Abstract
The nuclear hormone receptor (NHR) superfamily is composed of a wide range of receptors involved in a myriad of important biological processes, including development, growth, metabolism, and maintenance. Regulation of such wide variety of functions requires a complex system of gene regulation that includes interaction with transcription factors, chromatin-modifying complex, and the proper recognition of ligands. NHRs are able to coordinate the expression of genes in numerous pathways simultaneously. This review focuses on the role of nuclear receptors in the central nervous system and, in particular, their role in regulating the proper development and function of the brain and the eye. In addition, the review highlights the impact of mutations in NHRs on a spectrum of human diseases from autism to retinal degeneration.
Introduction
The nuclear hormone receptor (NHR) superfamily is a group of transcriptional regulators that play a key role in numerous pathways, including development, growth, metabolism, and maintenance. These receptors belong to a large family of genes that regulate gene expression. NHRs are activated by ligands, such as hormones and vitamins, and a subset of these NHRs fall under the category of orphan receptors with unidentified ligands. NHRs share a common structure characterized by dimeric zinc fingers that can bind to a lipid-soluble hormone and form a complex with other proteins, such as cofactors and histones, to regulate gene expression. NHRs modulate numerous processes, such as central nervous system (CNS) development, homeostasis, reproduction, differentiation, metabolism, circadian functions, steroidogenesis, cell differentiation, and lipid metabolism.1, 2 The goal of this review is to discuss the role of NHRs in the development of the CNS, with an emphasis on their function in the brain and the eye and their impact on human disease.
While being involved in a wide range of functions, NHRs have very conserved structure of functional domains (Fig. 1A). There are two activation function (AF) domains that bind to DNA. The variable NH2 terminal (A/B) contains an activation function AF-1 domain, and the AF-2 domain is found in the E domain also known as the ligand-binding domain (LBD). Modifying the A/B domain produces different isoforms of the receptors. A conserved DNA-binding domain (C) binds to specific DNA sequences known as the hormone response element (HREs). In addition, the DNA-binding domain is composed of two zinc fingers and is the most conserved region of the NHR. A P-box region is found on the first zinc finger and functions to recognize the Schematic Representation of the Functional Domains of NHRs in Each Classification Cluster. The A/B Domain is the Most Variable Region, and it Contains the Activation Function-1 (AF-1) Site. The Activation Functions AF-1 And -2 are Important in the Regulation of Receptor Transcriptional Activity. The C Domain is One of the Most Highly Conserved Regions Across the Superfamily on Nuclear Receptors and Consists of the DNA-Binding Region. The D Domain, Also Known as the Hinge Region, is Involved in the Conformational Changes that Occur After Ligand Binding Allowing the Coactivators or Corepressors to Bind. LBD is the Second Most Conserved Region and is in Charge of the Recognition and Binding of the Ligands. The F Region is a Carboxyl Terminal that is not present in all receptors.2, 4, 73.
Regulation
Hormone Response Elements’ (HREs) Sequences and their Coactivators and Cosuppressors.
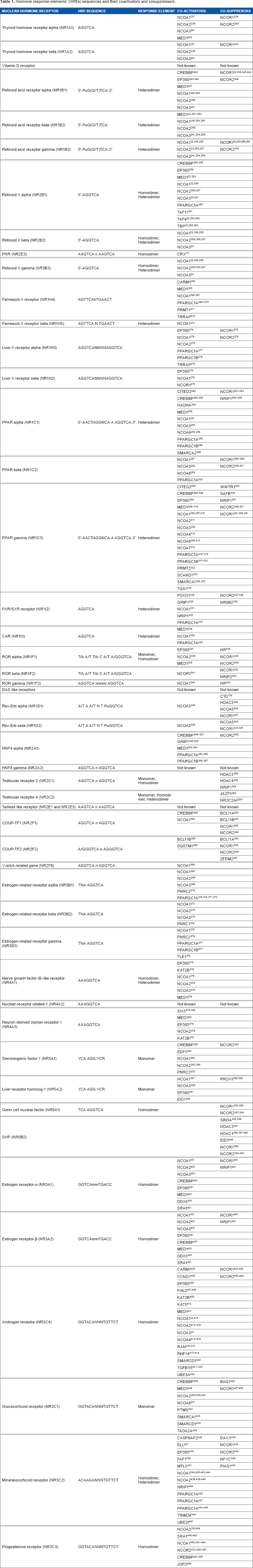
Regulation by HREs
HREs are small sequences of DNA commonly located near the promoter region of a gene, yet they may also be found up to several hundred kilobases upstream of the promoter and even within the first intron of a gene.
