using Superscript (Invitrogen, Carlsbad, CA). Rabbit VL and VH sequences were combined with human CL and CH1 sequences by overlap PCR, as previously described. 51 PCR fragments encoding a library of antibody fragments (Fab) were digested with SfiI, purified and cloned into pComb3X 50 which contains an amber stop codon, 6x histidine (His6) for purification, and hemaglutinin (HA) for detection. 50 Fab fragments were expressed in fusion with protein III of M13 phage and screened against LANA1 proteins. 88 anti-LANA1 Fabs were expressed, purified and screened by ELISA against LANA11000 and LANA12000 proteins. For the conversion of a LANA1-specific Fab into a scFv, specific oligonucleotide primers covering all known rabbit antibody family sequences were used to amplify VH and VL gene segments. 50 The purified PCR products were assembled by overlapping PCR and the resulting PCR products encoded different scFvs in which the VL region was linked with the VH region through an 18-amino acid peptide linker (SSGGGGSGGGGGGSSRSS). DNA fragments were gel purified, digested with SfiI, and cloned into pComb3X. LANA1 binding activity of the corresponding scFv was confirmed and the genes encoding the scFv were cloned into pCDNA3.1/Zeo (+) (Invitrogen, Carlsbad, CA). 50 ScFv proteins were purified using BD TALON Metal Affinity columns (Clontech, Palo Alto, CA) as previously described. 47 Purified scFv proteins were analysed by immunoblotting using standard methods. 4BL is an irrelevant scFv used as control. 47
Mapping LANA1 epitopes
LANA1 epitopes were mapped using 171 cleavable, biotinylated (N-terminus SGSK) 17-mer peptides offset by 5 amino acids as previously described. 45 Briefly, streptavidin ELISA plates (Roche Applied Science, Indianapolis, IN) were coated with 1.5 μg of each peptide. After blocking with 3% BSA/PBS, peptides 5 to 73 (1–357 aa) were incubated with purified scFvs BM8, BM9, BM10 and BM11 and peptides 74–179 (346–1162 aa) were incubated with S2, S3 and S5. Detection was performed by HRP conjugated anti-HA monoclonal antibody (high affinity 3F10; Roche Applied Science, Indianapolis, IN) at a dilution of 1:3000. Absorbance was measured on a spectrophotometer (Biophotometer, Eppendorf) at 405 nm.
B cell transfection by electroporation-nucleofection
24 to 48 hours before transfection cells were spun down at 1000 rpm for 5 min and ressuspended in fresh RPMI 1640 complete media. An aliquot of culture medium was pre-warmed at 37 °C in a 50 ml tube (500 μl per sample). The Nucleofector solution (Amaxa) was pre-warmed to room temperature. Twelve well plates were prepared by adding 1.5 ml of media and pre-incubated at 37 °C/5% CO2 incubator. Cells were spun, washed with PBS, and cell density was calculated by counting on a hemacytometer after staining cells with trypan blue (stains dead cells). Appropriate volume of cells (1 x 10 6 cells per transfection) was put in an eppendorf, spun and supernatant was completely discarded. Pellet was ressuspended in 100 μl of Nucleofector solution and 1–5 μg of plasmid DNA (in 1–5 μl of TE or water) was added. Sample was transferred to an Amaxa cuvette and nucleofected using the appropriate program. 500 μl of pre warmed media was added to the cuvette, carefully transferring the sample into the 12 well plate. For transfection of BJAB cells solution R and program T-01 was used; for BC-1, BCBL-1 and BC-3 cells, solution V and program T-01. Transfection efficiency was measured by FACS (Flow activated cell sorting) using a GFP plasmid provided by Amaxa. Antibiotic selection or harvesting of cells was done 48 h post-transfection.
Outgrowth assays
48 h after transfection FL-9/8TR cells were selected for LANA1 with hygromycin 200 U/ml, for Z6 with G418 at 400 μg/ml and for scFv with 200 μg/ml of zeocyn (antibiotic concentrations were optimized after viability and growth assays were performed using untransfected cells). PEL cell lines were selected only with zeocyn for the different scFv and controls. Cell counts with trypan blue (colometric staining of dead cells) or using cell proliferation reagent WST-1 (Roche) measuring absorbance at 450 nm with reference at 655 nm were done. This assay is based on the cleavage of the tetrazolium salt WST-1 by mitochondrial dehydrogenases in viable cells. The activity of mitochondrial dehydrogenases, an indicator of cell viability, was measured in 24 replicas every 2 days, by the conversion of the tetrazolium salt WST-1 to the red dye formazan.
Immunofluorescence assays (IFAs)
IFAs were performed 48 hours after Amaxa transfection using different cell lines. Cells were washed twice with PBS, counted in a hemacytometer and diluted to around 25000 cells per spot in a total volume of 200 μl PBS. Cells were cytospun at 1000 rpm for 10 minutes onto superfrost/Plus slides. Slides were air dried, and cells were fixed in 4% paraformadehyde for 20 min at RT and washed for 1 min in water. Cells were permeabilized in 1% Triton-X/PBS for 10 min at RT, washed in PBS for 1 min and blocked for 30 min at RT with 10% normal goat serum (NGS). Cells were incubated with the different antibodies, for 1h at RT in a humidified chamber, washed 3 x 5 min with 0.1% Tween-20/PBS, incubated with the secondary antibodies for 1h at RT and washed 3 x 5 min with 0.1% Tween-20/PBS. Slides were mounted with vectashield and DAPI vectashield (3:1) to stain nuclei.
Metaphase spreads-IFAs and G1 arrest
To arrest cells in metaphase, these were incubated in appropriate medium containing 0.1 μg/ml of colcemid (stock-10 μg/ml in Hank's Balanced Salt Solution from Irvine Scientific) for 24 hours (250000 cells), 24 hours after Amaxa transfection. Cells were washed twice with PBS and ressuspended at 1,25 x 10 5 cells/500 μl 0,075M KCl for 12 minutes to swell nuclei. IFAs were performed same as above. To arrest cell in G1, cells were incubated in appropriate medium containing 200 μm of mimosine for 24 h. After washing twice in PBS, cells were cytospun at 1000 rpm for 10 min (no KCl). IFAs were performed same as above.
Abbreviations
single-chain variable domain fragment;
bone marrow;
spleen;
Kaposi's Sarcoma Associated Herpesvirus;
Latency Associated Nuclear Antigen.
Disclosures
This manuscript has been ready and approved by all authors. This paper is unique and is not under consideration by any other publication and has not been published elsewhere. The authors report no conflicts of interest.
Footnotes
Acknowledgments
We would like to thank Dr. Patrick Moore and Dr. Yuan Chang for the valuable and outstanding help during the development of this work. This work was supported by grants from the Fundação para a Ciência e Tecnologia, Portugal (POCTI/33096/MGI/2000 and PSIDA/MGI/49729/2003) and NIH/NCI grants. S.C.R was a recipient of a doctoral fellowship from Fundação para a Ciência e Tecnologia, Portugal.