7
Each receptor subfamily has a preferred HRE motif it binds, although significant variation has been observed in the ability of NHRs to bind to particular HRE sequences (Table 1). For example, steroids predominantly bind to the motif AGAACA, while thyroid hormones bind to AGG/TTCA.
8
The motif AGGTCATGACCT is recognized by both the thyroid hormone and the retinoic acid (RA) receptor (RAR). In addition to the motif, the number of nucleotides between motifs and the direction of the motif contribute to binding capacity. For example, the arrangement of direct repeats (DR) AGGTCAnxAGGTCA, inverted repeats AGGTCAnxT GACCT, and reverted repeats (ER) TGACCTnxAGGTCA has anything from 0 to 8 nucleotides as spacers.
9
The different combination of spacing and orientation ensures specificity. Nuclear receptors can bind to more than one half-site configuration, allowing coregulation and cross talk between nuclear receptors.
10
The optimal binding HRE site for the orphan hormone receptor Nr2e3, identified by evaluating a series of synthetic duplex DNA fragments acquired from a library of random segments, is a hexamer core binding site of AG(G/T)TCA.
11
Receptors can bind to the HREs as monomers as in the case of many orphan receptors, as homodimers as in the case of the majority of steroid receptors, or as heterodimers as in the case of several nonsteroid receptors, which usually bind to the promiscuous retinoid X receptor (RXR) (Fig. 2A).3, 12, 13 There are nonpermisive heterodimer complexes that are activated only by the partner receptor and not by RXR, and permissive heterodimer complexes that can be activated by either of the receptors.14, 15
HREs’ Potential Patterns Associated with NHRs. Schematic Representation of (
HREs are classified by their ability to repress (negative HREs [nHREs]) or activate (positive HREs) gene expression. HRE sequences have been identified near the 5′ or downstream of the TATA box and even been within the 3′-untranslated region.6, 7 nHREs have been identified in genes that are expressed in the pituitary gland highlighting the importance of the feedback mechanism in hormone regulation. 16 nHREs differ from pHREs in (1) the nucleotide sequence of the half-site, (2) in the orientation of the half-site, and (3) in the number of nucleotides that separates them (Fig. 2B). 17 It is interesting to note that unoccupied nHREs increase the transcription of a gene, while binding to the ligand has the opposite effect of repressing gene expression. Some sequences of the nHREs overlap with the binding site of different activation factors, therefore, creating a physical competition between the receptors and the transcription factors in the regulation gene expression. 18 HREs are one of the several mechanisms by which receptors regulate genes. Another important mechanism is the regulation through coregulators.
Coregulators
Accessory proteins, known as coregulators, can repress (corepressors) or activate (coactivators) the expression of a given gene (Fig. 3 and Table 1). Coactivators have several mechanisms of action. For example, they can directly interact with the nuclear receptor activation domain in the absence of an antagonist and do not directly bind to the promoter, therefore, preventing them from initiating transcription on their own.19, 20 Another way in which coactivators operate is by interacting with components of the transcription machinery and other transcription factors or directly with the promoter changing chromatin conformation via histone acetyltransferase (HAT) proteins (Fig. 3A).
20
Coactivators bind predominantly to the AF-2 and AF-1 domains and interact with transcription factors, such as transcription factor II B and TATA box-binding protein. A coactivator can contain more than one motif allowing the molecule to interact with receptors found in dimeric form. One of the most common known motifs is the Leucine LXXLL motif; however; not all coactivators have this motif. For example, the coactivator SUG-1, an ATPases of the 26s proteasome, lacks the LXXLL motif and, therefore, does not bind to the AF-2 region.6, 21, 22 Several coactivators have been identified, including the steroid hormone receptor coactivator group (SRC-1, -2, and -3), the E3 ubiquitin protein ligase E6-AP, and the RFP-1 coactivator. It is important to highlight that coactivators can also be found in complexes, such as the TRAPs/DRIPs and TRAP220/PBP that also interact with the transcription factors.
23
Expression of genes can be repressed in the presence of an antagonist or in the absence of the ligand.
Ranscription Repression or Activation Mediated by RAR-RXR Nuclear Receptors. Coregulator Binding can Alter Chromatin Structure, Aiding Gene Transcription or Blockage via Histone Acetylation or Deacetylation. (
Corepressors are known for their ability to bind to a receptor as well as to elements of the transcription machinery, thus reducing the promoter activity (Fig. 3B and Table 1). 22 Examples of corepressors includes the nuclear receptor corepressor (NCoR), also known as the silencing mediator for the retinoid and thyroid hormones (SMRT) (Fig. 3B). Steroid hormones, such as glucocorticoid receptor (GR), mineralocorticoid receptor (MR, Nr3c2), and estrogen receptor (ER), do not interact with NCoR; they instead bind to the corepressors in the presence of an antagonist molecules silencing the expression of a gene. 24 The mechanism of action of the coactivators has been defined largely by crystallographic studies that illustrate the molecular conformation changes of NHRs upon ligand binding, thereby releasing the corepressor complex and leaving the site open for the recruitment of a coactivator complex. 25
Classification
The classification of NHRs has proven to be as complex as the myriad of interactions that they regulate. Another way of grouping the more than 50 members of this group is by focusing on the dimerization and DNA-binding properties of each receptor, grouping the NHRs into four classes. Receptors that belong to Class I bind as homodimers to a palindromic HRE sequence of two half-sites separated by a variable DNA sequence. Class II receptors commonly bind as heterodimers to RXR direct repeats. Class III receptors bind as homodimers to direct repeats. Finally, Class IV receptors bind as monomers to a single half-site.26, 27 This form of grouping does not reflect the great number of functions in which the receptors are involved. Taking into considerations the function they perform, the nuclear receptors are divided into three major clusters. The first cluster contains the families of the glucocorticoid receptors, estrogen receptors, mineralocorticoid receptors, androgen receptors, and progesterone receptors. The second cluster includes the families of the thyroid receptors (THRs), vitamin D receptors (VDRs), retinoic acid receptors (RARs), peroxisome proliferator-activated receptors (PPARs), farnesoid X receptor (FXRs), liver x receptor (LXR), pregnane X receptor/steroid and zenobiotic receptor (PXR/SXR), RXR, and constitutive androstate receptor (CAR). The third cluster is composed of the receptors whose ligands have not yet been discovered and are commonly known as the family of the orphan nuclear receptors (ONRs).28–30 Figure 1 illustrates the functional domains of NHRs in each classification cluster. Interestingly, the phylogeny of NHRs depicted in Figure 4 shows the cluster of receptors independent of their expression profile or dimerization and DNA-binding properties, highlighting the importance to put in place another classification system. This classification leads to a different nomenclature where a number from 0 to 6 identifies each clade. In this review, we focus on the major families of NHRs that play important roles in the CNS development. Table 2 describes NHRs expressed in adult mice and in the human brain. It is important to highlight that the receptors involved in the CNS (Fig. 4, blue colored) are common and can be found throughout all the clades.
Phylogenetic Depiction of the Nuclear Receptor Families. Receptors Expressed in the CNS are Highlighted in Blue.
122
. Nuclear Receptors Expressed in the Different Areas of the Human or Mouse Brain.

Nuclear Retinoid Receptors Family and the CNS
There are two families of nuclear retinoid receptors: RARs and R XRs. Both families of receptors can bind to target genes as either heterodimers or homodimers allowing them the capacity to target genes in different pathways. 6 Humans have the following three types of RAR genes: RARα, RARβ, and RARγ; there are the following three types of RXR: RXRα, RXRβ, and RXRγ. Thirteen different proteins can be produced from the six different genes identified thus far as a consequence of alternative splicing and different promoters. 31 An important characteristic that differentiates the two classes of retinoid receptors is the unique amino acid sequence found in LBD conferring each receptor with specific individual characteristics, thereby allowing them to modulate different biological pathways. 32
The unique LBD sequence establishes varying affinity toward a particular ligand. For example, RARs bind with high affinity to 9-
When RXR binds to a THRs or VDR it increases the affinity of DNA biding. 37 Similarly, the interaction of THRs with RARs also increases the affinity of the DNA binding. A study performed in monkey kidneys showed that the interaction of both receptors had a positive effect in the transcriptional rate of the gene whose promoter contains a response element for the thyroid hormone, while it also represses those genes that did not have it. This result suggests that the formation of the heterodimers allows for a better control of the transcription rate, and it also hints to the idea that both RAR and THR control the expression of genes in overlapping networks.32, 38 Retinoid receptors are involved in a wide range of biological processes that include embryonic development, growth, differentiation, apoptosis, and spermatogenesis. Their functions are not only limited to development, but they also play important roles in adults to maintain proper function of the lungs, liver, and neuronal and immune systems. 39 In brain, RARs and RXRs are expressed in multiple stages (Table 2) and play a major role in the development and function of the CNS.
During brain cortex development, RA is synthesized in a tissue-specific manner by the aldehyde dehydrogenase (ALDH) family of enzymes40–42 that is composed of the following three members: ALDH1A1, ALDH1A2, and ALDH1A3. ALDHs have distinct and specific expression patterns that correlate with RA activity as detected by reporter transgenes. 43 In brain, ALDH1A3 is primarily expressed in the lateral ganglionic eminence, and ALDH1A1 is expressed in the dopamine-containing mesostriatal system and meninges, while ALDH1A2 is present only in the meninges. Consistent with the presence of ALDH1A3 and ALDH1A1 in the lateral ganglionic eminence and its derivative striatum, where high levels of RA and other retinoids are detected in the developing striatum.40–42 RA is derived from the liposoluble vitamin A and is critical for vision, since its derivative (retinaldehyde) is the light-sensitive molecule, which triggers the phototransduction process in photoreceptor cells in the retina. 44 Vitamin A and RA are essential for eye development and maintenance and serve a similar role in brain cortex development. Young animals raised on a vitamin A-deficient diet grow poorly, become blind, and have a low survival rate.45, 46 The teratogenic effect of RARs was demonstrated in a study using the antagonist of RARs (AGN193109). An-or microphthalmia was observed in 75% of treated mice. 47 Additionally, a recent study revealed mutations in human ALDH1A3 is associated with microphthalmia and anophthalmia.48–53
RA biological action is to bind to RARs, switching them from transcription repressive elements to transcription activator elements, while they are bound to RAR elements (RAREs).54, 55 In contrast, RARβhas been involved in brain structure development and maintenance; especially in ventral and dorsal telencephalon development, hippocampal plasticity, and brain barrier development.56–59 In addition, RARβis expressed with other RARs and RXRs in adult mice and human brains (Table 2). Other studies revealed that Rarβexpression is inducible by adding external RA at stage E15 to rat brain cortex cultures. Therefore, increases in Rarβexpression by RA in brain cortex may be due to a partial depressive mechanism as
Knockout Mice Models Excising that Effect the CNS Function.
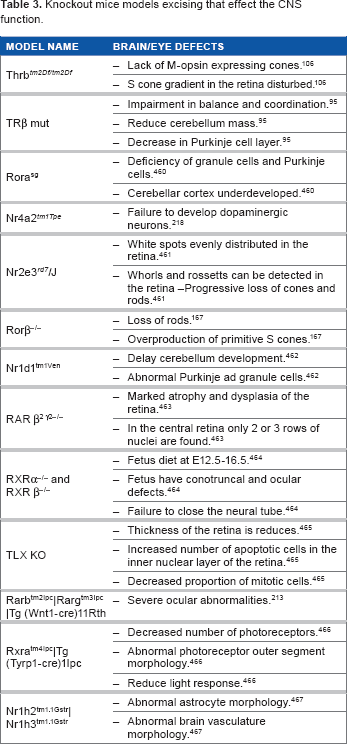
Thyroid Hormone Receptor Family and the CNS
The thyroid hormone receptor (THR) family plays a major role in the development of brain, regulation of metabolic rate, and the development of photoreceptor cells in the vertebrate retina. THRs are involved in a great number of biological activities that are both hormone dependent and independent. Some of the activities in which this receptor is involved include regulation of the
TRs are members of Class II, meaning that they predominantly bind to RXR. However, THRs do not bind exclusively to RXR, and they can also form heterodimers with VDR and PPAR.73, 74 The receptors of this family are coded by two genes, such as thyroid hormone receptor α (
Thyroid hormone receptors regulate several developmental events in the CNS, including neuronal differentiation and migration, synaptogenesis, and myelination.78–80 T3, T4, and TRs are present in the developing cortex, prior to the onset of fetal thyroid gland activity, before week 12 in humans. This suggests that maternal thyroid hormone (TH) concentration is important during brain development.81, 82 Thyroid hormone influence in the CNS is well documented through studies of TH in the cerebellum.
83
Two genes encode at least three high-affinity THRs in mammals, THRβ1 (
Several isoforms of the thyroid nuclear receptors are expressed in the eye. THR form complex with activators, such as SMART and histone deacetylases (HDACs) to regulate the transcription activity of the target genes.
102
The receptor THRα is found in the progenitor layer, while THRβ1 is found in the inner nuclear layer, and THRβ2 is expressed in the photoreceptor layer in particular in the developing cones.85, 102, 103 TRs play an important role in the development of the retina by participating in the signaling system by which the progenitor cells are differentiated. It has been determined that the abundance of T3 in the environment can induce the proliferation of cone photoreceptor cells.
104
In general, thyroid hormone receptors act as coactivators in the presence of T3, and in the absence, they repress the expression of their target gene.
105
It has been demonstrated that in
PPARs Family and the CNS
The PPAR family was first discovered for its role in increasing the quantities of peroxisomes in different animal species and increasing the transcription of genes that encode for fatty acids. 107 Since this discovery, several additional roles have been discovered for these receptors, such as lipid metabolism, inflammation, glucose homeostasis, and cell differentiation. It has become clear that PPARs have an important role in body's response toward exogenous ligands. 108 Three different isoforms have been identified for PPAR: α, β/Δ, and γ. Each isoform has a unique pattern of expression and has affinity to different ligands. PPARα is an important factor regulating lipid metabolism and is mostly found in the heart, kidney, intestine, and fat tissue. 12 In contrast, PPARβ/Δ is found predominantly in different areas of adult brain (Table 2), kidney, and small intestine, is involved in cell survival, and is often present in the formation of tumors in the colon. For this reason, several studies have focused on understanding how PPAR might be an important therapeutic target. It was discovered that the nonsteroidal anti-inflammatory drugs compete with PPARβ/Δ for its responsive elements inhibiting the action of PPARβ/Δ and eventually reducing tumorigenesis. 109 PPARγ is mostly found in the spleen, intestine, and adipose tissue. 12 PPARs have the ability to bind to different types of fatty acids as well as synthesized compounds. Their ability to bind to different types of molecules is most likely due to the spacious ligand-binding pocket that is observed in the crystallographic structure. 110 While PPARs can be found in different tissues, they are predominantly expressed in tissues that regulate energy homeostasis. The fact that the majority of ligands are fatty acids and other metabolic molecules is evidence that PPARs have an important role in the regulation of metabolism. For example, the signal molecule eicosanoids is a product of fatty acid metabolism and is also known for being an inflammatory agent. This led to the discovery of a second function of PPARs, which is inflammatory signaling regulated in particular by PPARα and PPARγ.111, 112 Since Alzheimer's disease has an inflammatory component to its pathology as a consequence of the neurodegeneration and the extracellular deposition of β-amyloid peptides, recent studies have focused on the protective effects of nonsteroidal anti-inflammatory drugs in delaying or reducing the risk of Alzheimer's disease. 113 PPARs as many other receptors form heterodimers with RXR. Therefore, there is evidence of competition between PPARs and receptors, such as TRs and vitamin D, which also partner with RXR. As an evidence of this competition, it was observed that the transcription of the genes that are regulated by PPARs was repressed in the presence of T3. 114 The fact that different receptors compete for their heterodimer partners is evidence of the complexity of gene regulation and the dual roles that the nuclear receptors play. The PPARs are an excellent example of previous orphans whose ligands were finally discovered; for all other receptors whose ligands still remain to be discern, they conform the ONR family. 115
ONRs and the CNS
Orphan receptors are a unique class of NHRs in that they do not have a ligand identified and often lack all of the conserved domains typical of NHRs. Furthermore, a high degree of diversification has been observed in this group of NHRs, as many of them are not evolutionary or functionally related. This is evidenced in a phylogenetic analysis when aligning the DBD and LBD sequences. The resulting evolutionary tree has six major branches with the orphan receptors distributed across all branches.116, 117 The degree of diversification of orphan NHRs is further illustrated by studies revealing that some orphan receptors in invertebrate species such as
Another striking difference of orphan NHRs is the size of the A/B region that can range from a minimal eight amino acids in the retinoid orphan receptor (ROR) βto 280 amino acids in the NGFI-B receptor. 30 Some orphan receptors lack the activation function AF-1, while others lack an AF-2 domain. Many orphan receptors form heterodimers with RXR, while the remaining orphan receptors bind to DNA as homodimers. 3 The classification system divides the ONRs into seven different groups starting with group 0. Group I is comprised of seven different families, Group II is comprised of 5 families while group III, IV, V and VI contain one family.2, 12, 120
The NHR Nr2e3 was first identified as photoreceptor-specific nuclear receptor. Unlike most NHRs that exhibit ubiquitous expression, expression of NR2e3 is restricted solely to the outer nuclear layer of the neurosensory retina (cones and rods; Fig. 5).121–123 During human eye development, Nr2e3 transcription is observed at week 11.7 in the putative immature human rods on the foveal edge.
124
Many studies performed using the Nr2e3rd7/rd7 mouse, which lacks a functional Nr2e3 gene, indicate that Nr2e3 has a dual regulation role regulating rod and cone development and function.60, 121, 125 Lack of Nr2e3 leads to abnormal proliferation of blue cones (the least populous of the photoreceptors) as a consequence of the loss of Nr2e3 in the mitotic progenitor cell program.
126
In rods, Nr2e3 does not affect rhodopsin expression per se, although it seems that rhodopsin expression needs both Nr1d1 and Nr2e3 potentiated by the orthodenticle-like cone-rod homeobox transcription factor (CRX)-and the neural retina basic motif-leucine zipper transcription factor (NRL)-driven activities.
127
NR2E3 works in conjunction with multiple transcription factors, including CRX, which is detected initially at day E12.5 in retinal progenitor cells and newly postmitotic cells,128–130 and NRL, which is detected at day E14.5 and directs the generation of rod photoreceptor cells.131, 132 Although previous studies have identified several genes that are differentially expressed between normal and knockout mice, the transcriptional network regulated by Nr2e3 is not fully understood. Some potential genes that are regulated by Nr2e3 include the following cone-specific phototransduction genes: blue opsin (Opn-SW), GnαaLt2 and GnβbT3 cone transducing subunits, the rod-specific gene guanine nucleotide-binding protein 1 ( List of NHRs Involved in the Differentiation of the Mitotic Progenitor Cells and the Retinal Cell Types. Genes Marked in Bold Correspond to NHRs Unique for Each Cell Type.
The RORs and the Rev-Erb are two of the receptor families that belong to Group I. The ROR family is made up of the following three members: RORα, RORβ, and RORγ. RORα is ubiquitously expressed and is involved in the development of Purkinje cells (Table 2), bone metabolism, immune response, development of cone photoreceptors of the retina, bone morphogenesis, angiogenesis, and lipid metabolism. 134 RORβis mostly found in the retina, 135 brain (Table 2),135–138 and spleen and functions in maintenance of the circadian rhythm.139–141 RORγ is found in the liver, kidney, thymus, and lungs, and its functions consist of lymph node organogenesis and thymopoiesis.1, 141–145 Each member of the family also has different isoforms that are formed by alternative splicing and alternative promoters. 146 The isoforms differ in the A/B domain: four ROR isoforms have been identified in humans and two have been identified in mice. RORα1 and 4 are localized in the cerebellum. 146 RORβhas two known isoforms; RORβ2 is restricted to the pineal gland and the retina (Fig. 5). 135
Both RORα and RORβhave an important role in the CNS, as observed by knockout mouse models that have severe cerebral abnormalities.147, 148 Cerebellum development is regulated by RORs and takes a large period during fetal development. The main types of cells in the cerebellum develop at different stages and positions. The Purkinje cells arise from the ventricular zone of the metencephalic alar plates at the fifth or sixth week of development. 149 Granule cells are added from the rhombic lip at the end of the embryonic period, and their migration is facilitated by Bergmann glia cells to their deeper definitive site. 150 During this process, RORα expression is restricted to Purkinje cells and not to the granule cell layer, as observed in mice at age E12.5.109–116
The staggerer mouse, which lacks functional RORα (RORαsg/sg or RORα-/-), presents with fewer Purkinje cells and a loss of cerebellar granule cells.147, 151–157 RORα heterozygous mice (RORα+/sg) with only one functional allele exhibit a milder phenotype and present with an accelerated Purkinje and granule cells loss in aged animals.154, 158–160 RORα regulates genes related to Purkinje cells maturity, in particular its dendritic differentiation. RORα-/--deficient mice show downregulation of several genes involved in signal-dependent calcium release, including the calmodulin inhibitors
RORβplays an important role in retina cell differentiation as studies in murine models have described. Differentiation of cone photoreceptor cells in mice is regulated by the nuclear receptors Rorβand Rxrγ (Fig. 5), which control the generation of short(S)-opsin-expressing cones (also referred to as blue cones).165, 166 RORβalso stimulates rod photoreceptor differentiation by activating the rod-selective transcription factor neural retina leucine zipper (
NHRs regulate gene expression in a common pool of multipotent retinal progenitor cells found in the neuroephithelium of the optic cup to form all retinal cell types (Fig. 5). 169 Retinal proliferation is a highly conserved dynamic process in vertebrates that occurs in the following two overlapping phases: (1) the first phase begins with the proliferation of ganglion cells followed by horizontal cells, cone photoreceptors, and amacrine cells and (2) the second phase overlaps with the generation of rod photoreceptors, bipolar cells, and MüuUller glial cells. The differentiation of retinal cells also has overlapping phases. 170 The formation and maintenance of the different types of cells in the retina requires specific gene regulation. The transcription and regulation of NHRs, including Nr2e3, RARβ, RORβ, Nr1d1, RXRγ, and Trβ (Fig. 5), in synergy with several transcription factors, including CRX, NRL, and Chx10, help determine retinal cell fate and pluripotent capacity of retinal progenitors.171, 172
The Rev-Erb members are characterized for having unique features, such as their transcription from the opposite strand of the T3 receptor gene. 173 They also lack the AF-2 domain and mostly function as repressors of gene expression by associating with the corepressors NCoR.24, 174–176 Rev-Erb has been associated with several functions, such as being a mediator of adiponogenesis, 177 the regulator of myogenesis, 178 involvement with cell proliferation, 34 the regulator of the circadian clock, 179 and effects on lipid metabolism. 180
The testicular nuclear receptors (TRs) and tailless (TLX) receptor are two of the five members of Group II of the orphan receptors. There are two members of the testicular nuclear receptors (TR2 and TR4). TR2 is highly expressed in the testes as well as in several other tissues, including brain, retina, kidney, and intestine, with the highest peak of expression during embryonic time points. 181 TR2 can bind as a homodimer or as a heterodimer, and it has the ability to act as a repressor of gene expression. 182 TR4 is expressed in several tissues from the CNS, adrenal gland, spleen, and prostate gland with the highest expression observed in the skeleton and testes, as indicated by the name of the family. 181 TR4 is expressed in a temporal and spatial manner during brain development, 183 and TR4 knockout mice show growth defects, and females exhibited poor maternal behavior skills. 184
These behavioral traits in TR4 knockout rodent animals seem to be the consequence of a poor cerebellum development. During this process, TR4 expression patterns correlate directly with cerebellum neurogenesis, suggesting a role in this stage.183, 185 During postnatal cerebellum development, several changes occur in the cerebellum laminar construction with engrossing of the external granule layer followed by granule cells migration toward the internal granule layer. Precursors of these cells migrate as well to the internal granule layer guided by Bergmann fibers, which expresses the astrocyte-specific glutamate transporter, GLAST.150, 186, 187 This migration process is affected fundamentally by the order of the Bergmann fibers. If these fibers are disorganized or atrophied, migration will not occur. Additionally, migration impairment can also be the consequence of defects in granule cells.188, 189 Mouse models lacking TR4 exhibit abnormal cerebellar development in lobules VI-VII, which correlate with poor granule cell migration and Bergmann glia maturation.190–195 These alterations are related to a lack of GLAST, causing a long-lasting negative impact on neural connectivity pathways during brain development. 196 As a consequence of TR4 knockout, aberrant neuronal connectivity has been observed due to the delay in granule cell migration and the disarrangement of the cerebellar Purkinje neurons. The result is an abnormal response to environmental changes and also problems with social contact, startle activity, and deficits in prepulse startle inhibition and cerebellar motor learning when no motor coordination seems to be present. 196
The TLX receptor is the human homolog of the tailless gene in
Group IV is composed of one receptor member with three different subtypes, α, β, and γ. This family, also known as Nr4a, is expressed in adult mice and human brain (Table 2). Nr4a receptors play an important role in the retina and cerebellum development and
NHRs Associated with Diseases in the Eye and Brain
Nuclear Hormone Receptor and Associated Diseases in Humans.
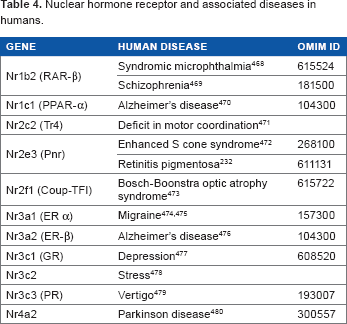
In the brain, RARs are expressed with different location patterns: RARα is found ubiquitously, and RARβand RARγ are found in the striated regions that have dopaminergic neurons.
215
Dopamine receptor 2 knockout mouse phenotype resembles the human Parkinson's disease.
216
Parkinson's disease is characterized by the degeneration of the dopaminergic pathway. Both
The most common disease associated with RARα mutations is acute promyelocytic leukemia (MIM: 612376), caused by gene rearrangements. Often, a gene fusion of promyelocytic leukemia (PML) and RARα occurs due to the translocation t(15,17)(q21;q22). 220 However, though most of the rearrangements involving RARα are associated with different types of leukemia, one individual diagnosed with Smith-Magenis syndrome presented a region translocation of chromosome 17 q11.2-q21.3 that was inserted into the short arm of the same chromosome at region p11.2. 221 Patients with Smith-Magenis syndrome have multiple congenital anomalies, including mental retardation, which are associated with a deletion involving band p11.2 of chromosome 17. 222 Though the report given by Park et al 221 did not expose RARα translocation as causative of Smith-Magenis syndrome, it shows how chromosome 17 q11.2-q21.3 region is a breakage and rearrangement hotspot, same as chromosome 17 p11.2 region, which may cause different related syndromes 221 not directly associated with the CNS.
Unlike RARα, RARβis important for the proper development of the brain cortex, and variants in this gene have been associated with locomotor disease and ocular disorders. RXRB and RARβexpression have been observed in the striatum. 223 Several polymorphisms in RARβare associated with microphthalmia in humans (MIM: 615524). Double knockout mutants for RARβand RXRγ have a 30%-40% reduction in D1 and D2 dopamine receptor expression, though they are the most abundant type of receptors in the striatum, 223 and exhibit reduction in forward locomotion. In the case of D2, its lower expression is possibly explained by the presence of RAREs in its promoter region. 224 This fact permits hypothesize that RARβand RXRγ are essential in developing the proper locomotion, and it is involved in controlling the dopaminergic system during development and adult stages in CNS.
RORα gene networks appear to play an important role in the neurodegenerative disease spinocerebellar ataxia-1. A microarray analysis from mutant and normal mouse brain showed that genes regulated by RORα were downregulated in the first stages of development of the Purkinje cells. Loss of RORα leads to an increase expression of the ataxia gene ATXN1. A further analysis by coimmunoprecipitation revealed that ATXN1, RORα, and the RORα coactivator Tip60 form a complex to modulate cerebellar development. The RORα1 knockout model lacks the LDB region and is characterized as having abnormal Purkinje cells and abnormal morphology and ataxic.225, 226 The staggerer mouse, which has a deletion in the RORα gene that blocks the translation of LBD, models cerebellar ataxia affecting the development of the Purkinje cells. RORα-/- also shows atrophy and loss of the dendritic cells and synaptic rearrangement of the Purkinje cells. 227 The phenotype of the staggerer mouse suggests that RORα is involved in the thyroid hormone signaling pathway in charge of the maturation of the Purkinje cells. 227 Different studies have demonstrated that RORα is also involved in the protection of neurons, suppresses inflammation, and is important in regulating circadian rhythm. 228
NR2E3 has been associated with several retinal diseases, including enhanced S-cone syndrome (ESCS), 229 Goldmann-Favre syndrome (GFS), and retinitis pigmentosa.230–232 ESCS and GFS are autosomal recessive diseases characterized by an increase in the sensitivity of the S (blue wavelength)-cone photoreceptor cells accompanied with a decreased in the rods and L/M (red/green) cones.11, 233, 234 GFS is a milder form of ESCS. Patients with autosomal dominant retinitis pigmentosa suffer from early onset loss of night vision followed by retinal spots in the peripheral field, and eventually loss of central vision. By analyzing the phenotypes of patients with different diseases caused by mutations in the Nr2e3, it is possible to conclude that the location of the mutation has a major impact on disease outcome. The diseases are viewed as a collection of Nr2e3-associated retinal diseases. 235 NHRs are involved in a myriad of functions that cover more than just the CNS normal functioning. Yet, it is still important to understand how NHRs participate in diseases, as NHRs play a key role in normal CNS development.
Perspectives and Conclusions
This review discusses the important role of NHRs in the development, function, and maintenance of the CNS. There is great variation in the types of isoforms, as well as the abundance and time of expression of NHRs throughout the CNS. Furthermore, it is not uncommon to see members of a particular family with multiple functions as illustrated by the ROR and TR families of NHRs. All members of the THR family, for example, are involved in the development and/or function of the CNS. In contrast, RARβsimply has one member involved in a specific tissue of the CNS, and even more restrictive is the orphan receptor NR2E3 that functions in only one type of retinal cell, the photoreceptor cell. Thus, while most NHRs are ubiquitously expressed, others have family members or isoforms with expression patterns limited to particular in organs or cell types. There is a clear overlap in NHRs signaling of the developing eye and brain, suggesting a common template for CNS development. The majority of NHRs have a dual role as both transcription activators and suppressors. Their function depends on their temporal and spatial expression, cellular context, as well as the interaction with other factors, ligands, and DNA sequences of HREs. Furthermore, NHRs can modulate gene expression, via HRE recognition, by several mechanisms, including ligand binding and cofactor recruitment. A comprehensive reconstruction of the complex, multimodal interacting gene networks and identifying the target genes they regulate will provide greater insight into overall neuronal signaling of the CNS.
It is well established that NHRs regulate numerous biological pathways and processes. NHRs interact with several factors, such as transcription factors, chromatin-modifying complexes, and hormones. NHRs often serve as master regulators simultaneously modulating several signaling pathways in a temporal/spatial/cell type-specific manner. The highly conserved nature of NHRs and their numerous isoforms throughout the CNS suggests functional redundancy and a possible mechanism to avoid disease. This is evidenced in data demonstrating that NHRs can serve as genetic modifiers that alter disease onset, progression, or severity. 236 This new role for NHRs as genetic modifiers of disease is important in understanding normal CNS development and has great therapeutic implications. NHRs that function as master regulators of key biological pathways may have a profound impact on disease treatment as they have the potential to impact multiple mutations and multiple pathways.
Author Contributions
Conceived and designed the experiments: AMO, OAM-R, and NBH. Analyzed the data: AMO, OAM-R. Contributed to the writing of the article: AMO, OAM-R, and NBH. Agreed the manuscript results and conclusions: AMO, OAM-R, and NBH. Jointly developed the structure and arguments for the article: AMO and OAM-R. Made critical revisions and approved the final version: NBH. All authors reviewed and approved the final article.
